Abstract
The interaction between chemokines and their receptors is crucial for inflammatory cell trafficking. CCL14 binds with high affinity to CCR5. In leporids, CCR5 underwent gene conversion with CCR2. The study of CCR5 ligands in leporid species showed that CCL8 is pseudogenized, while CCL3, CCL4 and CCL5 are functional. Here, we study the evolution of CCL14 in mammals with emphasis in the order Lagomorpha. By employing maximum likelihood methods we detected six sites under positive selection. Some of these sites are located in regions crucial for CCL14 activation and binding to receptors. Sequencing of CCL14 in Ochotona species showed that O. princeps, O. pallasi, O. alpina and O. turuchanensis have a mutation at the start codon (Met > Thr), while O. hoffmanni, O. mantchurica, O. dauurica and O. rufescens present the mammalian conserved Met. Ochotona hyperborea has the two alleles. In O. pusilla, CCL14 is a pseudogene due to a seven base pair insertion. Like CCL3, CCL4 and CCL5, CCL14 is functional in all leporids but in the Ochotonidae family it underwent a pseudogenization process. This suggests that CCL14 has an important biological role in other mammals by evolving under positive selection that has been lost in Ochotonidae (subgenera Pika and Lagotona).
Introduction
Chemokines are proteins that have several functions in the immune system and are found only in vertebrates. 1 These proteins are encoded by a multigene family that emerged by gene duplication events. 1 Gene duplication is considered a powerful force in the adaptive evolution of genes, in which one gene gives rise to new copies.2–4 These new copies can evolve by pseudogenization, neofunctionalization or subfunctionalization.3,5,6 Pseudogenization is the process in which a redundant functional copy passes through mutational decay and loses its ability to produce a functional protein.6,7 In the remaining processes, both copies escape to mutation as either one copy emerges into new functions (neofunctionalization) or either divides the functions with the other copy (subfunctionalization).3,5,8
Chemokines can only exert their functions through binding to specific receptors, having the ability to bind and activate different receptors; for example, CCL14 binds with high affinity to the chemokine receptors CCR1, CCR3 and CCR5. 9 CCR5 is used as co-receptor by HIV type-1 (HIV-1) for infection and, despite controversy, it has been also associated with myxoma virus infection in rabbits.10–17 In some mammalian species, CCR5 underwent gene conversion with CCR2.18–23 Several studies have been conducted in the CCR5 ligands to determine the consequences of the CCR5–CCR2 gene conversion and its putative association with the myxoma virus infection. Indeed, the study of CCL8, a potential CCR5 ligand, revealed that in some leporid species, such as Oryctolagus and Bunolagus, this ligand is pseudogenized, whilst in Lepus and Sylvilagus is intact. 12 A study on CCL3, CCL4 and CCL5 showed that, unlike CCL8, these genes are all functional in leporids. 24 Although mouse CCL12 is the true ortholog to human CCL8, 25 in leporids, the gene identified as CCL8 is the true ortholog, as supported by their phylogenetic position within the mammalian CCL8 cluster. 12
In leporids, other CCR5 ligands remain to be studied, including CCL14. CCL14, also known as hemofiltrate CC-chemokine 1 (HCC-1), is found in high concentrations in blood plasma and acts as an inflammatory chemokine after N-terminus cleavage.15,26,27 In humans, this protein is located in the macrophage inflammatory protein (MIP) region of CC chemokine genes,1,28 and is expressed in numerous tissues.9,15,27,29 In humans, CCL14 encodes a protein with 93 aa with three predicted cleavage sites (Thr25, Glu31 and Ser33) that are necessary for the protein to become active. 30 There are two described variants: one encoded by three exons in which the active CCL14 consists of a 74 aa protein after the signal peptide cleavage, and another variant encoded by four exons, which has an insertion of 48 nucleotides between exons 1 and 2 encoding 16 aa.1,15,17,27 These forms have no effect against HIV-1 infection; however, the cleaved form CCL14 (9–74) has the ability to block HIV-1 entry and replication. 16 CCL14 (9–74) is an agonist of CCR1, CCR3 and CCR5, which promotes calcium influx and migration of T lymphocytes, monocytes and eosinophils; through the interaction with the second external loop of CCR5, it has also the ability to inhibit this receptor.15–17,27 In CC chemokines there are two main interactions important for receptor binding and activation, both located in the N-terminus close to the CC motif. In CCL14, the first interaction corresponds to the high-affinity binding to the receptor and includes residues located after the CC motif, and the second interaction comprises the residues located before the CC motif and that are essential for the receptor activation.15,31–34 The N-terminus is also important for CCL14 degradation. 35 Degradation occurs following the binding of D6 to the N-terminus of CCL14 and then through the interaction with the residue Pro53 of CCL14.35,36
The order Lagomorpha includes two families, Ochotonidae (pikas) and Leporidae (hares and rabbits) that diverged ∼35 million years ago. 37 The Ochotonidae family is composed of only one genus, Ochotona, which is divided into four subgenera, Pika, Ochotona, Conothoa and Lagotona. 38 The Leporidae family is divided into 11 genera. 37 Rodents and lagomorphs form the superorder Glires that is considered a sister group of the superorder Euarchonta that includes primates.39,40 In rodents, CCL14 was shown to be absent in mouse and rat, while present in squirrel and guinea pig; 28 in lagomorphs, information is only available for the European rabbit (Oryctolagus cuniculus), where a functional gene exists.
By comparing the O. princeps CCL14 coding sequence (CDS) available in GenBank/Ensembl with the remaining mammalian species it was possible to observe that the initiation codon typically present in mammals is mutated into a Thr (ACG) (Figure 1). Following this observation, and considering the apparent ‘random’ absence of CCL14 in rodents and its role as a ligand for CCR5 which in some lagomorphs underwent gene conversion with CCR2, we aimed at determining the presence and function conservation of CCL14 in lagomorphs. In addition, we studied the evolution of CCL14 in mammals by identifying codons that are evolving under positive selection.
(A) Alignment of CCL14 for the different mammalian species (GenBank, Ensembl and Uniprot accession numbers are indicated for the retrieved sequences). Positively selected amino acids are boxed and the three possible cleavage sites are shaded in dark grey. (*) Represent stop codons, (–) represent deletions, *1 and *2 represent alleles. (B) Alignment of CCL14 for the different Ochotona species studied. The characteristic mammalian initiation codon is boxed. (C) Alignment of the CCL14 for O. princeps and O. pusilla with the insertion of the seven base pairs shaded in grey.
Material and methods
Mammalian CCL14 sequences were retrieved from public databases (accession numbers are given in Figure 1). Sequences were aligned using MUltiple Sequence Comparison by Log-Expectation (MUSCLE) available at http://www.ebi.ac.uk/. 41 Similar results were obtained using Multiple Alignment using Fast Fourier Transform (MAFFT); however, MUSCLE alignment introduced less gaps, leading to a more conserved alignment.
In order to determine the impact of positive selection on CCL14 sequence evolution, we estimated ω, i.e, the ratio of nonsynonymous (dN) to synonymous (dS) differences in functional CCL14 orthologs (final dataset of 55 sequences). The codon-based maximum likelihood (ML) method (CODEML) implemented in PAML v4.4 was used. 42 An un-rooted neighbor-joining tree was constructed using MEGA6, 43 with p-distance as substitution model and the pairwise deletion option for gaps/missing data. The topology of the phylogenetic tree obtained is in accordance with the accepted mammalian phylogeny. Two pairs of site-based models were compared: M1 (nearly neutral) vs. M2 (selection) and M7 (neutral, beta) vs. M8 (selection, beta and ω), where M1 and M7 correspond to the null hypothesis and M2 and M8 to the alternative hypothesis by allowing positive selection. A likelihood ratio test with two degrees of freedom determined if a selection model fitted better the data than a neutral model.44,45 Codons under positive selection were identified by using a Bayes Empirical Bayes approach with probability >95%. Codon-based ML methods available in the HYPHY package implemented in the DataMonkey webserver were also used:46,47 Single Likelihood Ancestor Counting (SLAC), Fixed-Effect Likelihood (FEL), Internal Branch Fel (iFEL), Mixed Effects Model of Evolution (MEME), Random Effect Likelihood (REL) and Fast Unconstrained Bayesian AppRoximation (FUBAR).47–49 For the first four methods the P-value was set to ≤ 0.05; for FUBAR we used a P-value ≥ 0.95; and for REL we used a Bayes factor >95. The best fitting model for nucleotide substitution was determined by the automatic model selection available in the webserver. As done previously for other immunity genes,50–55 only the codons detected by more than one method were considered as being under positive selection.
Primers and conditions used in CCL14 PCR amplification.
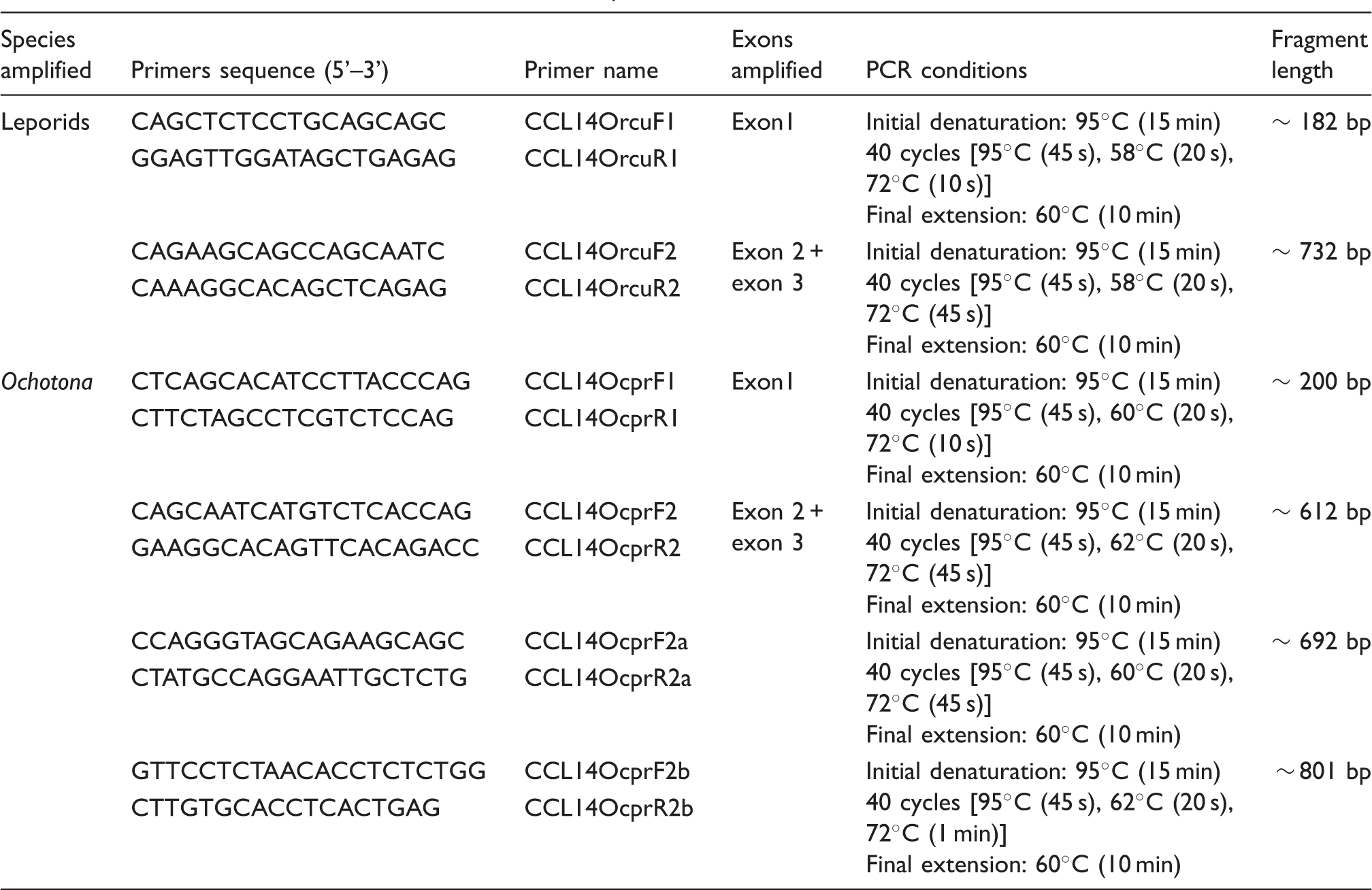
The program PHASE, built into the DnaSP software, 56 was used to reconstruct the haplotype phases of the obtained sequences. These sequences were aligned and translated using BioEdit. 57
The Tajima’s Relative Rate Test was conducted in order to check the evolutionary rate of CCL14 between lagomorphs using MEGA6.43,58
Results and discussion
In our study, six aa were identified as being under positive selection: Ala22, Thr25, Thr62, Lys67, Met74 and His96 (aa residues were numbered from the first methionine residue in human CCL14. The signal peptide and indels, indicated as (–), were included in the numbering) (Figure 1). Ala22 and Thr25 are located in the pre-peptide, which is cleaved in order for the protein to become active; 17 however, the modifications observed at residue 22 are not expected to alter the CCL14 protein. In contrast, Thr25 is located within the region where CCL14 cleavage occurs. As cleavage is required for protein activation, alterations in this site might negatively affect the CCL14 protein. Thr62 and Lys67 are in the region reported as to be essential to the binding of CCL14 to the several receptors;15,31–34 thus, changes in this region may not only affect both the ligand–receptor binding, but also CCL14 degradation.35,36 Met74 and His96 are located in the β1 and β3 sheets, respectively. Alterations in these sites may lead to alterations in the protein conformation.
The comparison of the O. princeps CCL14 CDS available in GenBank and Ensembl with the remaining mammalian species showed that in O. princeps the initiation codon, in contrast to the other mammalian sequences, is mutated into a Thr (ACG) (Figure 1A). Following this observation, we attempted the amplification of CCL14 from gDNA of O. princeps. Our results indicated that, as for the sequences publicly available, the amplified fragment also presents the ACG mutation. Next, we attempted to amplify CCL14 from cDNA of O. princeps. Despite having used different Taq polymerases and different PCR conditions (e.g. including the addition of DMSO in the reaction, increasing extension times) and different pairs of primers designed according to the O. princeps CDS, and having previously successfully amplified other CDS of other genes from the same sample, 59 we were unable to amplify the CCL14. This may suggest that CCL14 is a pseudogene in O. princeps. We amplified and sequenced the CCL14 gene for eight leporid species: O. cuniculus cuniculus and O. c. algirus, Brachylagus idahoensis, Pentalagus furnessi, Sylvilagus bachmani and S. floridanus, Lepus europaeus, Romerolagus diazi.
Results obtained in Tajima’s Relative Rate Test.
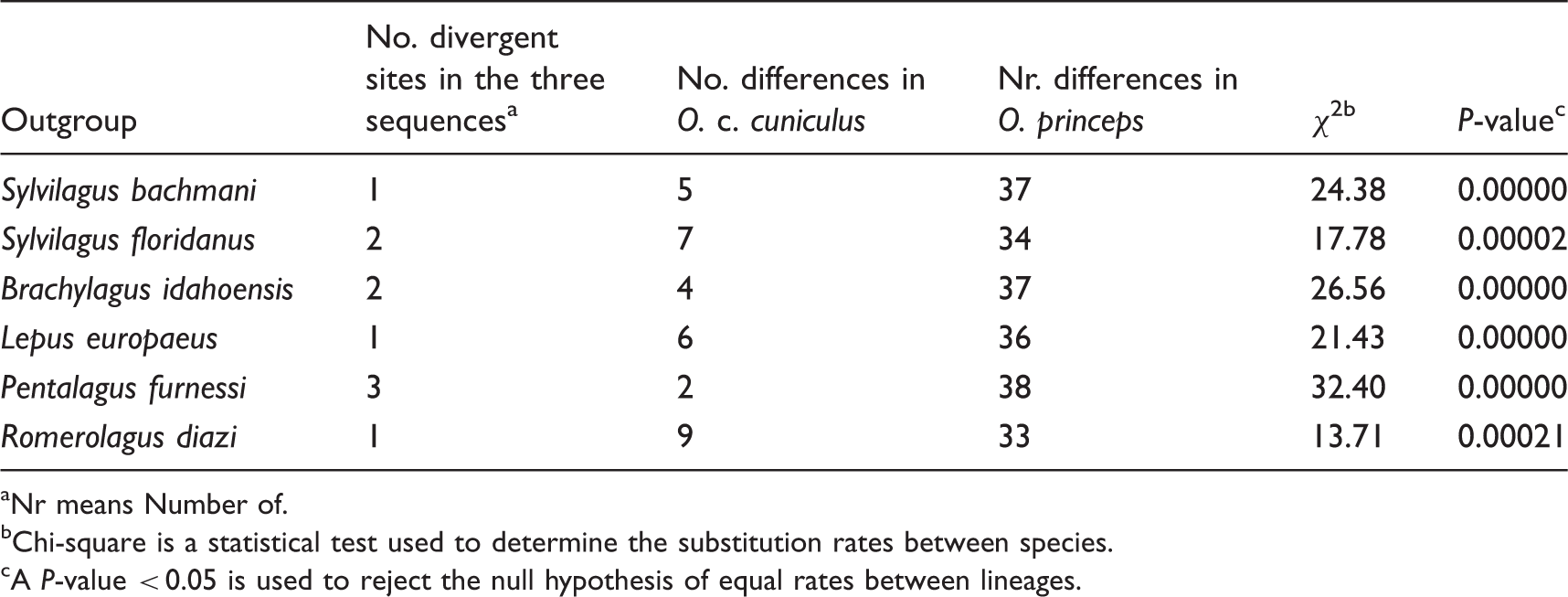
Nr means Number of.
Chi-square is a statistical test used to determine the substitution rates between species.
A P-value < 0.05 is used to reject the null hypothesis of equal rates between lineages.
We attempted to amplify the CCL14 gene from gDNA of several pika species that encompass the different Ochotona subgenera: O. alpina, O. dauurica, O. hyperborea, O. hoffmanni, O. mantchurica, O. pallasi, O. pusilla, O. rufescens and O. turuchanensis. Interestingly, we found different results within the Ochotonidae family (Figure 1B, C). Indeed, O. alpina, O. pallasi and O. turuchanensis, share the ATG > ACG mutation with O. princeps; in contrast, in O. dauurica, O. hoffmanni, O. mantchurica and O. rufescens, as in all the other mammals examined, the putative initiation codon is conserved. In addition, in O. hyperborea, we could define two alleles, one with the putative initiation codon and the other with ACG. In O. pusilla, despite presenting the putative start codon, CCL14 seems to be a pseudogene due to an insertion of seven base pairs (base pairs 43–49) that leads to a frameshift that disrupts the CDS (Figure 1C). Our results seem to indicate that in the Ochotonidae family subgenus Pika CCL14 is under a process of pseudogenization with inactivation of the gene in some, but not all, species; this pseudogenization, however, does not reflect the taxonomic relationships in the Ochotona genus (Figure 2). Indeed, the subgenus Pika where the mutation in the start codon emerged encompasses species from different lineages that present both the ATG and the ACG.38,60 Ochotona pusilla is the only species studied where CCL14 is obviously a pseudogene (Figure 1C), and belongs to a different subgenus (Lagotona).38,60–63
Evolutionary topology showing the molecular phylogeny within the Ochotonidae family (adapted from Melo-Ferreira et al.
60
). In the AYG found in O. hyperborea, Y is a pyrimidine indicative that this species has two alleles, one with ATG and other with ACG.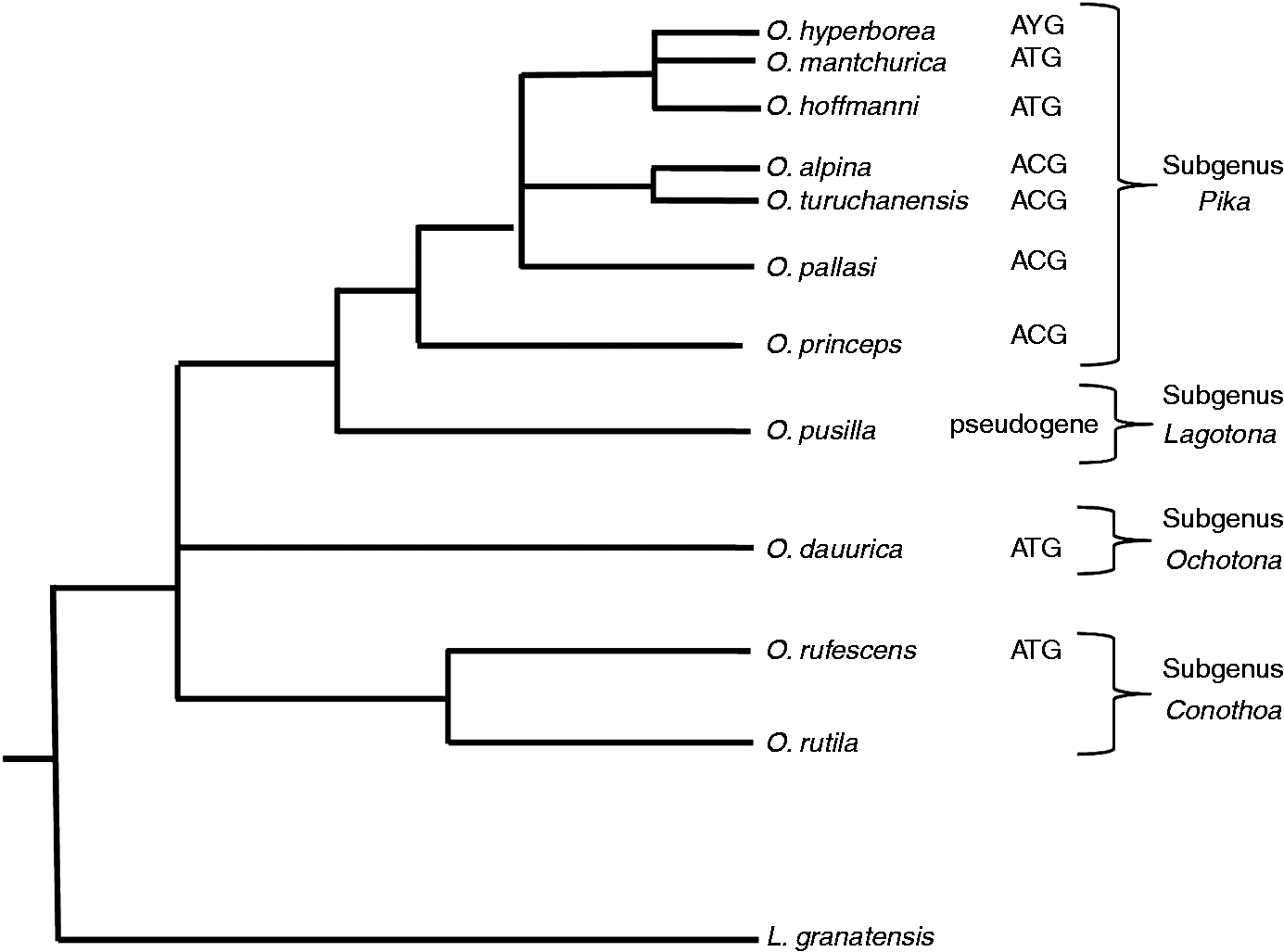
Taking into account the CCL14 alterations described in the Ochotona species, we also determined if those alterations could have had any influence in the receptors. For this we sequenced the CCR5 and the CCR1 genes (data not shown). However, these sequences did not reveal alterations that could explain the pseudogenization of CCL14.
In the human genome, pseudogenes are quite common, with high prevalence in multigene families such as chemokines, 64 but little is known for other mammals. For CCL14, it was previously shown that in Rodentia, CCL14 is absent in rat and mouse of the Muridae family, 28 but is a functional gene in the families Dipodidae (Jaculus jaculus), Spalacidae (Nannospalax galili), Sciuridae (Spermophilus tridecemlineatus), Bathyergidae (Heterocephalus glaber), Chinchillidae (Chinchilla laniger) and Caviidae (Cavia porcellus). In addition, in the Trichechus manatus latirostris (Florida manatee) that belongs to the superorder Afrotheria, the CCL14 predicted sequence available in GenBank (accession number XM_004385435.1) seems to be a pseudogene due to an early stop codon, while for the other species within this order for which the CCL14 sequences are available (Loxodonta africana and Echinops telfairi), the gene seems to encode a functional protein. This indicates that although CCL14 is present as a functional gene in the common ancestor of these species, it was later inactivated or deleted in some species, even within the same clade. This suggests that CCL14 might have functions that overlap with the function of other genes, making its presence unnecessary and leading to its pseudogenization.
Conclusions
In most of the mammals we observed that CCL14 encodes a functional gene that is evolving under positive selection, with positively selected codons located in regions important for ligand–receptor binding. In contrast, in two of the Ochotonidae family subgenera (Pika and Lagotona), CCL14 is a pseudogene due to distinct disabling mutations and the detection of an acceleration of the mutation rates. While in some mammals CCL14 retained its biological significance in the Ochotonidae family (subgenera Pika and Lagotona) it seems to have been lost.
Footnotes
Acknowledgements
We would like to thank Jeff Wilcox and Dr Michael Hamilton from Blue Oak Ranch Reserve, University of California, Berkeley, for providing brush rabbit (Sylvilagus bachmani) tissue samples. The authors are grateful to Dr Janet Rachlow, Dr Lisette Waits and Dr Caren Goldberg from Department of Fish and Wildlife Sciences, University of Idaho, USA, for providing Brachylagus idahoensis tissue samples.
Funding
This work was partially funded by FEDER (Fundo Europeu de Desenvolvimento Regional) funds through the Programa Operacional Factores de Competitividade (COMPETE program; FCOMP-01-0124-FEDER-028286) and Portuguese national funds through FCT (Fundação para a Ciência e a Tecnologia; research project PTDC/BIA-ANM/3963/2012) – Quadro de Referência Estratégico Nacional (QREN) funds from the European Social Fund and Portuguese Ministério da Educação e Ciência. FCT also supported the doctoral grants of Fabiana Neves (ref.:SFRH/BD/81916/2011) and the FCT Investigator grant of Joana Abrantes (ref.: IF/01396/2013). ‘Genomics Applied To Genetic Resources’ co-financed by North Portugal Regional Operational Programme 2007/2013 (ON.2 – O Novo Norte), under the National Strategic Reference Framework (NSRF), through the European Regional Development Fund (ERDF), also supported this work.
Conflict of interest
The authors do not have any potential conflicts of interest to declare.
