Abstract
We observed persistent peritoneal bacteria despite a transient early innate immune response to intraperitoneal (IP) Klebsiella pneumoniae. Pretreatment with LPS prior to peritonitis induced a tolerant pattern of pro-inflammatory cytokine protein production over 72 h, but not at the mRNA level. MicroRNAs (miRNAs) regulate inflammatory cytokines and may explain this paradox. After pretreatment with IP LPS or saline, C57BL/6 mice were given 103 CFU of K. pneumoniae IP. Total RNA was isolated from peritoneal exudate cells (4 h, 24 h and 48 h following infection). mRNA and miRNA expression levels were detected and bioinformatics pathway analysis was performed, followed by measuring TNF-α, IL-1β, IL-6 and High-mobility Group Box 1 (HMBG1) protein levels. Of 88 miRNAs studied, 30 were significantly dysregulated at all time points in the LPS-pretreated group, including MiR-155, -146a, -142-3p, -299, and -200c -132 and -21. TNF-α, regulated by miR-155 and miR-146a, was decreased in the LPS-pretreated group at all time points (P < 0.05), as were HMGB1, a key alarmin regulated by miR-146, -142-3p, -299 and -200c (P < 0.05), and IL-1β and IL-6, both regulated by miR-132and miR-21 respectively (P < 0.05). Specific dysregulation of miR-155, -146a, -142-3p, -299, and -200c -132 and -21 with their corresponding effects on the TLR and NF-κB signaling pathways during inflammation, suggests a plausible mechanism for tolerance in this novel chronic model with persistent peritoneal infection.
Introduction
Secondary bacterial peritonitis is an invasion of the normally sterile peritoneal cavity with microorganisms after mechanical breach of the gastrointestinal tract, which is typically caused by diverticulitis, appendicitis, bowel infarction or trauma. 1 Despite current treatment strategies involving antibiotic therapy combined with intensive care support and surgical intervention, patients with peritonitis are at high risk of developing sepsis, which can, ultimately, lead to organ damage and eventual mortality. 2 In response to severe infections, including peritonitis, there may be significant alterations in the immune response of circulating leukocytes, described as immune paralysis. 3
Interestingly, endotoxin tolerance mirrors this clinical condition of immune paralysis. First described in the late 1940s by Beeson, the subsequent administration of typhoid vaccine produced less of a fever response than the initial administration. 4 In another study, Greisman et al. injected bacterial endotoxin intravenously into healthy patients and reported diminished fever responses upon continuous or repeated endotoxin administration; this response was coined endotoxin tolerance. 5 Endotoxin tolerance is defined as a pre-exposure to bacterial endotoxin or LPS that leads to a diminished pro-inflammatory response upon subsequent stimulation. 6 Endotoxin tolerance can be divided into two stages. The first stage is an Ab-independent phase that occurs hours following LPS insult. This stage may last for several days, and is considered to be of great importance in septic patients who have impaired host defense against virulent and persistent pathogens. The second stage is Ab-dependent and appears weeks after the initial LPS administration. 1
While the exact mechanism of LPS tolerance is unclear, it is clear that the innate immune system recognizes pathogen associated molecular patterns (PAMPs) through pattern recognition receptors (PRRs), including the TLRs, which are members of the IL-1 receptor superfamily. 1 LPS is recognized by TLR4, whereas other bacterial components are sensed by different TLRs. The interaction between TLRs and their ligands induces tolerance through transduction pathways controlling the expression of immune-response genes. 7 Multiple studies have shown that endotoxin tolerance is associated with impaired activation of the NF-κB and MAPK pathways. 8 This also includes a decrease in cytokines released by activated macrophages, including TNF-α, IL-10, IL-6, IL-1 and IL-12. 1 In addition, several reports have shown that diminished secretion of these cytokines is beneficial and decreases mortality in cases of LPS-induced endotoxic shock, acute infection and ischemia/reperfusion; however, this effect may be detrimental for bacterial clearance. 9
We have previously reported microbial tolerance at the cytokine protein and mRNA levels in a novel Klebsiella model of persistent secondary murine peritonitis. In this model, mice were injected intraperitoneally (IP) with 103 CFU Klebsiella pneumoniae, a Gram-negative, anaerobic organism in secondary peritonitis. 1 This induced death in all animals within 3 d of infection. However, if animals received antibiotic treatment with gentamicin (5 mg/kg BID) subcutaneously (SC), survival was significantly enhanced and some mice survived the 14-d observation period. 10 The use of antibiotics protected animals from severe life-limiting bacterial overgrowth, but bacteria were never fully eliminated from the peritoneal cavity. Within 48 h of injection, peritoneal TNF-α and IL-6 concentrations diminished significantly despite peritoneal bacteria concentrations exceeding 106 CFU. 10 The relatively steady concentrations of peritoneal bacteria in our model induced microbial tolerance at the macrophage level, which likely accounted for diminished peritoneal cytokine concentrations and late mortality in vivo. 1 However, we showed previously that microbial tolerance could not be attributed to dysregulation at the mRNA level in this model. 11 This is in line with previous studies, which found that mRNA expression correlates with sepsis and an increased pro-inflammatory response, but is less indicative of tolerance. As the regulation of endotoxin tolerance involves receptors, negative modulators and signaling molecules, it is speculated that gene reprogramming and immune modulation, rather than a global down-regulation of gene expression, could explain such phenomena. Post-transcriptional modifications of the expressed cytokines could possibly fit into a mechanistic framework that could explain the molecular basis of microbial tolerance.
Recent studies have shown that microRNAs (miRNAs), an evolutionarily conserved class of endogenous 22-nucleotide, non-protein-coding RNAs, contribute to the regulation of inflammation by repressing gene expression at the post-transcriptional level. 12 miRNAs are inhibitory and repress the translation of a complementary mRNA leading, ultimately, to a decrease in protein levels associated with the bound mRNA. 13 Wightman et al. demonstrated the importance of the complementary sites for regulation of lin-14 mRNA by miR-lin-4, also showing that this regulation reduces the amount of LIN-14 protein levels substantially without noticeable change in the levels of lin-14 mRNA. 14
Based on our previous findings at the cytokine–protein and mRNA level, we sought to explain the mechanism behind microbial tolerance at the miRNA level using our previously-described model. We hypothesized that microbial tolerance was the result of dysregulation in the production of specific miRNAs related to inflammation leading to inhibition of cytokine production and, thereby, could account for the diminished immune response, bacterial persistence and, thus, late mortality.
Therefore, the aim of this study was to (i) investigate the differential expression of miRNAs by peritoneal exudate cells (PECs) during microbial tolerance and (ii) correlate the miRNA expression with the target mRNAs and cytokine levels indicative of microbial tolerance.
Materials and methods
Animals
Male 6–8-wk-old C57BL/6 animals were obtained from Jackson Laboratory (Bar Harbor, ME, USA). Animals were housed in a facility approved by the American Association for Accreditation of Laboratory Animal Care and were provided with food and water ad libitum. Studies were carried out according to the National Institutes of Health guidelines and under the supervision of a veterinarian. Mice were allowed to acclimate for at least 1 wk prior to experimentation.
Experimental protocol
Experimental mice were pretreated with saline (saline group) or LPS (LPS group) 24 h prior to inoculation with K. pneumonia. Then, these mice received gentamicin BID until harvest at 4 h, 24 h and 48 h. Control mice were harvested without receiving pretreatment, K. pneumonia inoculation or antibiotics (Figure 1).
Experimental protocol for chronic peritonitis model.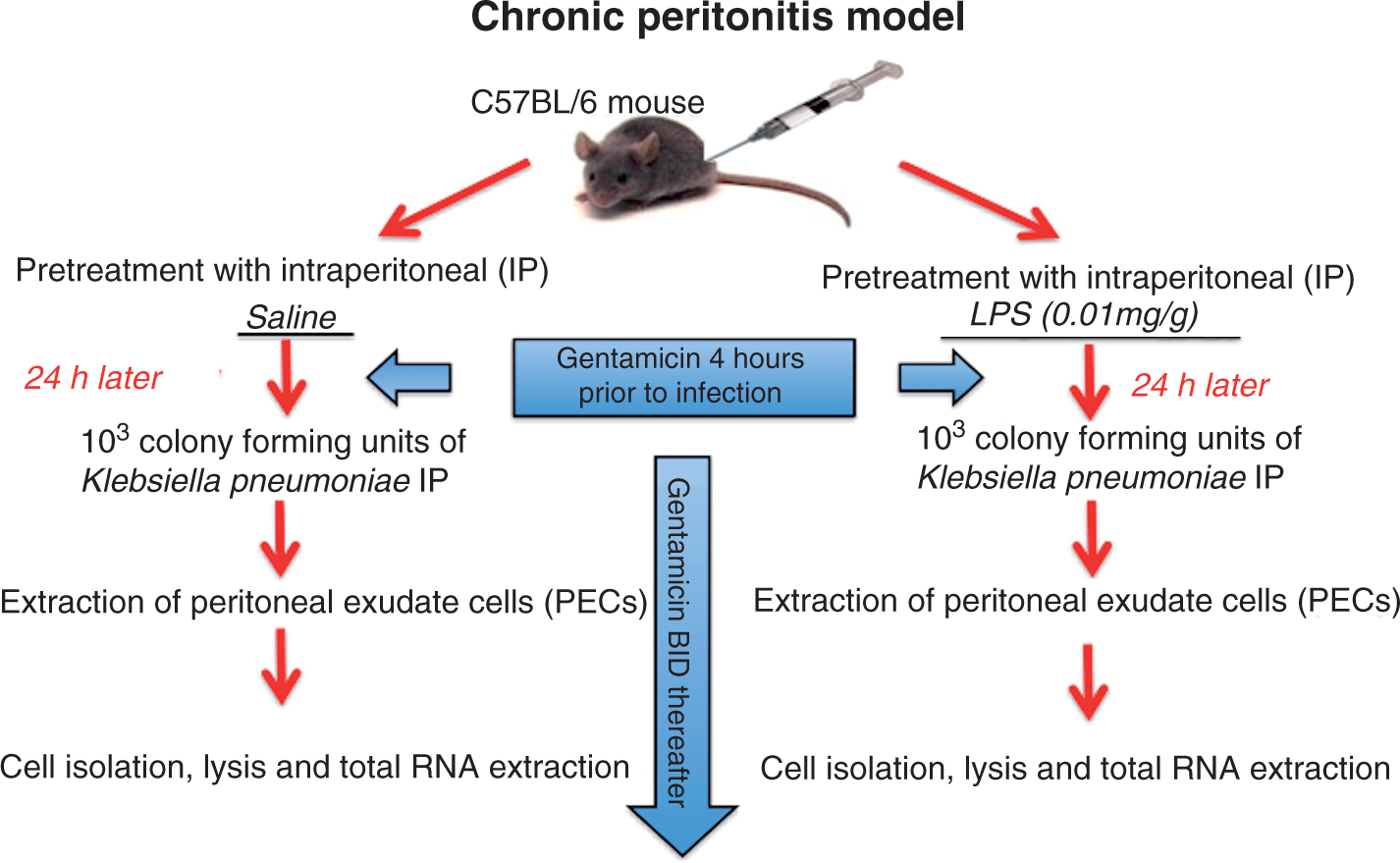
Pretreatment
Animals were pretreated 24 h prior to inoculation with IP 0.2 ml saline or 10 mg/kg LPS (Ultra Pure Escherichia coli LPS, 0111:B4 strain; InvivoGen, San Diego, CA, USA), in a final volume of 0.2 ml saline. Uninfected, non-pretreated animals served as controls. Twenty-four hours later, pretreated animals were inoculated with K. pneumonia.
K. pneumoniae inoculation
Mid-log growth phase of K. pneumoniae strain 43816 serotype 2 (American Type Culture Collection, Rockville, MD, USA) were obtained by growth in nutrient broth (Difco, Detroit, MI, USA) for 18 h at 37°C, followed by inoculation into Lennox L broth (Difco) for an additional 4 h growth. Bacterial concentration of the broth was quantified by measuring the absorbance at 655 nm and comparing it with a previously determined standard curve of concentrations. Bacteria were then diluted with endotoxin-free sterile saline to provide bacterial numbers of 103 CFU in a final volume of 0.2 ml. Animals were injected with a final volume of 0.2 ml IP. Injected bacterial numbers were confirmed by colony counts of nutrient agar plate dilutions after a 24-h incubation period at 37°C.
Antibiotic treatment
Animals received 5 mg/kg/d (BID) subcutaneous gentamicin (Sigma-Aldrich, St Louis, MO, USA) in 0.2 ml of sterile saline. The first dose was given 4 h prior to IP bacterial administration and antibiotic treatment was continued BID throughout the observation period.
PEC collection
PECs were harvested at 4 h, 24 h and 48 h post-IP inoculation. PECs were recovered by peritoneal lavage with 4 ml of ice-cold heparinized RPMI 1640 medium (GIBCO/BRL, Bethesda, MD, USA). PECs were quantified manually using a hemocytometer with trypan blue exclusion to determine total PEC number and cell viability. PEC lavage media was diluted and plated for peritoneal bacteria determination. PEC samples were then centrifuged to pellet cells for RNA extraction and to collect supernatants. Supernatants were aliquoted and frozen at −80°C for cytokine analysis. The cell pellets were re-suspended in recommended volumes of RNAlater Stabilization Buffer and frozen until RNA extraction. Resulting peritoneal CFU concentrations were multiplied by four to account for earlier dilution by the peritoneal lavage fluid and were expressed as log CFU/mouse. Limit of detection was peritoneal cavity 1.4 log CFU/mouse.
Cell lysis, RNA extraction, mRNA and miRNA plate runs
Stabilized PEC cells were thawed and pelleted, and white cells were re-suspended in an appropriate lysis buffer. Total RNA extraction was performed using the Qiagen Kit Animal Spin protocol (Qiagen, Valencia, CA, USA). Total RNA was quantified using the Nanodrop N-1000 by Agilent Biosystems (Santa Clara, CA, USA). RNA quality control was maintained through measurements of RNA purity by UV spectrophotometry, and assessment of RNA quality and integrity. The first-strand mRNA-cDNA and miRNA-cDNA syntheses were performed on an Eppendorf Mastercycler with the appropriate cDNA Synthesis kit (SuperArray Bioscience, Frederick, MD, USA) and 100 ng of total RNA. cDNA was processed according to the manufacturer’s protocol for mRNA and miRNA plates. The cDNA template was combined with RT2 Real-Time™ SYBR Green/Rox Master Mix and RNAse-free water (SuperArray Bioscience). A final reaction volume of 25 μl was added to each well of the mRNA TLR signaling Pathway PCR Array (SuperArray Bioscience) and the immune-pathology RT2 miRNA PCR Array (MAM-104 A) (SABiosciences). Finally, pathway-focused mRNA and miRNAplates were amplified on a 7300 Fast Real-Time PCR system (Applied Biosystems, Foster City, CA, USA), following the manufacturer’s protocol. 15 The mRNA plates tested for 88 different mRNAs that have been shown previously to be involved in the TLR signaling pathways (supplementary Table S1). 15 The miRNA plates tested for 88 different miRNAs involved in the immune pathology process in mice (supplementary Table S2). 15 Housekeeping genes, as well as reverse transcription and positive controls, were included in both the mRNA and miRNA formats.
RT-PCR data analysis
Alterations in mRNA and miRNA transcript levels at 4 h, 24 h and 48 h in both the saline and LPS groups were analyzed initially using SABiosciences webportal software. Fold changes were calculated and P-values were calculated using student’s t-test for significant mRNAs. A P-value < 0.05 was considered significant.
Cytokine assays
TNF-α, IL-1β, IL-6 and HMGB1 levels were measured in peritoneal lavage fluid supernatants using ELISA. TNF-α, IL-1β and IL-6 ELISA murine Ready-Set-Go kits were purchased from e-Biosciences (San Diego, CA, USA) and a HMGB1 ELISA kit was purchased from IBL International Corp (Toronto, ON, Canada); these were then performed according to the manufacturer’s instructions. Where necessary, samples were diluted in assay diluent to achieve cytokine concentrations within the range of the standard curve. Cytokine concentrations were multiplied by four to account for dilution by lavage media and expressed in pg/mouse. Sensitivities for TNF-α, IL-1β, IL-6 and HMGB1 were 8 pg/ml, 8 pg/ml, 4 pg/ml and 0.2 ng/ml respectively.
Pathway analysis
In order to understand the genetic significance of mRNA and miRNA expression, fold changes and dysregulation, pathway analysis was done using Ingenuity Pathway Analysis (IPA 9.0; Redwood City, CA, USA). IPA is a software application used to identify biological mechanisms, dysregulation in signaling or metabolic pathways, and functions most relevant to their experimental data sets or genes of interest. 16 Official permission to use figures and analyses performed by this software was obtained. All RNA results were normalized against uninfected controls.
Results
Microbial tolerance in peritoneal macrophages
We first established that our selected pretreatments actually resulted in the induction of microbial tolerance. The animals appeared healthy at all time points—moving spontaneously around the cage, self-grooming, and had healthy coats and eyes. We have previously noted that they do experience a small percentage of mass loss post-Klebsiella inoculation which is regained after 48 h.
Following LPS pretreatment, peritoneal bacterial levels decreased significantly between 4 h and 24 h (P < 0.05). At 48 h, bacterial numbers stabilized at 102–103 CFU (Figure 2). Despite 102–10 Peritoneal bacterial levels in the saline and LPS pretreated groups at 4 h, 24 h and 48 h.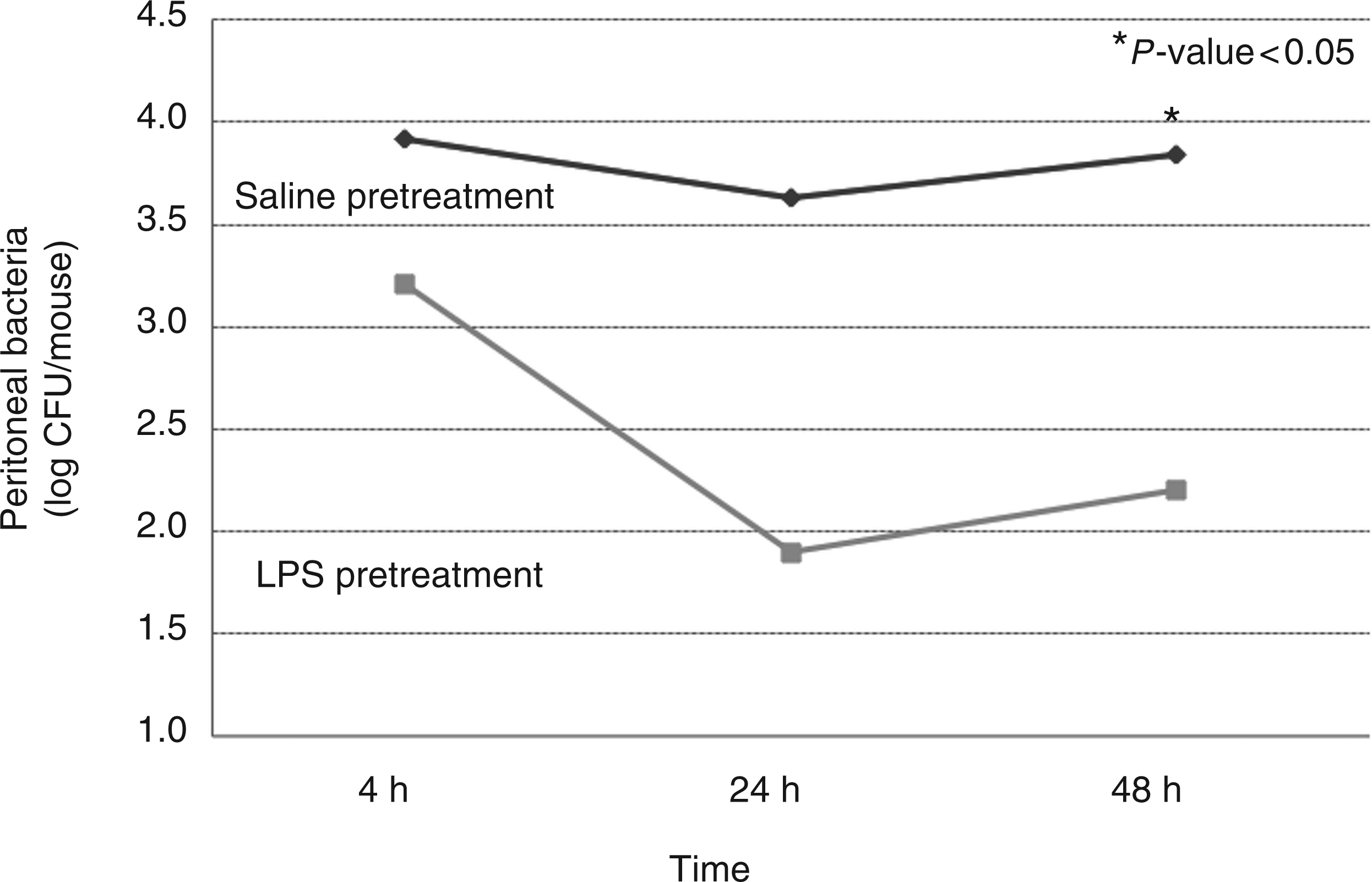
Significantly dysregulated miRNAs in the LPS-pretreated group compared with the saline-pretreated group
We first identified significantly dysregulated miRNAs in the LPS and saline pretreated groups compared with uninfected controls. Then, we identified the significantly dysregulated miRNAs in the LPS pretreated group compared with the saline pretreated group. At 4 h, 18 miRNAs were significantly upregulated, namely: miR-155, miR-383, miR-19, miR-493, miR-147, miR-154, miR-370, miR-140, miR-142-3p, miR-205, miR-130, miR-132, miR-574, let-7f, miR-17, miR-146a, miR-200c and miR-150. At 24 h, 24 miRNAs were significantly upregulated, namely: miR-155, miR-383, miR-19, miR-493, miR-147, miR-154, miR-370, miR-140, miR-142, miR-205, miR-130, miR-132, miR-574, let-7f, miR-17, miR-146a, miR-150, miR-145, miR-363, miR-335, miR-17, miR-8, miR-9, and miR-299. At 48 h, 12 miRNAs were significantly upregulated, including miR-155, miR-146a, miR-574, miR-8, miR-9, miR-299, miR-21, miR-125, miR-183, miR-325, miR-27 and miR-154.
Pathway analysis of miRNA and mRNA data
According to IPA analysis, and based on our miRNA expression data, miR-155, miR-146a, miR-142-3p, miR-299, miR-200c, miR-132 and miR-21 were predicted to have the most significant direct effects on the TLR and NF-κB signaling pathways, and were thus evaluated along with their protein targets. After applying IPA analysis to our mRNA data, 6 main canonical pathways were constantly dysregulated and, in the same order of significance, in both the saline and LPS groups at 4 h, 24 h and 48 h (supplementary Figure S1). These were TLR and NF-κB signaling, hepatic cholestasis, IL-6- and LPS-mediated MAPK signaling pathways, and PRR of the bacterial pathway.
miR-155 and miR-146a dysregulation, and TNF-α mRNA and protein levels
There was no difference in TNF-α mRNA expression between the saline and LPS pretreated groups at all time points despite differences in TNF-α protein thus indicating a potential role for miRNAs as post-transcriptional fine tuners. TNF-α protein levels were significantly decreased at all time points in the LPS pretreated group (Figure 3). MiR-155, shown to target SHIP1/SOCS1, both inactivators of the TLR pathway, was significantly down-regulated at 24 h and 48 h in the LPS pretreated group, possibly explaining the decreased TNF-α levels at the same time points (Figure 3). MiR-146a, which targets key molecules of the TLR/IL-1 receptor pathway, such as IRAK 1/2 and TNF receptor–associated factor (TRAF) 6, and thereby inhibiting the inflammatory response, was significantly up-regulated at 24 h and 48 h. The inhibitory effect of miR-146a on TNF-α production would explain the decreased TNF-α levels (Figure 3).
miR-155 and miR-146a were up-regulated in the LPS group at all time points (P < 0.05). TNF-α, a common target of the aforementioned microRNAs (miRNAs), was, correspondingly, significantly downregulated at the protein level in the LPS group at 4 h, 24 h and 48 h as compared with the saline pretreated group.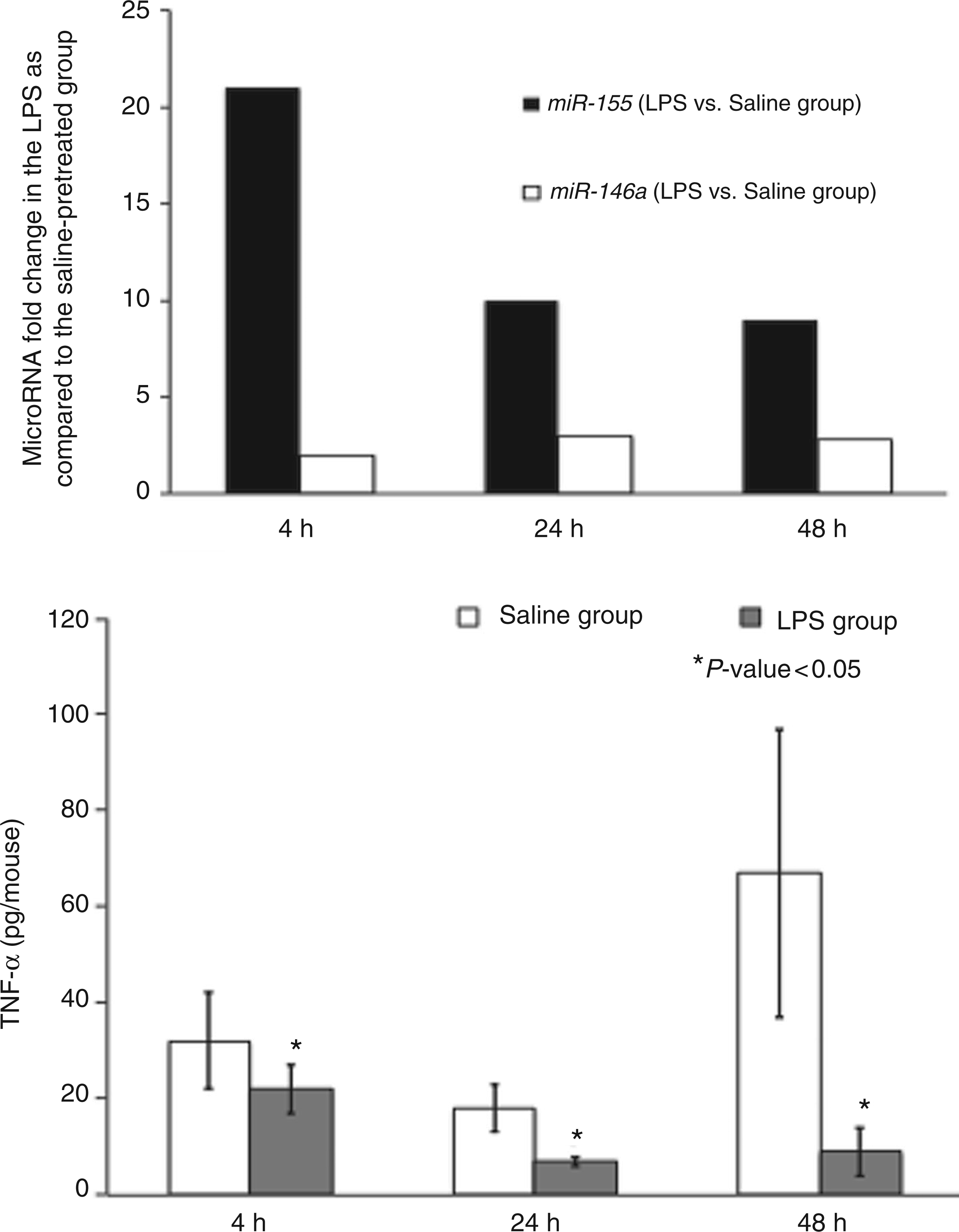
miR-146, -142-3p, -299 and -200c, and HMGB1 mRNA and protein levels
We found miR-146, -142-3p, -299 and -200c to be significantly up-regulated in the LPS pretreated group. The protein levels of HMGB1, a key target of the aforementioned miRNAs and involved in inflammation after monocyte stimulation,
17
was decreased at 24 h and 48 h in the LPS-pretreated group in our study (Figure 4).
miR-142-3p, -146a, -299 and -200c were up-regulated in the LPS group at all time points (P < 0.05). HMGB1, a common target for all these microRNAs (miRNAs), was, correspondingly, significantly down-regulated at the protein level in the LPS group at 24 h and 48 h in the LPS pretreated group compared with the saline pretreated group.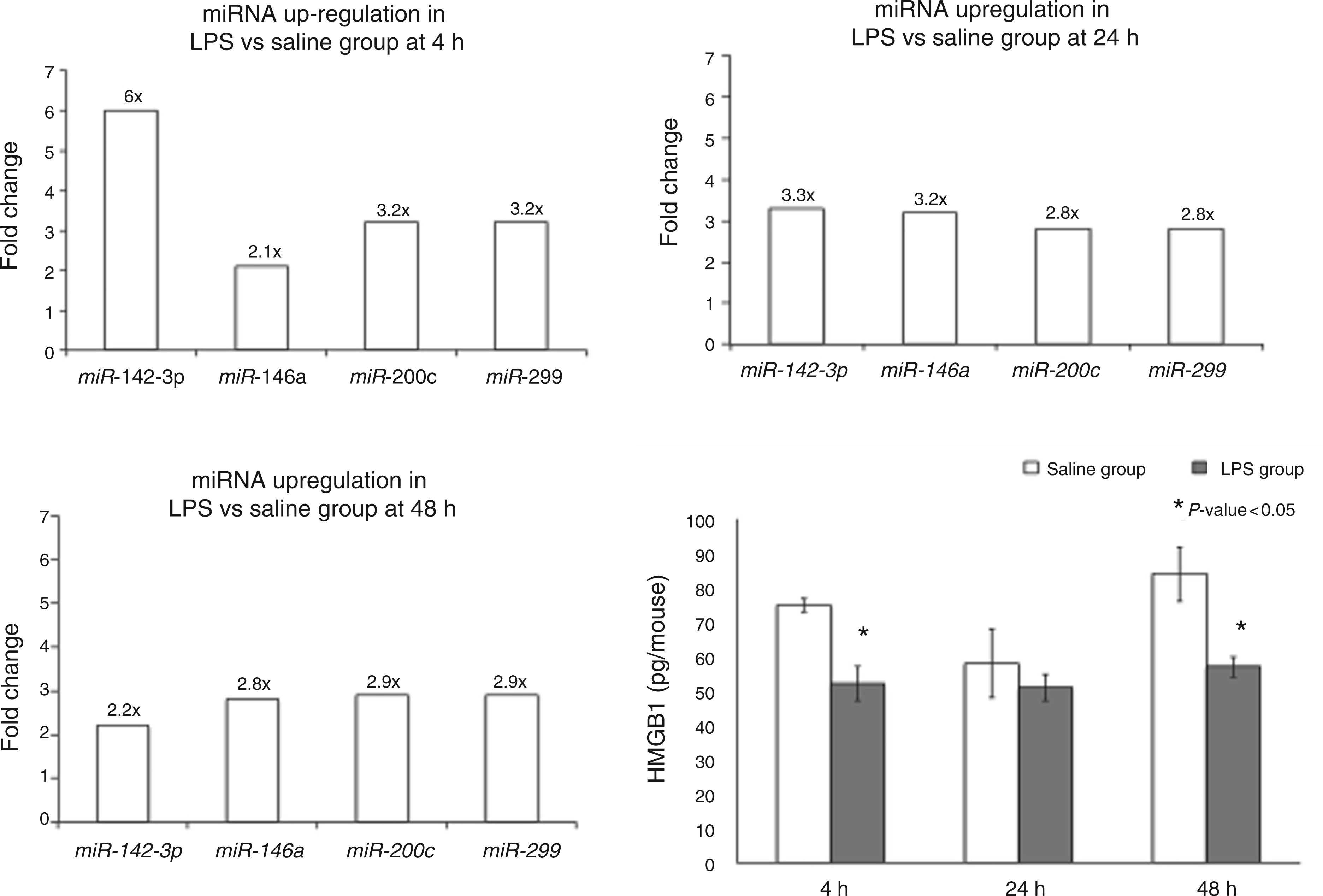
MiR-132, miR-21, and IL-1β and IL-6 mRNA and protein levels
miR-132 and miR-21, predicted to target IL-1β and IL-6, respectively, were both up-regulated significantly in the LPS group. Compared with the saline pretreated group, IL-1β levels decreased at 48 h (P < 0.05), whilst IL-6 levels decreased at 24 h and 48 h (P < 0.05) (Figure 5). miR-142-3p also targets IL-6 and the up-regulation of miR-142-3p corresponds to the downregulation of IL-6 in LPS pretreatment.
miR-132 was up-regulated in the LPS group (P < 0.05). IL-1β, a target of the aforementioned miRNA, was, correspondingly, significantly downregulated at the protein level in the LPS group at 48 h compared with the saline pretreated group. miR-21 was also up-regulated in the LPS group (P < 0.05). IL-6, a target of the aforementioned microRNA (miRNA), was, correspondingly, significantly down-regulated at the protein level in the LPS group at 24 h and 48 h compared with the saline pretreated group.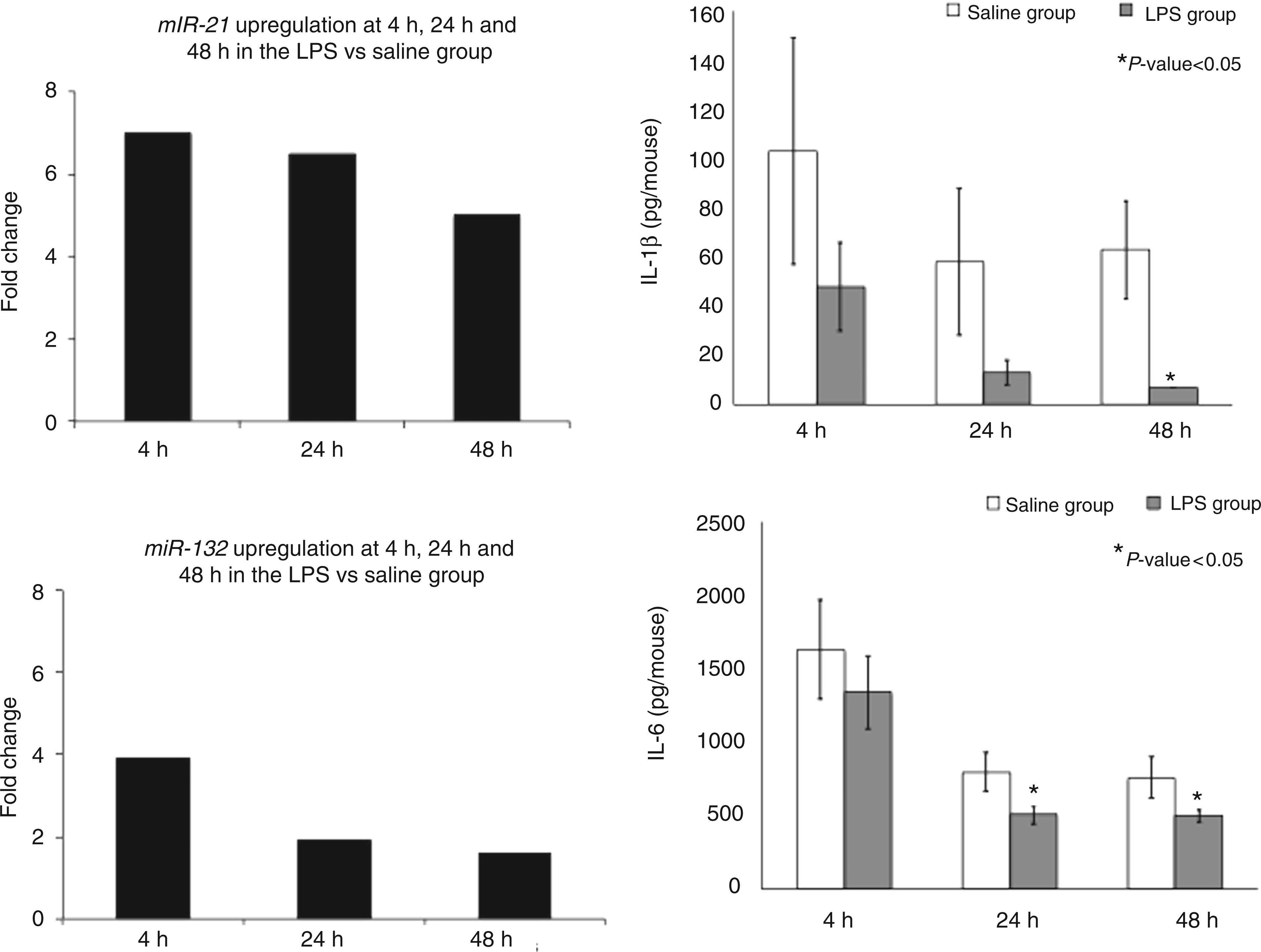
Discussion
miRNAs and microbial tolerance
miRNAs are non-protein coding RNAs that downregulate gene expression via their effect on mRNA. They play a crucial role in regulating the expression of proteins through specific interaction with complementary mRNA 18 and are now widely accepted as major regulatory gatekeepers of coding genes within the human genome. A single miRNA can bind to, and regulate, many different mRNA targets and, conversely, several different miRNAs can bind to and cooperatively control a single mRNA. The regulation of gene products by miRNA is therefore complex, with diverse roles affecting many aspects of transcription and translation that can play a role in overall cellular homeostasis.
Whilst the role of miRNAs in cell fate decisions linked to proliferation, differentiation and apoptosis has been studied extensively and recognized, the significance of these non-coding small RNAs in immune system development and response has only recently been investigated in detail. 19 Several miRNAs have been attributed significant roles in the development of the immune system and during the formation of an immune response. For example, the activation of the innate immune response has been associated with changes in the expression of selected miRNAs (namely miR-155, 20 miR-146a, 21 miR-142-3p, 22 miR-132 23 and miR-21 2 4 ). However, the role of most of these miRNAs in the development of an adequate immune response remains insufficiently elucidated. Herein, we report the profiles of miRNAs induced by inflammatory stimuli in PECs after LPS stimulation and during the development of microbial tolerance. We identify a significant synchronous correlation between miR-155, miR-146a, miR-125b, miR-142-3p, miR-299, miR-200c, miR-132 and miR-21 levels, and the development of a tolerant immune response through inhibition of specific targets in key pro-inflammatory pathways. These inhibit the inflammatory response and may lead, eventually, to the development of microbial tolerance.
The 48-h time point corresponds to the peak bacterial concentration observed in the animals, which would be similar to the clinical condition for generalized secondary bacterial peritonitis. Bacterial concentrations remain similar after this time point in all animals treated with antibiotics over at least 2 wk, as noted in previously-published experiments. 25
miR-155 and miR-146a dysregulation correlates with TNF-α protein levels
O’Connell et al. initially demonstrated that LPS treatment in human monocytic cells augments the expression of two microRNAs, miR-146a and miR-155. 20 miR-155 is up-regulated early in endotoxin shock after TLR4 activation and is essential for a proper immune response. 20 Once up-regulated, miR-155 targets negative regulator proteins of the inflammatory response, such as SOCS1 and SHIP1, which act on the signaling transducers of the TLR family and enhance the pro-inflammatory signals. 26 Later, miR-155 attenuates TLR4 signaling by targeting IKKe or MyD88, an essential signaling protein of all TLRs except TLR3, 27 directly; however, it was shown to increase TNF-α production. 28 Costinean et al. further elucidated that miR-155 knock-in mice are highly susceptible to LPS shock owing to high levels of TNF-α. 29 The anti-inflammatory cytokine IL-10 downregulates miR-155, supporting the primary pro-inflammatory role of miR-155. 30
In the LPS pretreated group in our study, the downregulation of miR-155 correlated with a significant TNF-α downregulation at 24 h and 48 h, whilst both were up-regulated in the saline pretreated group at the same time points. The downregulation of miR-155 and TNF-α could contribute to tolerance in the LPS-pretreated group, despite persistent high peritoneal bacterial levels.
miR-146a is a well-described miRNA in the delayed anti-inflammatory phase in an inflammatory response.
31
miR-146 is first
miR-142-3p, -146a, -299 and -200c dysregulation correlates with HMGB1 protein levels
Alarmins have been described to serve as danger signals that promote activation of the innate immune system pathways and have been shown to participate in the activation of several immune receptors, including the TLRs. 33 HMGB1, an alarmin itself, is usually actively released by immune cells, such as monocytes, macrophages and dendritic cells. 34 The stimuli for secretion of HMGB1 from immune cells are diverse and include PAMPs, cytokines and certain states of cellular stress. 35 Normally, LPS-activated macrophages and monocytes secrete HMGB1 as a cytokine mediator of inflammation. 17 In our study, LPS pretreatment might have led to early secretion of HMGB1, which may have led to a perpetual activation of the miRNAs and, eventually, a late downregulation of HMGB1 at the mRNA and protein level.
According to IPA, targetscan.org and microRNA.org, miR-142-3, miR-146a, miR-299 and miR-200c are predicted to target HMGB1 (www.ingenuity.com, accessed 2 December 2011). In our study, these 4 miRNAs were significantly up-regulated at 24 h and 48 h in the LPS pretreated group compared with the saline pretreated group. In correlation, HMGB1, a common target for these 4 miRNAs, was significantly downregulated at the mRNA and protein levels at 24 h and 48 h.
miR-132 and miR-21 dysregulation correlates with IL-1β and IL-6 protein levels
Taganov et al. identified, by means of expression profiling, several miRNA genes, including miR-132, among others, to be induced in the human monocytic cell line THP-1 by LPS, as well as other microbial components and pro-inflammatory mediators.23,36 Early expression of miR-132 with LPS-pretreatment in our study correlated with downregulation of IL-1β levels, a presumed target of miR-132, contributing to the development of microbial tolerance.
The tumorigenic function of miR-21, along with many of its predicted targets has been reported extensively and confirmed in the literature. 36 Recently, miR-21 has been proven to affect the innate immune response by inducing IL-10 production via a complex negative feedback loop. 36 Mice deficient in PDCD4, a confirmed miR-21 target, 37,38 exhibit lower LPS-induced mortality rates, lower IL-6 production and increased IL-10 protein levels. miR-21 up-regulation, in our study, correlated with decreased IL-6 production and lower IL-1β levels at 24 h and 48 h compared with the saline pretreated group, contributing to the development of microbial tolerance. Furthermore, IL-6 is targeted by miR-142-3p. Sun et al. demonstrated convincingly that miR-142-3p is a major, if not the only, regulator of IL-6 production. 22 The up-regulation of miR-142-3p in our study correlated with the consecutive downregulation of IL-6 and might thereby contribute to the observed tolerance pattern.
This study has several limitations. We chose to use a monobacterial model for peritonitis instead of a polymicrobial model, which is clinically more relevant. However, the purpose of this model was to study the development of tolerance in an in vivo model and we believed that this question might be better addressed with a monobacterial, Gram-negative bacteria model rather than a polymicrobial one. Furthermore, we did not measure the level of downregulation of the target proteins of miR-155 and miR-146a, which would strongly support the proposed action of these two miRNAs, nor did we transfect or knockdown any of the significant miRNAs to prove their action. However, for most of the miRNAs mentioned in this study, their proposed effect has been demonstrated and proven by others. For example, miR-155 and miR-146a have been shown to control TNF-α production in the macrophage response by several different groups.23,32,39 The effect of miR-142-3p on IL-6 has been demonstrated thoroughly by Sun et al. 40
Despite these limitations, we have shown that microbial tolerance can be induced in vivo and that miRNAs might play a pivotal role in the development of tolerance by fine-tuning the macrophage response, by targeting central inflammatory signaling pathways and by modulating cytokine production.
Conclusion
The molecular mechanisms underlying microbial tolerance remain very complex. As discussed previously, microbial tolerance is a biologic phenomenon of gene reprogramming and immune modulation rather than a global downregulation of gene expression and function. 8 As the regulation of endotoxin tolerance involves multi-leveled components of the inflammatory pathway, we believe that post-transcriptional modification through miRNA regulation of pro-inflammatory cytokines could contribute to the molecular basis of microbial tolerance. According to our study, the specific dysregulation of miRNAs, including miR-155, -146a, -142-3p, -299, and -200c -132 and -21, along with their effect on major inflammatory pathways like the TLR and NF-κB signaling pathway, but also on cytokine production directly, suggests a plausible mechanism for tolerance in this novel model with persistent peritoneal infection. Further in vivo studies using miRNA mimics and inhibitors could possibly be useful to further elucidate the effects of these miRNAs on survival in microbial tolerance.
Footnotes
Funding
This research received funding from the Department of Veterans Affairs Merit Review Program.
Acknowledgements
The authors would like to thank James C. Peyton for technical assistance with bacterial cultures and animal harvests.
Conflict of interest
The authors do not have any potential conflicts of interest to declare.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
