Abstract
TLRs are key innate immune receptors that recognize conserved features of biological molecules that are found in microbes. In particular, TLR2 has been reported to be activated by different kinds of microbial ligands. To advance our understanding of the interaction of TLR2 with its ligands, the recombinant human TLR2 ectodomain (hTLR2ED) was expressed using a baculovirus/insect cell expression system and its biochemical, as well as ligand binding, properties were investigated. The hTLR2ED binds synthetic bacterial and mycoplasmal lipopeptides, lipoteichoic acid from Staphylococcus aureus, and synthetic lipoarabinomannan precursors from Mycobacterium at extracellular physiological conditions, in the absence of its co-receptors TLR1 and TLR6. We also determined that lipopeptides and glycolipids cannot bind simultaneously to hTLR2ED and that the phosphatidyl inositol mannoside 2 (Pim2) is the minimal lipoarabinomannan structure for binding to hTLR2ED. Binding of hTLR2ED to Pim4, which contains a diacylglycerol group with one of its acyl chains containing 19 carbon atoms, indicates that hTLR2ED can bind ligands with acyl chains longer than 16 carbon atoms. In summary, our data indicate that diacylglycerol is the ligand moiety of microbial glycolipids and lipoproteins that bind to hTLR2ED and that both types of ligands bind to the same binding site of hTLR2ED.
Introduction
TLRs have emerged as one of the most important group of pattern recognition receptors (PRRs) in innate immunity. TLRs are type I transmembrane receptors that are important for the initiation of immune responses against microbes. Ten different human TLRs are known, some of which are located in the plasma membrane (TLR1, TLR2, TLR4, TLR5, TLR6), while others are sequestered in internal compartments, including endosomes (TLR3, TLR7, TLR8, TLR9).1,2
TLR2 has been reported to recognize different kinds of microbial ligands, which include synthetic triacyl-lipopeptides, 3 surfactant protein A, 4 meningococcal porin b, 5 mycoplasmal lipopeptides and mycobacterial lipoarabinomannan. 6 This promiscuity in the recognition of ligands has been partially explained by the association of TLR2 with other TLRs, such as TLR1 and TLR6, 7 but in other cases, the activation of TLR2 was the result of the presence of contaminants in the ligand preparations. 8 The class B scavenger receptors, CD36 and CD14, also contribute to the ligand recognition process by TLR2.9–11 The crystal structure of a chimera between the human TLR2/TLR1 ectodomain fused to leucine-rich repeats (LRRs) of a variable lymphocyte receptor (VLR) in complex with the triacyl-lipopeptide Pam3CSK4 revealed that a hydrophobic pocket composed of the central LRRs 9–12 of TLR2 is the binding site for Pam3CSK4. 12 The crystal structures of a mouse TLR2 ectodomain (mTLR2ED)/VLR hybrid in complex with TLR6 and Pam2CSK4, as well as mTLR2ED/VLR in complex with PE-DTPA, a synthetic derivative of phosphatidylethanolamine, and lipoteichoic acid (LTA) from Streptococcus pneumonia also uncovered similar findings. 13 While these are extremely important steps in determining the recognition of ligands by TLR2, our knowledge of TLR2 interactions with its different ligands is far from complete. 14
In order to investigate the binding of human TLR2 with the synthetic diacyl-lipopeptide fibroblast stimulating lipopeptide-1 (FSL-1) and LTA from Staphylococcus aureus and lipoarabinomanan precursor, hTLR2ED was expressed employing a baculovirus/insect cell expression system. The biochemical properties of hTLR2ED and the interaction with its ligands were investigated.
Materials and methods
Cell culture and cell line
Sf9 (Invitrogen, Carlsbad, CA, USA) and High Five (Invitrogen) insect cells were employed for baculovirus generation and protein expression, respectively. Both cell lines were cultured in suspension cultures in serum-free HyQ media (Thermo Scientific Hyclone, Logan, Utah, USA) at 25℃ under shaking conditions.
Generation of hTLR2ED profold ER1 baculovirus
hTLR2ED (residues 19–590, National Center for Biotechnology Information accession number NP_003255) with a C-terminal His6-tag was amplified by PCR using pCDNA3.1 CMV TLR2 flag plasmid as template. The PCR product was cloned in TopoXL PCR (Invitrogen), and subcloned into the BamHI and NotI sites of the pAcGP67A transfer vector (BD Biosciences Pharmingen, San Diego, CA, USA). A baculovirus, which we call hTLR2ED Profold ER1, was generated by homologous recombination of hTLR2ED pAcGP67A with Profold ER1 linearized baculovirus vector (AB Vector, San Diego, CA, USA). After five rounds of viral amplification, a titer of 1 × 108 viruses/ml was achieved. Fluorescence from GFP encoded in the Profold ER1 linearized baculovirus vector was monitored using an Axioplan Zeiss fluorescent microscope (Zeiss AG, Oberkochen, Germany).
Purification of recombinant hTLR2ED protein
Six-liter cultures of 1 × 106 High Five cells/ml were infected with hTLR2ED Profold ER1 at a multiplicity of infection of 1. The infected cells were incubated at 27℃ for 5 d under shaking conditions. The supernatant collected after centrifugation was buffer exchanged into PBS and concentrated to 100 ml with a Pall Filtron Centramate concentrator device (EMD, Billerica, MA, USA) using membranes with a 30-ku molecular mass cut-off (GE Healthcare, Uppsala, Sweden). Next, 5 ml Ni (II) nitrilotriacetic acid (Ni-NTA) beads and 20 mM imidazole were added and incubated overnight (12 h) at 4℃ with agitation. Subsequently, Ni-NTA beads were separated from the supernatant by filtration and washed three times with 20 mM imidazole and 300 mM NaCl (pH 8). Bound proteins were eluted from the Ni-NTA beads with 100 mM imidazole and 300 mM NaCl. Anionic exchange chromatography using a Mono Q column (GE Healthcare) followed by size exclusion chromatography using a Superdex 200 10/30 column (GE Healthcare) were employed to purify hTLR2ED to homogeneity.
Identification of recombinant hTLR2ED by liquid chromatography-mass spectrometry and Western blot analysis
hTLR2ED was concentrated to 5 mg/ml and its purity was analyzed by SDS-PAGE. The band between 50 and 75 ku was excised, reduced with dithiothreitol (10 mM), and digested with trypsin overnight before being analyzed by nano LC-MS/MS (The Scripps Research Institute Center for Mass Spectrometry, La Jolla, CA, USA). Peptides were identified using MASCOT. Protein expression of hTLR2ED was also confirmed by a Western blot probing for the presence of the C-terminal His-tag of the hTLR2ED construct. After proteins were electroblotted from SDS-PAGE to a polyvinylidene fluoride (PVDF) membrane, the PVDF membrane was blocked for 1 h at room temperature with 3% BSA, 0.5% Tween 20. Subsequently, the membrane was incubated overnight with a 1/1000 dilution of a mouse IgG anti-penta His-tag Ab (Qiagen, Venlo, the Netherlands), washed three times with 0.5% Tween 20 in PBS, and incubated for 1 h at room temperature with a 1/5000 dilution of rabbit peroxidase-conjugated anti-mouse IgG Ab (Thermo Scientific Pierce, Rockford, IL, USA). After three washes with 0.5% Tween 20 in PBS, the Western blot was developed using the Super Signal West Pico ECL substrate (Thermo Scientific Pierce).
Presence of disulfide bonds in hTLR2ED
Purified hTLR2ED was incubated for 10 min in a reducing buffer containing β-mercaptoethanol or in a non-reducing protein gel loading buffer. The samples were then run on SDS-PAGE to determine any difference in electrophoretic mobility that would indicate the presence of disulfide bonds in hTLR2ED. The presence of disulfide bonds was also monitored in hTLR2ED deglycosylated with endoglycosidases Endo H and PNGase F. Ten μg of purified hTLR2ED were digested with 50 units of PNGase F (New England Biolab, Ipswich, MA, USA) or Endo H (New England Biolab) in native conditions overnight at 37℃. The samples were examined by SDS-PAGE under reducing and non-reducing conditions.
Analysis of hTLR2ED glycosylation
The hTLR2ED band was excised from an SDS PAGE gel, cut into 1-mm cubes, and frozen overnight. The gel pieces were washed with acetonitrile to remove any reagents and then dried in a vacuum centrifuge. The dried gel pieces were rehydrated and incubated with PNGase F (Boehringer Mannheim, Mannheim, Germany) for 16 h at 37℃. The supernatant was collected. Distilled water was added to the gels, which were sonicated for 30 min, and the supernatant retained. This procedure was repeated twice with acetonitrile, followed by sub-boiling point distilled (sbpd) water, finishing with acetonitrile to maximize the recovery of glycans. Samples were then treated with an AG 50 × 12 (Biorad, Hemel Hempstead, UK) anion-exchange resin to eliminate sodium. The solution was filtered through a filter syringe to eliminate the resin and then labeled with 2-aminobenzamide using the Ludger Tag 2-AB labeling kit (Ludger, Oxford, UK). Excess label was removed by ascending chromatography on 3-mm Whatman paper with acetonitrile as the solvent, and the glycans were eluted with sbpd water. Glycans were examined by normal-phase HPLC (NP-HPLC), as previously reported, 15 and by MALDI-TOF MS. The HPLC system consisted of a 2690 Alliance separation module (Waters, Milford, MA, USA) with a 4.6 × 250 mm TSK Amide-80 column (Anachem, Luton, UK) equipped with a fluorescence detector set at 420 nm and was calibrated against a dextran standard. The retention time of each glycan was converted into Glc units (GU) by comparison with the elution times of a dextran ladder. The experimental GU values were then referred to Glycobase, a relational database (http://glycobase.ucd.ie/cgi-bin/public/glycobase.cgi) that stores information for glycan structure according to its GU values. Preliminary structure assignments were made for each peak. Exoglycosidase array digestions further confirmed these assignments. The 2-AB labeled glycans were digested with α-glucosidase II (EC 3.2.1.20) and jack bean α-mannosidase (EC 3.2.1.24) at 37℃ for 18 h. Digested samples were passed through a protein-binding filter (Micropure-EZ centrifugal devices; EMD Millipore) to remove the enzymes from the glycans, which were washed thoroughly with sbpd water. Finally, the glycans were separated by NP-HPLC to determine the changes in GU values as a result of the enzyme digestions.
MALDI-TOF MS
Samples were cleaned with a Nafion membrane and MALDI-TOF mass spectra were recorded with a Waters TofSpec 2 E mass spectrometer (Waters MS-Technologies, Manchester, UK) in positive ion reflectron mode with delayed extraction. The acceleration voltage was 20.0 kV and the pulse voltage was 3.0 kV. Samples in 0.3 µl of water were mixed with 0.3 µl of matrix (a saturated solution of 2,5-dihydroxybenzoic acid in acetonitrile) on the MALDI target and allowed to dry under ambient conditions. The dry sample was then recrystallized from ethanol.
Electrospray ionization MS
Negative ion electrospray ionization (ESI) MS/MS spectra were recorded with a Waters Q-TOF Ultima Global mass spectrometer. Samples (about 50 pmoles/µl), cleaned with a Nafion membrane as above, in 1:1 (v:v) methanol:water were infused with a Proxeon (ProxeonBiosystems, Odense, Denmark) borosilicate capillary. The ion source was maintained at 120℃, the infusion needle potential was 1.1 kV, the cone was at 100 V and the RF-1 voltage was 180 V. Spectra (2-s scans) were acquired with a digitization rate of 4 GHz. Collision-induced fragmentation data (CID) acquisition used a 4-u mass window for parent ion selection and argon at approx 0.5 mBar as the collision gas. The collision cell voltage was 80–120 V, depending on the parent ion mass, and other voltages were as recommended by the manufacturer. The instrument was externally calibrated with N-glycans released from bovine fetuin, and monoisotopic masses are cited to one decimal place. Instrument control, data acquisition and processing were performed with a MassLynx data system (version 4.0).
Circular dichroism analysis of purified hTLR2ED
Circular dichroism (CD) measurements of hTLR2ED were performed in 50 mM NaCl, 10 mM Tris-HCl buffer at a concentration of 3.5 µM on an AVIV 202 spectropolarimeter equipped with a thermostat (Hellma, Mullheim, Baden, Germany). The purified hTLR2ED solution was placed in a 1-mm path length quartz cuvette and heated from 25 to 95℃ at a rate of 1℃/min. Spectra were recorded every 5℃ after equilibration of 5 min at the corresponding temperature. The scans were performed in duplicate from 260 to 200 nm with 0.5-nm resolution. The spectra from duplicate scans were averaged and baseline corrected by subtracting the corresponding buffer spectrum obtained under identical conditions. Results were expressed as a mean residue ellipticity (Θ) at a given wavelength.
Native PAGE experiments
The following synthetic lipopeptides were employed for native PAGE experiments: Cys-Ser-Lys4 (CSK4), N-palmitoyl Cys-Ser-Lys4 (PamCSK4), S-(2,3-bis-palmitoyloxypropyl)-Cys-Ser-Lys4 (Pam2CSK4), Pam2CSK4 fluorescein, N-palmitoyl-S-(2,3-bis-palmitoyloxypropyl) -Cys-Ser-Lys4 (Pam3CSK4), and S-(2,3-bis-palmitoyloxypropyl)-Cys-Gly-Asp-Pro-Lys-His-Pro-Lys-Ser-Phe-K-Aca-Aca-Fluorescein (FSL-1–fluorescein) activator. All of these synthetic lipopeptides were obtained from EMC Microcollection (Tübingen, Germany). LTA and the human TLR9 ligand CpG-ODN2006 was obtained from Invivogen (San Diego, CA, USA). Synthetic phosphatidylinositol mannosides (Pim2 and Pim4) were synthesized as described previously.16,17 The gp120 from HIV-1 clade B was a gift from Dr. Robert Pejchal of our laboratory. One microgram of each of the mentioned compounds was separately incubated with 10 µg of hTLR2ED overnight at 37℃ (pH 7.4). In case of the simultaneous incubation with LTA and Pam2CSK4-fluorescein, 1 µg of each compound was mixed and incubated with 10 µg of hTLR2ED. For the lipid ligands, an excess of ligands was added with respect to hTLR2ED [ratio ligand/hTLR2ED: 11.3 (CSK4), 8 (PamCSK4), 5.41 (Pam2CSK4), 4.55 (Pam3CSK4), 6.53 (FSL-1), 3.7 (Pim2), 4.55 (Pim4), 2.9 (LTA)] because the lipid alone forms aggregates due to hydrophobic interactions and thus it is difficult to calculate an accurate ratio of monomeric ligand to hTLR2ED. The molar ratio of hTLR2ED to gp120 was 1.7:1. All samples were run for 3 h at 100 volts in a 4–20% native PAGE. Fluorescence was determined with a Versadoc imaging system equipped with an emission filter with a 60 nm bandwidth (from 500 to 560 nm). Subsequently, the gel was stained with Coomassie Blue and de-stained with a solution containing 20% methanol, 10% acetic acid and 1% glycerol. Gel images were taken with a Versadoc gel imaging system.
Modeling of hTLR2ED–FSL-1, hTLR2ED–LTA, hTLR2ED–LTA C19 hTLR2ED–Pim2 and hTLR2ED–Pim4 complexes
Molecular modeling was performed on a Silicon Graphics Indigo 2 workstation using Insight II and Discover software (Accelrys, San Diego, CA, USA). Figures were produced using Molscript.
18
The models of hTLR2ED in complex with various ligands were based on the crystal structures of the hTLR2ED/VLR hybrid in complex with Pam3CSK4 [Protein Data Bank (PDB) 2Z7X]
12
and the Nogo receptor (PDB 1OZN).
19
Residues 27–508 were taken from 2Z7X, while residues 509–585 are based on residues 236–309 of 1OZN. The model was created by overlaying the backbone atoms of residues 500–509 of 2Z7X with residues 227–236 of 1OZN. Five disulfide bonds were added to the model, one in the N-terminal cap, two in the LRR domain (LRR12-13 and LRR15-16) as present in 2Z7X, and two C-terminal disulfide bonds, which are conserved in Nogo receptor and hTLR2ED (Figure 1). High-mannose (Man) glycans were attached to each of the four glycosylation sites (Asn-114, Asn-199, Asn-414, and Asn-442) (Figure 1). All four potential glycosylation sites were solvent accessible. N-Glycan structures were generated using the database of glycosidic linkage conformations
20
and in vacuo energy minimization to relieve unfavorable steric interactions. The Asn–GlcNAc linkage conformations were based on the observed range of crystallographic values,
20
the torsion angles around the Asn Cα-Cβ and Cβ-Cγ bonds then being adjusted to eliminate unfavorable steric interactions between the glycans and the protein surface. The complexes involving ligands with C16 acyl chains were modeled simply by replacing the ligand head group of the crystallographic Pam3CSK4 as appropriate. All of the modeled head groups could be accommodated with no need to adjust any of the rest of the structure. The complexes of hTLR2ED with a C19 ligand (LTA C19) and Pim4 were modeled by keeping the head group position constant and allowing the protein structure to alter to accommodate the increased acyl chain length. This process was carried out by increasing the chain length one carbon at a time and using molecular dynamic simulations to allow relaxation of the local protein structure, while keeping the ligand head group and the regions of the protein that do not interact with the acyl chains fixed. This procedure resulted in three of the outer-face loops changing their conformation slightly.
Protein sequence of hTLR2ED employed for modeling with its ligands. hTLR2ED has 20 predicted LRR repeats that are capped at its N-terminus (LRR-NT) and C-terminus (LRR-CT). The individual LRRs of hTLR2ED were aligned based on the hTLR2ED/VLR crystal structure solved by Jin et al.
12
The canonical N-terminal, central and C-terminal LRRs are indicated in red, blue and green, respectively. Cysteines that form disulfide bonds in the crystal structure solved by Jin et al.
12
and cysteines from the C-terminal cap that are predicted to form disulfide bonds are color-matched. Asparagines that are the potential glycosylation sites are highlighted in red, and the N-linked glycosylation consensus sequences are highlighted in yellow. The LRR consensus sequence is depicted, and the position of the β-strand is indicated by an arrow
Results
hTLR2ED is expressed as a monomer in insect cells with internal disulfide bonds and high-Man sugars
The hTLR2ED, consisting of an N-terminal cap, an LRR domain, a C-terminal cap and a C-terminal His-tag, was successfully expressed using a baculovirus/insect cell expression system. Baculovirus for expression of hTLR2ED was generated by homologous recombination of a plasmid encoding hTLR2ED and linearized baculovirus DNA. Baculovirus amplification was monitored by fluorescent microscopy to detect expression of GFP encoded by the baculovirus DNA. After three steps of purification, highly purified hTLR2ED was obtained (Figure 2A). Moreover, hTLR2ED ran as an apparent monomer on size exclusion chromatography by comparison with molecular mass (MM) standards (Figure 2B). Digestion of hTLR2ED with PNGase F under native conditions indicated the presence of N-linked glycosylation (Figure 2C). The decreased electrophoretic mobility of native and deglycosylated hTLR2ED under reducing conditions in the presence of β-mercaptoethanol was indicative of the presence of disulfide bonds (Figures 2D,E). This finding is consistent with the reported crystal structure of hTLR2ED, which reveals disulfide bonds in the N-terminal cap and in the LRR domain and C-terminal cap.
12
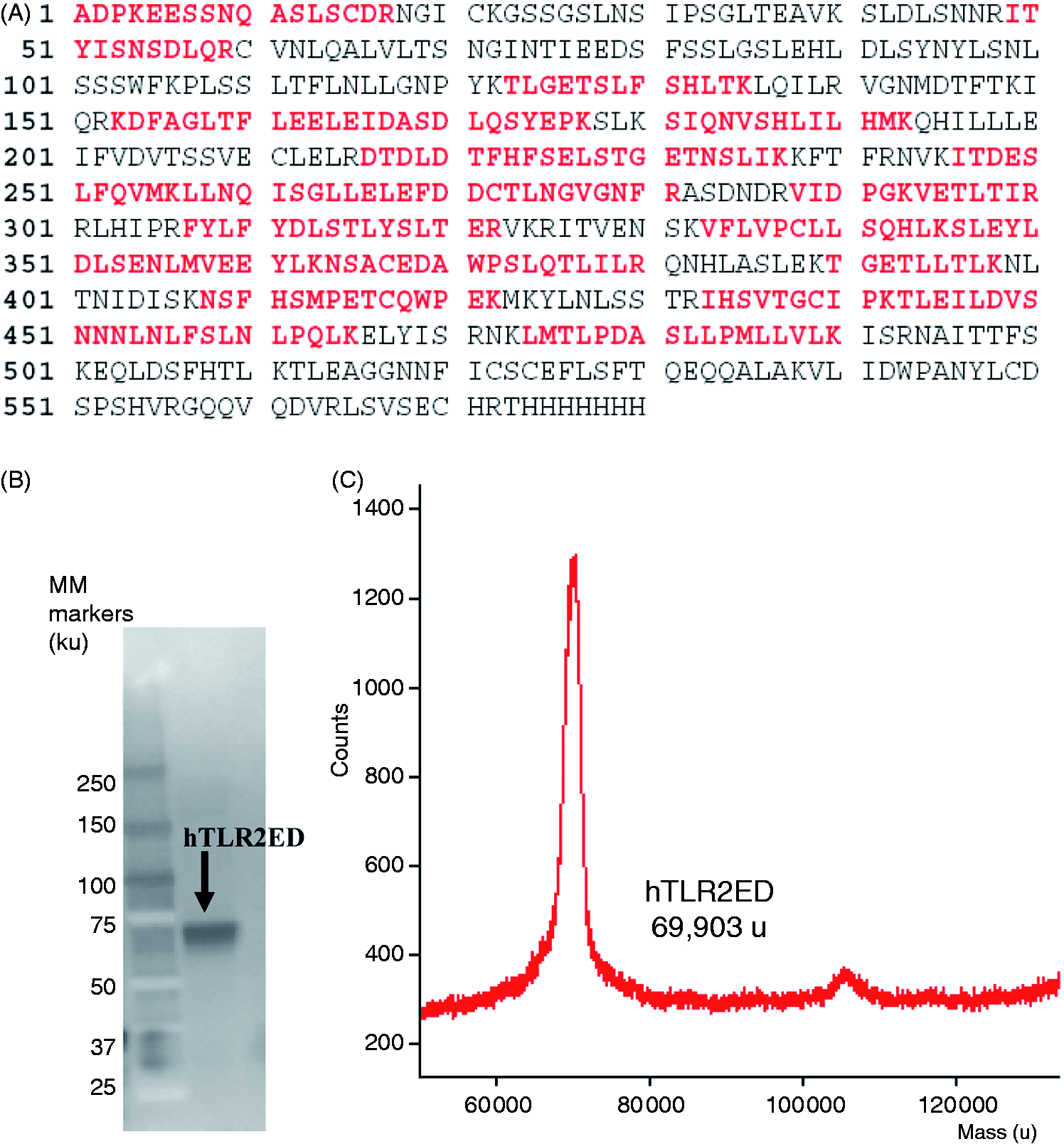
The identity of hTLR2ED was confirmed by anti-His-tag Western blot and by LC-MS peptide fingerprinting (Figure 3A,B). The theoretical MM of the hTLR2ED protein is 65,734 u (ProtParam, http://web.expasy.org/protparam/), while the MM of the hTLR2ED determined by MALDI TOF MS is 69,903 u (Figure 3C). Digestion of hTLR2ED with PNGase F under native conditions indicated that this difference in mass is due to the presence of N-linked glycosylation (Figure 2C). The hTLR2ED has four N-linked glycosylation sites, which are necessary for its secretion,
21
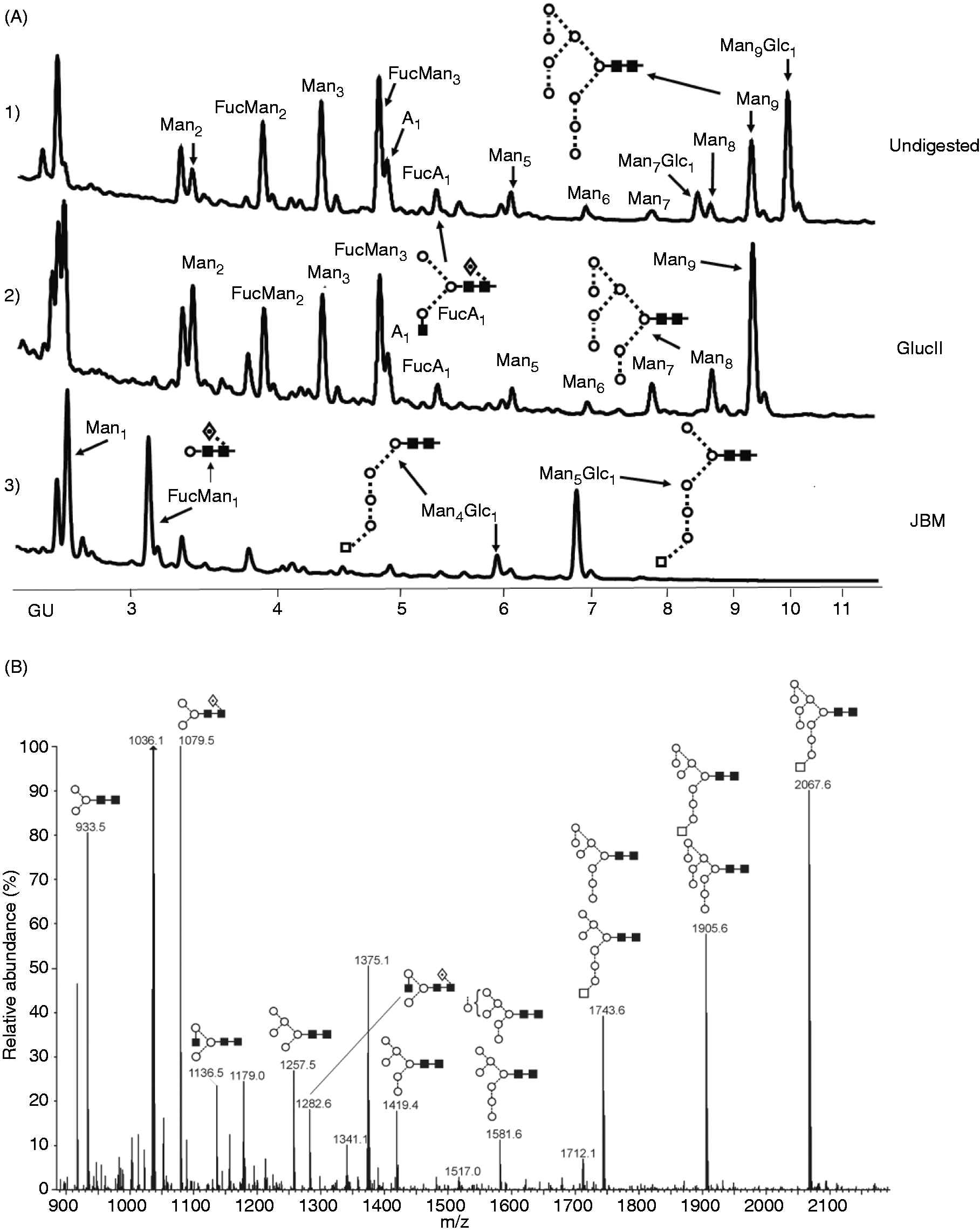
highlighting the importance of N-linked glycosylation in correct folding and stability.22–24 Furthermore, we determined the type of glycosylation by releasing the N-linked glycans from recombinant hTLR2ED using in-gel digestion with PNGase F. We labeled the glycans with a fluorescent tag, digested them with glucosidase II and α-jack bean mannosidase, and analyzed their composition by HPLC with fluorescence detection (Figure 4A). The N-linked glycans were also identified by MALDI-TOF (Figure 4B) and negative ion CID MS. These data indicate that hTLR2ED glycosylation is predominantly of the high-Man or pauci-Man type, which is consistent with the absence of secondary glycan processing in insect cells.25,26 This result was further confirmed by glucosidase II and α-jack bean mannosidase digestions. Thus, the main glycan structures found in hTLR2ED were Fuc1Man3GlcNAc2, Man3GlcNAc2, Man9GlcNAc2 and Man9Glc1GlcNAc2 (Table 1).
hTLR2ED expression and purification from insect cell culture. (A) Culture supernatants of insect cells infected with hTLR2ED Profold ER1 were concentrated and purified to homogeneity by Ni-NTA affinity, anion exchange and gel filtration chromatography. Samples were analyzed on 4–20% SDS-PAGE and stained with Coomassie blue. (B) hTLR2ED is a monomer in solution. Purified hTLR2ED and MM standards were run on a Superdex 200 10/30 gel filtration column. Purified hTLR2 ED elutes at 14.2 ml, which corresponds to the elution volume of a protein of 70 ku, which is approximately the molecular mass of the hTLR2ED monomer (note that the MM of the hTLR2ED monomer as determined by MALDI TOF mass spectrometry is 69,903 u). (C) hTLR2ED has N-linked glycosylation. hTLR2 undigested (1) and digested overnight with PNGase F in native conditions (2). (D,E) hTLR2ED contains intramolecular disulfide bonds. (D) Purified hTLR2ED was run under reducing (1 R) and non-reducing (2 NR) conditions in an SDS-PAGE gel. The difference in electrophoretic mobility indicates the presence of disulfide bridges. (E) hTLR2ED was digested overnight at 37℃ with PNGase F and Endo H, and then run in SDS-PAGE under reducing (1 R) and non-reducing (2 NR) conditions (E). MM markers are shown on each gel in ku. hTLR2ED expression using a baculovirus/insect cell expression system. (A) The identity of the recombinant hTLR2ED was confirmed by LC-MS/MS. Nineteen peptides, which cover 49% of hTLR2ED protein sequence, were identified using MASCOT and are depicted in red. (B) The identity of the hTLR2ED was also confirmed by Western blot against its C-terminal His tag. (C) The MM was determined by mass spectrometry (The Scripps Research Institute mass spectrometry core). hTLR2ED expressed in insect cells is glycosylated with high-Man N-linked sugars. (A) HPLC chromatogram showing the profile of hTLR2ED glycans. The glycans were labeled with 2-AB and digested for 18 h at 37℃ with α-glucosidase II (Gluc II) and jack bean α-mannosidase (JBM), and run on a normal phase HPLC. An alternative nomenclature for naming the glycan structures was employed, which does not mention the two core GlcNAc2 that are present in all the glycan structures (e.g. Man2 corresponds to Man2GlcNAc2). (B) MALDI-TOF mass spectrum of N-linked glycans released from hTLR2ED by PNGaseF treatment. The spectrum has been processed by the MaxEnt 2 algorithm from MassLynx (Waters). Ions at 1036, 1179, 1341, 1375 and 1712 u are contaminants. Man (open circle), Glc (open square), GlcNAc (filled square). Fuc: core fucose (diamond with central dot); A1: mono-antennary. Percentages and GU of the N-linked glycans of hTLR2ED expressed in insect cells. The most abundant glycan structures are Man9Glc1GlcNAc2, Man9GlcNAc2, fucosylated-Man3GlcNAc2 and Man3GlcNAc2.The alternative nomenclature of glycan structures, depicted in the HPLC chromatogram, is shown in parentheses. Man: open circle; Glc (open square); Gal (diamond); Fuc (diamond with central dot); GlcNAc (filled square); A1: mono-antennary. (1) Listed in order of GU value. (2) Not detected by mass spectrometry. (3) Not detected by ESI MS; structure not confirmed by fragmentation. (4) Not detected by HPLC. (5) Trace amount by mass spectrometry; structure not confirmed by fragmentation. (6) As measured by HPLC.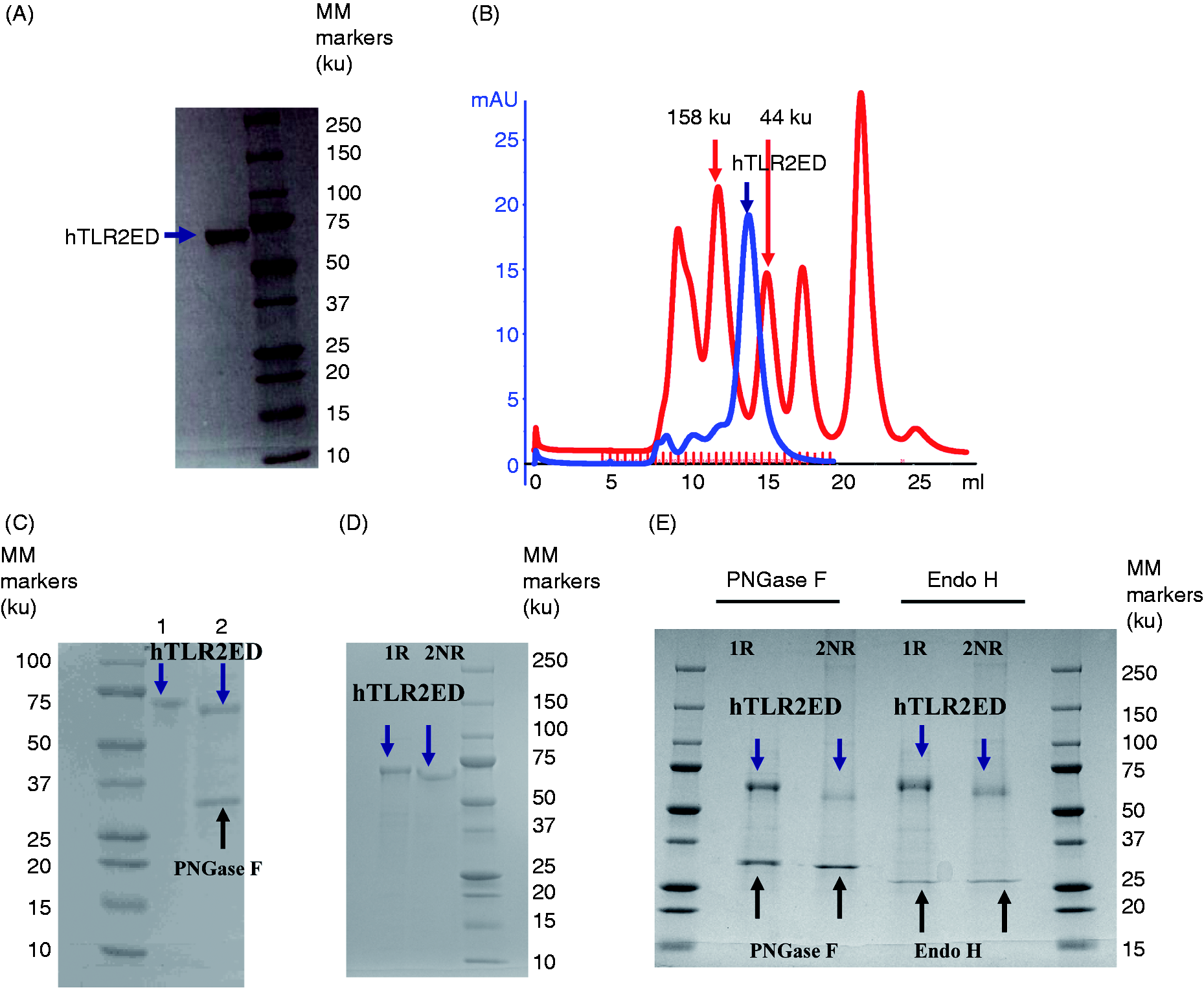

Thermal denaturation of hTLR2ED is an irreversible two-stage process
The CD spectrum of hTLR2ED at 25℃ revealed a minimum between 215 and 220 nm (Figure 5A), which is indicative of a protein fold dominated by β-sheet secondary structure.
27
Furthermore, thermal denaturation of hTLR2ED, as monitored by CD spectroscopy, uncovered a two-stage process (Figures 5A,B). The first stage occurred between 45℃ and 55℃, and another transition occurred between 90° and 95℃ (Figure 5A,B). After denaturation at 95℃, hTLR2ED was slowly cooled back down to 25℃ and another CD spectrum was recorded. This spectrum showed no difference to the spectrum recorded at 95℃, demonstrating that thermal denaturation of hTLR2ED appears to be an irreversible process (Figure 5C).
CD analysis of hTLR2ED. (A) CD spectra of hTLR2ED recorded at various temperatures. The CD spectrum of hTLR2ED at 25℃ reveals a minimum around 215 nm, which is indicative of a folded protein with a significant portion of β-sheet secondary structure. At temperatures greater than 50℃, the β-sheet character of the hTLR2ED CD spectra is lost and the shape of the spectra indicate that the hTLR2ED is now mostly unfolded. (B) Thermal denaturation of hTLR2ED is a two-stage process. The CD signal at 208 nm was plotted against the corresponding temperatures. hTLR2ED undergoes two unfolding transitions: at the first Tm of 45-55℃ and a second Tm of 90-95℃ (the protein becomes mostly unfolded). (C) The thermal denaturation of hTLR2ED is irreversible. After the CD spectra of hTLR2ED were recorded at 95℃, hTLR2ED was slowly cooled down to 25℃ and new CD spectra were recorded (TLR2 post-dT). There were no differences between the spectra at 95℃ and after re-cooling, indicating that no secondary structure is regained and that the thermal denaturation of hTLR2ED is irreversible.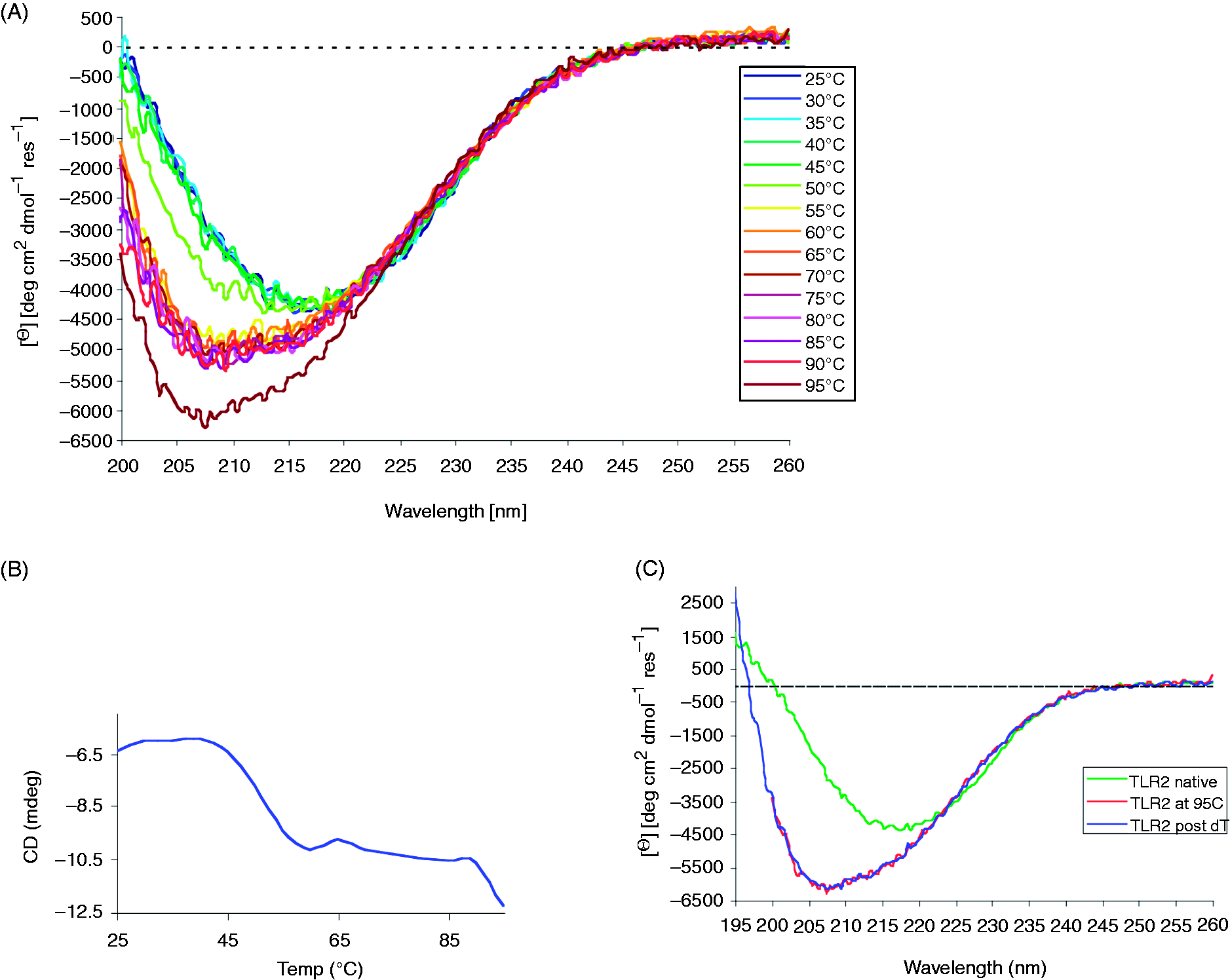
hTLR2ED does not bind gp120 from HIV-1
The hTLR2ED was employed for binding to gp120 from HIV-1, different lipopeptides and glycolipids (Figure 6). Because levels of soluble TLR2 have been shown to correlate with HIV progression and increase after anti-retroviral treatment,
28
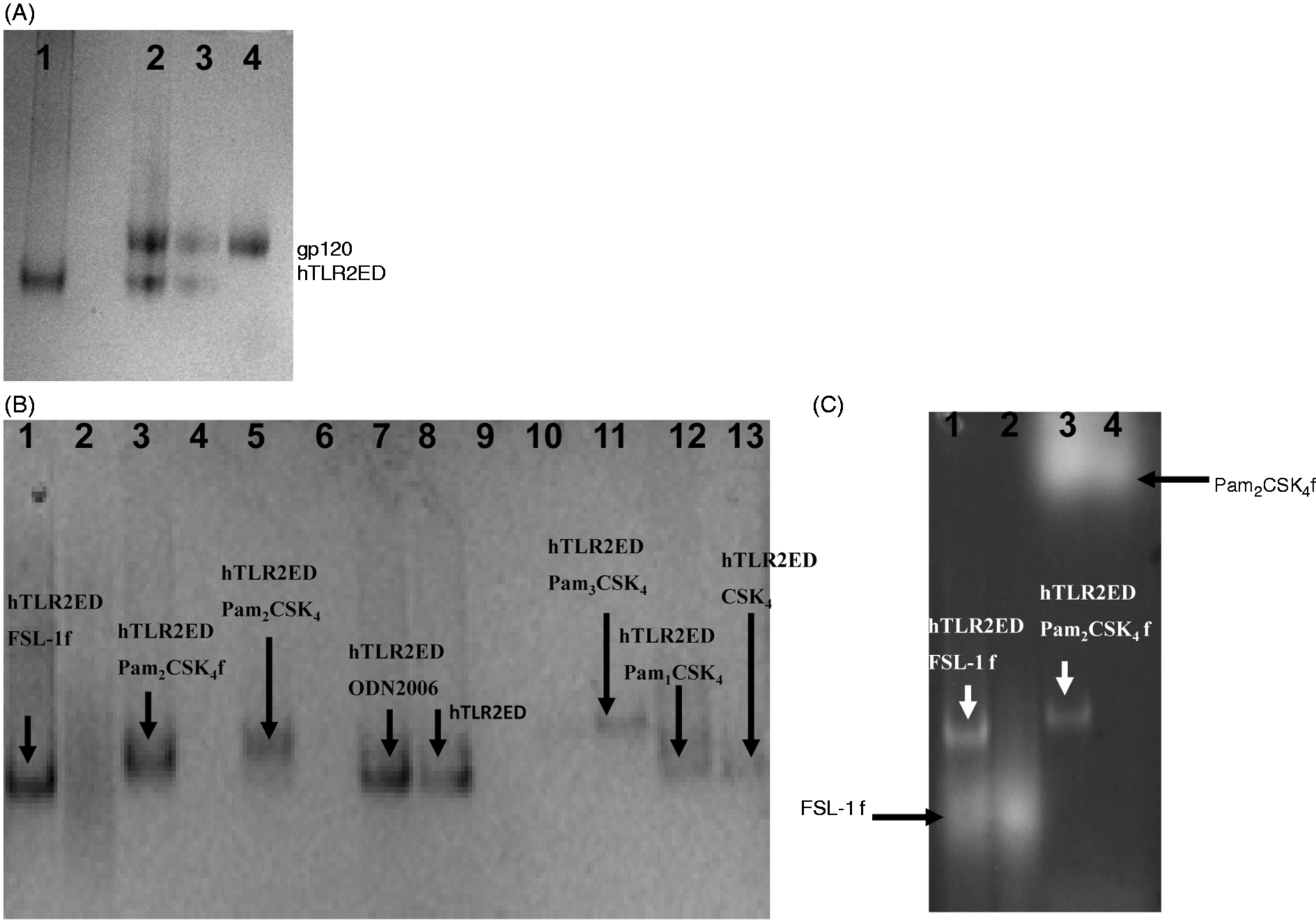
we tested whether hTLR2ED is able to recognize the surface viral glycoprotein gp120 from HIV-1. We determined that hTLR2ED does not bind gp120 as no complex was observed in native PAGE (Figure 7A).
The chemical structures of lipopeptides and glycolipids employed for binding studies with hTLR2ED. The diacylglycerol moieties that are involved in the interaction with hTLR2ED are highlighted by boxes. hTLR2ED binds diacyl and triacyllipopeptides, but not HIV-1 gp120. Samples were incubated overnight at 37℃ and run for 4 h at 100 volts on a native 4–20% native PAGE. Native gels stained with Coomassie blue. (A) hTLR2ED incubated with a clade B HIV-1gp120. Lane 1, 10 µg of hTLR2ED; lane 2, 10 µg hTLR2ED with 10 µg gp120; lane 3, 1 µg hTLR2ED with 1 µg gp120; lane 4, 10 µg gp120. (B,C) hTLR2ED incubated with lipopeptides. Native gel stained with Coomassie blue (B) and corresponding fluorescence determined using a Versadoc imaging system (C). Lane 1, 10 µg hTLR2ED with 1 µg FSL-1 fluorescein (FSL-1f); lane 2, 1 µg FSL-1f; lane 3, 10 µg hTLR2ED with 1 µg Pam2CSK4 fluorescein (Pam2CSK4f); lane 4, 1 µg Pam2CSK4f; lane 5, 10 µg hTLR2ED with 1 µg Pam2CSK4: lane 6, 1 µg of Pam3CSK4); lane 7, 10 µg hTLR2ED; lane 8, 10 µg hTLR2ED with 1 µg ODN2006 (human TLR9 activator; lane 9, 1 µg ODN2006; lane 10, 1 µg Pam2CSK4; lane 11, 10 µg of hTLR2ED with 1 µg Pam3CSK4; lane 12, 10 µg hTLR2ED with 1 µg Pam1CSK4; lane 13, 10 µg hTLR2ED with 1 µg CSK4. Fluorescence of Pam2CSK4f-hTLR2ED and FSL1f-hTLR2ED is indicated by arrows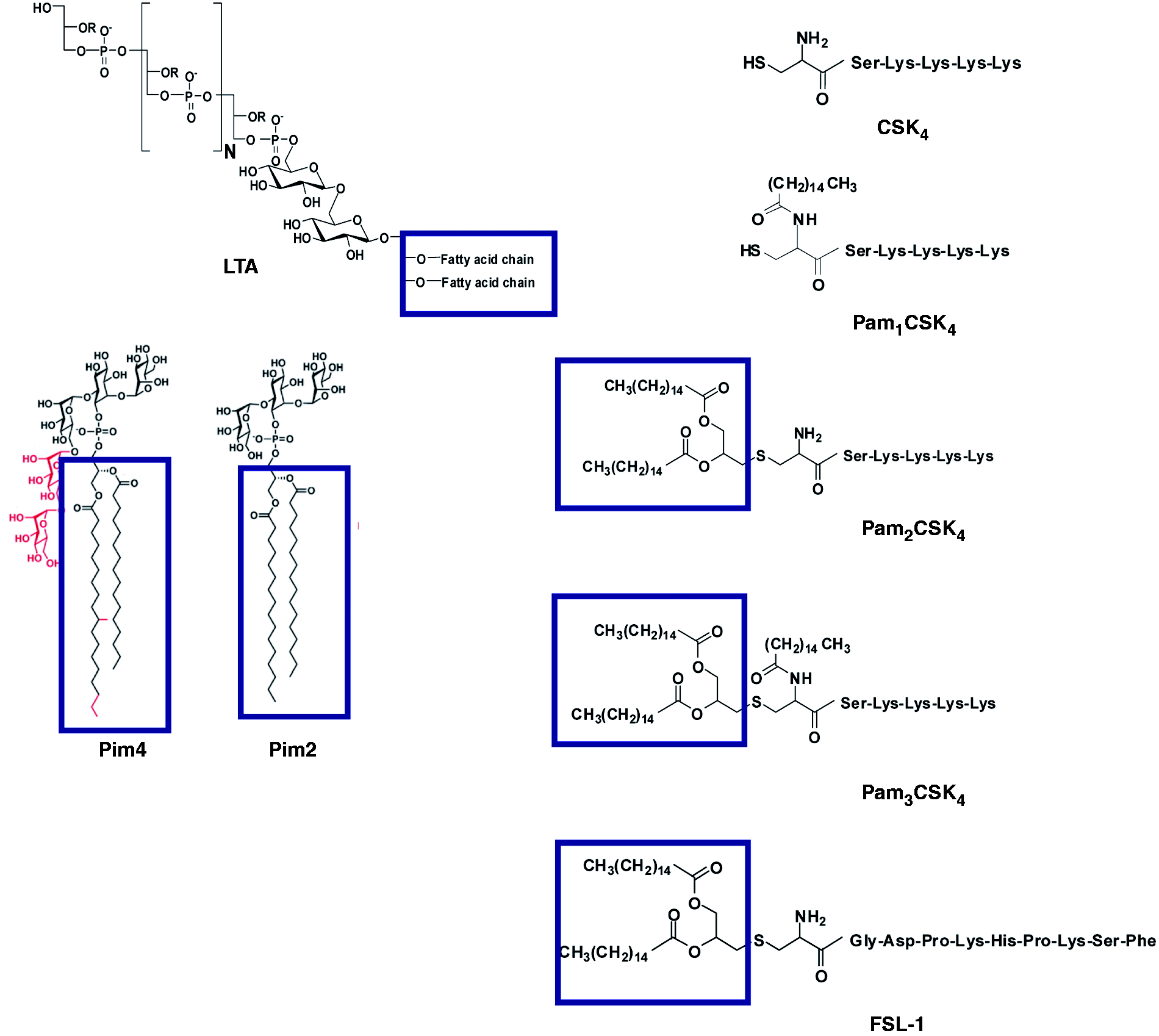

hTLR2ED binds synthetic diacylglycerol lipopeptides, but not monoacylated lipopeptides
Native PAGE experiments showed that hTLR2ED binds synthetic lipopeptides (Figure 7B,C). A gel shift in the presence of Pam2CSK4 and Pam2CSK4-fluorescein (Pam2CSK4f), but not with CSK4 and Pam1CSK4, indicated that hTLR2ED binds the two fatty acid chains linked to cysteinyl S-glycerol (Figure 7B,C) in accord with the crystal structure of hTLR2ED-Pam3CSK4. 12 Likewise, the gel shift of hTLR2ED incubated with fibroblast stimulating factor-1 fluorescein (FSL-1f) also reflects recognition of the palmitic acid chains connected to the cysteinyl S-glycerol of FSL-1 (Figure 7B). Binding of FSL-1f to hTLR2ED was confirmed by co-localization of hTLR2ED with the fluorescence, as with Pam2CSK4f (Figure 6C). As expected, hTLR2ED binds the synthetic lipopeptide Pam3CSK4, but not the human TLR9 ligand CpG-ODN200629–31 (Figure 7B).
hTLR2ED binds LTA from S. aureus, but cannot bind Pam2CSK4 and LTA simultaneously
A considerable increase in the electrophoretic mobility was observed for the hTLR2ED–LTA complex (Figure 8A), consistent with the presence of the polyglycerophosphate moiety in LTA. Interestingly, native PAGE of hTLR2ED co-incubated with both LTA from S. aureus and synthetic Pam2CSK4f showed that hTLR2ED preferentially binds to LTA rather than to Pam2CSK4f (Figure 8A,B).
hTLR2ED binds LTA from S. aureus and mycobacterial Pims. (A,B) hTLR2ED incubated with LTA and/or Pam2CSK4f. (A) Native gel stained with Coomassie blue and (B) fluorescence determined using a Versadoc imaging system. Lane 1,10 µg hTLR2ED with 1 µg LTA; lane 2, 1 µg hTLR2ED with 1 µg of LTA; lane 3,10 µg hTLR2ED; lane 4, 10 µg hTLR2ED with 1 µg Pam2CSK4 fluorescein (Pam2CSK4f); lane 5, 10 µg hTLR2ED with 1 µg Pam2CSK4f and 1 µg LTA; lane 6, 1 µg LTA; lane 7, 1 µg Pam2CSK4f plus 1 µg LTA; lane 8, 1 µg Pam2CSK4f. (C) hTLR2ED incubated with Pims. Native gel stained with Coomassie blue. Lane 1, 10 µg hTLR2ED with 1 µg Pim4; lane 2, 10 µg hTLR2ED; lane 3, 10 µg hTLR2ED with 1 µg of Pim2; lane 4, 1 µg Pim2: lane 5, 1 µg Pim4. The chemical structures of Pim4 and Pim2 are shown in Figure 6, with the two extra Man and extra three carbon atoms of Pim4 highlighted in red.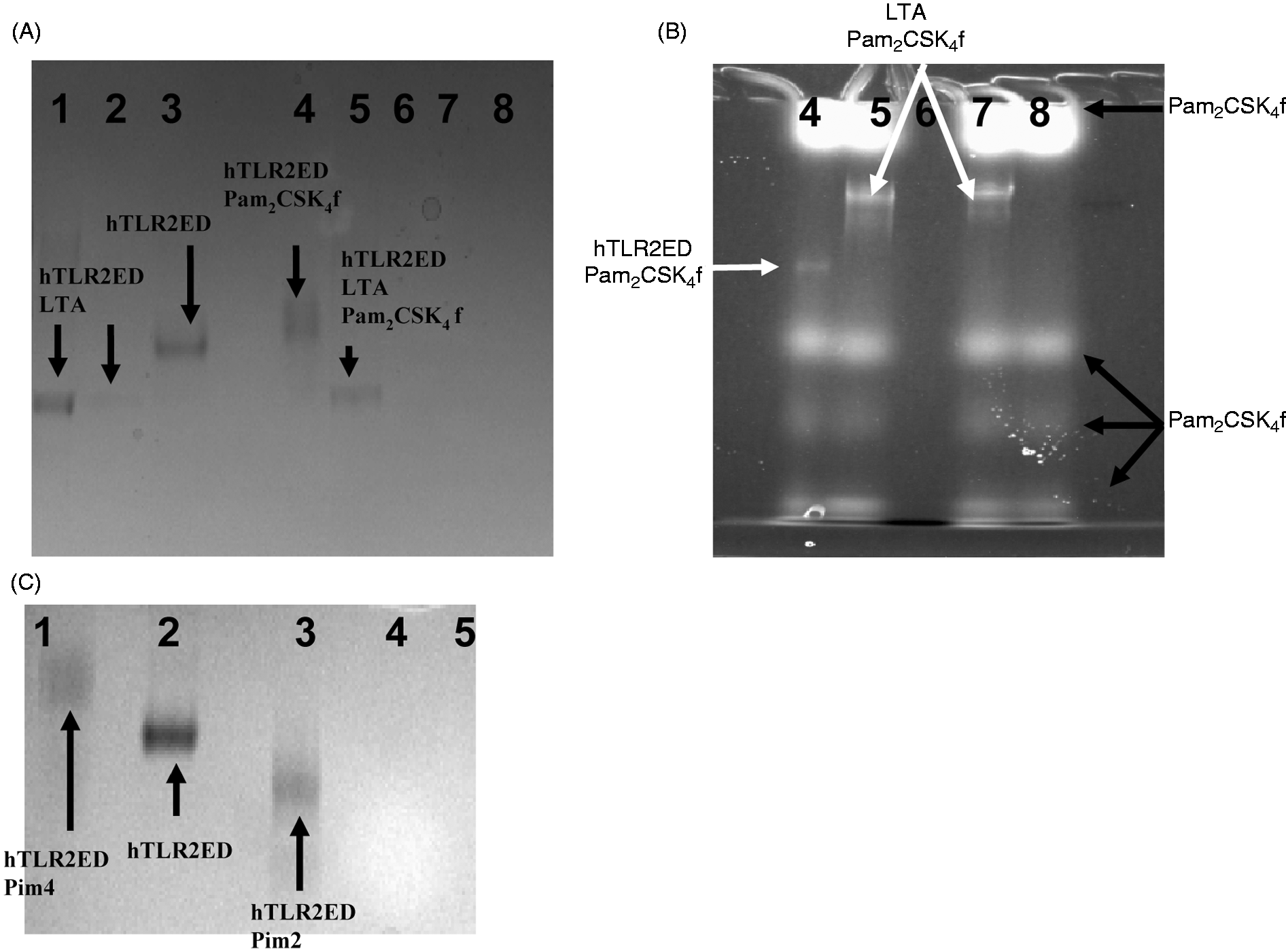
hTLR2ED binds synthetic phosphatidylinositol mannosides (Pim2 and Pim4)
We also determined whether hTLR2ED binds other important glycolipids, such as mycobacterial Pims, which were previously shown to be ligands for the TLR2/TLR1 heterocomplex.
32
For these binding assays, we employed synthetic Pims, which we had previously shown to bind CD36, but not induce TNF-α secretion by macrophages.
10
We also found that hTLR2ED binds to lipoarabinomannan precursors (Pim2 and Pim4) (Figure 8C). The negative charge of Pim2 increased the mobility in the native gel of the complex hTLR2ED–Pim2. In contrast, the extra two Man and the extra three carbons in one of the acyl chains of Pim4 (Figure 6) decreased the mobility of hTLR2ED-Pim4 (Figure 8C). Previously published results showed that human TLR2 has an impaired recognition of lipopeptides with acyl chains shorter than 16 carbons atoms (C16)
33
and can recognize longer acyl chains than C16.
34
We report here that human TLR2 can bind glycolipids with an acyl chain of 19 carbon atoms (C19), demonstrating that human TLR2 can, indeed, accommodate acyl chains longer than C16 in its hydrophobic binding pocket. Taken together, these binding assays demonstrate that the presence of two acyl chains linked to glycerol is the essential ligand moiety recognized by TLR2 ligands, and that both glycolipids and lipopeptides bind to the same binding site in hTLR2ED. Based on the crystal structure of hTLR2ED–Pam3CSK4,
12
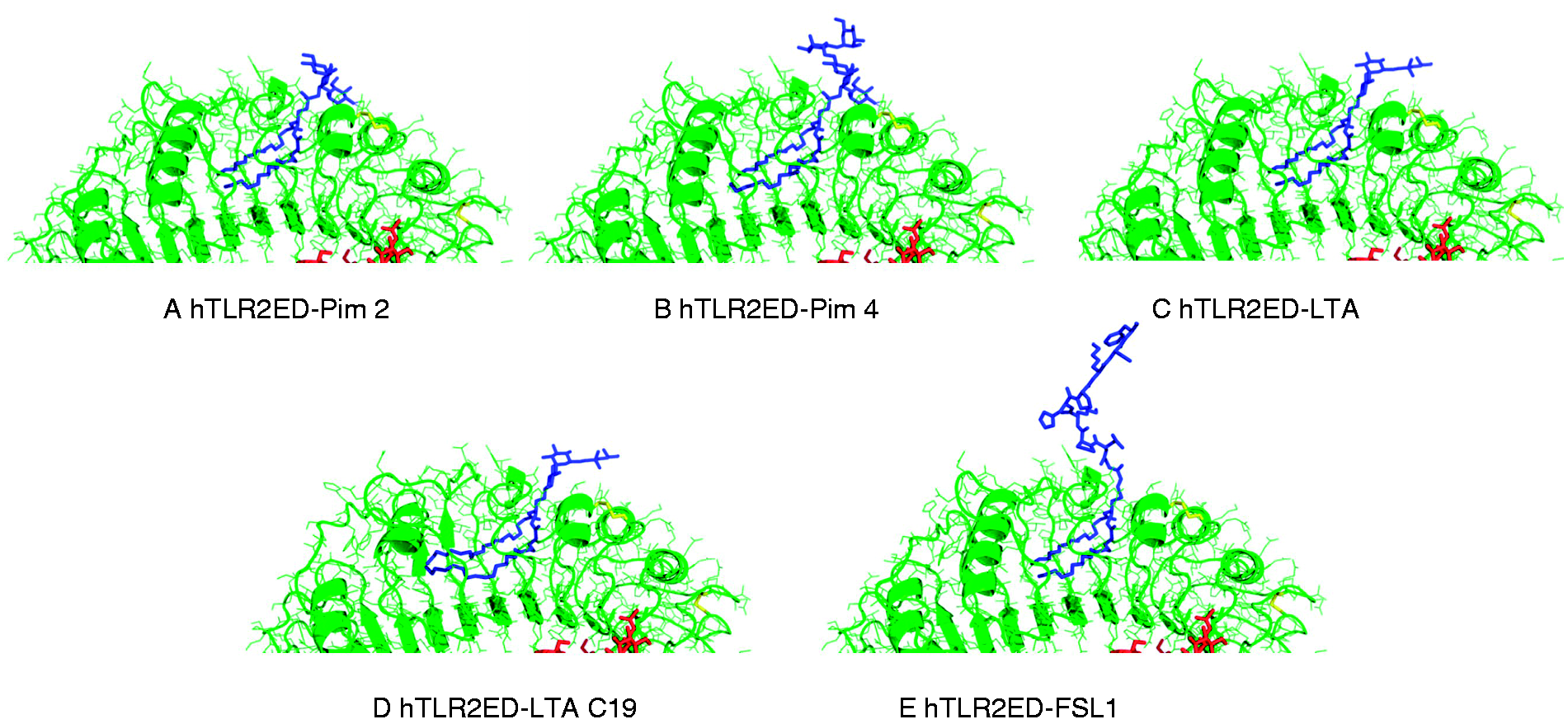
we modeled the hTLR2ED–FSL-1, hTLR2ED–LTA, hTLR2–LTA (C19), hTLR2ED–Pim2, hTLR2–Pim4 complexes (Figures 9 and 10). No clashes were found between any of the ligand head groups and hTLR2ED, indicating that the binding pocket of hTLR2ED might accommodate all of these ligands without major structural rearrangements. In addition, our models suggest that the hTLR2ED could also accommodate ligands with longer acyl chains than C16 without displacement of the ligand head group and only slight conformational adjustments of some of the loops on the outer face of hTLR2ED.
Modeling of hTLR2ED in complex with (A) Pim2, (B) Pim4, (C) LTA, (D) LTA 19 C and (E) FSL-1. Modeling is based on the crystal structures of hTLR2ED in complex with Pam3CSK4 (PDB 2Z7X)
12
and the Nogo receptor (PDB 1OZN).
19
The most abundant N-linked glycans from the glycan analysis were added to the hTLR2ED models: Man3GlcNAc2 (Asn114), Man9GlcNAc2 (Asn199), Man3Fuc1GlcNAc2 (Asn414), Man9Glc1GlcNAc2 (Asn442). All ligands have two acyl chains with 16 carbon atoms, except for LTA 19 C, which has two acyl chains with 19 carbon atoms and Pim4 which has one acyl chain of 19 carbon atoms and another of 16 carbon atoms. Binding sites of hTLR2ED. Close-up views of the modeled binding sites for hTLR2ED in complex with (A) Pim2, (B) Pim4, (C) LTA, (D) LTA 19 C and (E) FSL-1.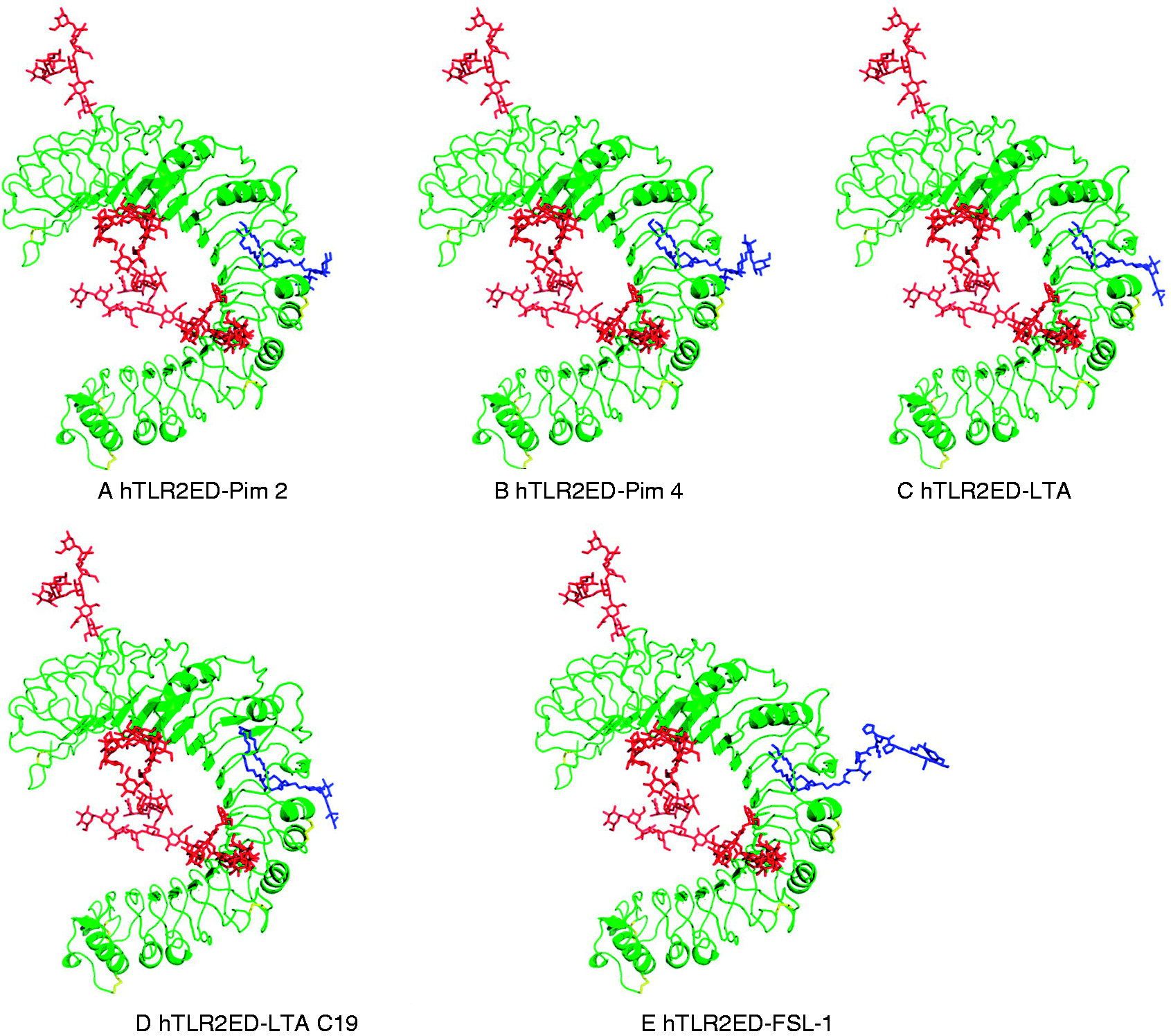
Discussion
TLRs are one of the most important groups of receptors responsible for the recognition of bacteria, virus, parasites and initiation of a host immune response. TLRs are PRRs because they recognize conserved structures in macromolecules in these microorganisms. The crystal structures of hTLR2ED–Pam3CSK4, mTLR2ED–Pam2CSK4 and mTLR2ED–Pam3CSK4 showed that the two acyl chains connected to the cysteinyl S-glycerol occupy the hydrophobic binding pocket of TLR2ED. 12
Our aim was to investigate binding of other relevant lipopeptides and glycopeptides to hTLR2ED. Therefore, we expressed and purified the complete ectodomain of human TLR2 (hTLR2ED) using a baculovirus expression system. Purified hTLR2ED is a monomer in solution with intramolecular disulfide bonds and N-linked glycosylation of the high-Man type. It is likely that, besides disulfide bonds in the N-terminal cap and LRR domain, two disulfide bonds are also present in the C-terminal cap domain (Figure 1). This assumption is based on the observation of disulfide bonds in similar positions in hTLR3ED35–37 and the Nogo receptor. 19 Employing CD spectroscopy, we followed the thermal stability of hTLR2ED. We employed different algorithms to determine the remaining secondary structure, but did not obtain a satisfactory result, as the theoretical curves were statistically far away from the experimental curves. Although some secondary structure remains at 50℃, the majority of the secondary structure appears to be lost at 95℃ as the curve shifts to a peak of 208 nm (random coil). Furthermore, although we did not measure the aggregation state, it is likely that the loss of secondary structure shown as the curve shifts to 208 nm would expose to the solvent the hydrophobic residues of the LRR motifs and result in aggregation. The relatively low value of ∼ 45–55℃ for the first melting point of hTLR2ED suggests that its secondary structure is thermally not very stable and might also indicate some structural flexibility. Asparagines (‘asparagine ladder’) and phenylalanines (‘phenylalanine spine’) in the consensus LRR repeats are known to be relevant for stabilization of LRR proteins through hydrogen bond formation and hydrophobic interactions, respectively. 38 The lack of those particular features in the central LRR repeats of hTLR2ED 12 (Figure 1) may render this domain more flexible and able to accommodate ligands with long acyl chains, thereby contributing to an explanation of its low melting point. We have previously tried other binding methods, such as surface plasmon resonance, to evaluate binding of ligands to TLR2, but did not achieve satisfactory results. Thus, we decided to employ native PAGE, which we have also successfully employed to study the binding to TLRs and accessory molecules such as TLR3 with dsRNA, 37 CD36 with LTA, Pims, FSL-1, 10 MD-1 with LPS 39 and TLR5 with flagellin. 40 Electrophoretic mobility shift assays have also been successfully used to study the binding between LTA–CD14 41 and TLR4–MD-2 with LPS. 42 Ligand binding alters the hydrodynamic size and/or charge of the protein, which then manifests itself in different electrophoretic mobilities of the liganded and unliganded forms. Here, we have demonstrated that hTLR2ED directly binds synthetic, diacyl-FSL-1 and Pam2CSK4, and triacyl-lipopeptides (Pam3CSK4), purified LTA from S. aureus, and synthetic Pim2 and Pim4, in the absence of TLR1 or TLR6. Soluble isoforms of hTLR2ED present in plasma, 43 breast milk,43,44 amniotic fluids, 45 saliva 46 and expressed by HEK cells 47 can act as a decoys of TLR2 and inhibit NFκB activation by transmembrane TLR2. 48 The hTLR2ED expressed in insect cells should have the same inhibitory effect as the hTLR2ED from natural sources as the only difference with them would be in the types of glycan structures, which should not be relevant for binding as the glycosylation sites are not located near the binding site of hTLR2ED. Therefore, our ligand-binding studies under physiological conditions suggest that ligand competition between secreted isoforms of hTLR2ED and the transmembrane hTLR2 receptor is likely to be responsible for the down-regulation of TLR2 activation. Soluble TLR2 has also been implicated in the pathogenesis of HIV.28,44 Thus, we investigated whether hTLR2ED could interact with HIV-1 gp120. In native PAGE experiments, we did not observe binding between hTLR2ED and gp120 from HIV-1, consistent with gp120 not being a lipoprotein.
One of the main classes of TLR2 ligands are microbial lipoproteins. Synthetic lipopeptides can mimic the activity of microbial lipoproteins; therefore, we investigated interaction of these peptides with hTLR2ED. CSK4 represents the amino acid moiety of synthetic lipopeptides, such as Pam1CSK4, Pam2CSK4 and Pam3CSK4. No appreciable differences were observed in the electrophoretic mobility of hTLR2ED with and without CSK4, demonstrating that hTLR2ED does not interact with the amino acid moiety of lipopeptides. Although no shift was seen for the hTLR2ED–Pam1CSK4 complex, a gel shift was observed when hTLR2ED bound Pam2CSK4. Although we did not detect binding of Pam1CSK4 to hTLR2ED, it is still possible that hTLR2ED may bind Pam1CSK4 with low affinity. The absence of the diacylglycerol moiety in Pam1CSK4 explains its low biological activity compared with Pam2CSK4. 49
The crystal structure of the hTLR2ED/hTLR1ED complex reveals that TLR2 binds the two acyl chains linked to the cysteinyl S-glycerol of Pam3CSK4 and not the N-acyl chain, which is bound by TLR1. The two cysteinyl S-glycerol fatty acid chains can establish a greater number of hydrophobic interactions and thus form a more stable complex with hTLR2ED.
FSL-1 is a synthetic diacyl-lipopeptide that mimics the N-terminal sequence of the 44-ku lipoprotein LP44 of Mycoplasma salivarium.50–52 FSL-1 is a potent activator of antigen presenting cells and is a useful tool for treatment of cancer and viral infections. 53 In native PAGE experiments, we observed that hTLR2ED binds Pam2CSK4 and FSL-1 fluorescent ligands. FSL-1 shares the cysteinyl S-glycerolipid of Pam2CSK4, but differs in the amino acids that are bound to the cysteine. Thus, as in the case of Pam2CSK4, hTLR2ED is likely to interact with the cysteinyl S-diacyl glycerol moiety of FSL-1.
Peptidoglycan was thought to be a ligand for TLR2, but a recent report clearly showed that a purified peptidoglycan shows no activity, and that earlier reported activity was caused by the presence of contaminants, such as LTA. 54 Recent reports55–58 also challenge the concept of LTA as a ligand for TLR2, suggesting that its reported activity is actually caused by contaminating lipoproteins. However, synthetic S. aureus LTA was reported to induce secretion of TNF-α from whole blood cells and macrophages in a TLR2-dependent manner.59,60
Although peptidoglycan does not have fatty acid chains, LTA has a diacylglycerol structure similar to FSL-1, Pam2CSK4 and Pam3CSK4. Here, we show a considerable increase in the electrophoretic mobility of hTLR2ED after incubation with LTA from S. aureus. LTA from S. aureus consists of up to 50 negatively charged glycerophosphate units linked to a gentiobiosyl diacylglycerolipid that anchors the LTA to the plasma membrane, 59 thereby explaining the increased electrophoretic mobility of the hTLR2ED–LTA complex compared with uncomplexed hTLR2ED. Similarly increased electrophoretic mobility was shown for CD14–LTA and LBP–LTA complexes as compared with unliganded proteins in native PAGE experiments. 41 We also found that LTA can form aggregates with synthetic Pam2CSK4f, but that these complexes are not recognized by hTLR2ED. In fact, the electrophoretic mobility of hTLR2ED incubated with LTA and Pam2CSK4f is identical to that obtained when hTLR2ED is incubated with LTA alone. The acyl chains of the diacylglycerol anchor, as in the case of synthetic lipopeptides, are likely to be the motif responsible for interaction with hTLR2ED. The crystal structure of mTLR2ED/VLR in complex with LTA from S. pneumoniae also demonstrates the relevance of the diacylglycerol of LTA as the ligand binding moiety recognized by TLR2. 13 However LTA from S. pneumoniae has been shown to have an activity independent of TLR2, 58 suggesting that binding to the hydrophobic pocket of TLR2 is necessary, but not sufficient, for a microbial ligand to activate TLR2.
The acyl chains of LTA from S. aureus have a length between C14 and C19,
59
while those in Pam2CSK4 have only 16 carbon atoms (palmitates). Our modeling, based on the published crystal structure of TLR2,
12
shows that the hydrophobic pocket can accommodate longer fatty acid chains than palmitic acid with minimal conformational changes to the outer loops of hTLR2ED. Thus, the extra carbon residues of some LTA molecules could establish additional hydrophobic interactions compared with Pam2CSK4 and could explain the preferential binding of hTLR2ED for LTA from S. aureus. The binding of Pim4, which has one of the two acyl chains composed of 19 carbon atoms, then confirms that TLR2 can accommodate ligands with acyl chains longer than 16 carbon atoms. Mycobacteria are intracellular pathogens that lives inside macrophages, inhibiting macrophage activation and phagosome maturation.61–63 Mycobacterium tuberculosis causes around two million deaths per year worldwide.
64
TLR2 has been shown to be important for controlling M. tuberculosis
65
and M. leprae
66
infections. Mycobacteria possess a unique cell wall structure rich in long fatty acids (mycolic acid), lipoglycans (lipoarabinomannan) and polysaccharides. Lipoarabinomannan is composed of three structures: a phosphatidyl-mannosyl-myo-inositol anchor, a polysaccharide backbone composed of
It is likely that hTLR2ED also binds to the acyl chains linked to the diacylglycerol moieties, as for lipopeptides. Although synthetic Pim2 and Pim4 were able to bind TLR2, they were not able to support TNF-α secretion from C57BL/6 mice. 10 Pims have also been shown to inhibit the activation of TLR2 in a CD14-independent manner, 68 decrease allergic response69,70 and affect dendritic cell maturation. 71 Lipoarabinomannan and Pims were reported to be sensed by TLR1 and TLR2. 41 The failure to activate the TLR2 signaling pathway by these synthetic Pims likely results from the lack of a third acyl chain, which is necessary for TLR1 binding.72,73 Therefore, the capacity of Pims to bind, but not activate, the TLR2 signaling pathway could explain their inhibitory properties. The binding site of hTLR2ED is situated on its convex face, while the concave face of hTLR2ED is covered by glycosylation, making it an unlikely site for binding ligands. The hydrophobic binding pocket of hTLR2ED, situated in the transition between the central LRR domain and the C-terminal LRR domain (LRR 9–12), is the only binding site large enough to accommodate diacylglycerol structures, 12 making it unlikely that ligands bind to a secondary site, even at the high concentration of ligands employed. However, mutations that abrogate ligand binding, such as the mutation of isoleucine 319 to aspartate in the LRR 11 domain of human TLR2 74 (Figure 1), could be tested, for example. We used in vitro conditions to test the binding of hTLR2ED with its ligands, where ligand concentrations are higher than in vivo conditions so that the experiments can be performed in the absence of accessory molecules, such as CD36 or CD14. In vivo, CD14 and CD36 are relevant to efficiently load ligands into TLR2 and activate the TLR2-dependent signaling. 10 Because of differences in the hydrophobic pocket of TLR2 between human and other species, such as murine 33 and equine, 75 other mammals may not be suitable models for the development of inhibitors for human TLR2. Instead, ligand binding assays, such as those using native PAGE, may be a better approach for evaluating inhibitors for human TLR2.
In summary, we have shown that hTLR2ED is able to bind synthetic FSL-1, Pim2, Pim4 and LTA from S. aureus without the presence of its co-receptors TLR1 and TLR6. Although the structures of ligands recognized by hTLR2ED are all different, they all share the presence of two fatty acid chains linked to glycerol that is recognized by hTLR2ED (Figures 6, 9 and 10). Overall, our data indicate the presence of a single binding site in hTLR2ED for microbial glycolipids and lipoproteins. Inhibitors with longer acyl chains than 16 carbon atoms, which bind, but do not activate, TLR2 signaling, could represent a novel strategy for the treatment of septic shock caused by Gram-positive bacteria.
Footnotes
Funding
This study was supported by NIH grant AI42266 (IAW) and by a Joint Scripps/Oxford Graduate Scholarship. We also thank the Wellcome Trust for grants to purchase the TofSpec and Q-TOF mass spectrometers, and the Oxford Glycobiology Bequest for additional funding.
Conflict of interest
The authors do not have any potential conflicts of interest to declare.
Acknowledgements
We thank Dr. Terry Butters for providing the α-glucosidase II as well as for helpful discussions, Julie Vanhnasy for technical support, and Drs. Robyn Stanfield and Adam Corper for comments and suggestions on the manuscript.
