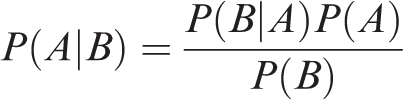Abstract
Breast cancer is the second most leading cancer occurring in women compared to all other cancers. Around 1.1 million cases were recorded in 2004. Observed rates of this cancer increase with industrialization and urbanization and also with facilities for early detection. It remains much more common in high-income countries but is now increasing rapidly in middle- and low-income countries including within Africa, much of Asia, and Latin America. Breast cancer is fatal in under half of all cases and is the leading cause of death from cancer in women, accounting for 16% of all cancer deaths worldwide. The objective of this research paper is to present a report on breast cancer where we took advantage of those available technological advancements to develop prediction models for breast cancer survivability. We used three popular data mining algorithms (Naïve Bayes, RBF Network, J48) to develop the prediction models using a large dataset (683 breast cancer cases). We also used 10-fold cross-validation methods to measure the unbiased estimate of the three prediction models for performance comparison purposes. The results (based on average accuracy Breast Cancer dataset) indicated that the Naïve Bayes is the best predictor with 97.36% accuracy on the holdout sample (this prediction accuracy is better than any reported in the literature), RBF Network came out to be the second with 96.77% accuracy, J48 came out third with 93.41% accuracy.
Introduction
The number and the size of databases recording medical data are increasing rapidly. Medical data, produced from measurements, examinations, prescriptions, etc., are stored in different databases on a continuous basis. This enormous amount of data exceeds the ability of traditional methods to analyze and search for interesting patterns and information that is hidden in them. Therefore, new techniques and tools for discovering useful information in these data depositories are becoming more demanding. 1 Analyzing these data with new analytical methods in order to find interesting patterns and hidden knowledge is the first step in extending the traditional function of these data sources.
Breast cancer
The organs and tissues of the body are made up of tiny building blocks called cells. Cancer is a disease of these cells. Although cells in each part of the body may look and work differently, most repair and reproduce themselves in the same way. Normally, cells divide in an orderly and controlled way. But if for some reason the process gets out of control, the cells carry on dividing and develop into a lump called a tumour. Breast tumours are usually caused by an overgrowth of the cells lining the breast ducts. They can be either benign or malignant. In a benign tumour, the cells grow abnormally and form a lump. But they do not spread to other parts of the body and so are not cancers. The most common type of benign breast tumour is called a fibroadenoma. This may need to be surgically removed to confirm the diagnosis. No other treatment is necessary. In a malignant tumour, the cancer cells have the ability to spread beyond the breast if they are left untreated. For example, if a malignant tumour in the breast is not treated, it may grow into the muscles that lie under the breast. It can also grow into the skin covering the breast. Sometimes cells break away from the original (primary) cancer and spread to other organs in the body. They can spread through the bloodstream or lymphatic system. When these cells reach a new area they may go on dividing and form a new tumour. The new tumour is often called a secondary or metastasis. Breast cancer occurs when cells within the breast ducts and lobules become cancerous. If caught at an early stage, breast cancer can often be cured. If the cancer has spread to other areas of the body it cannot usually be cured, but it can normally be effectively controlled for a long time.
Risk factors associated with breast cancer
Every woman wants to know what she can do to lower her risk of breast cancer. Some of the factors associated with breast cancer
2
:
Genetics: About 5% to 10% of breast cancers are thought to be hereditary, caused by abnormal genes passed from parent to child. Personal history of breast cancer: If you have been diagnosed with breast cancer, you are three to four times more likely to develop a new cancer in the other breast or a different part of the same breast. This risk is different from the risk of the original cancer coming back (called risk of recurrence). Radiation to chest or face before age 30: If you had radiation to the chest to treat another cancer (not breast cancer), such as Hodgkin’s disease or non-Hodgkin’s lymphoma, you have a higher-than-average risk of breast cancer. If you had radiation to the face at an adolescent to treat acne (something that is no longer done), you are at higher risk of developing breast cancer later in life. Certain breast changes: If you have been diagnosed with certain benign (not cancer) breast conditions, you may have a higher risk of breast cancer. There are several types of benign breast conditions that affect breast cancer risk. Race/ethnicity: White women are slightly more likely to develop breast cancer than African American, Hispanic, and Asian women. But African American women are more likely to develop more aggressive, more advanced-stage breast cancer that is diagnosed at a young age. Being overweight: Overweight and obese women have a higher risk of being diagnosed with breast cancer compared to women who maintain a healthy weight, especially after menopause. Being overweight also can increase the risk of the breast cancer coming back (recurrence) in women who have had the disease. Pregnancy history: Women who have not had a full-term pregnancy or have their first child after age 30 have a higher risk of breast cancer compared to women who gave birth before age 30. Breastfeeding history: If a woman breastfeeds for longer than one year this may reduce breast cancer risk. Menstrual history: Women who started menstruating (having periods) younger than age 12 have a higher risk of breast cancer later in life. The same is true for women who go through menopause when they are older than 55. Using HRT (hormone replacement therapy): Current or recent past users of HRT have a higher risk of being diagnosed with breast cancer. Drinking alcohol: Research consistently shows that drinking alcoholic beverages – beer, wine, and liquor – increases a woman’s risk of hormone-receptor-positive breast cancer. Having dense breasts: Research has shown that dense breasts can be six times more likely to develop cancer and can make it harder for mammograms to detect breast cancer. Lack of exercise: Research shows a link between exercising regularly at a moderate or intense level for 4 to 7 h per week and a lower risk of breast cancer. Smoking: Smoking causes a number of diseases and is linked to a higher risk of breast cancer in younger, premenopausal women. Research also has shown that there may be link between very heavy second-hand smoke exposure and breast cancer risk in postmenopausal women. Low of vitamin D levels: Research suggests that women with low levels of vitamin D have a higher risk of breast cancer. Vitamin D may play a role in controlling normal breast cell growth and may be able to stop breast cancer cells from growing. Light exposure at night: The results of several studies suggest that women who work at night – factory workers, doctors, nurses, and police officers, for example – have a higher risk of breast cancer compared to women who work during the day. DES (diethylstilbestrol) exposure: Women who took DES themselves have a slightly higher risk of breast cancer. Women who were exposed to DES while their mothers were pregnant with them also may have slightly higher risk of breast cancer later in life. Eating unhealthy food: Diet is thought to be at least partly responsible for about 30% to 40% of all cancers. No food or diet can prevent you from getting breast cancer. Exposure to chemicals in cosmetics: Research strongly suggests that at certain exposure levels, some of the chemicals in cosmetics may contribute to the development of cancer in people. Exposure to chemicals in food: There is a real concern that pesticides, antibiotics, and hormones used on crops and livestock may cause health problems in people, including an increase in breast cancer risk. There are also concerns about mercury in seafood and industrial chemicals in food and food packaging. Exposure to chemicals for lawns and gardens: Research strongly suggests that at certain exposure levels, some of the chemicals in lawn and garden products may cause cancer in people. But because the products are diverse combinations of chemicals, it is difficult to show a definite cause and effect for any specific chemical. Exposure to chemicals in plastic: Research strongly suggests that at certain exposure levels, some of the chemicals in plastic products, such as bisphenol A (BPA), may cause cancer in people. Exposure to chemicals in sunscreen: While chemicals can protect us from the sun’s harmful ultraviolet rays, research strongly suggests that at certain exposure levels, some of the chemicals in some sunscreen products may cause cancer in people. Exposure to chemicals in water: Research has shown that the water you drink – whether it is from your home faucet or bottled water from a store – may not always be as safe as it could be. Everyone has a role in protecting the water supply. Exposure to chemicals when food is grilled/prepared: Research has shown that women who ate a lot of grilled, barbecued, and smoked meats and very few fruits and vegetables had a higher risk of breast cancer compared to women who did not eat a lot of grilled meats.
The main purpose of this research work involves methodology that starts with understanding the domain, locating proper data sources, preparing the raw data, applying advanced analysis techniques, and extracting and validating the resulting knowledge for breast cancer survivals.
Related work
Several studies have been reported that have focused on breast cancer survivals. These studies have applied different approaches to the given problem and achieved high classification accuracies. Details of some of the previous research works are given in the following:
Liu et al. 3 used decision table (DT)-based predictive models for breast cancer survivability, concluding that the survival rate of patients was 86.52%. They employed the under-sampling C5 technique and bagging algorithm to deal with the imbalanced problem, thus improving the predictive performance on breast cancer.
Tan and Gilbert 4 demonstrated the usefulness of employing ensemble methods in classifying microarray data and presented some theoretical explanations on the performance of ensemble methods. As a result, they suggest that ensemble machine learning should be considered for the task of classifying gene expression data for cancerous samples.
Chaurasia and Pal 5 compare the performance criterion of supervised learning classifiers, such as Naïve Bayes, SVM-RBF kernel, RBF neural networks, Decision Tree (Dt) (J48), and simple classification and regression tree (CART), to find the best classifier in breast cancer datasets. The experimental result shows that SVM-RBF kernel is more accurate than other classifiers; it scores at the accuracy level of 96.84% in the Wisconsin Breast Cancer (original) datasets.
Chaurasia and Pal 6 offered three popular data mining algorithms: CART, ID3 (iterative dichotomized 3), and DT for diagnosing heart diseases, and the results presented demonstrated that CART obtained higher accuracy within less time.
Chaurasia and Pal 7 conducted an experiment to identify the most common data mining algorithms, implemented in modern Medical Diagnosis, and evaluate their performance on several medical datasets. Five algorithms were chosen: Naïve Bayes, RBF Network, Simple Logistic, J48 and Decision Tree. For the evaluation two Irvine Machine Learning Repository (UCI-UC) databases were used: heart disease and breast cancer datasets. Several performance metrics were utilized: percent of correct classifications, True/False Positive rates, area under the curve (AUC), precision, recall, F-measure, and a set of errors.
Li et al. 8 discovered many diversified and significant rules from high-dimensional profiling data and proposed aggregation of the discriminating power of these rules for reliable predictions. The discovered rules are found to contain low-ranked features; these features are found to be sometimes necessary for classifiers to achieve perfect accuracy.
Kaewchinporn et al. 9 presented a new classification algorithm tree bagging and weighted clustering (TBWC) combination of decision tree with bagging and clustering. This algorithm is experimented on two medical datasets: cardiocography1, cardiocography2 and other datasets not related to medical domain.
Delen et al. 10 had taken 202,932 breast cancer patients records, which then pre-classified into two groups of “survived” (93,273) and “not survived” (109,659). The results of predicting the survivability were in the range of 93% accuracy.
Cao et al. 11 proposed a new decision tree-based ensemble method combined with feature selection method backward elimination strategy with bagging to find the structure activity relationships in the area of chemometrics related to pharmaceutical industry.
Methodology
This paper uses three popular data mining algorithms each on breast cancer dataset, Naïve Bayes, RBF Network, and J48. One of the reasons for choosing Naïve Bayes classification algorithm is because it is a simple yet powerful model and it returns not only the prediction but also the degree of certainty, which can be very useful. RBF Network is used due to their advantages over traditional multilayer perceptrons (MLPs), namely faster convergence, smaller extrapolation errors, and higher reliability. Radial basis function network (RBFN) is a class of single hidden layer feed forward network where the activation functions for hidden units are defined as radially symmetric basis functions phi such as the Gaussian function. J48 is an extension of ID3. The additional features of J48 are accounting for missing values, decision trees pruning, continuous attribute value ranges, derivation of rules. These classification algorithms are selected because they are very often used for research purposes and have potential to yield good results. Moreover, they use different approaches for generating the classification models, which increases the chances for finding a prediction model with high classification accuracy.
Naïve Bayes
Naïve Bayes is a machine learning algorithm for classification problems. It is based on Thomas Bayes’s probability theorem. It is primarily used for text classification which involves high-dimensional training datasets. A few examples are spam filtration, sentimental analysis, and classifying news articles. It is not only known for its simplicity but also for its effectiveness.12–14 It is fast to build models and make predictions with Naïve Bayes algorithm. Naïve Bayes algorithm is the algorithm that learns the probability of an object with certain features belonging to a particular group/class. In short, it is a probabilistic classifier. The Naïve Bayes algorithm is called “naïve” because it makes the assumption that the occurrence of a certain feature is independent of the occurrence of other features. The “Bayes” part refers to the statistician and philosopher Thomas Bayes and the theorem was named after him, Baye’s theorem, which is the base for Naïve Bayes algorithm. Naïve Bayes algorithm is Baye’s theorem or alternatively known as Baye’s rule or Baye’s law. It gives us a method to calculate the conditional probability, i.e. the probability of an event based on previous knowledge available on the events. More formally, Baye’s theorem is stated as the following equation
RBF network
An RBFN is a particular type of neural network. Neural networks or “Artificial Neural Networks” they are referring to the MLP. Each neuron in an MLP takes the weighted sum of its input values, that is each input value is multiplied by a coefficient and the results are all summed together. A single MLP neuron is a simple linear classifier, but complex non-linear classifiers can be built by combining these neurons into a network. The RBFN approach is more intuitive than the MLP. An RBFN performs classification by measuring the input’s similarity to examples from the training set. Each RBFN neuron stores a “prototype”, which is just one of the examples from the training set. When we want to classify a new input, each neuron computes the Euclidean distance between the input and its prototype 15 If the input more closely resembles the class A prototypes than the class B prototypes, it is classified as class A.
Figure 1 shows the illustration of the typical architecture of an RBF network. It consists of an input vector, a layer of RBF neurons, and an output layer with one node per category or class of data.

RBF Network architecture.
J48 Decision Tree
It is based on Hunt’s algorithm. Hunt’s algorithm grows a decision tree in a recursive fashion by partitioning the training records into successively purer subsets. Let Dt be the set of training records that reach a node t. The general recursive procedure is defined as below
16
:
If Dt contains records that belong to the same class yt, then t is a leaf node labeled as yt. If Dt is an empty set, then t is a leaf node labeled by the default class, yd. If Dt contains records that belong to more than one class, use an attribute test to split the data into smaller subsets.
It recursively applies the procedure to each subset until all the records in the subset belong to the same class. The Hunt’s algorithm assumes that each combination of attribute sets has a unique class label during the procedure. If all the records associated with Dt have identical attribute values except for the class label, then it is not possible to split these records any further. In this case, the node is declared a leaf node with the same class label as the majority class of training records are associated with this node.
J48 handles both categorical and continuous attributes to build a decision tree. In order to handle continuous attributes, J48 splits the attribute values into two partitions based on the selected threshold such that all the values above the threshold as one child and the remaining as another child. It also handles missing attribute values. J48 uses gain ratio as an attribute selection measure to build a decision tree. It removes the biasness of information gain when there are many outcome values of an attribute. At first, calculate the gain ratio of each attribute. The root node will be the attribute whose gain ratio is maximum. J48 uses pessimistic pruning to remove unnecessary branches in the decision tree to improve the accuracy of classification. 17
Breast Cancer Wisconsin dataset
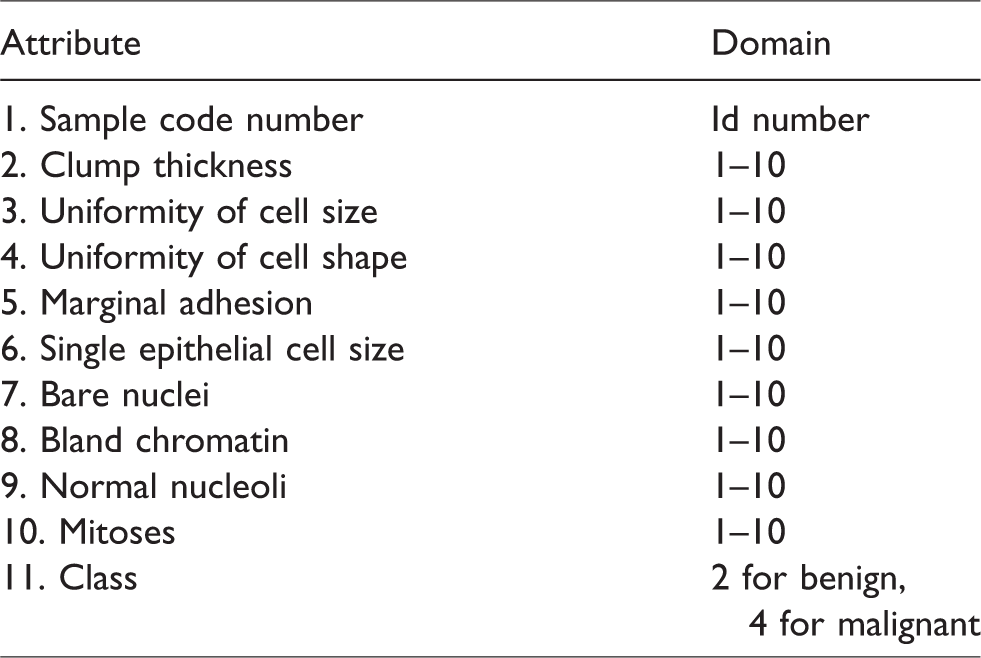
The data used in this study are provided by the UC Irvine Machine Learning repository located in Breast Cancer Wisconsin sub-directory, filenames root: breast-cancer-Wisconsin having 699 instances, 2 classes (malignant and benign), and 9 integer-valued attributes. We removed the 16 instances with missing values from the dataset to construct a new dataset with 683 instances (see Table 1). Class distribution: benign: 458 (65.5%), malignant: 241 (34.5%). Here, we have taken benign as 65.5% and malignant as 34.5% because it is better to take prevention than to cure, and therefore, large instances of benign patients have been taken for the study.18,19
Breast cancer dataset.
Experimental tool
This topic presents Waikato Environment for Knowledge Analysis (WEKA) version 3.6.9, the tool which is chosen in experiment to analyze medical datasets and evaluate the performance of data mining techniques applied to these sets. The selected data mining methods are presented with detailed description of parameters they use for analyses. Furthermore, the measures of model performance are presented which are the basis for the comparison of methods’ effectiveness and accuracy. Finally, the visualization of each algorithm’s performance is shown for medical datasets. This is based on own experience with WEKA environment supported with information included in.
Results and discussion
The breast cancer database consists of nine conditional attributes. The decisional attribute takes the values 0 or 1. As presented in Figure 2, the distributions of almost all values of attributes are even. In case of almost all the attributes, the number of instances in which the attributes take the lowest values is the greatest. All conditional attributes are multi-valued.

Distribution of the attributes of the breast cancer data.
The results of the comparison of the algorithms are presented in Table 2. The table shows the ranking of the algorithm in case of each of the performance measures and databases. The unquestionable leader in majority of cases is the Naïve Bayes. Nevertheless, overall performance was always better in comparison to other algorithms. When it comes to the RBF Network, it wins the second place in terms of the performance. For most of the databases and metrics the results gained by this algorithm were slightly worse than for the Naïve Bayes in most of the cases. Finally, the worst results were yielded by the J48. The reason for this may the nature of medical data. Its complexity and heterogeneity of values of attributes can hinder data mining.
Training and simulation error.

The results from Table 2 are also presented (for better visualization) in Figure 3. These graphs confirm high performance of the Naïve Bayes in case of the breast database. However, overall best algorithm is the Naïve Bayes, with the RBF Network being the second.
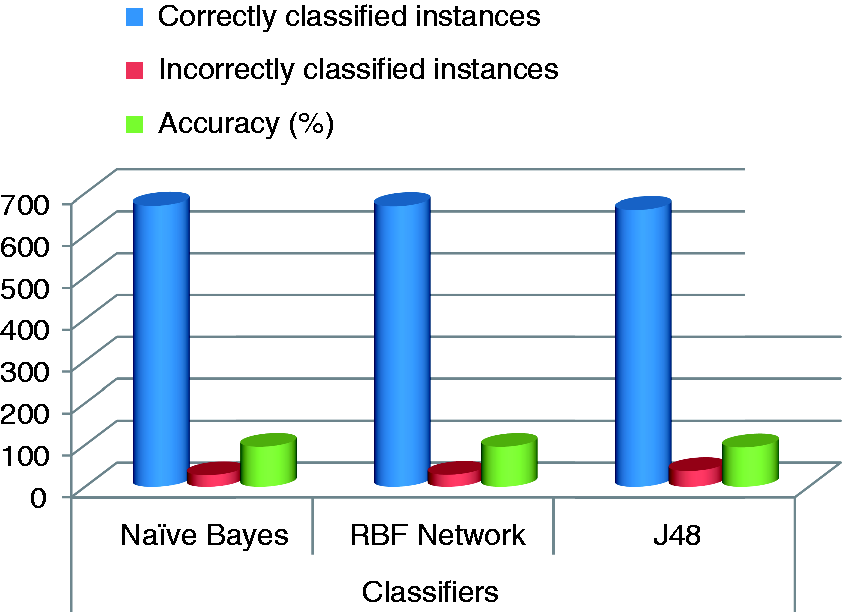
Comparative graph of different classifiers showing at different evaluation criteria.
In following predictive analytics, is a table of confusion matrix with two rows and two columns that reports the number of False Positives (FP), False Negatives (FN), True Positives (TP), and True Negatives (TN). This allows more detailed analysis than mere proportion of correct classifications (accuracy). Accuracy is not a reliable metric for the real performance of a classifier, because it will yield misleading results if the dataset is unbalanced. Using WEKA the confusion matrix is shown in Table 3.
Confusion matrix.

Now we calculate the accuracy, sensitivity and specificity for the methods Naïve Bayes, RBF Network, and J48 using the formulae



The calculated values for the three methods Naïve Bayes, RBF Network, and J48 are listed in Table 4.
Accuracy, sensitivity, specificity of different methods.

The analysis delivered interesting results. The best classifier is the Naïve Bayes. Its overall performance turned out to be the highest in case of most of the databases. This may be caused by the nature of data being complex could have caused overtraining of the other algorithms. The second place was won by the RBF Network. Its general performance was only slightly worse than Naïve Bayes’. On the third was the J48.
Conclusions
In this paper, we applied three prediction models for breast cancer survivability on two parameters: benign and malignant cancer patients. Here, we used three popular data mining methods: Naïve Bayes, RBF Network, and J48. We acquired a dataset (683 instances) from the UCI Machine Learning repository. We applied data selection, preprocessing, and transformation to develop the prediction models. In this research, we used a binary categorical survival variable, which was calculated from the variables in the raw dataset, to represent the survivability where malignant is represented with a value of ‘‘1’’ and benign is represented with ‘‘0’’. In order to measure the unbiased prediction accuracy of the three methods, we used a 10-fold cross-validation procedure, that is we divided the dataset into 10 mutually exclusive partitions (k-folds) using a stratified sampling technique. We repeated this process for each of the three prediction models. This provided us with a less biased prediction performance measures to compare the three models. The obtained results indicated that the Naïve Bayes performed the best with a classification accuracy of 97.36%, RBF Network came out to be second best with a classification accuracy of 96.77%, and the J48 came out to be the third with a classification accuracy of 93.41%. In addition to the prediction model, we also conducted sensitivity analysis and specificity analysis on Naïve Bayes, RBF Network, and J48 in order to gain insight into the relative contribution of the independent variables to predict survivability. The sensitivity results indicated that the prognosis factor ‘‘Class’’ is by far the most important predictor.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.