Abstract
Recognition of pathogen-associated molecular patterns that activate IL-1β is regulated by inflammasomes, predominantly of the nucleotide-binding oligomerization domain-like receptor (NLR) family. NLRP3 inflammasome is involved in the innate immune responses in periodontal disease. This is an inflammatory condition that destroys the tooth-supporting (periodontal) tissues, initiated by the subgingival formation of multi-species biofilms, frequently including the Gram-negative species Porphyromonas gingivalis. The aim of this study was to investigate the relative effect of P. gingivalis as part of subgingival biofilm, on the expressions of NLRP3 inflammasome, absent in melanoma (AIM)2 (a non-NLR inflammsome) and IL-1β by human gingival fibroblasts. The 10-species subgingival biofilm model, or its 9-species variant excluding P. gingivalis, were used to challenge the cells for 6 h. Gene expression analysis for various inflammasome components and IL-1β was performed by TaqMan real-time PCR. The 10-species subgingival biofilm reduced NLRP3 and IL-1β, but did not affect AIM2 expression. Exclusion of P. gingivalis from the biofilm partially rescued NLRP3 and IL-1β expressions. In conclusion, subgingival biofilms down-regulate NLRP3 and IL-1β expression, partly because of P. gingivalis. These dampened host innate immune responses may favour the survival and persistence of the associated biofilm species in the periodontal tissues.
Introduction
Periodontal diseases are characterized by the inflammatory destruction of the tooth-supporting (periodontal) tissues and they are the most common chronic inflammatory diseases in humans. This cluster of diseases is attributed to resident oral bacteria colonizing the tooth surfaces in the form of polymicrobial biofilm communities. 1 Interaction of biofilms with the juxtaposing periodontal tissues triggers an inflammatory response, which aims to prevent bacterial colonization and establishment. 2 However, if the inflammatory response becomes excessive, it will damage the periodontal tissues, rather than being protective. 3 Periodontitis is a form of periodontal disease in which the inflammatory response has progressed enough to destroy the tooth-supporting alveolar bone, eventually leading to tooth loss, if left untreated. The causative factor of periodontitis is the development of a ‘subgingival’ biofilm, in other terms of a biofilm that forms below the gingival margin within a periodontal pocket, consisting of characteristic bacterial species.4–6 Porphyromonas gingivalis is a black-pigmenting Gram-negative anaerobe, very frequently detected in subgingival biofilms from sites with periodontitis. 7 Interestingly, even at low colonization levels, P. gingivalis has a key role in altering the composition of the local oral commensal microbiota, which is also required for P. gingivalis-induced bone loss. 8 This species is considered notorious for its capacity to manipulate host cell signalling, either by promoting or by dampening inflammatory innate immune responses. 9 Hence, P. gingivalis can orchestrate a deregulation of the physiological host-microbial homeostasis. 10
IL-1 cytokines are key modulators of the inflammatory responses in periodontal diseases. 11 Clinical studies indicate that IL-1β, the best known member of this family, is detected at higher levels in gingival crevicular fluid 12 or gingival tissues 13 of patients with periodontal disease compared with healthy subjects. On the cellular level, activation and production of IL-1β is controlled by the ‘inflammasomes’, which are intracellular protein complexes that can sense pathogen-associated molecular patterns (PAMPs). 14 The nucleotide-binding oligomerization domain-like receptor (NLR) inflammasomes are intracellular pattern recognition receptors (PRRs) that detect PAMPs. Through the recruitment of cysteine proteinase caspase-1, NLRs activate intracellularly stored pro-IL-1β, which is subsequently released from the cell. The NLRP3 inflammasome, in particular, consists of three components: the NLRP3 ‘sensor’, the caspase-1 ‘effector’ and the apoptotic speck protein containing a C-terminal caspase recruitment domain (ASC) ‘adaptor’ that links the former two molecules. NLRP3 is activated by cell stresses 15 and bacteria or viruses.16–19 Recent clinical evidence demonstrates that the expression of this inflammasome is higher in periodontally diseased tissues compared with healthy ones. 13 Absent in melanoma (AIM)2 is a non-NLR inflammasome that senses double-stranded DNA from various sources, including bacteria, viruses or host cells. 20 A shared feature of the NLRP3 and AIM2 inflammasomes is that they both include ASC and caspase-1. 21
Gingival fibroblasts (GFs) are the major population of the gingival tissue, which is one of the constituents of periodontal tissues. They are responsible for the synthesis and degradation of the extracellular matrix, and respond to PAMPs by producing mediators of inflammation. 22 In doing so, they are crucial for regulating the homeostasis of the periodontal tissues in health and disease. It was recently shown that GFs express NLRP3 and AIM2, which are differentially regulated by in vitro supragingival and subgingival biofilms supernatants. Subgingival biofilm supernatants caused a down-regulation of NLRP3 expression but had a bi-phasic effect on AIM2 and IL-1β expression. 23 This ‘dampened’ host sensing elicited by subgingival biofilms is perceived as a strategy for evading immune surveillance, which may promote pathogen survival. Although it is postulated that P. gingivalis, as part of polymicrobial subgingival biofilms, is involved in this effect, this has not yet been demonstrated. Therefore, the present in vitro study aims to investigate the involvement of P. gingivalis in the regulation of NLRP3 and AIM2 inflammasomes in GFs by subgingival biofilms.
Materials and methods
In vitro biofilm model
Bacterial composition of the subgingival biofilms.
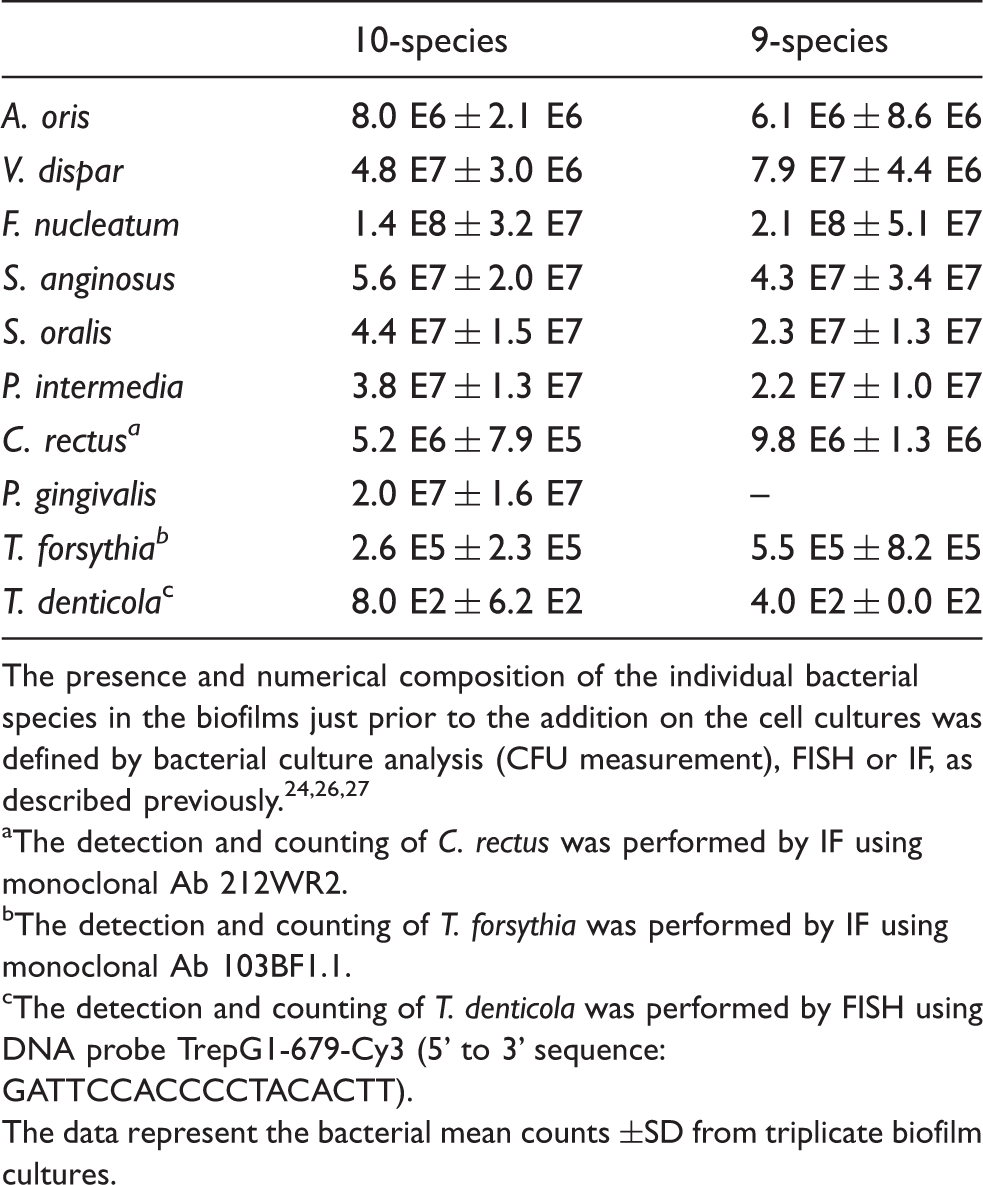

The presence and numerical composition of the individual bacterial species in the biofilms just prior to the addition on the cell cultures was defined by bacterial culture analysis (CFU measurement), FISH or IF, as described previously.24,26,27
The detection and counting of C. rectus was performed by IF using monoclonal Ab 212WR2.
The detection and counting of T. forsythia was performed by IF using monoclonal Ab 103BF1.1.
The detection and counting of T. denticola was performed by FISH using DNA probe TrepG1-679-Cy3 (5’ to 3’ sequence: GATTCCACCCCTACACTT).
The data represent the bacterial mean counts ±SD from triplicate biofilm cultures.
Cell cultures
Primary human GF cell lines were established as described previously. 28 Briefly, gingival tissue biopsies used were obtained from a healthy young individual, who had their first premolar removed during the course of orthodontic treatment. Ethics approval was granted by the Human Studies Ethical Committee of Umeå University, Sweden, and informed consent was given by the donor. The cells were passaged and cultured in Minimum Essential Medium Alpha (Gibco), supplemented with 5% heat-inactivated FBS (Sigma), 50 U/ml penicillin, and 50 µg/ml streptomycin (Sigma). For the experiments, GFs at passage 4 were seeded at a concentration of 1 × 104 cells/cm2 in antibiotic-free culture medium supplemented with 5% FBS. The cells were allowed to attach overnight, maintaining a sub-confluent status and then cultured for 6 h in the presence or absence of either biofilm.
RNA extraction and cDNA synthesis
Upon completion of the experiments, after 6 h of challenge, the culture supernatants were removed and the cell monolayers were washed twice in PBS before being lysed. Total RNA was extracted by the RNeasy Mini Kit (QIAGEN) and its concentration was measured by a NanoDrop spectrophotometer. One µg of total RNA was then reverse transcribed into single-stranded cDNA by M-MLV Reverse Transcriptase, Oligo(dT)15 Primers and PCR Nucleotide Mix (Promega), at 40°C for 60 min and 70°C for 15 min. The resulting cDNA was stored at −20°C.
Quantitative real-time PCR
Gene expression analysis was performed by quantitative real-time PCR (qPCR) in an ABI Prism 7000 Sequence Detection System and software (Applied Biosystems). For the amplification reactions, TaqMan Gene Expression Master Mix and Gene Expression Assay kits (Applied Biosystems) were used (assay IDs: NLRP3: Hs00918085-m1, ASC: Hs01547324-m1, caspase-1: Hs00354836-m1, AIM2: Hs00915710-m1, IL-1β: Hs00174097-m1, GAPDH: Hs99999905-m1). The standard PCR conditions were 10 min at 95°C, followed by 40 cycles at 95°C for 15 seconds and 60°C for 1 min. GAPDH was used as a housekeeping gene. The expression levels of the target transcripts in each sample were calculated by the comparative Ct method (2−ΔCt formula) after normalization to GAPDH.
Measurement of IL-1β by ELISA
The concentrations of IL-1β secreted by GFs into the culture supernatant were measured by a commercially-available ELISA kit (DY201, DuoSet, R&D Systems, Abingdon, UK). Absorbance was measured at 450 nm using a microplate reader (Epoch, BioTek, Luzern, Switzerland). A wavelength correction of 570 nm was used for the subtraction of the background. A standard curve was created using known concentrations of rhIL-1β provided in the kit. The concentration of IL-1β in each sample was calculated by a four-parameter logistic (4-PL) equation. The lowest detection limit of the assay was 1.7 pg/ml. The results represent the mean ± SEM from four independent cell cultures in each group.
Statistical analysis
ANOVA was used to analyse the statistical significances of the data. The Bonferroni post-hoc test was employed for the comparisons between individual groups. The data were considered significant at P < 0.05.
Results
The GF cultures were challenged for 6 h with either the 10-species subgingival biofilm, or its 9-species variant, which excluded P. gingivalis. The effect of these two biofilms on the gene expression of the NLRP3 and AIM2 inflammasomes, as well as IL-1β, was then investigated. The 10-species biofilm caused a significant down-regulation of NLRP3 expression by 33% (Figure 1). However, it did not affect the expressions of adaptor molecule ASC (Figure 2) or the effector molecule caspase-1 (Figure 3). Moreover, the expression of AIM2 was not affected by the biofilm challenge (Figure 4). Nevertheless, IL-1β expression exhibited a significant down-regulation by 35% (Figure 5).
Regulation of NLRP3 gene expression in response to biofilm challenge. GF cultures were challenged for 6 h with the 10- or 9-species (excluding P. gingivalis) subgingival biofilm. NLRP3 gene expression was calibrated against GAPDH and presented as the 2−ΔCT formula. Bars represent mean values ± SEM from four independent cell cultures in each group. * represents statistically significant difference compared with the control group. Regulation of ASC gene expression in response to biofilm challenge. GF cultures were challenged for 6 h with the 10 - or 9-species (excluding P. gingivalis) subgingival biofilm. ASC gene expression was calibrated against GAPDH and presented as the 2−ΔCT formula. Bars represent mean values ± SEM from four independent cell cultures in each group. Regulation of caspase-1 gene expression in response to biofilm challenge. GF cultures were challenged for 6 h with the 10- or 9-species (excluding P. gingivalis) subgingival biofilm. Caspase-1 gene expression was calibrated against GAPDH and presented as the 2−ΔCT formula. Bars represent mean values ± SEM from four independent cell cultures in each group. Regulation of AIM2 gene expression in response to biofilm challenge. GF cultures were challenged for 6 h with the 10- or 9-species (excluding P. gingivalis) subgingival biofilm. AIM2 gene expression was calibrated against GAPDH and presented as the 2−ΔCT formula. Bars represent mean values ± SEM from four independent cell cultures in each group. Regulation of IL-1β expression in response to biofilm challenge. GF cultures were challenged for 6 h with the 10- or 9-species (excluding P. gingivalis) subgingival biofilm. IL-1β gene expression was calibrated against GAPDH and presented as the 2−ΔCT formula. Bars represent mean values ± SEM from four independent cell cultures in each group. * represents statistically significant difference compared with the control group.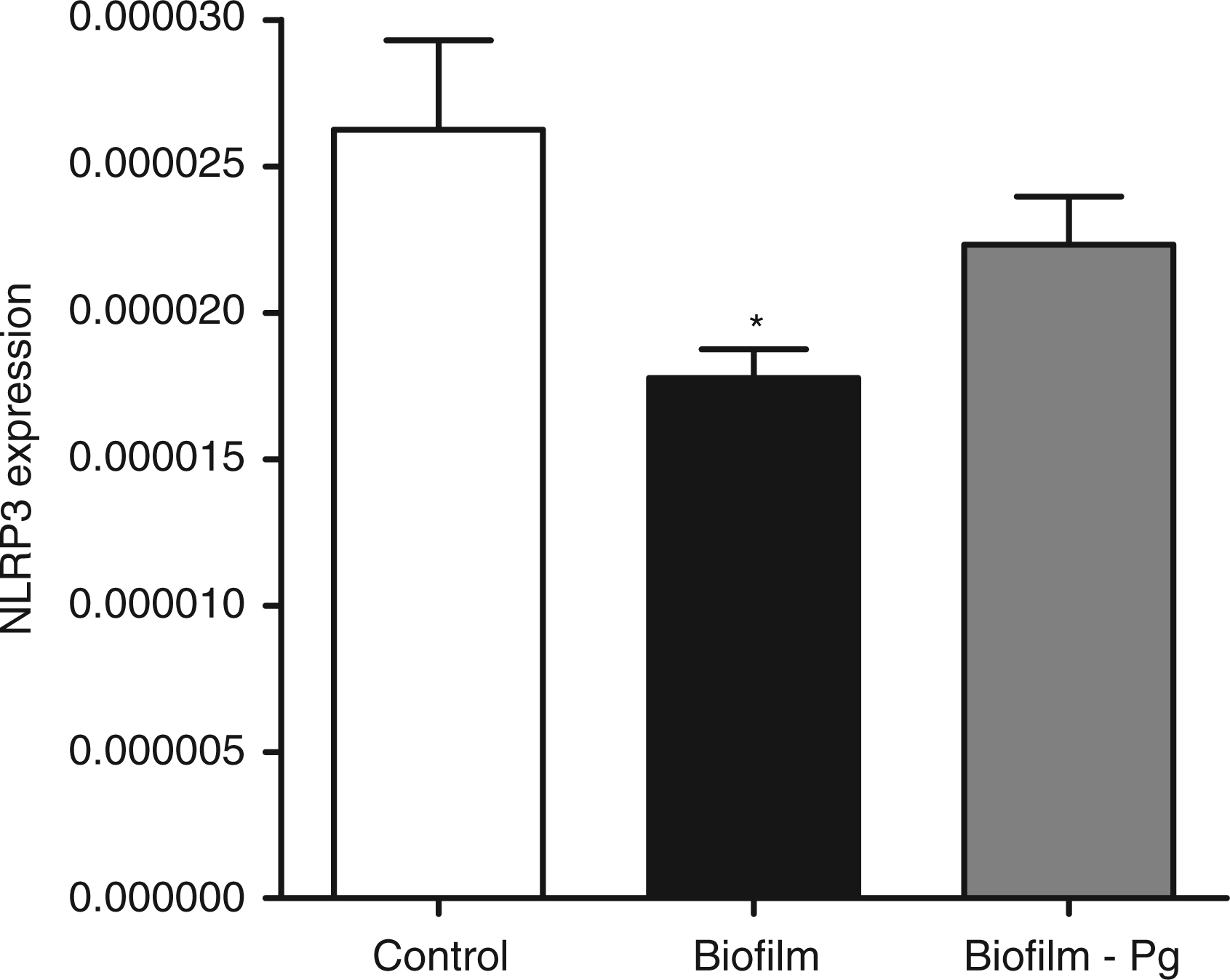
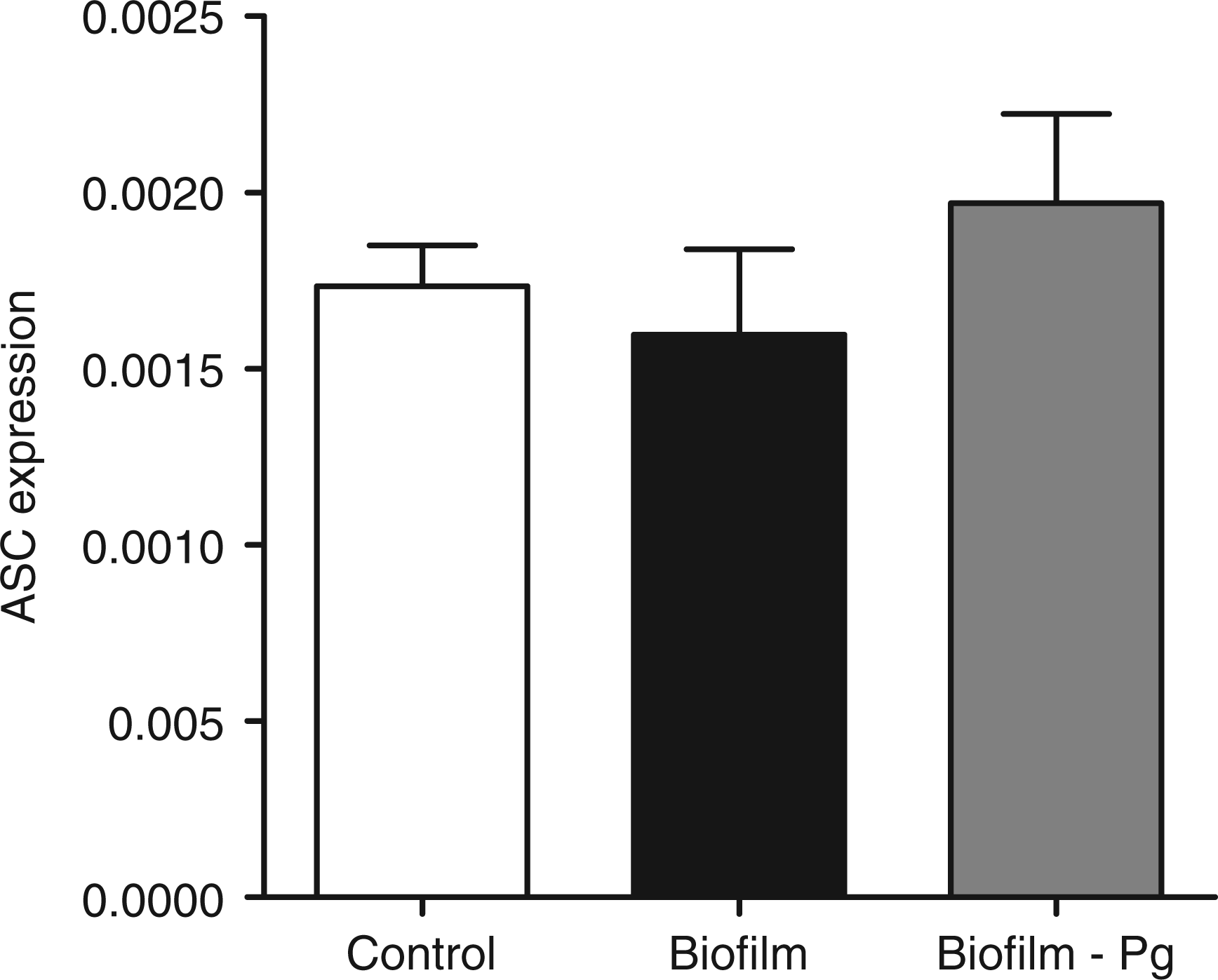
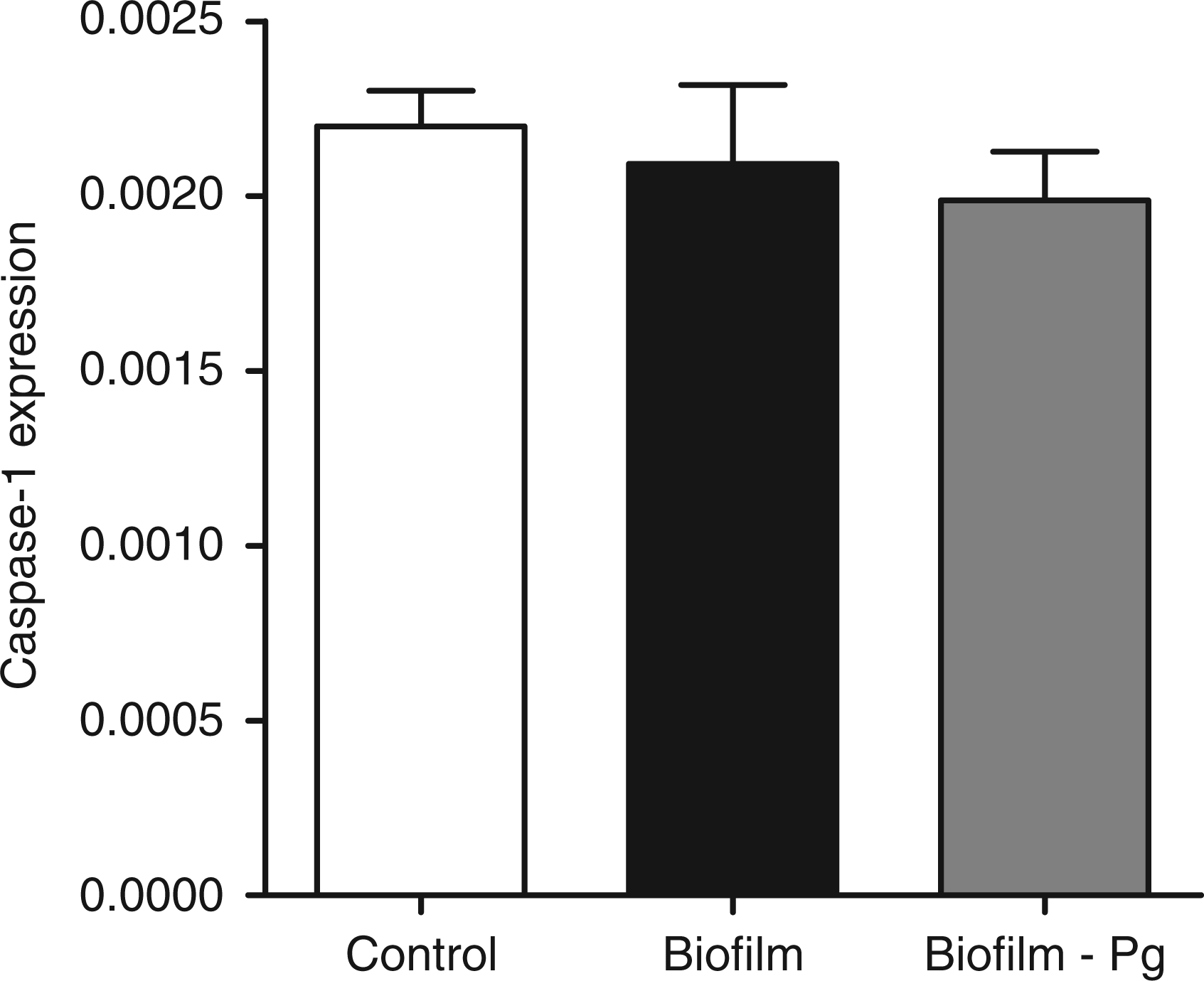
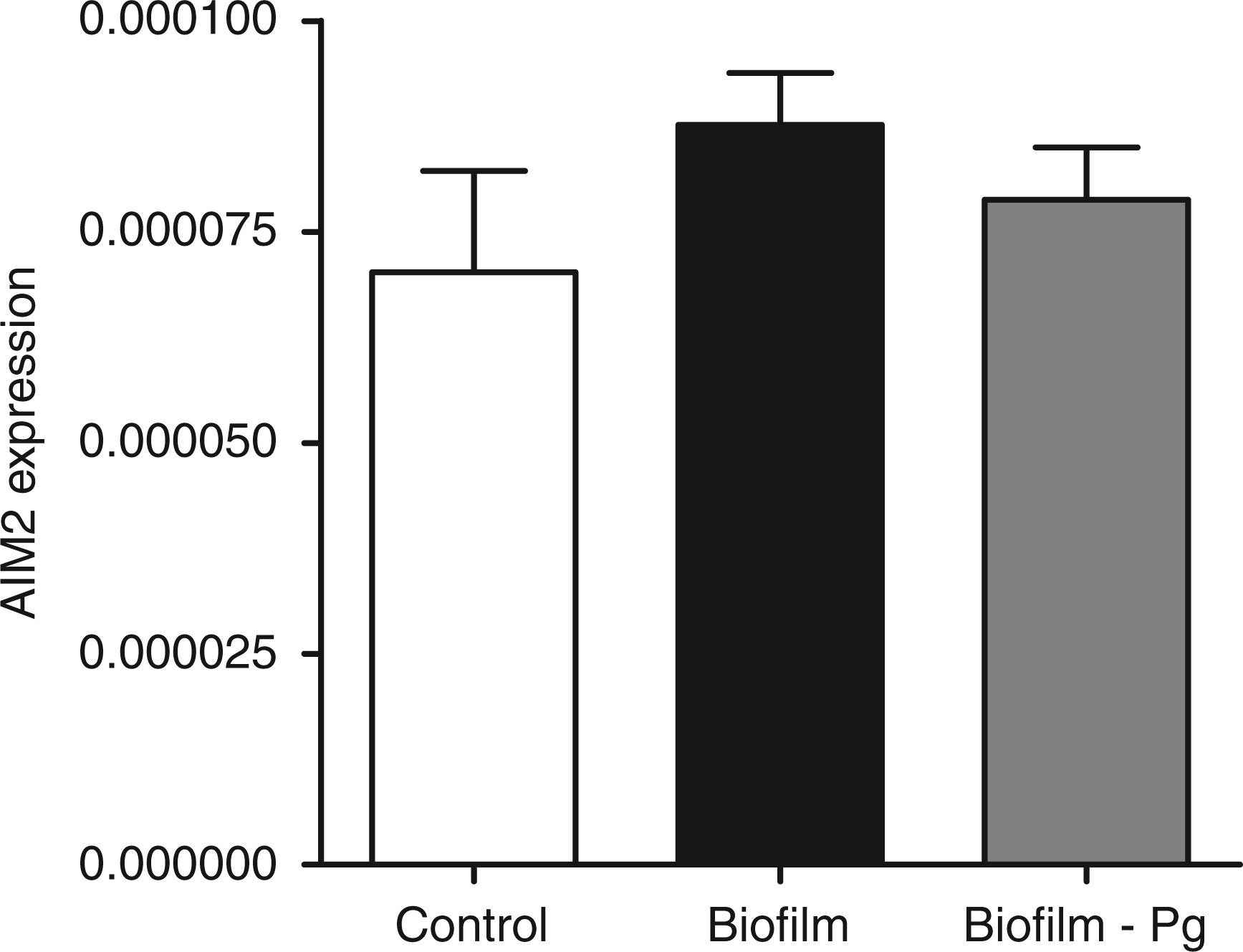
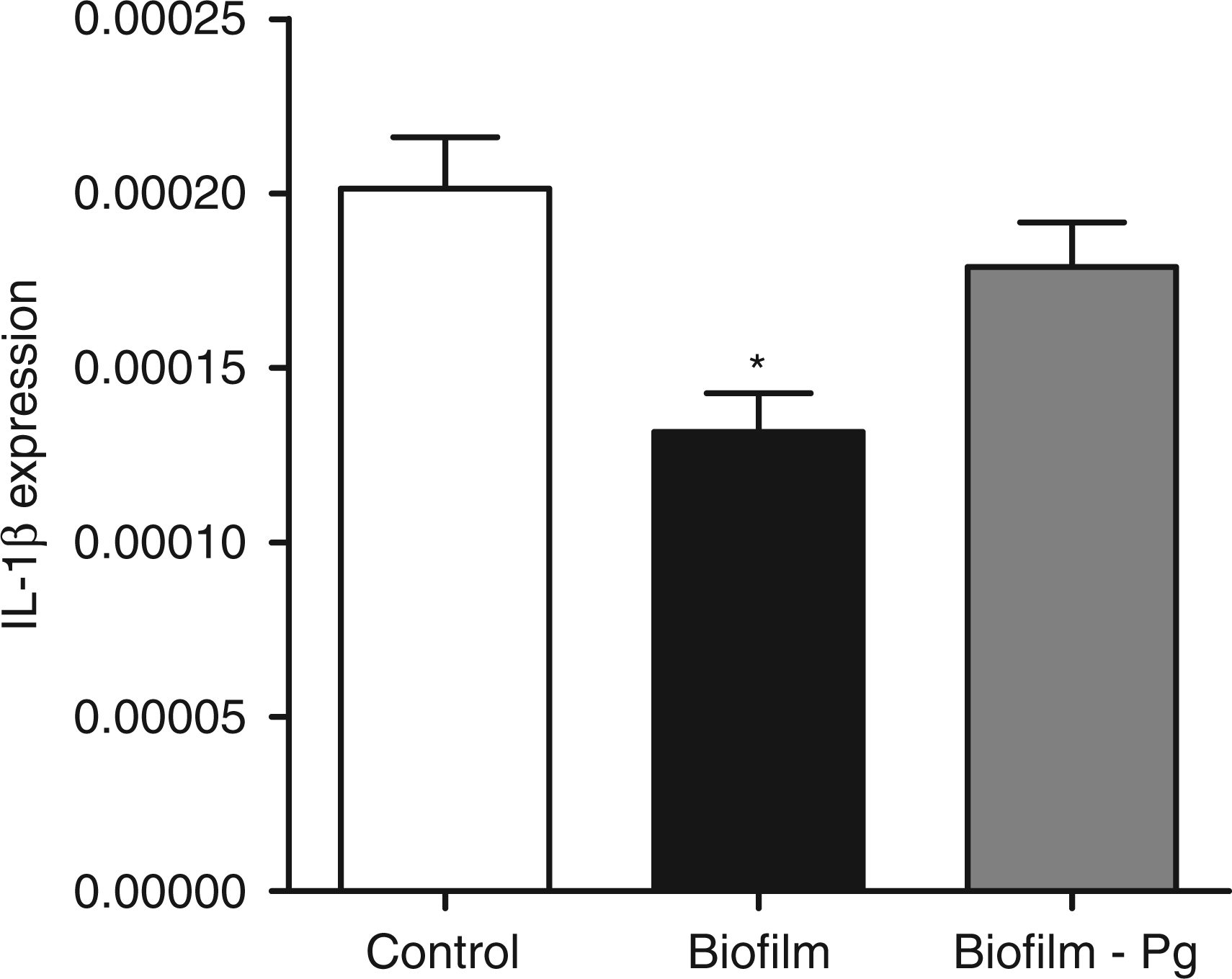
The effects elicited by the 10-species biofilm were compared with those caused by the 9-species biofilm where P. gingivalis was excluded. The bacterial composition of the two biofilms was otherwise comparable, as there were no log-scale differences (Table 1). With regards to the expression of the various inflammasome components, exclusion of P. gingivalis from the biofilm partially rescued NLRP3 expression to 85% of control levels (Figure 1), whereas it did not affect the expression of adaptor ASC (Figure 2) or effector caspase-1 (Figure 3). Accordingly, the expression of AIM2 was not affected (Figure 4). Nevertheless, the expression of IL-1β was partially rescued to 88% of the control levels (Figure 5), similarly to NLRP3 expression.
The secretion of IL-1β by the cells in response to the 6-h biofilm challenge was also investigated. IL-1β was detected at particularly low levels. In the control group, IL-1β concentration in the culture supernatant was 5.5 ± 0.4 pg/ml, whereas the presence of the 10-species biofilm reduced this to 4.3 ± 0.3 pg/ml (23% reduction). In the presence of the 9-species biofilm, which excluded P. gingivalis, the IL-1β concentration in the culture supernatant was 3.3 ± 0.1 pg/ml (41% reduction compared with control). The concentration of IL-1β in both biofilm groups was significantly lower than the control group. However, the difference between the two biofilm groups did not prove to be statistically significant.
Discussion
This study investigated the effect of in vitro subgingival biofilms on inflammasome expression by GF and evaluated the relative role of P. gingivalis. The results indicate a down-regulation of NLRP3 expression by subgingival biofilms, but no effect on the expressions of adaptor ASC and effector caspase-1. These responses are comparable with the effects elicited by subgingival biofilm supernatants, which, at higher concentrations, down-regulated NLRP3 but did not affect ASC or caspase-1 expression. 23 However, lower- and mid-range biofilm supernatant concentrations did increase ASC and caspase-1 expressions, rendering this effect bi-phasic. 23 Overall, the down-regulation of NLRP3, as a PRR sensor, may compromise the capacity of the cells to sense PAMPs, implying a dampening of the endpoint innate immune responses.
The observed effects of the subgibival biofilm on the expression of the NLRP3 inflammasome matches the well established capacity of P. gingivalis to manipulate host signalling.9,29 This may enable this species to evade host surveillance, 30 offering a survival advantage to all co-habiting organisms of the biofilm. 9 For this reason, the present study considered the role of P. gingivalis in the observed effects by establishing in parallel a 9-species biofilm that lacked P. gingivalis and comparing its effects with the 10-species biofilm. The two biofilms did not exhibit quantitative differences in composition other than the absence of P. gingivalis. The data demonstrated that the lack of P. gingivalis indeed rescued NLRP3 expression almost to control levels, implying a key role of this species in subgingival biofilms in down-regulating specific gene expression. Importantly, as a single species, P. gingivalis may either up-regulate13,31 or down-regulate 32 NLRP3 expression, depending on the cell type. Hence, this indicates that when part of a microbial biofilm community, P. gingivalis may selectively orchestrate the innate immune response, in line with its role as a ‘keystone’ species. 8
The relative effect of P. gingivalis as member of a biofilm community was also investigated on IL-1β expression by GFs. So far, there is evidence that co-infection of host cells with P. gingivalis and other putative periodontal pathogens decreases the IL-1-inducing capacity of the latter.33,34 It was recently demonstrated that subgingival biofilm supernatants in which P. gingivalis was present may exert a bi-phasic effect on IL-1β expression, with an increase by lower concentrations and a decrease to control levels by higher ones. 23 In the present experimental system with viable biofilms, IL-1β expression was decreased when the cells were challenged with the 10-species subgingival biofilm, but was partially rescued when P. gingivalis was absent (9-species biofilm). Hence, the regulation of IL-1β expression followed a similar pattern to that of NLRP3 expression. Although the expression of these genes is not necessarily interdependent, these results denote that they can be regulated in a parallel manner by subgingival biofilms, which is also in line with the recent observations. 23
Changes in NLRP3 expression would be expected to result in changes of IL-1β secretion by the cells in various experimental systems. In the present study, it was confirmed that IL-1β secretion was, indeed, lower in the biofilm-challenged cell cultures, which is in agreement with a lower NLRP3 expression. In the case where P. gingivalis was absent from the biofilm, NLRP3 and IL-1β expressions were partially rescued to control levels. Nevertheless, this was not accompanied by a concomitant resumption of IL-1β secreted protein levels. The reason for this is not clear, but in such a complex experimental system several factors may impose limitations in studying IL-1β protein secretion. For instance, GFs produce low levels of IL-1β35,36 compared to inflammatory cells; therefore it is difficult to evaluate efficiently differences in production, particularly at the lower end of the detection spectrum. Moreover, it is established that in vitro subgingival biofilms can degrade pro-inflammatory cytokines, including IL-1β, 24 an effect which may compromise the endpoint measurements.
The expression of AIM2, the sensor of the homonymous inflammasome that oligomerizes with ASC and caspase-1 14 was also investigated. AIM2 recognizes different PAMPs than NLRP3, including cytosolic double-stranded DNA.20,21 The present data do not identify significant changes in AIM2 expression, in the presence of either biofilm, although subgingival biofilm supernatants were earlier shown to have a bi-phasic effect. 23 Collectively, it is likely that high concentrations of biofilm supernatants elicit equivalent cell responses to viable biofilms, as seen in this study.
In conclusion, down-regulation of NLRP3 inflammasome expression may constitute a strategy to manipulate the local innate immune inflammatory responses. 37 In the case of subgingival biofilms, this could lead to the evasion of host surveillance by the associated species, with the obvious benefit of their survival into the tissue environment.10,23 The present study affirms that subgingival biofilms down-regulate NLRP3 and IL-1β gene expression in GF in vitro and further identifies P. gingivalis as a key species in these events. This supports the notion that, as part of polymicrobial communities, P. gingivalis can disrupt the host-microbial homeostasis, 8 in this case by dampening the pathogen-sensing capacity of the cells via inflammasomes.
Footnotes
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Acknowledgements
The authors would like to thank Mr Andre Meier, Mrs Helga Lüthi-Schaller and Mrs Verena Osterwalder for their excellent technical assistance, and Dr Anders Johansson (Institute of Odontology, Umeå University, Sweden) for providing the GFs.
