Abstract
Antibiotic-resistant bacterial pathogens threaten public health. Because many antibiotics target specific bacterial enzymes or reactions, corresponding genes may mutate under selection and lead to antibiotic resistance. Accordingly, antimicrobials that selectively target overall microbial cell integrity may offer alternative approaches to therapeutic design. Naturally occurring mammalian α- and θ-defensins are potent, non-toxic microbicides that may be useful for treating infections by antibiotic-resistant pathogens because certain defensin peptides disrupt bacterial, but not mammalian, cell membranes. To test this concept, clinical isolates of methicillin-resistant Staphylococcus aureus (MRSA), including vancomycin heteroresistant strains, and ciprofloxacin-resistant Pseudomonas aeruginosa (CipR-PA) were tested for sensitivity to α-defensins Crp-4, RMAD-4 and HNPs 1-3, and to RTD-1, macaque θ-defensin-1. In vitro, 3 μM Crp-4, RMAD-4 and RTD-1 reduced MRSA cell survival by 99%, regardless of vancomycin susceptibility. For PA clinical isolates that differ in fluoroquinolone resistance and virulence phenotype, peptide efficacy was independent of strain ciprofloxacin resistance, site of isolation or virulence factor expression. Thus, Crp-4, RMAD-4 and RTD-1 are effective in vitro antimicrobials against clinical isolates of MRSA and CipR-PA, perhaps providing templates for development of α- and θ-defensin-based microbicides against antibiotic resistant or virulent infectious agents.
Introduction
The economic costs of antimicrobial resistant infections has reached nearly $30 billion per year in the USA. 1 In 2002, of health care-associated infections that resulted in 99,000 deaths, approximately 16% were reported to the Centers of Disease Control as resistant to antibiotics.2–5 The increase in antibiotic resistance narrows the options for the treatment of infections caused by multidrug-resistant bacteria.5,6
The Gram-negative bacterium Pseudomonas aeruginosa (PA) is a leading cause of nosocomial infections for which the fluoroquinolone antibiotics, for example ciprofloxacin, are commonly prescribed. Over the last decade, the prevalence of ciprofloxacin-resistant (CipR) PA strains has increased threefold in parallel to the trend of prescribing this antibiotic class. 7 Upwards of 30% of clinical PA strains are multidrug-resistant, and some are resistant to all available antibiotics.5,8–10 In addition, PA strains have an arsenal of virulence factors, including a type III secretion system (TTSS) that induces cytotoxicity and expression of ExoU or ExoS effector proteins, which are virulence factors that influence disease severity by phagocyte evasion during acute infections. 11
Methicillin-resistant Staphylococcus aureus (MRSA) are Gram-positive bacteria with resistance to all β-lactam compounds except one, accounting for nearly 60% of all clinical isolates from intensive care unit (ICU) patients. 12 Limited to the healthcare setting in the past, new, more virulent MRSA strains have emerged in the community, and they are now responsible for infections across both the community and healthcare settings. In addition, MRSA strains have increasingly developed varying degrees of resistance to vancomycin, the accepted treatment standard, 13 and hospital-associated healthcare costs for patients with MRSA-related infections are nearly double those of patients with methicillin-sensitive S. aureus. 14
Multidrug resistant PA and MRSA are treated with antibiotics that target specific bacterial reactions or enzymes to inhibit cell replication and cell wall biosynthesis, or they may kill bacteria directly. Antibiotic exposure enables bacteria to acquire resistance through mutations that allow for target drug degradation, reduced drug affinity to target sites, altered metabolic pathways and/or reduced drug accumulation.15–17 For example, ciprofloxacin inhibits bacterial replication by inhibiting DNA gyrase and topoisomerase IV, thereby blocking bacterial cell division.
18
On the one hand, CipRPA acquire mutations to DNA gyrase and/or topoisomerase to escape its lethal antibiotic effects and by active removal of ciprofloxacin via broad substrate efflux pumps to expel the drug and prevent its accumulation within cells.
17
On the other hand, vancomycin inhibits bacterial cell wall biosynthesis by binding to the C-terminal
Mammalian defensins are 2–5 kDa, broad spectrum, cationic antimicrobial peptides, with structures that are defined by specific tridisulfide arrays. 23 The α-, β- and θ-defensins compose the mammalian defensin family, and each subfamily differs with respect to structural features, which are imposed by specific disulfide connectivities, and they have distinct primary sites of expression. 24 For example, the α-defensin tridisulfide array is characterized by conserved CI-VI, CII-IV, CIII-V cysteine pairings, and β-defensins are characterized by a CI-V, CII-IV, CIII-VI disulfide connectivities. 25 α-Defensin genes are expressed by promyelocytes and mainly accumulate in neutrophil azurophil granules, and those expressed by Paneth cells are secreted into the small intestinal lumen. Certain β-defensins, however, are expressed widely by epithelia at diverse mucosal barrier interfaces.26,27 θ-Defensins are the only macrocyclic peptides known in the animal kingdom and are expressed in the bone marrow of only Old World monkeys. The peptides derive from truncated α-defensin genes, assemble from two hemi-precursors and contain a tridisulfide array that is arranged in the form of a parallel ladder. 28
The mechanisms of defensin microbicidal activity have been investigated extensively. 29 Based on in vitro studies, micromolar levels of α-defensins may disrupt microbial membranes selectively by inducing either stable or transient defects of variable size in model membranes composed of microbial phospholipids.30,31 Induction of membrane defects leads to target cell permeabilization, K+ efflux, depolarization, dissipation of electrochemical gradients, leakage and eventual cell death.32–35 Analyses of defensin-bilayer interactions by small angle X-ray scatter showed that mouse Paneth cell α-defensin cryptdin-4 (Crp-4), rhesus myeloid α-defensin RMAD-4 and the rhesus θ-defensin RTD-1 induce negative Gaussian, or saddle splay, curvature to create pores in model membranes, facilitating membrane disruption. 36 However, at lower peptide concentrations defensins can also inhibit bacterial peptidoglycan synthesis by lipid II binding,37,38 and defensins may use a lipid II binding mechanism to exert antimicrobial effects.
Because α- and θ-defensins kill bacteria by these general mechanisms, which differ from those of ciprofloxacin and vancomycin, we reasoned that vancomycin-heteroresistant MRSA and CipR PA may be susceptible to the microbicidal effects of exposure to these defensin peptides. To test this hypothesis, the survival of clinical isolates of vancomycin-heteroresistant MRSA and CipR PA exposed to Crp4, RMAD-4, RTD-1 and human neutrophil α-defensins (human neutrophil peptides; HNP) HNPs 1–3 were determined. Under the conditions of the in vitro bactericidal assays, nearly all MRSA and PA strains were sensitive to all peptides except for HNPs, irrespective of antibiotic resistance. In addition, PA sensitivity to α-defensins was not related to site of isolation, degree of ciprofloxacin resistance, TTSS effector genotype or cytotoxic potential. Because Crp-4, RMAD-4, and RTD-1 are non-hemolytic, resistant to proteolytic degradation and among the most potent known defensins, they may offer promise for development of novel antimicrobial therapeutics.
Materials and methods
Peptide preparation
Peptides (Figure 1) were purified to homogeneity by reverse phase (RP)-HPLC, and their identities were confirmed by MALDI-TOF MS and by acid–urea (AU)-PAGE, as described previously.39,40 Recombinant Crp-4 and RMAD-4 peptides were expressed in Escherichia coli as N-terminal His6-tagged fusion proteins using the pET28a expression system (Novagen, Madison, WI, USA).32,41,42 Crp-4 and RMAD-4 templates were cloned in pCR-2.1 TOPO, verified by DNA sequencing, subcloned into pET28a plasmid DNA (Novagen) and transformed into E. coli BL21(DE3)-CodonPlus-RIL cells (Stratagene, Santa Clarita, CA, USA) for recombinant expression.32,41 His6-tagged Crp-4 fusion peptides were purified using nickel-nitrilotriacetic acid (Qiagen, Valencia, CA, USA) resin affinity chromatography.
43
After cyanogen bromide (CNBr) cleavage of the His6tag fusion, peptides were purified by sequential C18 RP-HPLC, and molecular masses of purified peptides were determined using MALDI-TOF MS on a Bruker Microflex LRF (Bruker, Fremont, CA, USA). Solution structures of recombinant peptides prepared by this approach have been determined by NMR.
44
Primary structures of α- and θ-defensins. The primary structures of peptides investigated are aligned; net overall charge is shown on the right. α-Defensins are aligned by their conserved Cys residues (in bold), and their CI-CVI, CII-CIV, and CIII-CV disulfide connectivities are depicted by brackets. The α-defensins from mouse Paneth cells and rhesus neutrophils are highly cationic compared with the human myeloid peptides, HNPs 1–3. The octadecapeptide RTD-1, a θ-defensin, is a macrocyclic peptide identified in rhesus macaque neutrophils and found only in Old World monkeys.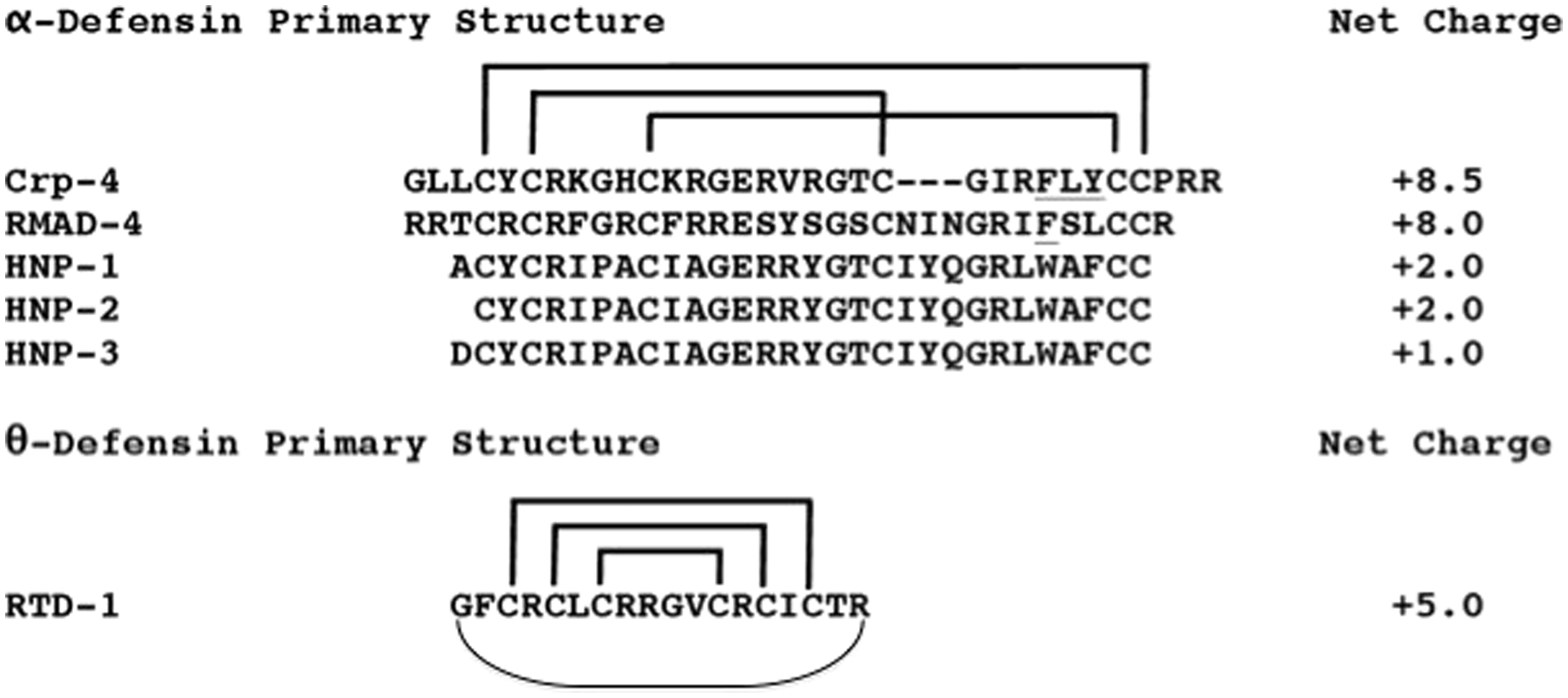
HNPs were isolated from samples enriched in neutrophil granules prepared from peripheral blood leukocytes.45,46 Briefly, after removing platelets from human blood by centrifugation at 200 g for 10 min and hypotonic lysis of red blood cells, the leukocyte-enriched cell fraction was re-suspended in 0.34 M sucrose, pH 7.4, homogenized and cellular debris deposited by centrifugation at 200 g for 10 min, leaving a granule-rich supernatant. Granules deposited by 30 min centrifugation at 27,000 g at 4℃ were extracted overnight (18 h) at 4℃ with 10% acetic acid, and protein extracts were clarified by centrifugation at 27,000 g for 30 min at 4℃. A mixture of natural HNPs 1–3 was purified by gel permeation chromatography using BioGel P10 (Bio-Rad Laboratories, Hercules, CA, USA). HNP-2 was purified further from pooled HNPs 1–3 by C18 RP-HPLC.
RTD-1 was synthesized as previously reported. 47 Synthetic RTD-1 is structurally and biologically indistinguishable from peptide isolated from rhesus monkey neutrophils. 28
Clinical isolates
Bacterial isolates were grown from culture specimens obtained from hospitalized patients at Huntington Hospital (Pasadena, CA, USA) as part of a longitudinal epidemiologic surveillance study of resistant pathogens at the institution, and were stored at –80oC until testing. All data were analyzed anonymously. Susceptibility of MRSA isolates to vancomycin was determined by an Etest-based method according to the manufacturer's instructions (bioMeriéux, Durham, NC, USA). Specifically, vancomycin heteroresistant phenotype was determined using the Etest glycopeptide resistance detection (GRD) method. 48 PA susceptibility to ciprofloxacin was performed by broth microdilution method, as recommended by the Clinical Laboratory Standards Institute, with ciprofloxacin resistance defined by a minimum inhibitory concentration of 2 µg/ml or greater. As for PA strains, genes encoding the TTSS effector proteins ExoU and ExoS were assayed by PCR, as described previously. 49 In vitro experiments were performed to determine PA cytotoxicity by infecting A549 lung epithelial cells at a multiplicity of infection of 10 and measuring lactate dehydrogenase release at 3 h post-infection using the CytoTox96 assay kit (Promega, Madison, WI, USA).
MRSA bloodstream isolates and PA strains that caused pneumonia, as well as bacteremia, wound and urinary tract infections were selected for this study to represent clinical isolates that exhibit varying degrees of resistance to vancomycin (MRSA) and ciprofloxacin (PA), respectively. The MRSA cohort included strains that caused persistent bloodstream infection and different molecular epidemiologic characteristics; PA strains included those with different virulence potential based on TTSS effector genotype, and the rate and extent of cytotoxicity observed in A549 cells. Two additional reference MRSA strains with heteroresistance (Mu3) and intermediate resistance (Mu50) to vancomycin also were studied.
In vitro bactericidal assays
The α- and θ-defensins were tested for bactericidal activity against clinical isolates of MRSA and PA in in vitro cell suspension assays.32,41 Bacteria grown to mid-exponential phase in trypticase soy broth were deposited by centrifugation at 10,000 g for 3 min, and washed three times with 10 mM piperazine-1,4-bis(2-ethanesulfonic acid) (PIPES), pH 7.4, supplemented with 1% (vol/vol) of trypticase soy broth (10 mM PIPES–TSB). In triplicate, 1–5 × 106 bacterial colony forming units (CFU)/ml were exposed to peptides in 50 μl 10 mM PIPES–TSB in 96-well polystyrene plates. Samples were incubated at 37℃ with shaking for 1 h, diluted 1:100 in 10 mM PIPES (pH 7.4) and plated on TSB agar plates using an Autoplate 4000 (Spiral Biotech, Bethesda, MD, USA). Bacterial cell survival as a function of peptide exposure was determined by counting CFU after overnight growth at 37℃. In these assays, after 1 h of peptide exposure replicate peptide–bacterial mixtures are plated with a plating stylus onto the agar plate surface. The assay enables bacteria that are not exposed or affected by peptide exposure to be enumerated, rather than produce lawns that cannot be counted. However, sample dilution limits the lower end of the assay, that is, when > 99.9% of exposed bacteria die. Because of the dilution factors involved, plates on which no CFU were detected after overnight incubation may actually have had between 1 and 999 viable CFU in the peptide–bacterial mixture before dilution and plating. Accordingly, the limit of detection is ≤ 103 CFU/ml, when no CFU are detected, and the ordinates of bacterial survival curves are labeled in that manner.
Results
Defensin sensitivity of vancomycin-heteroresistant MRSA strains
MRSA clinical isolates with varied resistance to vancomycin were exposed to Crp-4, RMAD-4 and RTD-1 to determine their sensitivities to the peptides. Against standard laboratory strains of S. aureus, Crp-4, RMAD-4 and RTD-1 are potent microbicides, reducing cell viability 1000-fold at low micromolar peptide concentrations.50,51 Here, Crp-4, RMAD-4 and RTD-1 similarly reduced viability of vancomycin-intermediate S. aureus (VISA) control strain Mu50 and heteroresistant VISA (hVISA) control strain Mu3 with minimal bactericidal concentration (MBC) values between 1.5 and 3 µM peptide (Table 1; Figure 2A, B). Clinical isolates characterized as hVISA by the Etest GRD method (see ‘Materials and methods’) were highly sensitive to all defensins tested (Figure 2C, D), with RMAD-4 had the greatest bactericidal activity against every VISA/hVISA strain tested with 1.5 μM MBC values for all strains (Table 1; Figure 2A–D).
Sensitivity of control MRSA strains to RMAD-4, Crp-4 and RTD-1 is independent of antibiotic resistance. Bactericidal peptide assays were performed in triplicate as described (see ‘Materials and methods’). Briefly, MRSA strains were exposed to peptides for 1 h, and surviving bacteria were counted as CFU/ml at each peptide concentration (see ‘Materials and methods’). Values ≤ 103 CFU/ml signify that no colonies were detected on plates after overnight growth. Strain Mu50 (A) is a VISA control, and Mu3 (B), DH290 (C) and DH280 (D) are classified as hVISA strains. Each condition was performed in triplicate, with error bars denoting SD from the mean. Symbols: Crp-4 (-•-), RMAD-4 (-○-) and RTD-1 (▾). Defensin MBC values against VISA and hVISA MRSA strains. Infection cleared by vancomycin administration. Persistent infection with vancomycin administration.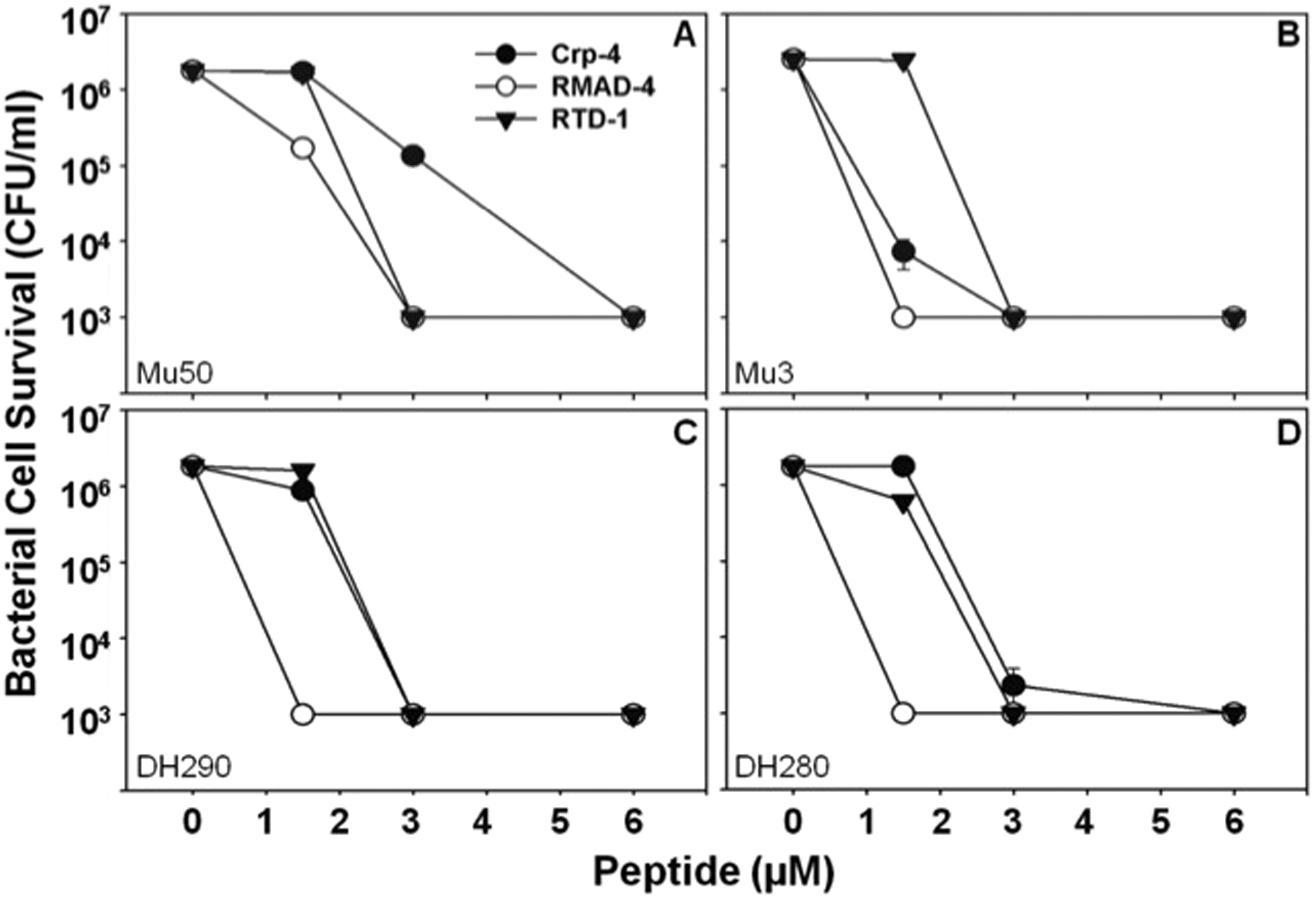
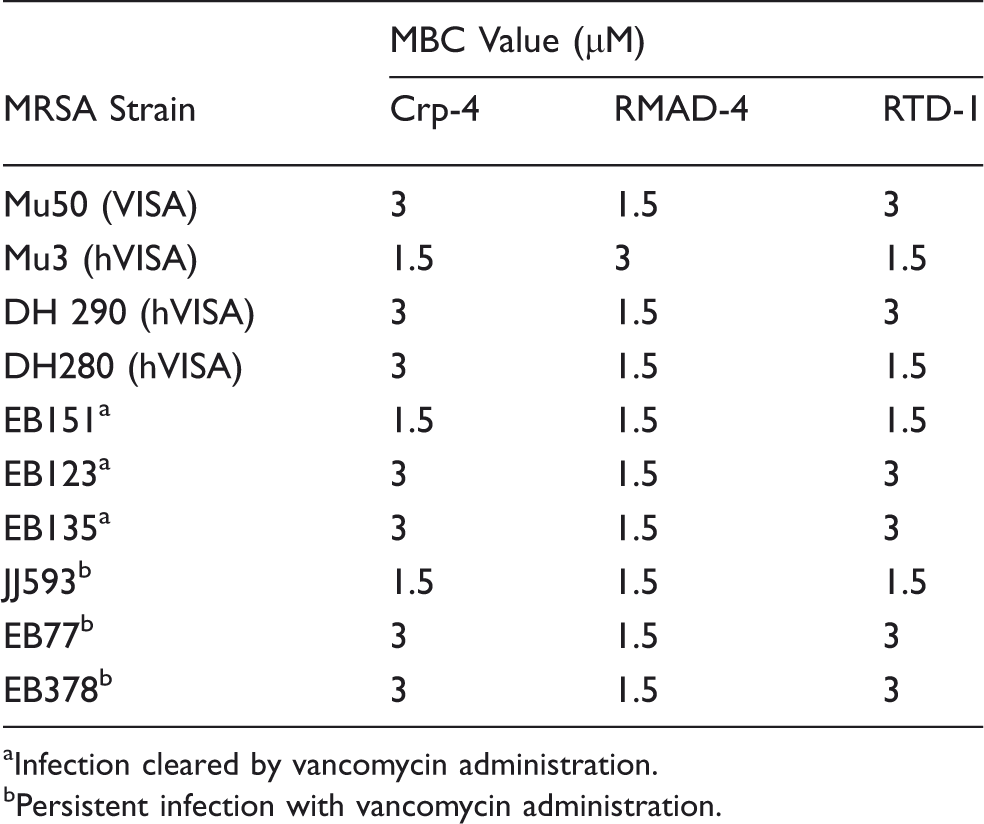
The potent microbicidal effects of Crp-4, RMAD-4 and RTD-1 against VISA/hVISA control strains prompted us to test whether MRSA that had persisted in bacteremic patients after extensive vancomycin treatment would be sensitive to these defensins. Clinical blood isolates of MRSA that were not cleared by vancomycin treatment were assessed for survival after Crp-4, RMAD-4 and RTD-1 exposure in vitro, and all MRSA clinical isolates were sensitive to Crp-4, RMAD-4 and RTD-1 in a peptide concentration-dependent manner, regardless of response to vancomycin treatment (Table 1; Figure 3A–F). For example, the MBC for RMAD-4 was 1.5 μM against vancomycin-resistant and sensitive MRSA strains (Table 1; Figure 3), and no association existed between antibiotic resistance and defensin susceptibility. Also, vancomycin-heteroresistant MRSA isolated from bacteremic patients that failed to resolve with antibiotic therapy were susceptible to the three defensins and as sensitive as culture-adapted S. aureus strains (Table 1).
MRSA sensitivity to RMAD-4, Crp-4 and RTD-1 is not associated with antibiotic resistance. Bactericidal peptide assays were performed in triplicate as described (Figure 2; ‘Materials and methods’). Strains EB151 (A), EB123 (B) and EB135 (C) are clinical isolates from patients whose infections were cleared by vancomycin administration, in contrast to strains JJ593 (D), EB77 (E) and EB378 (F), which were isolated from persistent infections after failure of treatment with vancomycin. All MRSA strains were sensitive to the defensins tested at low micromolar concentrations (Table 1). Each condition was performed in triplicate with error bars denoting SD from the mean. Symbols: Crp-4 (-•-), RMAD-4 (-○-), and RTD-1 (-▾-).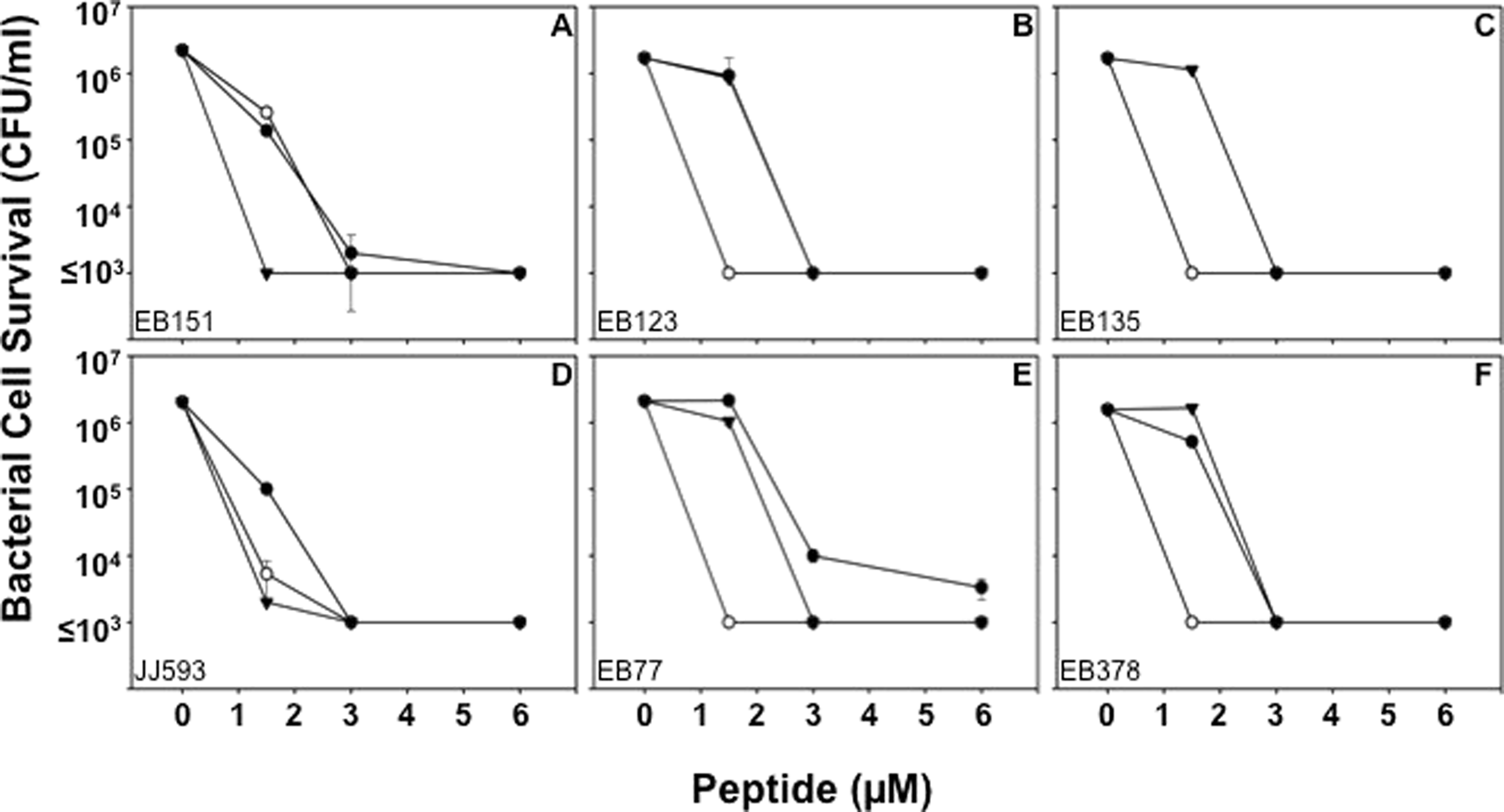
Sensitivity of ciprofloxacin-resistant PA to defensins
PA develops resistance to ciprofloxacin by mutations in DNA gyrase, topoisomerase IV or both, and via efflux pumps that remove the drug and prevent its accumulation. Because the α-defensins in this study are membrane-disruptive microbicides and peptide accumulation within target cells is not required for activity, we tested whether CipR PA are sensitive to Crp-4 and RMAD-4. Bactericidal assays against ciprofloxacin-sensitive (CipS) and CipR PA isolated from patient sputum showed that most PA sputum isolates were sensitive to both Crp-4 and RMAD-4 at low micromolar peptide levels, regardless of antibiotic resistance (Table 2; Figure 4; see Supplementary Figure S1). In contrast to the variability of the relative activities of Crp-4 and RMAD-4 against MRSA (Table 1; Figure 3), Crp-4 was consistently more active than RMAD-4 against most PA strains isolated from sputum (Table 2; Figure 4). Also, sputum isolates displayed more variability than MRSA strains regarding susceptibility to Crp-4 and RMAD-4. Consistent with the defensin sensitivities of vancomycin-heteroresistant MRSA strains, Crp-4 and RMAD-4 were bactericidal against PA sputum isolates irrespective of strain ciprofloxacin resistance, there being no association between PA isolate sensitivities to Crp-4 or RMAD-4 and antibiotic resistance.
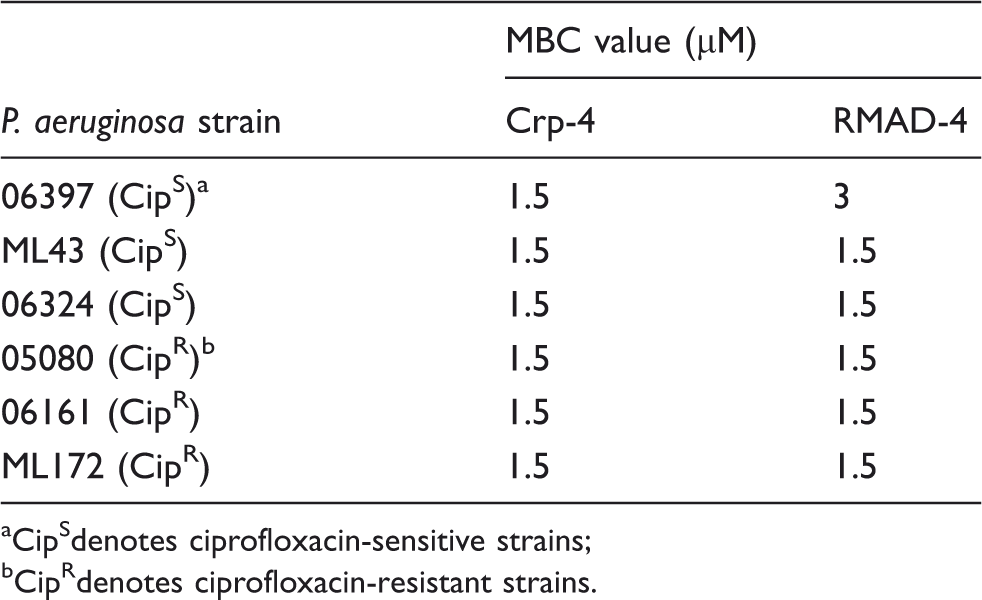
Activities of Crp-4 and RMAD-4 against CipR and CipS strains of PA. Bactericidal peptide assays were performed in triplicate against clinical isolates from patient sputum. Isolates 06397 (A), ML43 (B) and 06324 (C) are CipR-PA, and isolates 05080 (D), 06161 (E) and ML172 (F) are CipS-PA. No correlation exists between antibiotic resistance and α-defensin activity (Table 2). Bars denote SD from the mean. Symbols: Crp-4 (-•-) and RMAD-4 (-○-). α-Defensin MBC values against CipR and CipS strains of Pseudomonas aeruginosa. CipSdenotes ciprofloxacin-sensitive strains; CipRdenotes ciprofloxacin-resistant strains.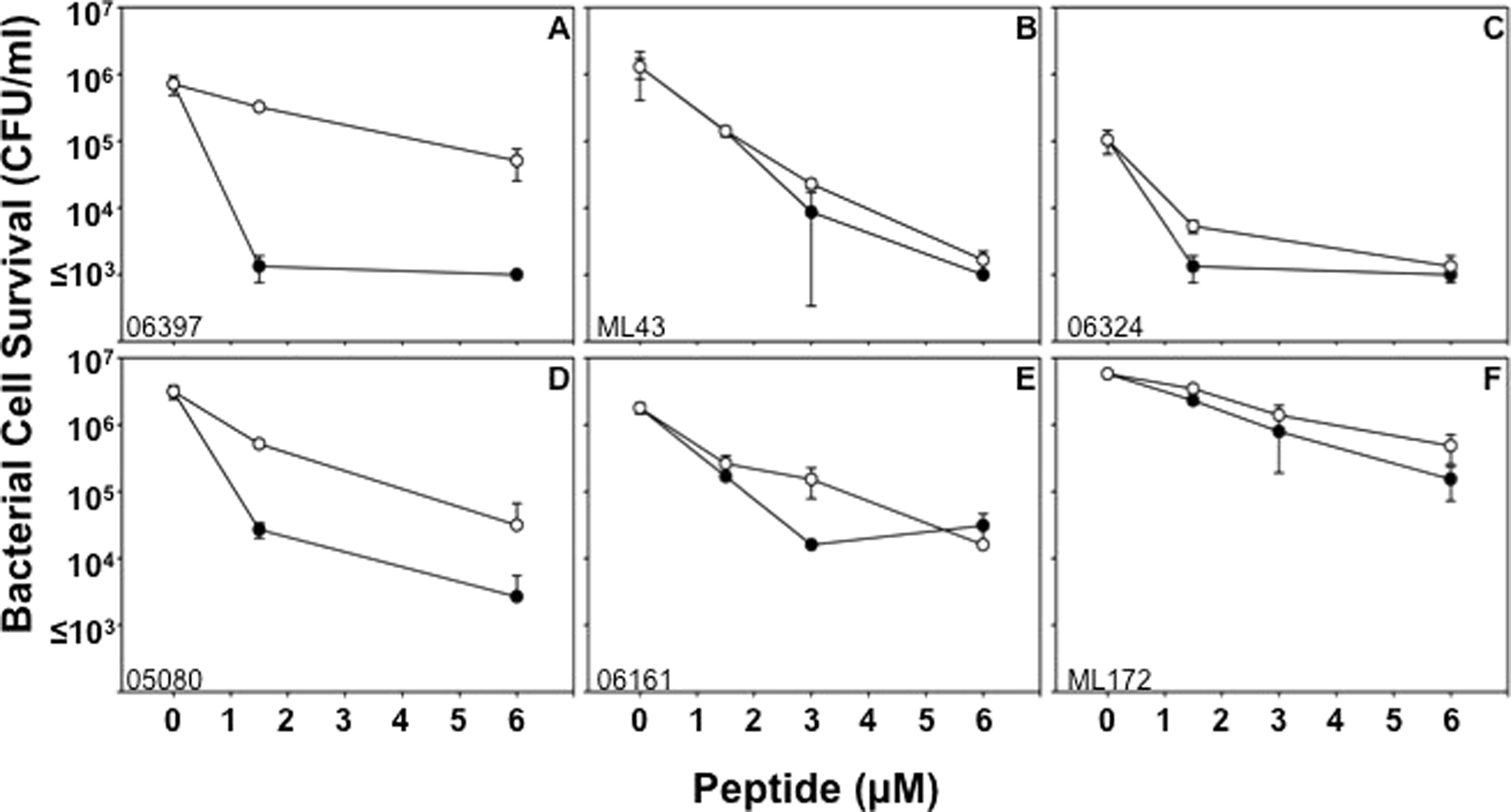
The activities of defensins against PA sputum isolates were tested further by determining the in vitro bactericidal effects of RTD-1, a mixture of natural HNPs 1–3, and purified HNP-2 against sputum PA isolates. RTD-1 at <1.5–6.0 μM reduced survival of the majority of sputum PA isolates to ≤ 103 CFU/ml, below the limit of detection for these assays (Figure 5). In contrast, HNPs 1–3 (Figure 4A–C) and HNP-2 (Figure 4D–F) did not affect survival of PA sputum isolates, even at peptide concentrations 10-fold greater than levels at which Crp-4 and RTD-1 are highly microbicidal. Thus, HNPs, endogenous α-defensins that occur at elevated serum concentrations during septicemia and certain infections in humans,
52
lacked bactericidal activity against CipR and CipS PA strains under these in vitro conditions. Although HNPs lacked activity against PA strains, they have been shown to have alternative innate host defense roles, including neutralization of anthrax lethal toxin and as chemoattractants.53,54 Because PA isolates were sensitive to Crp-4 and RMAD-4 irrespective of antibiotic resistance, we focused on characterizing the activities of those molecules, rather than optimizing the activities of HNPs. Perhaps, the non-human peptides, Crp-4, RMAD-4 and RTD-1, may offer promise as adjunct therapies for treatment of bacterial infections that fail to resolve with conventional treatment.
Attenuated activities of HNPs against PA clinical isolates. Survival of CipS P. aeruginosa strains 06397 (A), ML43 (B) and ML121 (C), and CipR-PA strains 06161 (D), 05038 (E) and ML172 (F) isolated from patient sputum was measured after exposure to Crp-4, RTD-1 and HNPs. In contrast to Crp-4 and RTD-1, 60 µM HNPs had little bactericidal effects against P. aeruginosa, irrespective of CipR. RTD-1 and HNPs assays were performed in triplicate, Crp-4 activity, assayed once at each peptide concentration, was consistent with Figure 3. Error bars denote SD from the mean.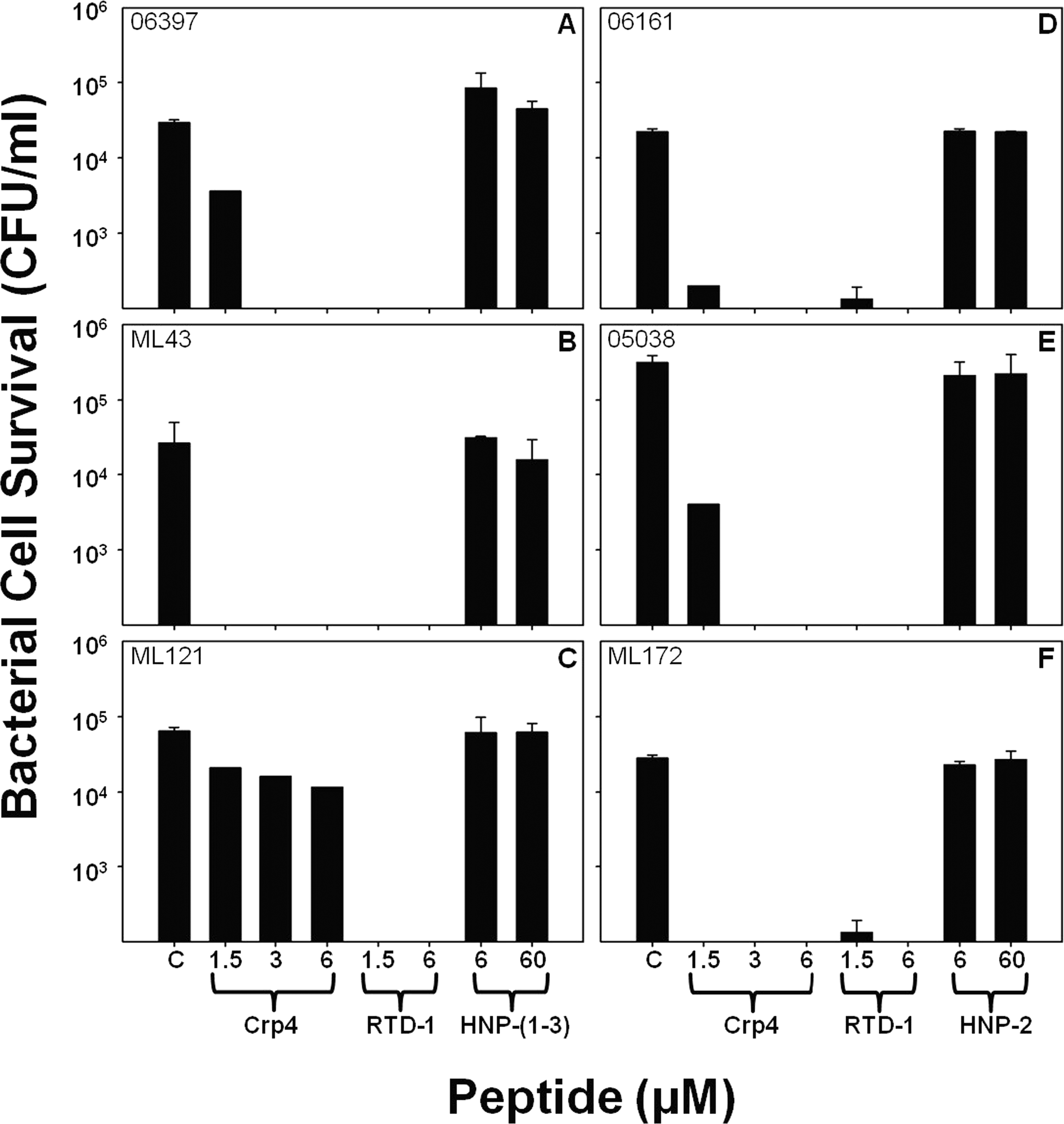
PA sensitivity to α-defensins is independent of the site of isolation
Major sites of PA infections include the urinary tract, lungs of cystic fibrosis patients and burn wounds, and PA strains may differ phenotypically, depending on the site of infection.
55
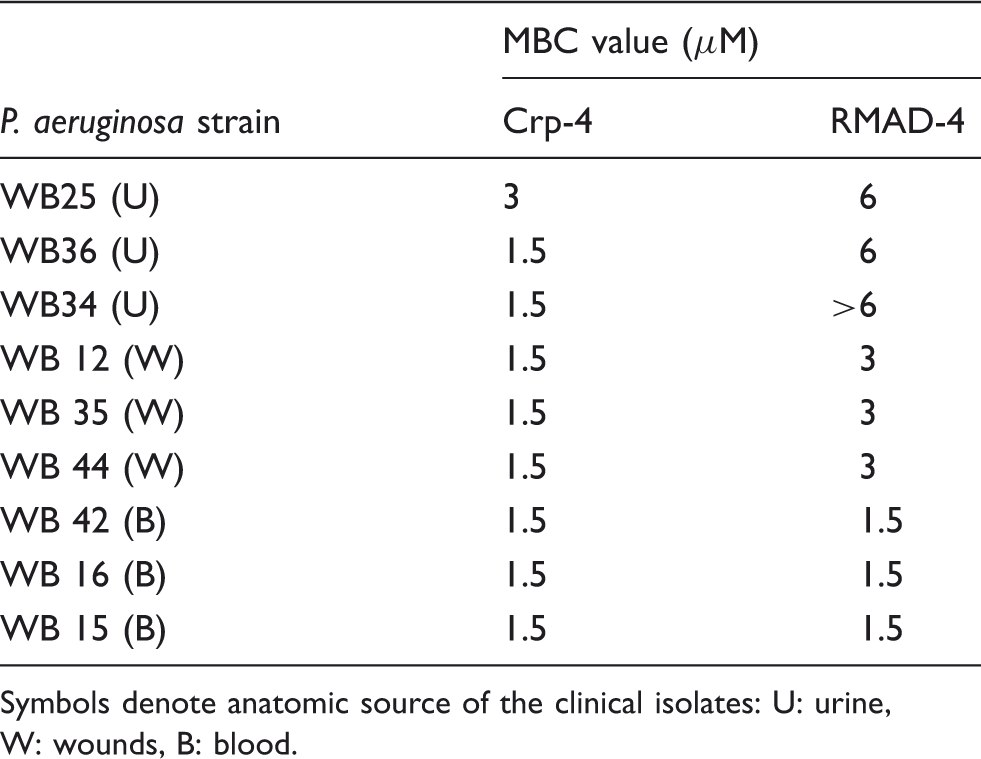
Accordingly, bactericidal peptide assays were performed against PA clinical isolates from patient urine (Figure 6A–C; see Supplementary Figure S2), burn wounds (Figure 6D–F; see Supplementary Figure S3), and blood (Figure 6G–I) to test their sensitivities to Crp-4 and RMAD-4. Crp-4 was highly bactericidal against almost every PA isolate, irrespective of the site of strain isolation. When exposed to ≥ 6 μM Crp4, no PA survivors were detected on plates, that is, ≤ 103 CFU/ml survived peptide exposure (Figure 6). RMAD-4 had similar activities against PA wound isolates, but PA strains from blood and urine were more variable in RMAD-4 susceptibility (Table 3). Therefore, little relation exists between the site of PA strain isolation and resistance to these α-defensins.
α-Defensin sensitivity is independent of the site of PA isolation. P. aeruginosa strains WB 25 (A), WB 36 (B) and WB 34 (C) isolated from the urine of patients, strains WB 12 (D), WB 35 (E) and WB 44 (F) isolated from patient wounds, and WB 42 (G), WB 16 (H) and WB 15 (I) isolated from patient blood were exposed to 1.5–6.0 µM peptides (Table 3). Assays were performed in triplicate; error bars denote SD from the mean. Symbols: Crp-4 (-•-) and RMAD-4 (-○-). α-Defensin MBC values against P. aeruginosa blood, urine, and wound isolates. Symbols denote anatomic source of the clinical isolates: U: urine, W: wounds, B: blood.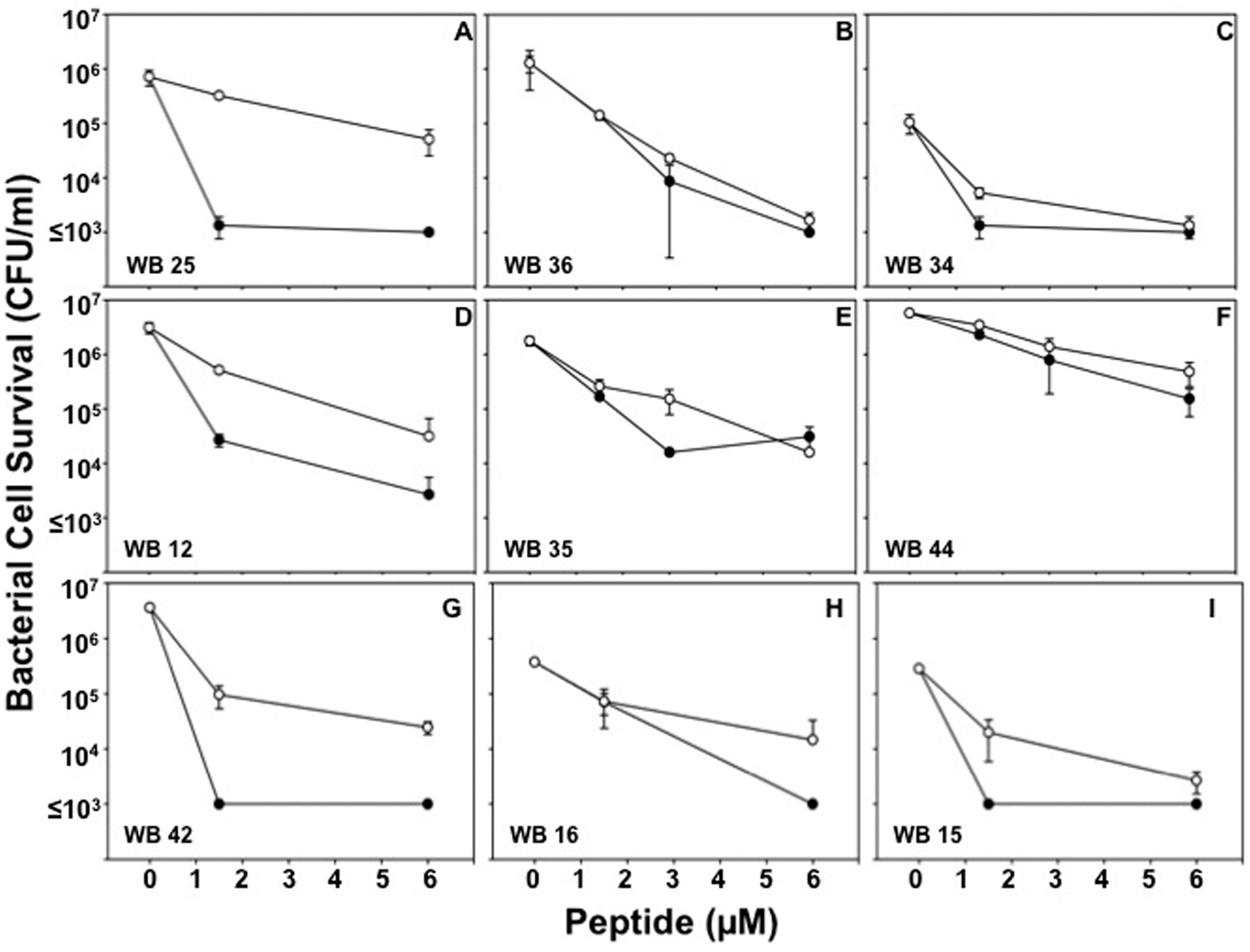
ExoU or ExoS PA genotype and cytotoxicity do not induce α-defensin resistance
Most wild type PA strains harbor the exoS gene encoding the TTSS effector protein, and CipRPA populations are enriched for the exoU gene that codes for a TTSS effector protein, which contributes to virulence in acute infection.49,56 Notably, ExoU expressing PA are more virulent than ExoS strains in a murine model of acute pneumonia, and they cause more severe disease in humans.11,57 To test whether these PA virulence genotypes have differential α-defensin sensitivities, we performed bactericidal assays and determined whether ExoU or ExoS genotype and α-defensin resistance are related significantly by constructing contingency tables and performing a Fisher's exact test. We also grouped the PA strains on the basis of the rate and extent of their cytotoxic effects on exposed A549 lung epithelial cells, and we compared strain α-defensin susceptibility to cytotoxic potential. For this purpose, PA sputum isolates were categorized as α-defensin resistant if < 99% killing occurred when exposed to 6 μM peptide. A subset of ExoU PA strains were resistant to Crp-4 and RMAD-4 by this definition (Figure 7), but no significant association existed between genotype and α-defensin resistance (P > 0.1). Reduced sensitivity to Crp-4 or RMAD-4, therefore, was not determined by TTSS effector genotype. Similarly, Crp4 and RMAD-4 bactericidal activities against sputum isolates were independent of PA cytotoxic potential because highly cytotoxic strains (05038 and ML172) or non-cytotoxic strains (06161 and ML75) were equally peptide sensitive (Figure 7). Collectively, these studies show that the bactericidal activities of Crp-4 and RMAD-4 against PA clinical isolates are independent of CipR status, site of isolation, TTSS effector genotype and cytotoxic potential.
Crp-4 or RMAD-4 bactericidal activity against PA are independent of ExoU or ExoS genotype and cytotoxic potential. PA sputum isolates, characterized as ExoU (black) or ExoS (white) are arranged in order of increasing bacterial survival following exposure to 6 μM of Crp-4 or RMAD-4. Cytotoxic potentials of each strain are also indicated by asterisks. PA strains were considered resistant if > 1% of bacteria survived α-defensin peptide exposure. TTSS effector genotypes and defensin resistance are not associated (P > 0.1), and bactericidal activities were independent of PA cytotoxic potential.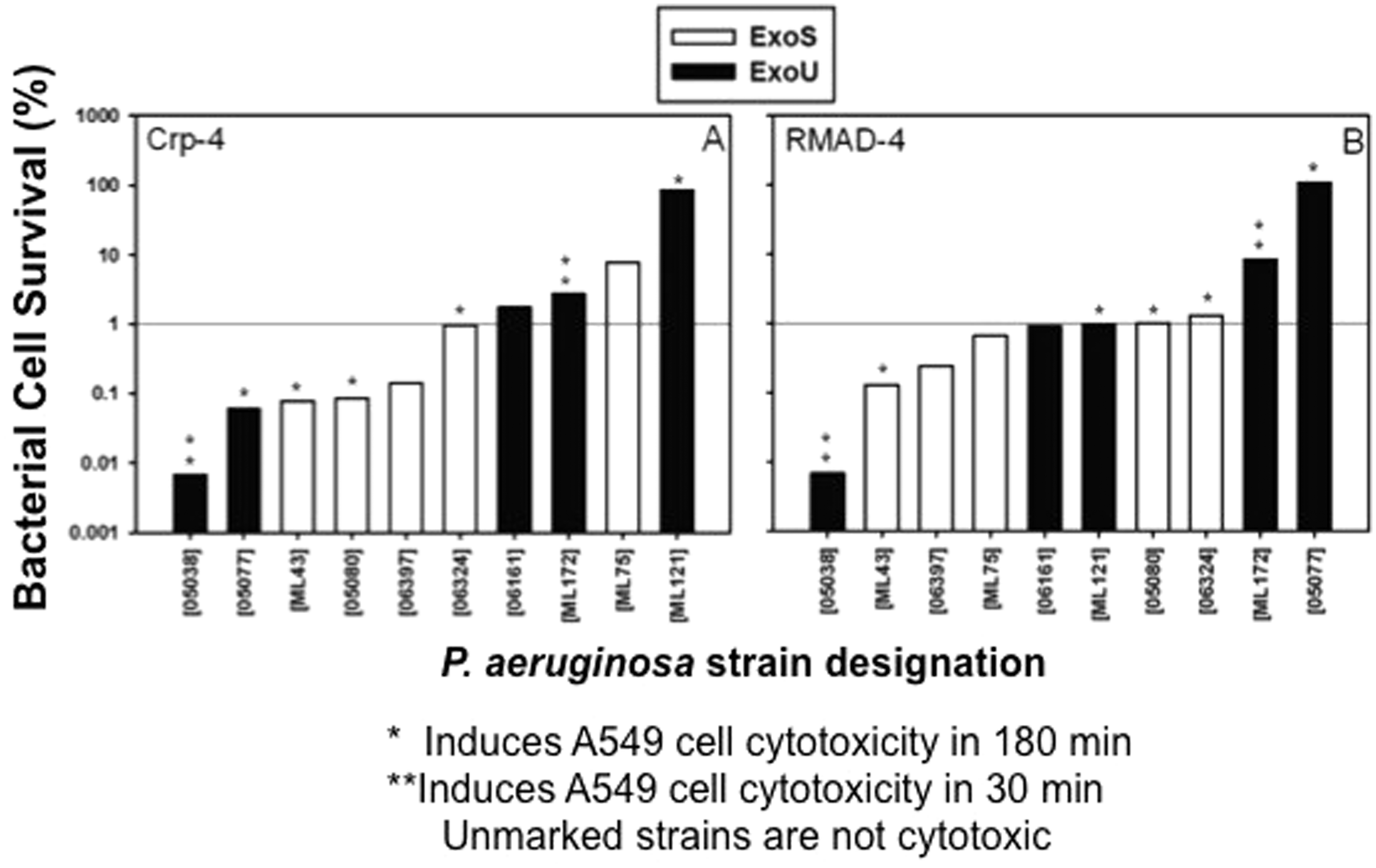
Discussion
Antimicrobial peptides have been proposed as templates for developing new therapeutics to combat multidrug resistant pathogens,58,59 yet the efficacy of α-defensins against antibiotic resistant clinical isolates has not been investigated extensively. Because general mechanisms of α-defensin microbicidal action differ from most antibiotics, they may be useful in combating infections by synergizing in combination with current antibiotic therapies. Because Crp-4, RMAD-4 and RTD-1 resist proteolysis and are non-cytotoxic, potent microbicides in vitro, they may provide new, peptide-based platforms for antibiotic development.39,60 To test the hypothesis that Crp-4, RMAD-4 and RTD-1 may be efficacious against clinically important bacteria that also are antibiotic resistant, we assayed the bactericidal activities of these defensins against isolates of MRSA with varying resistance to vancomycin and CipR-PA. Under the in vitro assay conditions, Crp-4, RMAD-4, and RTD-1 were highly active against MRSA and exhibited differential activities against PA, irrespective of antibiotic resistance.
The continuing emergence of MRSA strains with vancomycin resistance, the treatment standard, underscores the need for new therapeutics. Against all strains of MRSA tested, Crp-4, RMAD-4 and RTD-1 were highly active with MBC values of 1.5–3.0 μM peptide, regardless of vancomycin resistance. In addition, RMAD-4 had a low MBC of 1.5 μM peptide, the lowest concentration assayed, against all but one MRSA strain. The α-defensins investigated have microbicidal mechanisms of action that disrupt cellular integrity. 61 For example, certain defensins kill bacteria by sequestering lipid II and inhibiting bacterial cell wall biosynthesis,37,62,63 HNPs 1–3 mediate non-oxidative microbial cell killing by sequential permeabilization of the outer and inner membranes and formation of stable pores, 64 and Crp-4, RMAD-4 and RTD-1 disrupt membranes by forming transient pores by induction of saddle-splay curvature. 36 At concentrations approximately 10-fold greater than the MBC values reported here, Crp4 induces rapid efflux of potassium ions from bacterial cells, a sensitive indicator of membrane disruption and cell death.34,65–68 We do not discount inhibition of cell wall biosynthesis as contributing to the defensin mechanism of action in certain settings. However, over the 1-h course of peptide exposure in the experiments we have presented, membrane disruption is the most likely mechanism of action, 36 a mechanism that differs from beta lactam inhibitors and vancomycin. Thus, in combination with antibiotics, these defensin peptides could synergize with current antibiotics to improve treatment of MRSA-related infections.
Gram-negative bacteria account for more than 30% of US hospital infections and are the predominant ICU infections.69–72 For example, PA is a leading cause of pneumonia, urinary tract and bloodstream infections, 70 and the incidence of multidrug resistant PA is rising. 73 Crp-4, RMAD-4, RTD-1 and HNPs 1–3 were tested against PA strains characterized by their ciprofloxacin resistance, site of isolation, TTSS effector genotype and cytotoxic potential to determine whether these traits correlated with defensin resistance. Under the conditions of the in vitro assays, PA strains exhibited variable sensitivity to the defensin peptides tested (Tables 2 and 3), but defensin sensitivity was not associated with either of the phenotypic characteristics. Cationic charge contributes to defensin bactericidal activity, but even though Crp-4 and RMAD-4 are equally electropositive, Crp-4 was more active against PA strains, but RMAD-4 was more potent against MRSA (Tables 1–3). Perhaps, the differential bactericidal effects are owing to differences in the distribution of surface charge or hydrophobicity in the two α-defensin peptides. 50 Although PA isolates from lung, blood and urine differ phenotypically, 55 the strains tested were equally sensitive to Crp-4 and RMAD-4, irrespective of their site of isolation.
The α- and θ-defensin peptides tested hold promise toward the eventual development of new therapeutic agents against the growing number of antibiotic resistant pathogens. Although Crp-4, RMAD-4 and RTD-1 are highly active against many of the PA strains tested in vitro, activity against PA strains may be diminished in the setting of chronic lung infections where biofilm formation is induced74,75 in that biofilm production enhances resistance to bactericidal peptides. Whether biofilms diminish the activities of peptides tested here is unknown. Also, the ionic strength of assay media inhibits bactericidal activities of Crp4 and RMAD-4, and binding of Crp-4, RMAD-4 and RTD-1 peptides to plasma proteins has not been studied and may limit efficacy in vivo. Against bacteria that are sensitive to these peptides, for example MBC ≤ 1.5 µM, bactericidal effects of Crp-4 and RMAD-4 are partially inhibited by 50 mM NaCl and completely inhibited at 100 mM NaCl. Thus, the inherent salt sensitivity of these native molecules may limit therapeutic application of native, full-length α-defensins, but reiterative selection of salt-insensitive variants of these peptides could lead to drug development based on these peptide scaffolds. In contrast to the α-defensins, however, cyclized RTD-1 is as active in 150 mM NaCl as in low salt media,28,47,51 an indication that the θ-defensins may provide a more robust platform for therapeutic development. Also, the disulfide array protects Crp-4 and RMAD-4 from proteolytic degradation, and α-defensins can be recovered from the protease-rich environment of mouse small and large bowel.60,76 Biophysical studies of Crp-4, RMAD-4 and RTD-1 have established amino acid criteria necessary for their selective membrane disruptive activities, 36 and Crp-4 and RMAD-4 are non-cytotoxic and non-hemolytic at peptide concentrations as high as 100 μg/ml (data not shown). 51
In addition to their broad spectrum in vitro microbicidal activities, rhesus θ-defensins 1–5 (RTDs 1–5), RTD-1 in particular, reduced the levels of TNF-α, IL-1α, IL-1β, IL-6 and IL-8 released by mixed blood leukocytes and by THP-1 monocytes stimulated by bacteria or LPS, respectively. 77 Also, systemic administration of RTD-1 to BALB/c mice was non-toxic and stable in serum and plasma, and intravenous delivery of 5 mg/kg RTD-1 significantly improved survival of BALB/c mice with E. coli peritonitis and cecal ligation-and-puncture-induced polymicrobial sepsis. 77 In contrast to HNPs, which are pro-inflammatory,78–82 Crp4 and RMAD4 also inhibit cytokine and TNF-α release in the same in vitro assays, although both are less inhibitory than RTD-1 (Kamdar et al., submitted). Thus, new antimicrobials that are developed to optimize both defensin-based bactericidal and immunomodulatory properties may prove to be efficacious in combination with current antibiotic therapies against antibiotic resistant bacteria, while promoting survival from systemic infections.
Footnotes
Funding
This work was supported by the National Institutes of Health grants DK044632, AI059346 (A. J. O), AI022931, AI058129, DE021341,Southern California Clinical Translational Science Institute UL1RR031986 (M. E. S), National Institute of Allergy and Infectious Diseases grant AI073467 (A. W.-B.). Also supported, in part, by the USC Norris Cancer Center Support Grant P30CA014089 from the National Cancer Institute; the content is solely the responsibility of the authors and does not necessarily represent the official views of the National Cancer Institute or the National Institutes of Health.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
