Abstract
Three basic amino acid–based cationic lipids bearing a fluorescent naphthalimide moiety and a reducible disulfide linkage are synthesized and applied as non-viral gene vehicles. Their DNA interactions are investigated by agarose-gel retardant and ethidium bromide replacement assays. The sizes and zeta potentials of the liposome/DNA complexes are measured by dynamic light scattering. The cytotoxicities of the liposome/DNA complexes are examined using HeLa and 7702 cell lines by MTT assays. The glutathione-responsive DNA release process is studied through time-dependent fluorescence assays. Luciferase gene expression showed the transfection efficiency of the liposome is dramatically increased in the presence of 10% serum. Confocal laser scanning microscopy studies corroborated that the liposome/DNA complexes are successfully uptaken into HeLa cells. These results demonstrate the promising use of amino acids and naphthalimide-containing lipids for safe and efficient gene delivery.
Reducible amino acid based cationic lipids with a naphthalimide moiety were synthesized and applied as non-viral gene vehicles

Introduction
Gene therapy offers a promising strategy to prevent or cure human diseases of genetic origin by introducing exogenous genetic materials, such as DNA, RNA, or small interfering RNA (siRNA) into cells. 1 However, its clinical use is impeded by a lack of safe and effective delivery carriers. 2 As an important type of non-viral gene vehicles, cationic lipids have been widely investigated for their facile synthesis, low toxicity, and excellent biocompatibility compared to viral vectors.3,4 The structures of cationic lipids are characterized by three components: a hydrophilic head group, a hydrophobic tail, and a linker between the former two moieties.5–7 All these components can seriously affect the toxicity and transfection efficacy of the final lipids.
Gene transfection with cationic lipids needs to satisfy the contrary requirements of adequate extracellular DNA protection and efficient intracellular DNA release. 8 Amino acids and synthetic peptides with desirable physicochemical properties and good biocompatibility are prospective substances for the construction of the multivalent polar domains which can complex negatively charged nucleic acids via strong electrostatic interactions, leading to improved DNA condensation and enhanced cellular uptake.9–11 In our previous studies, several amino acid–based amphiphiles with multicharge character were found to have the ability to achieve efficient gene delivery with low cytotoxicity.11–15 To facilitate DNA release in the cell, gene delivery vectors with stimuli-responsive ability have attracted increasing interest. 16 Among diverse stimuli, the significant difference in the concentration of the reducing tripeptide glutathione (GSH) between extracellular and intracellular environments makes disulfide bonds unique in biology and in gene delivery systems. 8 In a characteristic tether group, disulfide can be degraded in the cytoplasm and facilitates the dissociation of vectors and DNA by specifically responding to the redox potential through thiol–disulfide exchange reactions.17,18 Many disulfide-bridged nanocarriers with high transfection efficiency (TE) and low toxicity have been synthesized.11,12,19–22 Aliphatic chain, cholesterol, and even tocopherol derivatives are commonly used as hydrophobic components to construct cationic lipids. 5 The design and synthesis of new lipid systems with alternative structural types are crucial for the development of potent synthetic gene vectors. 14 The rigid naphthalimide moiety with native hydrophobic properties can be used to functionalize cationic lipids.4,23–26 Moreover, it has high DNA complexing ability, strong yellow-green fluorescence, and good photostability. 27 Naphthalimide factionalized lipids with good fluorescence properties have been used to monitor the process of cellular uptake, DNA translocation, and release.4,23–26 Although a series of naphthalimide-modified lipids has been synthesized, naphthalimide-containing cationic lipid gene vehicles with reduction-responsive ability have been seldom studied.
Based on the above considerations, herein we have designed and synthesized three lipidic gene vectors with a multivalent amino acid head group, a reduction-responsive disulfide linker, and a rigid naphthalimide tail (Scheme 1). Their DNA interaction, physicochemical properties, cytotoxicities, gene transfection activity as well as cellular uptake have been systematically investigated.

Synthesis of lipids
Results and discussions
Syntheses of new amino-based cationic lipids
Lipids
UV-absorption and fluorescence of lipid 5c
The photophysical properties of lipid

(a) UV-Vis absorption spectra of
Preparation and characterization of liposomes and liposome/DNA complexes (lipoplexes)
The cationic liposomes were prepared by the thin-film hydration method and a lipid/DOPE (1,2-Dioleoyl-sn-glycero-3-phosphoethanolamine) ratio of 1:1 was used herein. To evaluate their DNA-binding abilities, both the agarose-gel retardant and ethidium bromide (EB) replacement assays were employed.12,18 The agarose-gel retardant assay shows that the liposomes formed from

Electrophoretic gel retardation assays (a) and ethidium bromide displacement assays (b) of pDNA binding for liposomes
To gain an insight into whether the disulfide linker is reducible by reducing molecules such as GSH, fluorescence assays were used to monitor the time-dependent degradation of liposomes
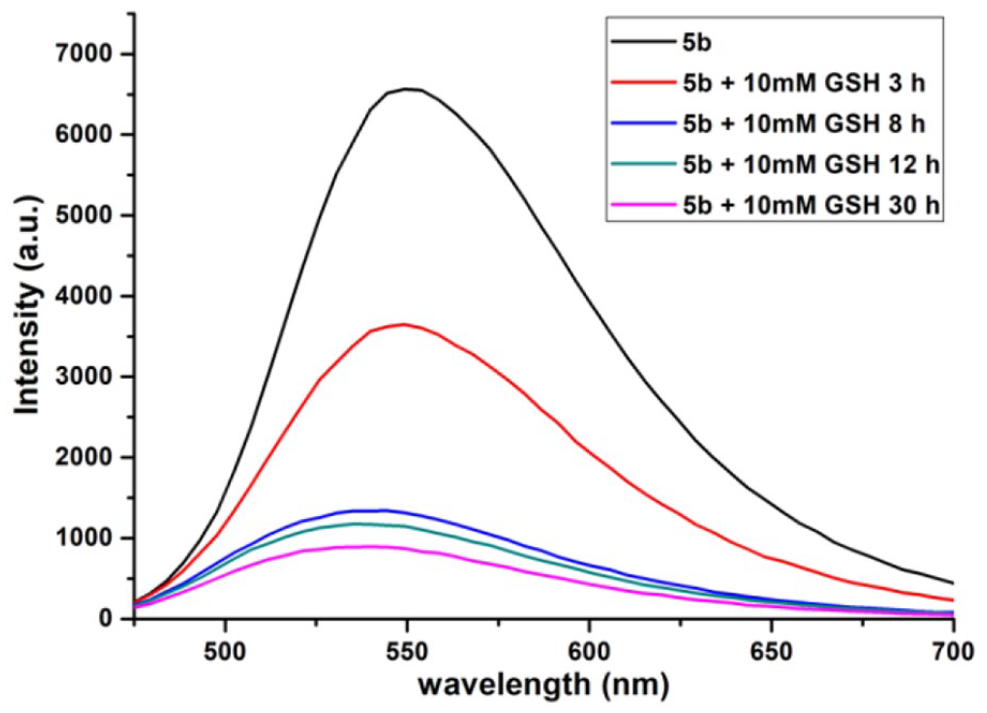
Fluorescence assays of liposomes
Sizes and zeta potentials of the formed lipoplexes
The particle sizes of different lipoplexes were measured by dynamic light scattering (DLS) assays, and the results are shown in Figure 4(a). Liposomes

Mean particle sizes (a) and surface charges (b) of the lipoplexes at various N/P ratios. Data represent mean value ± SD (n = 3).
The surface potentials of the lipoplex nanoparticles were also measured by DLS, and the results are shown in Figure 4(b). The zeta potentials of all the lipoplexes showed a similar trend on changing the N/P ratio: they increased from −38 to 36 mV when the N/P ratio increased from 1 to 16. The zeta potentials increased and turned positive with an increase in cationic materials.
Cytotoxicity
The cytotoxicities of the prepared lipoplexes were examined using the HeLa (human cervical cancer epithelia) and 7702 (normal human Liver cell lines) cell lines in serum-free medium by employing the MTT assay, with lipofectamine 2000 and PEI 25K as the control. The results shown in Figure 5 revealed that the chemical structure of the cationic headgroup significantly affected the cytotoxicity of the lipoplexes. Both HeLa and 7702 cells remained at a survival rate of over 85% for lipoplexes

Cell toxicities of the lipoplexes
In vitro transfection
The in vitro gene transfection efficiencies of the liposomes were studied with HeLa and 7702 cells using the pGL3 plasmid as a luciferase reporter gene. Figure 6a-b exhibit TEs at different N/P ratios in serum-free medium; lipofectamine 2000 and PEI 25K were used as controls. For both cell lines, liposomes

Luciferase expression induced by liposomes
It was found that the serum inhibits the transfection of cationic vectors.
30
Due to the better transfection performance of liposomes

Luciferase expression induced by liposomes
Cellular uptake of the amino acid–based cationic liposomes
Cellular uptake studies were performed in HeLa cells using Cy5-labeled dsDNA. Cy5-labeled DNA emitted red fluorescence under fluorescence microscopy conditions, while liposomes

Confocal laser scanning microscopy (CLSM) images of HeLa cells transfected with Cy5-labeled DNA by liposome
Conclusion
Three amino acid–based amphiphilic molecules bearing a naphthalimide moiety and a biodegradable disulfide linkage have been designed and synthesized. The liposomes formed from these lipids can condense stable lipoplex nanoparticles and deliver pDNA into cells. The structure of the hydrophilic headgroup significantly affected the TE values and cytotoxicities. Arginine- and lysine-containing lipids
Experimental section
Materials
Nα,Nβ-di-Boc-
Synthesis of lipids 5a–c
Preparation of compound 3
Triethylamine (101.2 mg, 1.0 mmol) was added drop wise to a solution of compound
Preparation of compounds 4
Compound
Compound 4a yield 44%
1H NMR (400 MHz, DMSO): δ 8.63 (d, J = 8.3 Hz, 1H), 8.39 (d, J = 7.1 Hz, 1H), 8.21 (d, J = 8.4 Hz, 1H), 7.97 (s, 1H), 7.89 (s, 1H), 7.51 (s, 1H), 6.72 (s, 1H), 4.10-4.06 (m, 1H), 3.96 (t, J = 8.0 Hz, 2H), 3.67 (d, J = 5.9 Hz, 2H), 3.03 (t, J = 6.6 Hz, 2H), 2.85-2.66 (m, 4H), 2.47 (s, 4H), 1.56 (s, 2H), 1.34-1.14 (m, 19H), 0.80 (d, J = 6.9 Hz, 3H). 13C NMR (100 MHz, DMSO): δ 172.10, 164.08, 163.28, 155.53, 150.50, 135.09, 134.53, 131.05, 129.73, 128.84, 124.79, 122.30, 120.56, 108.59, 104.22, 78.52, 54.94, 42.50, 38.43, 37.44, 36.35, 31.66, 30.05, 29.17, 29.03, 28.57, 28.03, 27.00, 22.50, 14.36. HRMS (ESI): [M + H]+ calcd for C35H48N6O5S2: 697.3200; found: 697.3206.
Compound 4b yield 41%
1H NMR (400 MHz, DMSO-d6): δ 8.62 (d, J = 8.4 Hz, 1H), 8.39 (d, J = 7.1 Hz, 1H), 8.00 (t, J = 6.0 Hz, 1H), 7.83 (s, 1H), 6.82 (d, J = 6.0 Hz, 1H), 3.98-3.90 (m, 2H), 3.88-3.86 (m, 1H), 3.72-3.67 (m, 2H), 3.21 (d, J = 5.2 Hz, 2H), 3.03 (t, J = 6.6 Hz, 2H), 2.86 (s, 2H), 2.77 (t, J = 6.6 Hz, 2H), 1.61-1.12 (m, 39H), 0.80 (t, J = 6.5 Hz, 3H). 13C NMR (100 MHz, DMSO): δ 172.51, 164.11, 163.53, 163.31, 162.74, 155.66, 152.47, 150.52, 134.57, 131.11, 129.76, 128.83, 124.85, 122.34, 120.59, 108.62, 104.26, 83.24, 78.51, 54.43, 42.50, 38.32, 37.56, 36.29, 31.68, 31.20, 29.08, 28.50, 28.01, 26.97, 22.49, 14.35. HRMS (ESI): [M + H]+ calcd for C45H69N7O9S2: 916.4171; found: 916.4677.
Compound 4c yield 46%
1H NMR (400 MHz, DMSO): δ 8.58 (d, J = 8.5 Hz, 1H), 8.36 (d, J = 7.3 Hz, 1H), 8.19 (d, J = 8.5 Hz, 1H), 7.97 (t, J = 5.6 Hz, 1H), 7.62 (t, J = 7.9 Hz, 1H), 3.93 (t, J = 8.0 Hz, 2H), 3.83-3.78 (m, 1H), 3.68-3.63 (m, 2H), 3.03 (t, J = 8.0 Hz, 2H), 2.88-2.75 (m, 4H), 1.53-1.17 (m, 36H), 0.78 (t, J = 8.0 Hz, 3H). 13C NMR (100 MHz, DMSO): δ 172.78, 164.06, 163.26, 155.98, 155.72, 150.46, 134.50, 131.03, 129.71, 128.77, 124.75, 122.29, 120.55, 108.61, 104.18, 78.37, 77.72, 54.79, 42.51, 37.56, 36.36, 32.11, 31.66, 29.64, 29.10, 28.64, 28.03, 27.00, 23.25, 22.49, 14.34. HRMS (ESI): [M + Na]+ calcd for C40H61N5O7S2 810.3905; found: 810.3912.
Preparation of lipids 5
Compounds
Lipid 5a yield 93%
1H NMR (400 MHz, DMSO): δ 8.62 (d, J = 8.3 Hz, 1H), 8.38 (d, J = 7.1 Hz, 1H), 8.22 (d, J = 8.4 Hz, 1H), 7.89 (s, 1H), 7.66 (t, J = 8.0 Hz, 2H), 7.50 (s, 1H), 6.75 (s, 1H), 4.09-4.05 (m, 1H), 3.96 (t, J = 8.0 Hz, 2H), 3.68-3.66 (m, 2H), 3.02 (t, J = 6 Hz, 2H), 2.85-2.70 (m, 4H), 2.45 (s, 4H), 1.57-1.54 (m, 2H), 1.25-1.20 (m, 10H), 0.79-0.81 (m, 3H). 13C NMR (100 MHz, DMSO): δ 172.10, 164.01, 163.24, 150.45, 135.12, 134.48, 131.01, 129.78, 128.86, 124.67, 122.30, 120.53, 108.52, 104.27, 54.94, 42.46, 38.46, 37.44, 36.35, 31.68, 30.02, 29.17, 29.03, 28.01, 27.00, 22.48, 14.10. HRMS (ESI): [M + H]+ calcd for C30H40N6O3S2: 597.2676; found: 597.2680.
Lipid 5b yield 88%
1H NMR (400 MHz, DMSO-d6): δ 8.66 (d, J = 8.4 Hz, 1H), 8.42 (d, J = 7.1 Hz, 1H), 8.02 (t, J = 6.0 Hz, 1H), 7.91 (s, 2H), 7.69 (t, J = 8.0 Hz, 1H), 6.85 (d, J = 8.6 Hz, 1H), 4.00-3.97 (m, 2H), 3.88-3.86 (m, 1H), 3.72-3.65 (m, 2H), 3.22 (d, J = 5.2 Hz, 2H), 3.06 (t, J = 8.0 Hz, 2H), 2.86 (s, 4H), 2.77 (t, J = 6.6 Hz, 2H), 1.60-1.21 (m, 14H), 0.85 (t, J = 6.5 Hz, 3H). 13C NMR (100 MHz, DMSO): δ 172.50, 164.08, 163.48, 150.45, 134.62, 131.10, 129.78, 128.81, 124.88, 122.32, 120.58, 108.62, 104.28, 54.43, 42.52, 38.30, 37.62, 36.42, 36.24, 31.70, 31.60, 31.20, 29.20, 29.08, 28.01, 26.97, 22.47, 14.20. HRMS (ESI): [M + H]+ calcd for C30H45N7O3S2: 616.3098; found: 616.3095.
Lipid 5c yield 94%
1H NMR (400 MHz, DMSO): δ 8.57 (d, J = 8.5 Hz, 1H), 8.36 (d, J = 7.3 Hz, 1H), 8.20 (d, J = 8.5 Hz, 1H), 7.97 (t, J = 5.6 Hz, 1H), 7.62 (t, J = 8.0 Hz, 1H), 3.93 (t, J = 8.0 Hz, 2H), 3.83-3.79 (m, 1H), 3.67-3.62 (m, 2H), 3.03 (t, J = 8.0 Hz, 2H), 2.84-2.76 (m, 4H), 1.50-1.15 (m, 18H), 0.80 (t, J = 8.0 Hz, 3H). 13C NMR (100 MHz, DMSO): δ 172.60, 164.10, 163.32, 150.46, 134.52, 131.03, 129.74, 128.79, 124.78, 122.26, 120.55, 108.63, 104.18, 54.75, 42.51, 38.35, 37.62, 36.36, 32.13, 31.68, 29.64, 29.20, 29.03, 28.02, 27.00, 23.25, 22.53, 14.32. HRMS (ESI): [M + Na]+ calcd for C30H45N5O3S2: 610.2856; found: 610.2851.
Preparation of the cationic liposomes
The cationic lipid (0.0025 mmol) and neutral colipid (DOPE, 0.0025 mmol) were dissolved in chloroform (2.5 mL) in autoclaved glass vials. The solvent was removed with a thin flow of moisture-free nitrogen gas, and the dried lipid film was then kept under high vacuum for 8 h. A portion of 2.5 mL of sterile deionized water was added to the vacuum-dried lipid film, and the mixture was allowed to swell overnight. The liposomes were vortexed for 1−2 min to remove any adhering lipid film and sonicated in a bath sonicator for 5 min at room temperature to produce multilamellar vesicles (MLVs). The MLVs were then sonicated in an ice bath using a probe sonifier to afford the corresponding cationic liposomes (1.0 mmol L−1).11–13
Agarose-gel retardation assays
Lipids
EB replacement assays
The ability of liposomes
Fluorescence assays
The fluorescence change of the liposomes in response to GSH was examined using fluorescence assays. The liposome solution (2.0 mM) in deionized water (250 μL) was added with GSH (250 μL, 20 mM). After incubation of the solution at 37 °C for different times (3, 8, 12 and 30 h), 30 μL of the above solution was diluted with deionized water (3 mL), and then the fluorescence spectra were obtained. Excitation wavelength at 445 nm, emission band centered at 550 nm.
Lipoplex particle sizes and zeta potentials
The sizes and zeta potentials of the liposome/pDNA lipoplexes at various N/P ratios were analyzed at room temperature using a DLS system (Zetasizer Nano ZS, Malvern Instruments Led, United Kingdom at 25 °C. The lipoplex particle solutions were first prepared by mixing
MTT cytotoxicity assays
The toxicities of the liposomes and lipoplexes toward 7702 and HeLa cells were determined using the MTT assay following literature procedures. 11 About 7000 cells/well were seeded into 96-well plates. For lipoplexes, after 24 h, optimized lipid/DOPE formulations were complexed with 0.2 μg of DNA at various N/P ratios. Lipoplexes prepared from lipofectamine 2000 and polyplexes prepared from PEI-25K were used as controls.11–13
In vitro transfection assays
7702 and HeLa cells were seeded into 24-well plates at densities of 80,000 cells/well in 0.5 mL of complete medium. Twenty-four hours prior to transfection experiments, the medium was replaced with 1 mL of fresh culture medium with or without FBS. Liposomes
Confocal laser scanning microscopy analysis
HeLa cells were seeded onto a 35-mm confocal dish (Φ = 15 mm) at a density of 2.5 × 105 cells per well. After 24 h, the medium was replaced with fresh complete culture medium. Next, complexes of liposomes and Cy5-labeled pDNA (1 μg DNA per well) at a known concentration were added into each well. After incubation for 2 h, the cells were washed three times with PBS buffer, and the cell nuclei were stained with DAPI for 15 min. The cells were subsequently rinsed three times with PBS buffer. The confocal laser scanning microscopy (CLSM) imaging was performed using a Zeiss, LSM 780 (Zeiss, Japan) with a 40× objective lens at excitation wavelengths of 488 nm for liposome
Statistical analysis
All the results are presented as the mean value ± SD. Paired or independent t-tests were performed to compare the differences between two groups. To meet the requirements (normal distribution and homogeneous variance) of the t-test, the data of luciferase expression were transformed by applying the base-10 logarithm. In addition, to determine whether the N/P was a factor affecting the cytotoxicity and luciferase expression, one-way analysis of variance (ANOVA) was performed. All statistical analyses were based on R 4.0.2; p < 0.05 was considered statistically significant.
Footnotes
Acknowledgements
W.-J.Y. thanks Xiao-Qi Yu and Ji Zhang from the Key Laboratory of Green Chemistry and Technology (Ministry of Education), College of Chemistry, Sichuan University for their support.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by the National Science Foundation of China (no. 21708032), the Science and Technology Department of Sichuan Province (no. 2019YJ0260; 2022NSFSC1252), and the Fundamental Research Funds for the Central Universities, Southwest Minzu University (no. 2019NQN14).
