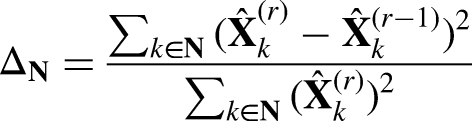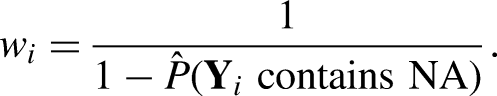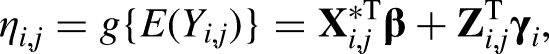Abstract
There is a significant public demand for rapid data-driven scientific investigations using aggregated sensitive information. However, many technical challenges and regulatory policies hinder efficient data sharing. In this study, we describe a partially synthetic data generation technique for creating anonymized data archives whose joint distributions closely resemble those of the original (sensitive) data. Specifically, we introduce the DataSifter technique for time-varying correlated data (DataSifter II), which relies on an iterative model-based imputation using generalized linear mixed model and random effects-expectation maximization tree. DataSifter II can be used to generate synthetic repeated measures data for testing and validating new analytical techniques. Compared to the multiple imputation method, DataSifter II application on simulated and real clinical data demonstrates that the new method provides extensive reduction of re-identification risk (data privacy) while preserving the analytical value (data utility) in the obfuscated data. The performance of the DataSifter II on a simulation involving 20% artificially missingness in the data, shows at least 80% reduction of the disclosure risk, compared to the multiple imputation method, without a substantial impact on the data analytical value. In a separate clinical data (Medical Information Mart for Intensive Care III) validation, a model-based statistical inference drawn from the original data agrees with an analogous analytical inference obtained using the DataSifter II obfuscated (sifted) data. For large time-varying datasets containing sensitive information, the proposed technique provides an automated tool for alleviating the barriers of data sharing and facilitating effective, advanced, and collaborative analytics.
Keywords
Introduction
Open science advocates for broad sharing of data, reproducible research, source code, and software tools. The benefits of open science on many aspects of our everyday lives are well documented.1–5 Along with the well-known exponential increase of the amount of newly acquired data, there is also an equally striking exponential decay of the value of data that is stored but not processed or shared.6,7 However, sharing data without loss of privacy is difficult, especially in medical and healthcare settings. In fact, 66% of the participants in the 2017 Health Information National Trends Survey were concerned about data privacy when health information is electronically exchanged. 8 For data sharing involving protected health information, organizations’ institutional review boards (IRBs) need to review the research before the required information can be retrieved from existing medical records and processed to extract valuable information. IRB’s initial review process may take up to 4 months. This process has significant variability depending on the type of review, for example, expedited, exempt, or full board reviews may take additional time, from 16 to 631 days. 9 In the United States, healthcare systems own the property rights for Electronic Health Records (EHR), and researchers have to bear the costs of data extraction and transfer under data use agreements. 10 Such regulation guarantees the protection of individual privacy rights but delays researchers’ ability to gain access to appropriate information, build models, and rapidly validate scientific discovery. This slows the knowledge transfer and impedes the translation of basic science discoveries into clinical practice. As a result, the restricted and significantly delayed access to data may limit the information utility for answering specific scientific questions. For example, in 2015, Keegan et al. 11 examined the relationship between ethnicity and short-term breast cancer survival using 2010 Kaiser Permanente Northern California EHR data. However, to obtain both the demographics and the cancer treatments for patients across facilities, they had to reduce the study cohort time frame from 8 to 3 years to have both datasets available and link them together, which significantly impacted the statistical power of this scientific investigation. Thus, there are enormous benefits in developing new statistical methods to facilitate secure and quick information exchange between data stewards and data science experts.
Three existing strategies provide secure mechanisms for modeling, processing, and interrogating sensitive cross-sectional data. These include secure enclave access, data encryption (e.g. fully homomorphic encryption), and synthetic data publishing. First, secure data enclave environments12,13 offer a platform for researchers to analyze sensitive data without compromising risks for misuse and other violations. Many health information storage solutions, for example, EHRs, rely on technology that provides managed data access for research in safe and controlled environments. 14 Several possible unmodified database management systems can be utilized to provide secure data enclaves.15,16 Second, data encryption methods, including fully homomorphic encryption (FHE), encode the data to allow computations directly on the resulting ciphertext.17–19 FHE relies on homomorphic computing (result-preserving property) on the ciphertext without exposing the sensitive raw data to independent researchers. The above two mechanisms provide secure channels for data transfer and storage, however, these approaches do not shorten the data sharing process and limit the type of advanced analytics that can be used to interrogate the data.
To address these limitations, the third strategy, synthetic data generation, provides “fake” records that closely resemble the real data for predictive and inference model constructions. Both the computer science and statistical science communities have developed various methods in this field. Most of the methods developed by computer science communities fall under the umbrella of differential privacy.20,21 For quantitative data types, this class of methods creates fully synthetic data for predictive purposes, where noisy conditional models generate each synthetic record.22–24 The most recent developments in this class are related to generative adversarial networks (GANs). 25 They generate candidate synthetic records and use a discriminator that accepts only differentially private data to provide privacy protection guarantee.23,26 To the best of our knowledge, there is less work related to generating data for model-based inference purposes in the differential privacy framework. A recent work 27 proposed HealthGAN for generating fully synthetic data for privacy preservation and inference purposes. They defined data utility and privacy using Euclidean distances between original and fully synthetic records, which is not guaranteed to be differentially private. The authors reported that seven out of 29 confidence intervals estimated using the synthetic data did not actually contain the original data. Moreover, the HealthGAN cannot handle time-varying data elements. In the statistical science community, synthetic data generation was first proposed by Rubin 28 with two classes of generating methods, as summarized by Reiter and Raghunathan. 29 The fully synthetic data sets are created by conditional distributions estimated from sensitive datasets. Popular methods for constructing these conditional distributions include multiple imputation (MI).28,30 Also, there are Bayesian data augmentation methods like SynSys, 31 which aims to enrich the existing data to train machine learning predictive models. However, since the distributions of subject characteristics are not considered in the data generation procedure, fully synthetic data may not represent the original patient population. The other class of methods creates partially synthetic data to alleviate this issue, which utilizes a set of multiple-imputed data replacing sensitive data value cells with imputations.32–34 This class of methods treats data obfuscation as a missing data handling problem, where they generate artificial missingness for sensitive values in the data set and impute the value with the remaining untouched data. As a result, partially synthetic data provide valid statistical inference. Yet, combined information from a set of multiple imputed datasets indicates the locations of true and obfuscated cells resulting in no privacy protection for the true cells. In practice, covering all possible sensitive values is barely achievable and selecting the obfuscation location is a subjective and critical step for data privacy protection.
The DataSifter technique (DataSifter I) was recently introduced 35 as an automated procedure for generating partially synthetic data. This approach offers better protection of participant privacy in the case of using high-dimensional sensitive cross-sectional data. The DataSifter framework is designed to help data governors (institutions who possess and manage data) safely share synthetic subsets of their sensitive data archives using customized levels of statistical obfuscation. It perturbs individual-level records and allows researchers to acquire population-level information that closely approximates the true signal securely. The core DataSifter technique relies on two processes supporting the critical statistical obfuscation of the data. First, it randomly and artificially generates missingness in the data, following the Missing Completely At Random (MCAR) mechanism, 36 and uses robust iterative imputation methods, for example, missForest, 37 to approximate the original information. After the first step, DataSifter I creates a partially synthetic dataset disguising the locations of true values, which provides better privacy protection than the MI method. Second, DataSifter I computed cluster neighborhoods using Euclidean and Gower distances for continuous and categorical variables, respectively. Then, within each neighborhood cluster, DataSifter I randomly swaps a subset of feature values between similar records. This second operation guarantees partial obfuscations within each record while preserving the holistic cohort distribution in the variable space.
The quality of generated partially synthetic data can be quantified by disclosure risk and the deviance of model-based inference. The disclosure risk describes the privacy-awareness of the synthetic data. There are two classes of disclosure risk criterion, namely
Time-varying correlated data, including longitudinal data, are ubiquitous and provide valuable information for many biomedical and health conditions. For example, in EHR databases, patient characteristics and disease progression variables are collected repeatedly across visits. Maintaining or preserving the within-subject covariance structure among time-varying measurements presents another layer of challenges. To the best of our knowledge, to date, there are no automated procedures that enable secure sharing of sensitive time-varying correlated data using partially synthetic data. The MI methods tilt the balance towards data utility, and the DataSifter I method cannot handle time-varying data elements. We propose a new algorithm, DataSifter with Time-varying Correlated Data (DataSifter II), that extends the DataSifter functionality in the case of dealing with large, cross-sectional, time-varying, and self-correlated data elements. DataSifter II introduces artificial missingness and embeds robust longitudinal imputation methods to handle high-dimensional sensitive data with time-varying measures. As illustrated in Figure 1, our proposed procedure operates on the time-varying data separately from the time-invariant (cross-sectional) data elements and then integrates the two parts to compile the obfuscated sifted dataset (output). Our extensive simulation and application studies show that compared to the MI method, the proposed method can better protect data privacy while preserving a proper level of data utility.

Graphical workflow depicting the organization of the DataSifter Time-varying Measurements (DataSifter II).
The contribution of our paper is twofold: (1) we propose an automated algorithm for time-varying partially synthetic data generation allowing user-specified secure level and protecting the original within-subject covariance structure, and (2) we formally define practical data privacy and data utility measurements to validate synthetic data before release. The rest of the manuscript is organized as follows: In section “Privacy and utility measurement for partially synthetic data,” we define the data privacy and utility measurement for partially synthetic data evaluation. Specifically, in section “Data structure and notations,” we define the disclosure risk and show that partially synthetic datasets generated by the DataSifter framework provide better privacy protection than that of the MI method. Section “DataSifter II technique” describes the DataSifter II protocol. Section “Simulation studies” validates the data utility preservation and privacy protection of the proposed algorithm under different simulation settings and compares the performance against the MI method. In section “Biomedical application,” we apply DataSifter II to the Medical Information Mart for Intensive Care III (MIMIC-III) clinical data and demonstrate its performance in maintaining a careful balance between protecting sensitive information and preservation of the data utility. We summarize the findings and discuss the expected impact and future developments in section “Discussion.”
Privacy and utility measurement for partially synthetic data
Data structure and notations
Time-varying correlated data are common in most biomedical and epidemiology studies. For example, in multi-center studies, we typically measure the target variables across all subjects at a single time point, but the subjects may be correlated within each center. In longitudinal data, the target variables are measured repeatedly at baseline and during follow-up, and thus we have intrinsic within-subject correlations. In this case, researchers take repeated measurements on the same subjects to reduce measurement errors, which may also involve within-subject correlations. The DataSifter II framework can be applied to any correlated data. For illustration purposes in this study, we investigate the use of DataSifter II on longitudinal data.
Consider a longitudinal EHR dataset with
Missing data occurs often in longitudinal observations. Missingness can come from a completely missing record or partially missing record, where patient
While generating partially synthetic data, we view data obfuscation as an artificial missing creation and imputation procedure. Here we focus on obfuscating time-varying data and further denote each row in partially synthetic time-varying data as
Data privacy measurement
In this section, we formally define data privacy in the form of disclosure risk and compare the disclosure risk between MI and DataSifter methods. Assume we have a
Data governors can quantify the privacy protection level of the synthetic longitudinal data using our disclosure risk defined above. Specifically, when calculating the maximal privacy loss for each record, we assume the intruder knows all other records in the raw dataset, that is,
Data utility measurement
Given a pre-specified model, we can obtain the desired utility of the partially synthetic data by comparing the model fitted with the original and synthetic data in terms of model inference and prediction accuracy. The data governor can consider a feasible parametric model regressing a summary variable on other covariates. For example, in EHR data, we can predict patients’ comorbidity scores over time, representing summary scores for the patients’ medical conditions. For model inference, to test the data utility based on a regression coefficient
Many alternative performance metrics can be defined to track the utility of the DataSifter-generated synthetic datasets. For example, the Akaike information criterion, which is commonly used to contrast the relative quality of statistical models for a given set of data, provides one strategy for comparing the overall quality of the models constructed from original and synthetic datasets. We will focus on the parameter-level utility measurement for the rest of the paper.
Evaluating the performance of partially synthetic datasets
We can assess the performance of multiple partially synthetic datasets or select the desired parameters for a synthetic data generation process based on the data privacy and utility measurements defined above. Practically, we can first define a minimal average PM level for selected or all data cells as the threshold for data privacy. Then, we compare the data utility among the qualified datasets in terms of the confidence interval coverage or the confidence interval overlap probabilities for different model parameters and model prediction accuracy. Datasets with higher PM, higher confidence interval coverage or confidence interval overlap, and lower model prediction accuracy reduction are preferable.
DataSifter II technique
The proposed DataSifter II procedure operates on static variables
The main assumptions of the DataSifter II include (A1) the possible missingness mechanisms in the original data include missing at random (MAR) or missing completely at random (MCAR); and (A2) the sensitivity of each variable is equally important. There are three possible missing mechanisms. MCAR assumes that missingness is unrelated to any observed or unobserved variables. Under this missing mechanism, the subset of complete records without missingness is representative of the study population without selection bias. However, MCAR happens rarely. MAR is more plausible to occur such that missingness can be accounted for with observed data. Otherwise, missing not at random may occur, where analysis performed on the complete portion of data can suffer from selection bias. We consider the (A1) assumption to hold in the original dataset to provide sufficient data utility for synthetic data generation. Under this assumption, MAR or MCAR guarantees unbiased missing imputation, ensuring the obfuscation quality of DataSifter II. (A2) allows indistinguishable obfuscation for each variable and greatly simplifies the obfuscation procedure. We can further adjust the procedure of missingness assignment for specific scenarios where the data governors believe a pre-defined set of variables can be more sensitive than the rest.
Sifting static variables with DataSifter I
We apply DataSifter I to obfuscate the static portion
The imputation procedures in DataSifter aim to create a single complete dataset disguising the original or artificial missing positions. We use the missForest technique that outputs a single imputed dataset and is proven to have smaller imputation errors than common methods, including MI.37,42 This non-parametric imputation technique sequentially imputes and updates the data by variable. In the first iteration, when imputing for the first target variable, it fills in the missing cells among other predictor variables with mean imputation. Then, it constructs random forest models (target variable versus all other variables) to provide imputations. In subsequent iterations, while imputing and updating by variable, the imputation accuracy for each target variable improves as the missing cells in all other variables are replaced with better predictions.
When imputing the original missing data, we employ the original stopping criterion in missForest. It stops to iterate when the difference between the latest and prior imputed data matrix is at least as great as the previous difference measured or the maximal iteration limit has been achieved. The difference between matrices in sets of continuous (
When imputing artificial missingness, the true missing value is known. We define the stopping criterion under a tolerance level
Iterative imputation algorithm for time-varying data
Although DataSifter I applies robust nonparametric imputation methods like missForest to impute static missing variables, effective missing imputation for time-varying data can be challenging. In this paper, we propose an iterative imputation algorithm for longitudinal data similar to the missForest algorithm. The proposed algorithm considers two types of missing mechanisms (MAR and MCAR) and two modeling options (linear mixed model and RE-EM tree). It handles missingness in time-varying variables
Before the imputation, we initiate all missing cells with the closest value from the same subject (last value carry forward or next value carry backward). If the subject has no observations of certain variables, we initialize such missing cells with mean imputations. Then, we sort the variables ascendingly based on missing percentage so that
Imputation model under missing at random assumption
Under different missing patterns, the algorithm utilizes different imputation models. When we have MAR, the missingness depends on observed data and the complete observations might be biased. We utilize inverse probability weighting to obtain an unbiased pseudo sample for imputation model fitting. By pseudo sample, we mean the weighted sample that creates balance by up-weighting the underrepresented population and down-weighting the over-represented population in the complete observations, where the weights can be calculated at the subject-level, or subject-and-time-level, to allow better imputation under different situations. For subject-level, the probability of missing for each subject denoted as
Accordingly,
Imputation model under MCAR
Under MCAR, we propose to employ two modeling options GLMM 43 or RE-EM tree 44 as imputation models within the iterative procedure. The two procedures are referred to as DataSifter II GLMM and DataSifter II RE-EM. Note that GLMM can handle various data types, including continuous, binary, and count data, whereas the RE-EM tree is an effective algorithm for continuous measurements. For the DataSifter II GLMM, variable selection is conducted separately with GLMM LASSO. Here we perform a grid search for the regularization parameter and use Bayesian information criterion to select the best model. Since an appropriate starting point is crucial for model convergence, DataSifter II incorporates two methods to initiate the parameters when fitting GLMM LASSO. The first method estimates all the parameters by glmmPQL, which is using pseudo-likelihood. GlmmPQL estimates the mean and variance parameters iteratively with maximum likelihood assuming normality. 45 It approximates the true likelihood with a strong normality assumption, but provides a computationally efficient way of estimating initial regression parameters. When the signal is sparse and the GLMM algorithm does not converge with the glmmPQL initial values, we may consider initialization using zeros or another user-specified initialization.
Selected variables are denoted as
After estimating
DataSifter II RE-EM provides an alternative robust obfuscation for continuous time-varying measurements. RE-EM tree model combines the tree-based estimation for fixed effects and parametric estimation for random effects.
44
RE-EM tree is a semi-parametric generalization of the linear mixed effect model
Create partially synthetic time-varying data
Using the proposed iterative imputation tool, we can create partially synthetic time-varying data by handling potential missing in
Similar to the preprocessing step for static variables, we intend to initiate the process with a complete dataset containing both static and time-varying data. The working complete static data
Following the initial imputation, we start Sifting by introducing artificial random missing values to the longitudinal variables in the working complete dataset. Such randomly generated missingness follows an MCAR missing mechanism, which guarantees that the unweighted complete-case analysis is bias-free. We then impute the missing variables one by one with the proposed imputation procedure under MCAR with a data-driven choice of either the parametric or semi-parametric imputation model.
Implementation of DataSifter II
We use Algorithm 1 to summarize the proposed imputation method for time-varying variables. The algorithm terminates the imputation for variable
Time-varying data missing imputation algorithm.

Outlines the complete DataSifter II implementation method.

Residual diagnostics
The residuals or errors introduced by DataSifter II obfuscated values follow a mixture distribution. When the final imputation model for variable
Simulation studies
Simulation setup
In this section, we conduct controlled simulation studies to evaluate the data privacy and utility protection of DataSifter and MI methods. The original simulation data is generated with
Under the linear association, 

We also consider cases with non-linear relationships. Similar to the linear setting, we construct models with a compound symmetry correlation structure. Our two longitudinal variables are derived by

The complete data generated by the above procedure will be used to examine different data obfuscation methods. To mimic real-world data, we also consider a scenario where some observations in 
We compared the performance of four types of synthetic datasets: DataSifter with GLMM generated on complete original data, DataSifter with RE-EM tree generated on complete original data, multiple imputed synthetic data generated on complete original data, and DataSifter with RE-EM tree generated on original data that contains missing. All the sifted data are generated without static data obfuscation (
Simulation results
Using the proposed data utility and data PM, we evaluate synthetic datasets generated by DataSifter and MI. Data utility is measured in terms of prediction accuracy and inference based on models trained on synthetic datasets. For prediction accuracy, we construct test sets with identical sample size as the training dataset (
The average prediction errors on test data are summarized in Table 1. Based on 100 replications, under most simulation settings, the average test error on
Mean absolute deviation (prediction error) for test dataset based on the model fitted on original and synthetic datasets. The test datasets are generated separately with the same sample size as the training sets.
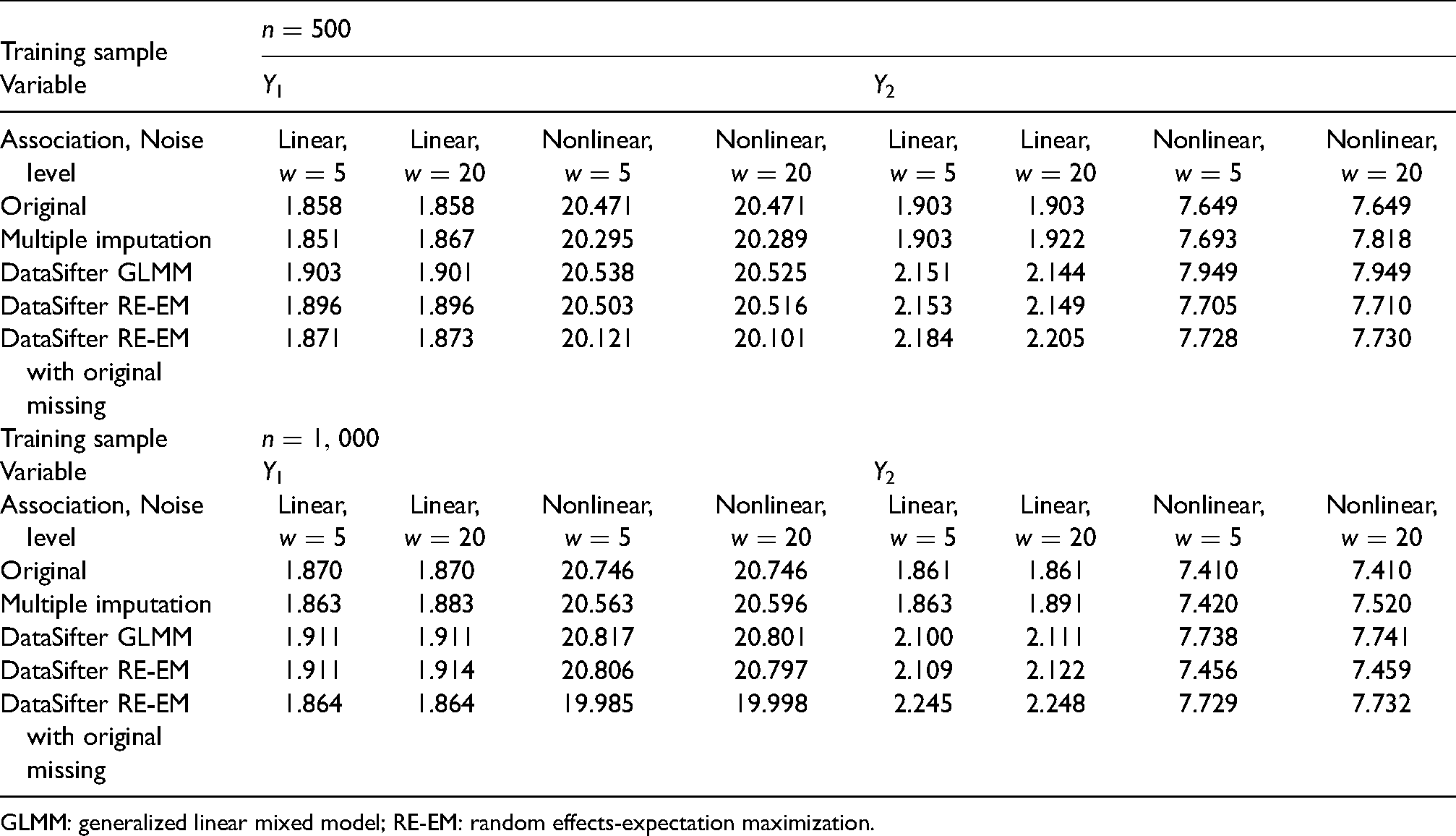
GLMM: generalized linear mixed model; RE-EM: random effects-expectation maximization.
Since the proposed imputation method is aimed at minimizing imputation error rather than accounting for the uncertainty of the missing values, the 95% confidence intervals constructed on sifted datasets are narrower than the original data. As shown in Table 2, for variable
Model confidence interval (95%) coverage among 100 replicates. The coverage records the percent of times that CIs from models trained on original and synthetic datasets cover the true parameter estimate.

CI: confidence interval; GLMM: generalized linear mixed model; RE-EM: random effects-expectation maximization.
The average PM for different synthetic datasets is illustrated in Figure 2. Each boxplot records the distribution of average PM for

Average privacy measurement among first 100 rows in the synthetic datasets. The scenario with small noise level contains
Biomedical application
The MIMIC-III represents a sizable single-center database that provides patients’ medical records in a large tertiary care hospital between 2001 and 2012. MIMIC-III data stores information related to patients’ admission, including vital signs, medications, laboratory measurements, length of stay, survival data, and more. 48 We consider a subset of 7080 patients who had at least two visits to the hospital who contributed 17,594 hospital admission records with demographic variables, including insurance type, gender, race, age, marital status, and death after admission. Admission information such as insurance type, admission type, and month of admission is also available. MIMIC-III contains de-identified or coded data that is considered free of protected health information. However, the data request process, including taking an online course and submitting an application with specific research topics and requested information, can still take more than three weeks. Using the data for any rigorous scientific investigation requires the researcher to go through a time-consuming data request procedure, while in the end, the investigation may find no significant results. DataSifter II allows a quicker turnaround for checking the potentials of research hypotheses. For example, we want to investigate the association between length of stay in tertiary care hospitals and Medicaid insurance type controlling other patient demographic variables using the MIMIC-III data. We illustrate how to use the sifted data to answer our initial research question and evaluate both utility and privacy protection performance in the sifted MIMIC-III data. We also compare the synthetic MIMIC-III datasets generated by DataSifter II with that of the MI method. A linear mixed effect model is used to regress the length of hospital stay on patient characteristics. The privacy protection effort is measured by the privacy measurement (PM) for age and length of hospital stay among the first 100 records.
First, we obfuscate the following longitudinal variables: (1) length of stay, (2) month of admission, (3) death after visit, and (4) age at visit. Next, we consider generating two types of Sifted data: with (
Next, we compare the model parameter estimates between models fitted on the original data and on the three different types of synthetic data, assuming the following linear mixed effect model:
Results in Figure 3 show that the DataSifter II provides much better privacy protection than the MI method with a small loss in data utility. Medicaid is not associated with the length of stay in any synthetic and original data fitted models. According to Figure 3(A), most of the mean PM by row (record) are below five among the 50 multiply imputed synthetic datasets. The average PM is 0.33 for age at admission and 1.53 for the length of hospital stay. The DataSifter method provides a significant improvement in terms of PM with 14.87 and 8.17 as the average PMs for age and hospital stay, respectively. We can also infer from Figure 3(A) that the mean PMs can vary considerably from row to row in sifted datasets without static obfuscation.

MIMIC III synthetic data privacy (A) and utility (B) evaluation. Plot A summaries the distribution of mean privacy measurement for age and length of hospital stay for the first 100 rows across 50 synthetic datasets generated by DataSifter (without static obfuscation using DataSifter I) and multiple imputation. Plot B compares the significant coefficient estimates (p-value < 0.05) among the models fitted with original data, and synthetic datasets generated by DataSifter II (with or without static obfuscation) and multiple imputation. The boxes illustrate the distribution of coefficients estimated on 50 synthetic datasets. The black dots and purple intervals are the parameter estimates and confidence intervals from the linear mixed model fitted by the original dataset.
Figure 3(B) illustrates the deviance of parameter estimates between the model fitted with three types of synthetic dataset and the original linear mixed model. Only significant parameter estimates are shown in the plot. The box plots represent the empirical CIs or the distribution of
Since none of the fitted models obtained from the sifted datasets show that
Discussion
The reported results shown above demonstrate that the DataSifter with Time-varying Correlated Data (DataSifter II) technique balances between maintaining the energy of the original data (preserves information or data utility) while simultaneously introducing a level of privacy protection safeguarding against re-identification of sensitive information contained in the archive. The simulation results based on introducing 20% artificial missingness suggest that data utility is better preserved for longitudinal variables that depend only on static variables (
The goal of the proposed algorithm is to support preliminary hypothesis testing on privacy-aware partially synthetic datasets. DataSifter II does not provide the best data privacy protection like differentially private algorithms or the best data utility preservation like MI. We aim to balance the level of perturbation introduced to the original data and the quality to provide statistical inference. This is to ensure maximal data utility under different levels of privacy protection needs. For example, a synthetic dataset for internal use requested by a trusted research institute might have a weaker privacy protection requirement than a synthetic dataset for educational purposes. Accurate quantification of both criteria is critical for achieving the goal. Hence, we proposed and applied the PM in this paper that provides record-level disclosure risk of the longitudinal synthetic data, assuming the intruder has maximal prior knowledge about the original data. Data utility corresponding to each level of obfuscation is then examined by the deviance in inference based on a pre-defined model.
Section “Data utility measurement” mentioned that the data governor could consider using a pre-defined parametric model for utility purposes, such as regressing summary variables like comorbidity score on other variables. One might argue that one model does not cover all inference-related use cases. In practice, we can propose multiple parametric models for better inference testing. It could be challenging to propose a comprehensive set of testing models for general EHR related to numerous diseases. However, when the data request is explicit about a specific disease, the data governor may consider models related to the context.
The data obfuscation for static variables was done by the original DataSifter I method, which includes the per record swapping operation ensuring record-level perturbation (some of the data cells among the static variables are altered) for any non-isolated subject. Our application shows that this extra level of protection on static variables only introduces a small bias in model-based statistical inference. It is almost impossible to apply the swapping operation on time-varying variables without damaging the within-subject correlation. As a result, some of the time-varying variables for a specific record might be untouched in the synthetic dataset due to chance when the percent of artificial missingness is small. We can consider multiple iterations of the DataSifter II operation and adaptively assign artificial missing data to guarantee obfuscation in each record’s time-varying variables.
Depending on the specific data characteristics, the DataSifter II performance may be impacted in terms of its efficiency and balance between privacy protection and utility reservation. We employed a linear (GLMM) model and a tree (RE-EM tree) structured model to approximate the distribution for each longitudinal variable conditional on other variables in the data. The utility preservation for each longitudinal variable may be affected by (1) the complexity of the relationship, (2) empirical variance of the target time-varying variable, (3) the data type of the predictors, and (4) alternative within-subject covariance structures.
The DataSifter II algorithm provides data governors and researchers with a semi-automated and reliable framework for sensible information exchange. Despite perturbing individual-level records, the overall sifted time-varying data shares similar population-level information with the original process. DataSifting allows data owners to create pseudo populations with a custom level of obfuscation, meeting different data sharing needs. This method provides a rapid and effective information exchange process facilitating research hypotheses testing (confirmatory analytics) and data-driven discovery (exploratory analytics). Many biomedical research and development partnerships may benefit from the DataSifter II technology to conduct advanced trans-disciplinary research and translate fundamental science advances into clinical practice. The proposed algorithm can also be useful beyond the health and biomedical domains, which in this study represent the primary target application areas. For instance, statistical obfuscation and generation of longitudinal synthetic data could be very useful in studies of insurance policies and claims, census records, geopolitical information, social justice, and environmental data. Many dynamic processes might contain protected personal, regulated governmental, or IP organizational information, which require guarding against inappropriate (mis)use. In such cases, DataSifter II can be utilized to simulate realistic records to meet different requirements for data privacy, security, and utility in various applications. We have shared the DataSifter II R package on our GitHub repository https://github.com/SOCR/DataSifterII.
Footnotes
Acknowledgements
We thank the Colleagues at the Michigan Institute for Data Science (MIDAS), the Michigan Neuroscience Graduate Program (NGP), and the Statistics Online Computational Resource (SOCR) who provided valuable suggestions and constructive recommendations. Special thanks to Lu Wei and Brandon Cummings at the University of Michigan for productive discussions, ideas, and contributions.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported in part by the National Science Foundation (NSF) grant nos. 1916425, 1734853, 1636840, 1416953, 0716055, and 1023115, and by the National Institutes of Health (NIH) grant nos. P20 NR015331, U54 EB020406, P50 NS091856, P30 DK089503, UL1TR002240, R01CA233487, R01MH121079, R01MH126137, and T32GM141746.
Supplemental material
Supplemental material for this article is available online.