Abstract
Defensins are an important family of natural antimicrobial peptides. Chlamydophila pneumoniae, a common cause of acute respiratory infection, has a tendency to cause persistent inflammatory diseases such as atherosclerosis, which may lead to cardiovascular disease or stroke. As endothelial cells are related to the physiopathology of stroke, the effects of in vitro C. pneumoniae infection on the expression of human β-defensin 2 (HBD-2) in brain capillary endothelial cells (BB19) was investigated. A time-dependent increase in HBD-2 mRNA was observed by means of real-time reverse transcription PCR (RT-PCR) in BB19 cells following C. pneumoniae infection, with a maximum increase at 24 h. A gradual induction of HBD-2 protein in the C. pneumoniae-infected endothelial cells was detected by immunoblotting. Immunofluorescence revealed the staining of HBD-2 in the cytoplasm of endothelial cells following C. pneumoniae infection. The secretion of HBD-2 (confirmed by ELISA) was significantly elevated 24 h after C. pneumoniae infection. These novel results indicate that HBD-2 is expressed and produced in the human brain capillary endothelial cells upon infection with C. pneumoniae, and provide evidence that HBD-2 plays a role in the early immune responses to C. pneumoniae and probably in the immunopathogenesis of atherosclerosis.
Introduction
Antimicrobial peptides are key components of innate immunity. Defensins, an important family of natural antimicrobial peptides, display antimicrobial activity against a variety of micro-organisms, such as bacteria, fungi and enveloped viruses. 1 Human defensins are categorized into α- and β-defensin subfamilies. Human α-defensins are produced by granulocytes and intestinal Paneth cells, whereas human β-defensins (HBDs) are expressed mainly in epithelial cells. To date, six human HBDs have been identified in human tissues. 2 The β-defensins are components of the innate immune system and also bridge innate and acquired immune responses by recruiting dendritic cells and T-lymphocytes to the site of microbial invasion. 3 While the expression of HBD-1 is generally constitutive, the levels of HBD-2, HBD-3 and HBD-4 are inducible by pro-inflammatory cytokines and bacteria. 2 Human β-defensin gene expression has been identified in various human epithelia, such as epithelium from skin, lungs, trachea, digestive tract or the urogenital system and its mRNA induction and protein secretion usually correlate. A moderate expression of β-defensin 2 has been observed also in human monocytes, macrophages and dendritic cells. 4 In addition to the antimicrobial activity, HBD-2 provides a link to the adaptive immune system by attracting immature dendritic cells and memory T-cells via the chemokine receptor CCR-6, suggesting that the tasks of defensins are more than simply antimicrobial.3,5– 7
Chlamydophila pneumoniae, an obligate intracellular parasite and a common cause of acute respiratory infection, has a tendency to cause persistent inflammatory diseases such as atherosclerosis, which may lead to cardiovascular disease or stroke. 9 – 11 Three principal cell types involved in the atherogenic process within the developing atheroma include endothelial cells, vascular smooth muscle cells, and monocytes/macrophages. Recent publications have indicated that C. pneumoniae infection induces HBD-2 expression in human mononuclear cells and in gingival fibroblasts.12,13
It has been reported that C. pneumoniae can infect human endothelial cells and induce the expression of cytokines, adhesion molecules, chemokines and molecules with procoagulant activity. 14 As endothelial cells are related to the physiopathology of stroke, 15 we considered it of interest to investigate the effects of in vitro C. pneumoniae infection on the HBD-2 expression in brain capillary endothelial cells using human brain capillary endothelial cell line BB19.
Materials and methods
Cell line
BB19 cells, human brain capillary endothelial cells immortalized with the E6E7 genes of human papilloma virus 16 were kindly provided by Dr J. Nelson (Department of Microbiology and Immunology, Oregon Health Sciences University, USA). The cell line was cultured as monolayer culture (cell culture medium RPMI supplemented with 10% FCS and 10% human AB serum, and 50 µg/ml endothelial growth factor [Sigma]). Cells were seeded onto culture surface precoated with 1% gelatine at a density of 100,000–150,000 cells/ml.
Chlamydophila pneumoniae propagation and inoculum preparation
Chlamydophila pneumoniae CWL-029 respiratory strain, obtained from ATCC, and C. pneumoniae CV6 cardiovascular strain, isolated from an atheroma of a patient with coronary artery disease 17 were propagated in Hep2 cells in MEM with Earle salts supplemented with 10% heat-inactivated fetal bovine serum (FBS), 0.5% glucose, 2 mmol/l l-glutamine, 1× non-essential amino acids (Sigma), 8 mmol/l HEPES, 25 µg/ml gentamycin, and 1 µg/ml cycloheximide. Infected cells were harvested on day 3 or 4 and then disrupted by two cycles of freeze–thawing and ultrasonication. Cell debris was removed by centrifugation at 300 g for 10 min, and bacteria were concentrated by centrifugation at 30,000 g for 30 min. Pellets were resuspended in sucrose-phosphate-glutamic acid buffer (0.22 mol/l sucrose, 10 mmol/l NaH2PO4, 3.8 mmol/l KH2PO4, and 5 mmol/l glutamic acid [pH 7.4]), formed into aliquots, and frozen at –80°C until use. The same stock was used for all experiments. For mock infection, uninfected Hep2 cells were processed as described above and stored at –80°C.
Chlamydophila pneumoniae titration was performed on Hep2 cells by use of immunofluorescent staining. Monolayers on 13-mm coverslips were infected with serial dilutions of concentrated bacteria, incubated for 48 h, fixed with acetone, and stained with mouse anti-C. pneumoniae major outer membrane protein (MOMP) monoclonal antibody (Dako). Fluorescein isothiocyanate (FITC)-labeled anti-mouse IgG (goat F[ab’]2 anti-mouse IgG; Sigma) was used as a secondary antibody. After the counting of chlamydial inclusions under a fluorescent microscope, titers were expressed as inclusion-forming units per millilitre (IFU/ml).
In vitro infection
BB19 cells were transferred to 6-well plates at a density of 1 × 106 cells/well, and infected with C. pneumoniae stock at a multiplicity of infection (MOI) of 5 IFU/cell. Control cells were not infected, or were treated with the same volume of mock stock. In addition to infection with viable C. pneumoniae, BB19 cells were stimulated with organisms that had been previously heat-inactivated (at 70°C for 45 min). Inoculated cells were centrifuged at 800 g for 60 min at room temperature (20–22°C). Medium was then changed for fresh medium without cycloheximide. The plates were incubated at 37°C under 5% CO2, and infected and control cells were harvested at the times indicated.
RNA isolation and PCR amplification
Total RNA was isolated with a Perfect RNA Eukaryotic M10.1177_1753425910kit (Eppendorf) according to the manufacturer’s instructions. The RNA concentration was determined by the A260 value of the sample. Complementary DNA (cDNA) was generated from 1 µg total RNA using the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems) in a final volume of 20 µl. Real-time reverse transcription PCR (RT-PCR) analyses were performed in a fluorescence temperature cycler (LightCycler; Roche Diagnostics GmbH, Mannheim, Germany) according to the manufacturer’s instructions. After reverse transcription, amplification was carried out using Light Cycler Fast Start DNA MasterPLUS SYBR Green I mix (Roche). Samples were loaded into capillary tubes and placed in the fluorescence thermocycler (LightCycler). Initial denaturation at 95°C for 10 min was followed by 55 cycles of 95°C for 5 s, the primer-specific annealing temperature for 10 s, and elongation at 72°C for 10 s. For HBD-2 (sense, 5′-ATC AGC CAT GAG GGT CTT GT-3′; antisense, 5′-GAG ACC ACA GGT GCC AAT TT-3′), the annealing temperature was set to 62°C. For hBD1- (sense, 5′-TTG TCT GAG ATG GCC TCA GGT GGT AAC-3′; antisense, 5′-ATA CTT CAA AAG CAA TTT TCC TTT AT-3′), the annealing temperature was set at 58°C Initial denaturation at 95°C for 10 min was followed by 50 cycles of 95°C for 10 s, the primer-specific annealing temperature for 7 s, and elongation at 72°C for 12 s. For the housekeeping gene GAPDH (glyceraldehyde-3-phosphate dehydrogenase); sense, 5′-AAG GTC GGA GTC AAC GGA TTT-3′; antisense, 5′-TGG AAG ATG GTG ATG GGA TTT-3′ primers were used. At the end of each run, melting-curve profiles were achieved by cooling the sample to 40°C for 15 s and then heating the sample slowly at 0.20°C/s up to 95°C with continuous measurement of the fluorescence to confirm the amplification of specific transcripts. Cycle-to-cycle fluorescence emission readings were monitored and analyzed using LightCycler software (Roche Diagnostics). Melting curves were generated after each run to confirm amplification of specific transcripts. Total RNA samples without reverse transcription were amplified in the same way in order to exclude the possibility of the DNA contamination of the RNA preparations. The amount of RNA was equivalent to that added to the reaction via the reverse transcription. reaction. Real-time PCR reaction and the melting point analysis were performed and the results were negative. All quantifications were normalized to the housekeeping GAPDH gene. Relative gene expression is given as a ratio between target gene and GAPDH gene expressions.
Western blot assays
Cells (1 × 107) were homogenized in ice-cold lysis buffer containing RIPA buffer and protease inhibitor cocktail (Sigma), and the mixture was then centrifuged at 10,000 g for 10 min to remove cell debris. Protein concentrations of cell lysates were determined using the Bio-Rad protein assay (Bio-Rad, Hercules, CA, USA). Supernatants were mixed with Laemmli’s sample buffer and boiled for 3 min. To detect β-defensin, aliquots of the supernatants containing 25 µg of total protein were resolved by SDS-PAGE and electrotransferred onto polyvinylidene difluoride membranes (Bio-Rad). Preblocked blots were reacted for 4 h with specific antibodies to human β-defensin 2 (RD Systems, Minneapolis, MN, USA) in PBS containing 0.05% (v/v) Tween-20, 1% (w/v) dried non-fat milk (Difco Laboratories, Detroit, MI, USA) and 1% (w/v) BSA (fraction V; Sigma). Blots were then incubated for 2 h with species-specific secondary antibody coupled to peroxidase (peroxidase-conjugated anti-goat antibody [Sigma]). Filters were washed five times in PBS-Tween for 5 min after each step and were developed using a chemiluminescence detection system (Amersham, Buckinghamshire, UK).
Human β-defensin 2 ELISA
For direct testing of the presence of the HBD-2 peptide in the infected and uninfected cells, and for measurement of the secreted HBD-2 protein into the tissue culture medium, a sensitive HBD-2 ELISA (Alpha Diagnostic, San Antonio, TX, USA) was used according to the manufacturer’s instructions. The detection limit of this ELISA kit was 0.8 pg/HBD-2 protein/ml.
Immunofluorescence
Expression s of HBD-2 peptide by C. pneumoniae infected and non-infected BB19 cells in Cytospin preparations were investigated by immunofluorescence analysis. Cells were fixed in 1% acetone for 15 min at room temperature. Cells were stained with goat anti-HBD-2 antibody (RD Systems) for 1 h. Bound antibody was detected with FITC-conjugated rabbit anti-goat IgG (Sigma) for 45 min. After each incubation step, the cells were washed three times with PBS containing 0.2% BSA. Fluorescence signals were analyzed via confocal microscopy.
Confocal microscopy and semiquantitative assessment of fluorescence intensities
Eight serial images of each immunostained sample were captured by Olympus FV1000 confocal laser scanning microscope with standard parameter settings. The immunofluorescence of control and C. pneumoniae-infected cells was quantitatively analyzed by ImageQuant software (Molecular Dynamics) as follows: 6-6 equally sized circular areas covering the cells were randomly selected on each image. The backgrounds of the selected areas were eliminated by threshold set up and the fluorescence intensities/pixel values of the randomly selected cells were quantified.
The mean and SD of the data gained from serial circular areas were calculated by means of Microsoft Excel. The level of statistical significance was determined by two-tailed t-test.
Statistical analysis
All values are expressed as mean ± SD. For the quantification of HBD-2 expression using RT-PCR and ELISA, results from C. pneumoniae infected samples were compared with non-infected controls. The data were subjected to two-tailed paired Student’s test. For immunofluorescence intensity analysis, intensity data from C. pneumoniae-infected cells were compared with those on the non-infected control. For all statistical evaluations, P < 0.05 was considered statistically significant. Data analyses were performed by Graph Pad Prism 5 (GraphPad Software Inc., San Diego, CA, USA) statistical program.
Results
Expression of human β-defensin 2 mRNA in BB19 cells and increase in expression by C. pneumoniae
To determine whether HBD-2 gene expression is induced in BB19 cells in response to C. pneumoniae infection, RT-PCR was performed. A time-dependent increase in HBD-2 mRNA was observed in BB19 cells (Fig. 1). The maximum increase in HBD-2 mRNA expression was observed at 24 h at a MOI of 5 IFU. The infection of endothelial cells by CV6, a cardiogenic strain, resulted in a more pronounced expression of HBD-2 mRNA.
Time-dependent increase in human β-defensin-2 (HBD-2) mRNA expression by C. pneumoniae. BB19 cells were infected with C. pneumoniae CWL 029 (grey bars) and with a cardiovascular strain of C. pneumoniae CV6 (black bars) or with heat-inactivated C. pneumoniae CV6 (hatched bars) at a multiplicity of infection (MOI) of 5 IFU for the time indicated. Open bars: BB19 cells alone (control cells). HBD-2 levels were normalized to GAPDH by using quantitative real-time PCR. Results are expressed as mean ± SD of the data from three experiments. Asterisks indicate a statistically significant difference between the data from the experimental test and the control test: *P < 0.01, **P < 0.001.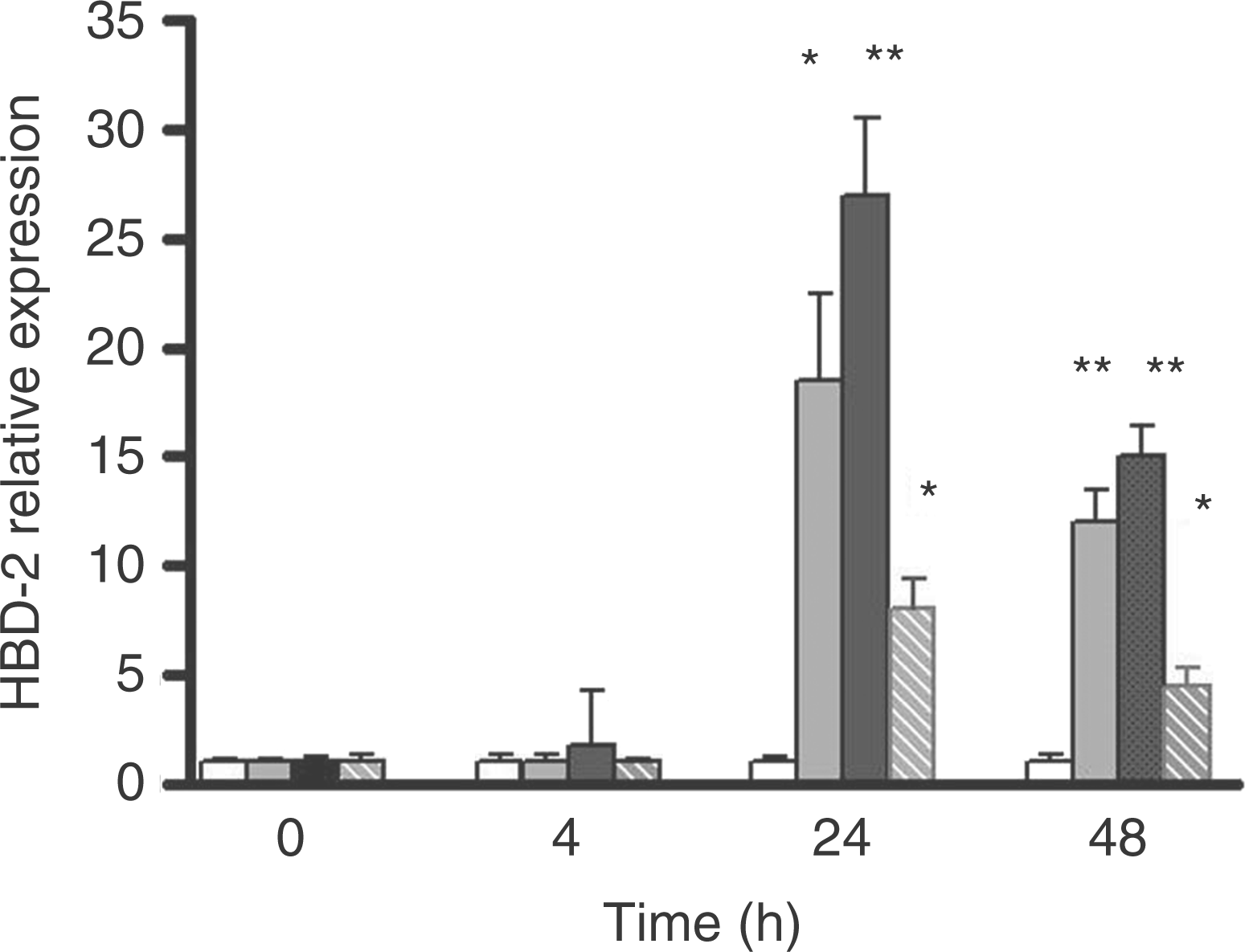
We compared the effects of heat-inactivated C. pneumoniae with those of viable C. pneumoniae. In cells stimulated with heat-inactivated C. pneumoniae, the expression of HBD-2 was induced, although at a reduced level (Fig. 1). Mock preparations did not result in any increase in HBD-2 mRNA level (data not shown).
We tested also the constitutive HBD-1 expression in BB19 cells. In contrast to HBD-2, there was no any increase in expression of HBD-1 mRNA following C. pneumoniae infection (data not shown).
Human β-defensin 2 protein expression in BB19 cells infected with C. pneumoniae
The expression of HBD-2 protein in C. pneumoniae-infected endothelial cells was determined by immunoblotting. Through use of the anti-HBD-2 antibody which binds to human HBD-2, a single band of about 4 kDa was detected. The signal was strong following the blotting of cell lysates from C. pneumoniae-infected cells. The level of HBD-2 was only barely detectable after the processing of the control cell lysate without C. pneumoniae infection (Fig. 2).
Immunoblotting analysis of HBD-2 peptide expression in C. pneumoniae-infected brain capillary endothelial cells. BB19 cell cultures were infected with C. pneumoniae at a MOI of 5 IFU for 24 h. Total cellular protein (25 g) from each culture was loaded on a 15% SDS-PAGE gel for immunblotting. After transfer, the membranes were incubated with anti-human goat HBD-2 antibody. Human β-defensin 2 was detected with ECL. Lane 1 relates to a culture without infection, Lane 2 to a culture with C. pneumoniae infection for 24 h. These results are representative of three experiments performed independently.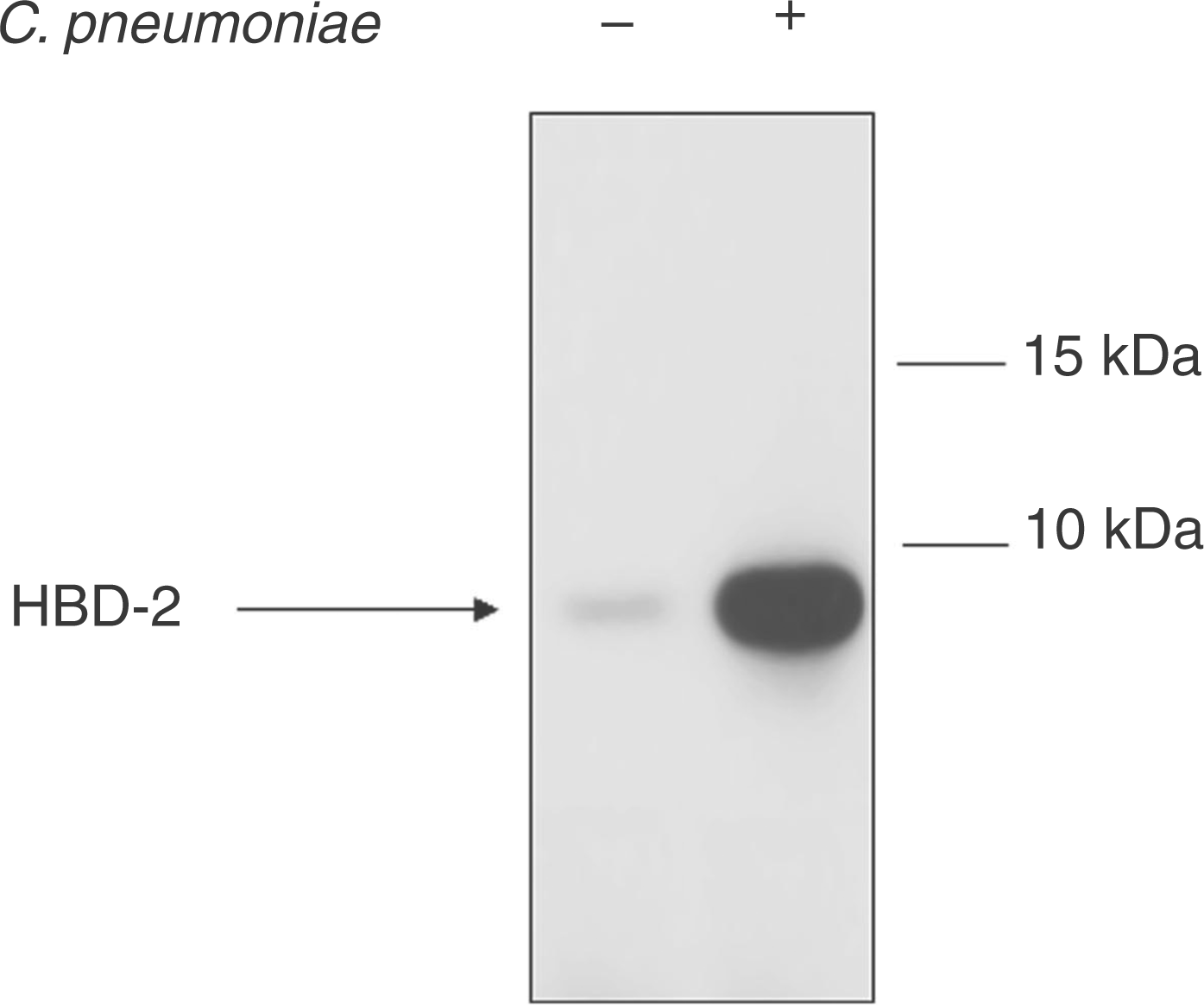
Immunofluorescent staining of BB19 cells for HBD-2
Immunofluorescent staining images of C. pneumoniae-infected BB19 cells revealed that HBD-2 was diffusely distributed as granules throughout the cytoplasm of the endothelial cells (Fig. 3). There was only a weak fluorescence in the uninfected controls. The calculated fluorescence intensity mirrored the ELISA (Fig. 4) and Western blotting results, indicating that the HBD-2 protein was highly inducible.
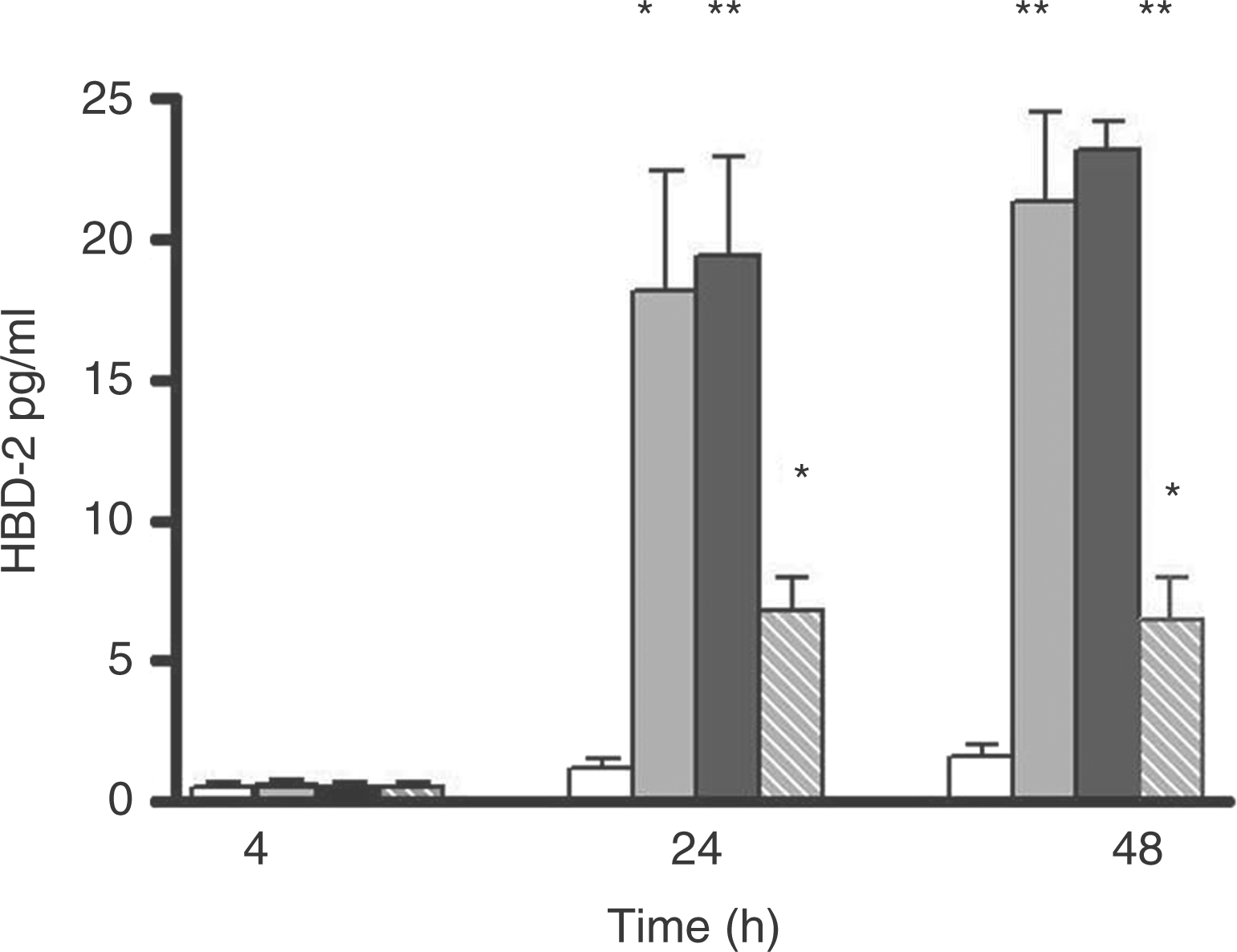
Immunofluorescence analysis of the HBD-2 peptide in BB19 cells. Cells were uninfected or infected with C. pneumoniae for 24 h. They were then harvested, and Cytospin preparations were stained for HBD-2 with goat anti-human HBD-2 antibody, and thereafter with secondary antibody (FITC labeled anti-goat antibody). Serial fluorescence images were captured by confocal microscopy and analyzed statistically. (A) Isotype control; (B) control cells without infection; and (C) cells infected with C. pneumoniae for 24 h. Kinetics of HBD-2 peptide production in culture supernatants from C. pneumoniae infected BB19 cells. BB19 cells were infected with C. pneumoniae CWL-029 (grey bars) or with C. pneumoniae CV6 (black bars) or with heat-inactivated C. pneumoniae CV6 (hatched bars) at a multiplicity of infection (MOI) of 5 IFU for the indicated periods and culture supernatants were harvested. Open bars: BB19 cells uninfected (control). The levels of HBD-2 were measured by ELISA. Values are mean + SD of the data from three independent experiments. Asterisks indicate a statistically significant difference between the data from the experimental test and the control test: *P < 0.01, **P < 0.001.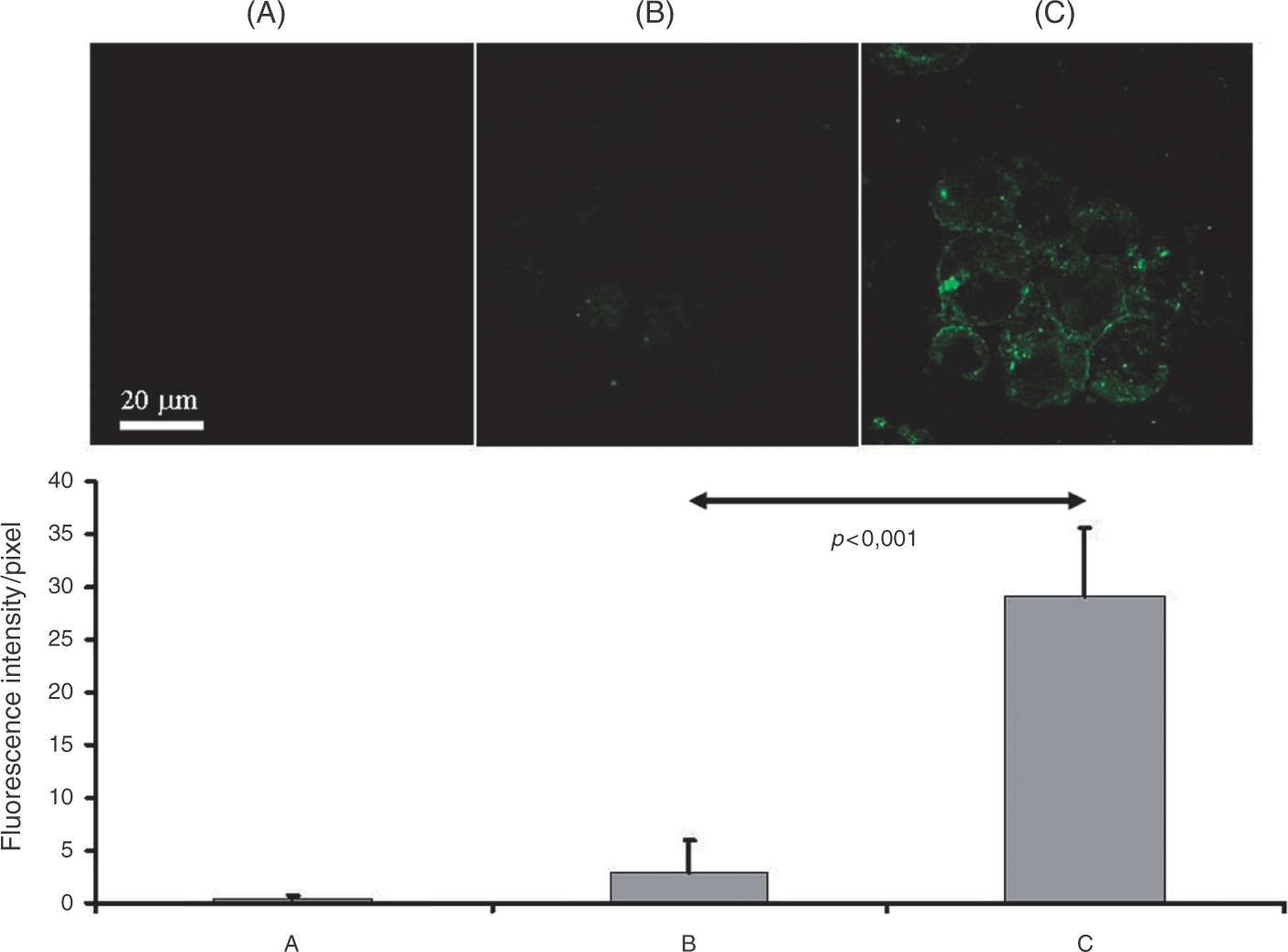
Chlamydophila pneumoniae induces HBD-2 secretion by BB19 cells
Experiments were performed to determine the HBD-2 protein secreted by BB19 cells into the tissue culture medium following C. pneumoniae infection. The amount of HBD-2 in the supernatants was determined by ELISA. Results of these experiments showed that the concentration of HBD-2 protein was significantly elevated in the supernatant 24 h after C. pneumoniae infection (Fig. 4). The supernatants from BB19 cells stimulated with heat-inactivated C. pneumoniae displayed a moderate release of HBD-2 peptide (Fig. 4). The BB19 cells incubated with mock stocks exhibited a similar level of HBD-2 release as that of the non-infected controls (data not shown). In parallel experiments, the concentration of HBD-2 was also determined in the cell lysates. A considerable increase in the amount of HBD-2 was detected in the cell lysates of cultures infected with C. pneumoniae CWL-029 or with C. pneumoniae CV6 for 24 h (35.2 ± 3.8 pg/ml and 48.4 ± 4.6 pg/ml, respectively, vs 5.5 ± 2.8 pg/ml in the control cell lysates).
These results imply that human endothelial cells stimulated by C. pneumoniae induce the expression of the HBD-2 gene and the release of HBD-2 protein into the medium.
Discussion
Expression of defensins in human endothelial cells in response to C. pneumoniae has not been studied previously. Sero-epidemiological studies in patients with stroke have provided evidence of an association of risk with prior infection with C. pneumoniae.10,11 The endothelial dysfunction observed in stroke patients has been related to the physiopathology of stroke, the stroke subtypes, the clinical severity and the outcome. 15 The present study was performed to analyze the expression of HBD-2 and the effects of C. pneumoniae on HBD-2 expression in the human brain capillary endothelial cell line BB19.
Human β-defensin 2, an antimicrobial peptide, has so far been found exclusively in a variety of epithelial cells. No data are available on HBD-2 expression in endothelial cells except for the very recent publication by Kawsar et al., 18 which reported the expression of HBD-2 in the intratumoral vascular endothelium and HUVECs (human umbilical vein endothelial cells).
In our study, the expression of HBD-2 mRNA in brain capillary endothelial cells was first assessed by RT-PCR. As mRNA may not always be translated to protein, we also investigated whether capillary brain endothelial cells express the HBD-2 peptide. The expression of the HBD-2 peptide in endothelial cells was confirmed by immunofluorescence staining for HBD-2 peptide and by Western blotting. The secretion of HBD-2 was confirmed by measuring the HBD-2 in the supernatants by ELISA. In good accordance with the inducible nature of HBD-2 in keratinocytes and epithelial cells, we observed a definite increase in the expression of HBD-2 upon exposure to C. pneumoniae.
In our experiments, heat-inactivated C. pneumoniae was still able to induce the expression and secretion of HBD-2, albeit at lower levels (Figs 1 and 4). This suggests that, among the potential products of bacteria, one candidate could be a heat-resistant lipopolysaccharide. On the other hand, heat–labile compounds, such as outer membrane proteins or other yet unidentified factors, might also be candidates for HBD-2 activation.
Infection of human endothelial cells with C. pneumoniae results in the stimulation of a wide variety of cytokines, adhesion molecules, chemokines and proteins with procoagulant activity,14,19 which furnishes evidence of the role of infection with this bacterium in the atherogenic process.20,21
The role of β-defensin peptide in endothelial cell biology is not clear. It might be that endothelial cells use defensins to exert antimicrobial activity. Defensins also exert a broad spectrum of immunological effects;7,8 human β-defensins may attract immature dendritic cells, naive T-cells and memory T-cells, 3 which suggests that these defensins are involved in orchestrating an immune response, 22 and it may be hypothesized that HBD-2 contributes to the possible innate immunity involved in atherosclerosis. Further studies are necessary to elucidate the role of HBDs in atherosclerosis or stroke.
The result demonstrating that HBD-2 is expressed and produced in the human brain capillary endothelial cells upon infection with C. pneumoniae is novel and provides evidence that HBD-2 plays a role in the early immune responses to C. pneumoniae and probably in the immunopathogenesis of atherosclerosis. This result might facilitate an understanding of the pathomechanism of the inflammation-mediated events of atherosclerosis or stroke in connection with C. pneumoniae infection.
Footnotes
Acknowledgements
The authors thank Mrs Györgyi Müller for expert technical assistance and Mrs Zsuzsanna Rosztoczy for skilful administration.
Funding
This work was supported by Hungarian Research Grant OTKA K67889/5K540 and ETT 019-05.
