Abstract
Background:
Valproate is a candidate for ischemic stroke prevention due to its anti-atherosclerotic effects in vivo. Although valproate use is associated with decreased ischemic stroke risk in observational studies, confounding by indication precludes causal conclusions.
Aims:
We applied Mendelian randomization to determine whether genetic variants that influence seizure response among valproate users associate with ischemic stroke.
Methods:
We derived a genetic score for valproate response using genome-wide association data of seizure response after valproate intake from the Epilepsy Pharmacogenomics Consortium. We then tested this score among valproate users of the UK Biobank for association with incident and recurrent ischemic stroke using Cox proportional hazard models. As replication, we tested found associations in an independent cohort of valproate users of the Mass General Brigham Biobank.
Results:
Among 2150 valproate users (mean 56 years, 54% females), 82 ischemic strokes occurred over a mean 12 year follow-up. Higher valproate response genetic score was associated with higher serum valproate levels (+5.78 µg/ml per 1 standard deviation (SD), 95% confidence interval (CI) (3.45, 8.11)). After adjusting for age and sex, higher valproate response genetic score was associated with lower ischemic stroke risk (hazard ratio (HR) per 1 SD 0.73, 95% CI (0.58, 0.91)) with a halving of absolute risk in the highest compared to the lowest score tertile (4.8% vs 2.5%, p trend = 0.027). Among 194 valproate users with prevalent stroke at baseline, a higher valproate response genetic score was associated with lower recurrent ischemic stroke risk (HR per 1 SD 0.53, 95% CI (0.32, 0.86)) with reduced absolute risk in the highest compared to the lowest score tertile (3/51, 5.9% vs 13/71, 18.3%, p trend = 0.026). The valproate response genetic score was not associated with ischemic stroke among the 427,997 valproate non-users (p = 0.61), suggesting minimal pleiotropy. In 1241 valproate users of the Mass General Brigham Biobank with 99 ischemic stroke events over 6.5 years follow-up, we replicated our observed associations between the valproate response genetic score and ischemic stroke (HR per 1 SD 0.77, 95% CI (0.61, 0.97)).
Conclusion:
These results demonstrate that a genetically predicted favorable seizure response to valproate is associated with higher serum valproate levels and reduced ischemic stroke risk among valproate users, providing causal support for valproate effectiveness in ischemic stroke prevention. The strongest effect was found for recurrent ischemic stroke, suggesting potential dual-use benefits of valproate for post-stroke epilepsy. Clinical trials will be required in order to identify populations that may benefit most from valproate for stroke prevention.
Data access statement:
UK Biobank participant data are available after approval of a research proposal. The weights of the used genetic scores are available in the Supplemental Tables.
Introduction
Valproate is a widely used antiepileptic drug that has been associated with decreased risk for ischemic stroke in observational studies.1 –4 Valproate is assumed to exert this preventive effect by inhibiting histone deacetylase 9 (HDAC9), 5 which in animal studies has been found to lead to a stabilizing and anti-inflammatory effect on atherosclerotic plaques.6 –8 Although genetic variants in the HDAC9 gene have been repeatedly associated with large-artery stroke in genome-wide association studies (GWAS),9,10 causal evidence for valproate’s role in human stroke prevention is still missing.
Germline genetic variation is not prone to confounding and can be leveraged to assess whether a drug causally contributes to a specific outcome. If there are genetic variants known to influence drug response, drug users of an observational study can be divided according to their predicted genetic response. Because neither prescribers nor patients are aware of this genetic response at the time of prescription, the drug users can be considered randomized and blinded by genetics. If the genetic variants that predispose to better drug response are associated with the outcome of interest, the results support a causal effect on the outcome. Pleiotropic effects of the genetic variants that have an independent effect on the outcome can be ruled out if no associations are found among non-drug users. We have recently applied this in silico simulation of a randomized controlled trial, a special case of Mendelian randomization, to show that statins causally contribute to intracerebral hemorrhage. 11
Utilizing this framework, we investigated the causal effect of valproate on ischemic stroke, and, to gain insight into clinically relevant effects on atherosclerosis, additionally on myocardial infarction. We leveraged data from a GWAS of clinical response to valproate, lamotrigine, and levetiracetam 12 to construct and validate a genetic score for valproate response in the UK Biobank (UKB) and study its association with the selected outcomes (Figure 1(a)). To rule out that detected associations are driven by an antiepileptic drug class effect, we also studied the effects of lamotrigine and levetiracetam on ischemic stroke. We showed the robustness of our findings in an independent replication cohort of the Mass General Brigham Biobank (MGBB).
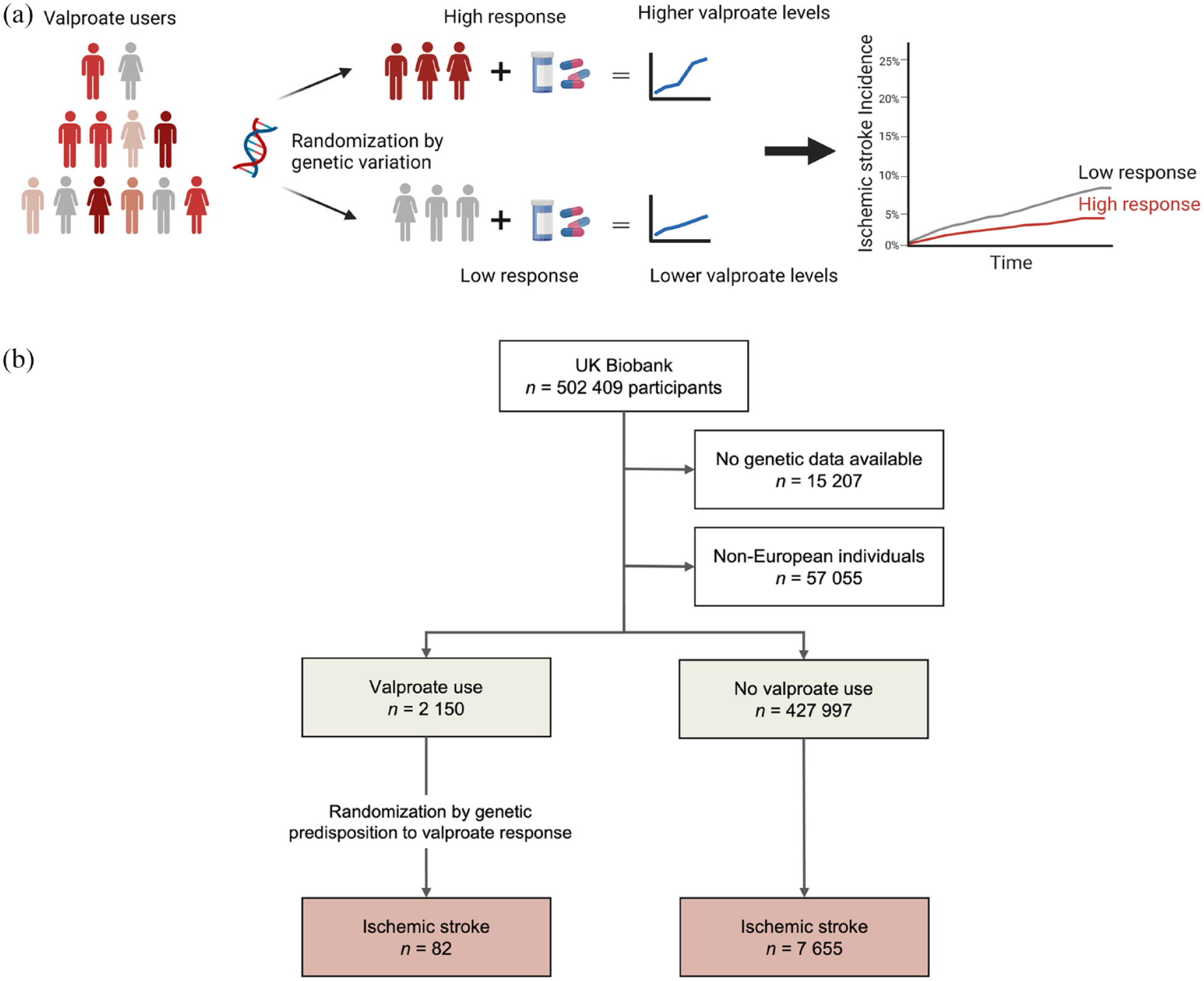
Study overview. (a) Concept of the in silico trial randomized by genetic variation. A genetic score consisting of genetic variants known to predispose to higher likelihood of seizure freedom after valproate intake was associated with serum valproate levels and incidence of ischemic stroke. (b) Study flow. After exclusion of non-European individuals and those without genetic data, 2150 valproate users and 427,997 valproate non-users were identified.
Methods
Study population
The UKB is an ongoing, population-based prospective cohort study of over 500,000 individuals recruited from 2006 to 2010 in 22 assessment centers across the United Kingdom. 13 We included only individuals with available genetic data. Because we used genetic variants 12 from individuals of European ancestry, we further restricted our cohort to European ancestry participants. The UKB has institutional review board approval from the Northwest Multi-Center Research Ethics Committee (Manchester, UK, approval 11/NW/0382). All participants provided written informed consent.
Mendelian randomization approach
Because the genetic variants that are used for the exposure (in our study, valproate, lamotrigine, and levetiracetam clinical response) are derived from a GWAS including only individuals exposed to those medications, but stroke and myocardial infarction outcome GWAS have been performed among drug users and non-users, we used an individual-level approach in the UKB to test for drug-specific and pleiotropic effects of our genetic instruments. We constructed a genetic score for response to each drug and tested each score for association with the outcomes among users and non-users of each medication. Because of the random assortment of common alleles in a population, genetically predicted drug response is randomly allocated, and thus an association of the genetic score with the outcome of interest in drug-users provides evidence of a causal drug effect. Pleiotropic effects of the genetic variants that modify the risk for chosen outcomes independent of the drug can be ruled out if there is no association among non-drug users. This approach has been described by us and others previously.11,14
Identification of antiepileptic drug users
We identified valproate, lamotrigine, and levetiracetam drug users via verbal baseline interview and primary-care prescription data. We have previously reported the details of our pipeline to extract medication data from UKB primary-care data. 11 Details are described in the Supplemental Methods.
Construction of the genetic scores for antiepileptic drug response
We used genome-wide association data from the Epilepsy Pharmacogenomics Consortium (EpiPGX) on seizure freedom after antiepileptic drug intake in European ancestry patients with generalized epilepsy. 12 In that study, treatment responders were defined as seizure-free patients under continuous treatment for at least 1 year, and treatment non-responders as patients with ⩾50% of pretreatment seizure frequency under adequate dosing of the drug. 12 Association tests were performed based on responder versus non-responder status for valproate (n = 565), lamotrigine (n = 387), and levetiracetam (n = 209). 12 There was no participant overlap with the UKB. Association results were available for single nucleotide polymorphisms (SNPs) associated with drug response at p < 0.05 (n = 162,242 for valproate, n = 162,666 for lamotrigine, and n = 162,430 for levetiracetam).
To construct the genetic scores to be used as instruments for valproate, lamotrigine, and levetiracetam response, we leveraged PRS-CS, 15 a novel genetic score construction method that assigns weights to the most relevant SNPs by using a high-dimensional Bayesian regression framework, placing a continuous shrinkage prior on SNP effect sizes, and models linkage disequilibrium. PRS-CS with default parameters generated 33,089, 33,300, and 32,736 SNP weights for valproate, lamotrigine, and levetiracetam, respectively. To test robustness of the discovered associations, we performed sensitivity analyses with an alternative genetic score for valproate response that was derived using a clumping and thresholding approach and included 20 SNPs (Supplemental Methods).
Validation of the genetic score on serum valproate response to normalized valproate dosing
We aimed to test the association of the genetic score for valproate response on valproate serum levels to determine whether valproate has an effect on ischemic stroke through serum level–dependent effects. If the genetic variants that are associated with seizure response after valproate intake predict valproate response through increased valproate serum levels, they are a proxy for genetically predicted drug exposure and thus can be used as an instrument for randomization. In this special case of Mendelian randomization, this is assertion of the relevance assumption. Valproate serum levels were gathered from the primary-care clinical data (Supplemental Methods). The association of the valproate dose and the genetic score with valproate serum levels was tested in a linear regression model, adjusted for age at the time of serum level measurement and sex. Levels for lamotrigine or levetiracetam were not available in the UKB.
Outcome ascertainment
Outcome events for ischemic stroke and myocardial infarction were gathered from inpatient hospital codes and death registry using International Classification of Diseases: Tenth Revision (ICD-10) codes that were aligned with the diagnostic algorithm in the UKB (Supplemental Methods).
Association of the genetic scores with outcomes
Cox proportional hazard models censored for death were used with the valproate-specific genetic scores, adjusted for age, sex, principal components 1–3, and genotyping assay in the cohort of valproate users. Stroke subtypes are not available in the UKB, ischemic stroke in the setting of atrial fibrillation was analyzed by adding an interaction term of the genetic score with prevalent atrial fibrillation in the model for ischemic stroke and performing subgroup analyses in individuals with and without a diagnosis of atrial fibrillation before or within 6 months after ischemic stroke. Main analyses were performed in all valproate users regardless of cryptic relatedness, and sensitivity analyses were performed in a cohort restricted to unrelated individuals (KING kinship coefficient < 0.0884). Chi-square test for trend in proportions was used to assess absolute risk differences across genetic score tertiles. To rule out pleiotropic effects of the genetic score, we tested the same associations among non-valproate users, thus assessing the independence and exclusion restriction assumptions of Mendelian randomization. To further rule out an antiepileptic drug class effect, all analyses were repeated with the genetic scores for lamotrigine and levetiracetam response among users of the respective drugs. To exclude drug interactions, we also performed a sensitivity analysis in patients on valproate monotherapy.
Replication analyses
We aimed to confirm our discoveries in the MGBB, an ongoing prospective clinical research cohort, using the same statistical approaches as in the UKB. For details about the cohort, identification of valproate users, outcomes, and statistical models, see Supplemental Methods.
Data availability
UKB participant data are available after approval of a research proposal. The weights of the genetic scores are available in the Supplemental Material.
Results
Baseline characteristics
We identified 2150 valproate users after exclusion of non-European ancestry individuals and those with unavailable genetic data (Figure 1(b), Table 1). The prescriptions ranged from 1987 to 2018, and the mean time between first and last prescription was 6.5 ± 6.9 years. Valproate users had a higher rate of vascular risk factors compared to non-users but no difference in the genetic score. The genetic score for valproate response was normally distributed among valproate users (Figure 2(a)). When comparing valproate users stratified by genetic score tertiles, no differences in baseline characteristics, vascular risk factors, antiplatelet and statin use, or approximated valproate doses were found across genetic score tertiles, indicating an appropriate randomization (Table 2). Most patients were on valproate monotherapy, and all tertiles had average valproate serum levels in the therapeutic range.
Baseline characteristics of valproate users compared to valproate non-users.
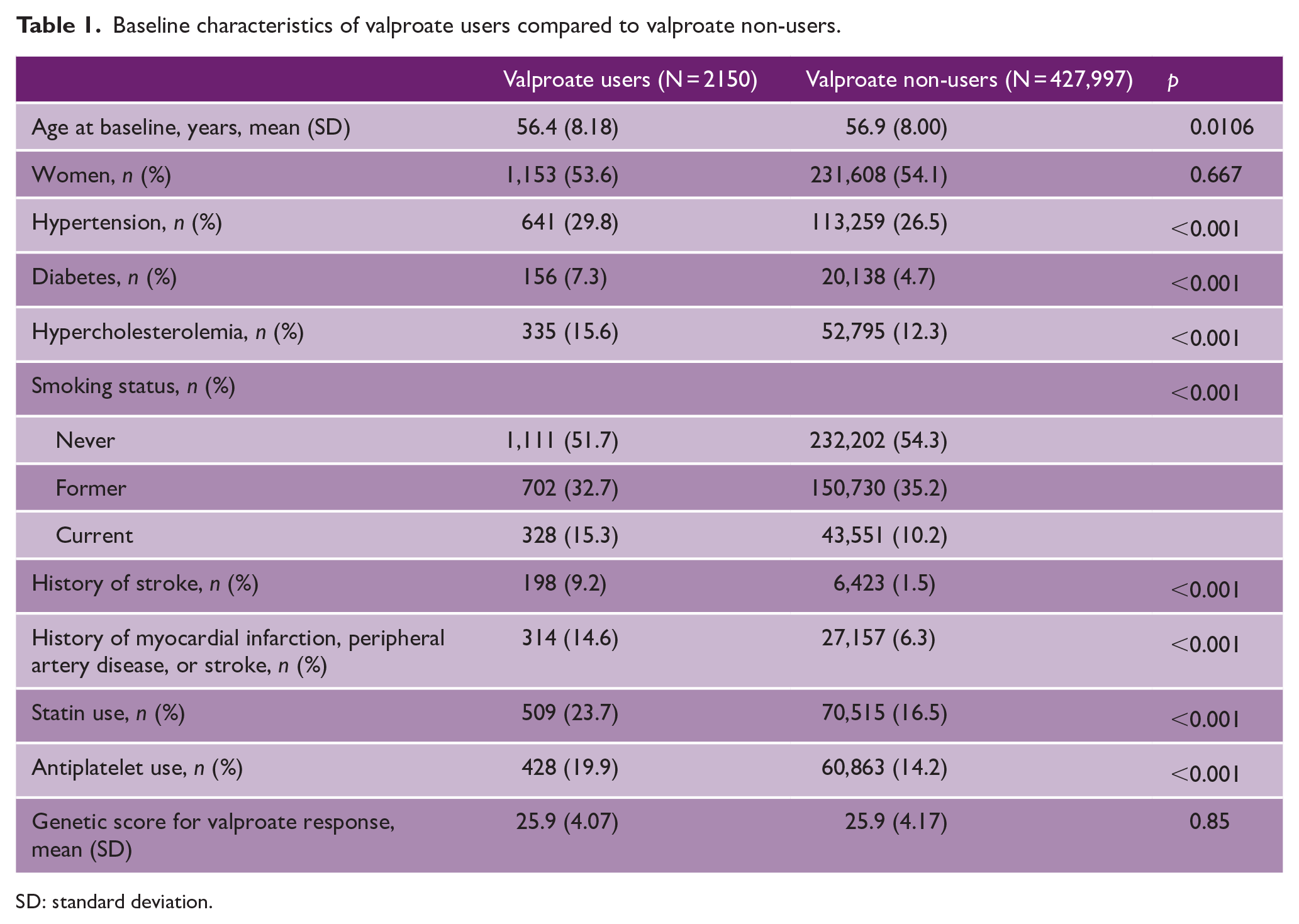
SD: standard deviation.
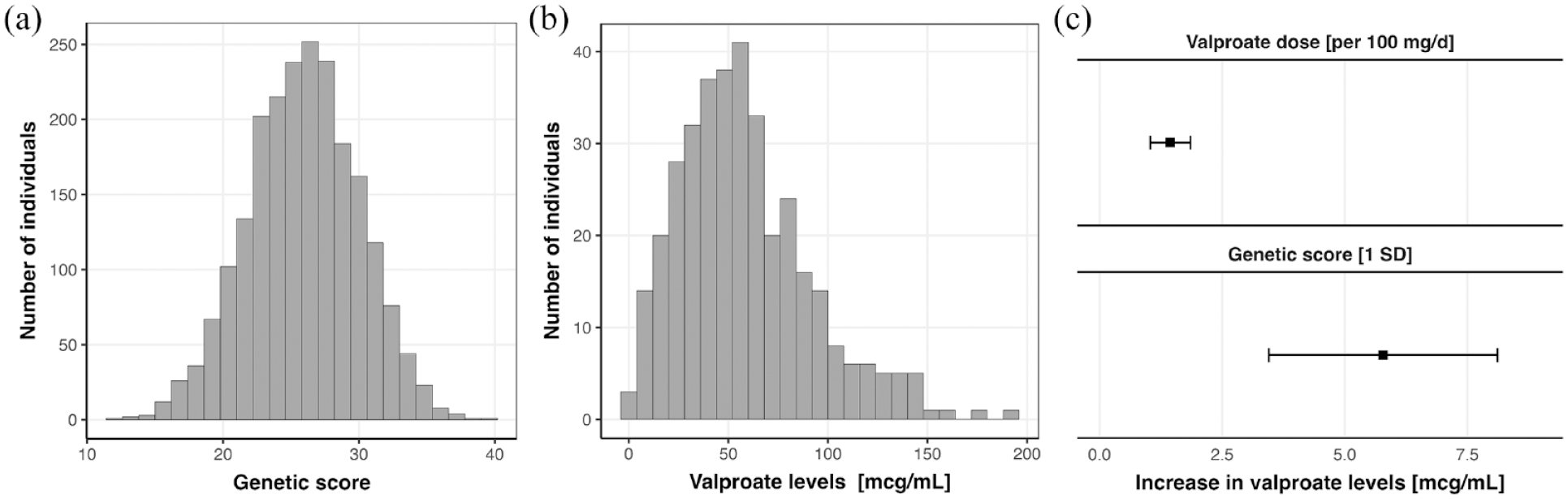
Valproate serum levels and their association with the genetic score for valproate response in the UK Biobank: (a) Distribution of the genetic score among the 2150 valproate users. (b) Distribution of the 549 valproate serum-level values among 202 valproate users. (c) Association of approximated daily valproate dose and the genetic score for valproate response with valproate serum levels in a linear regression model adjusted for age and sex. On the top is the coefficient for valproate dose, on the bottom is the genetic score for valproate response.
Baseline characteristics of valproate users stratified by the genetic score for valproate response.
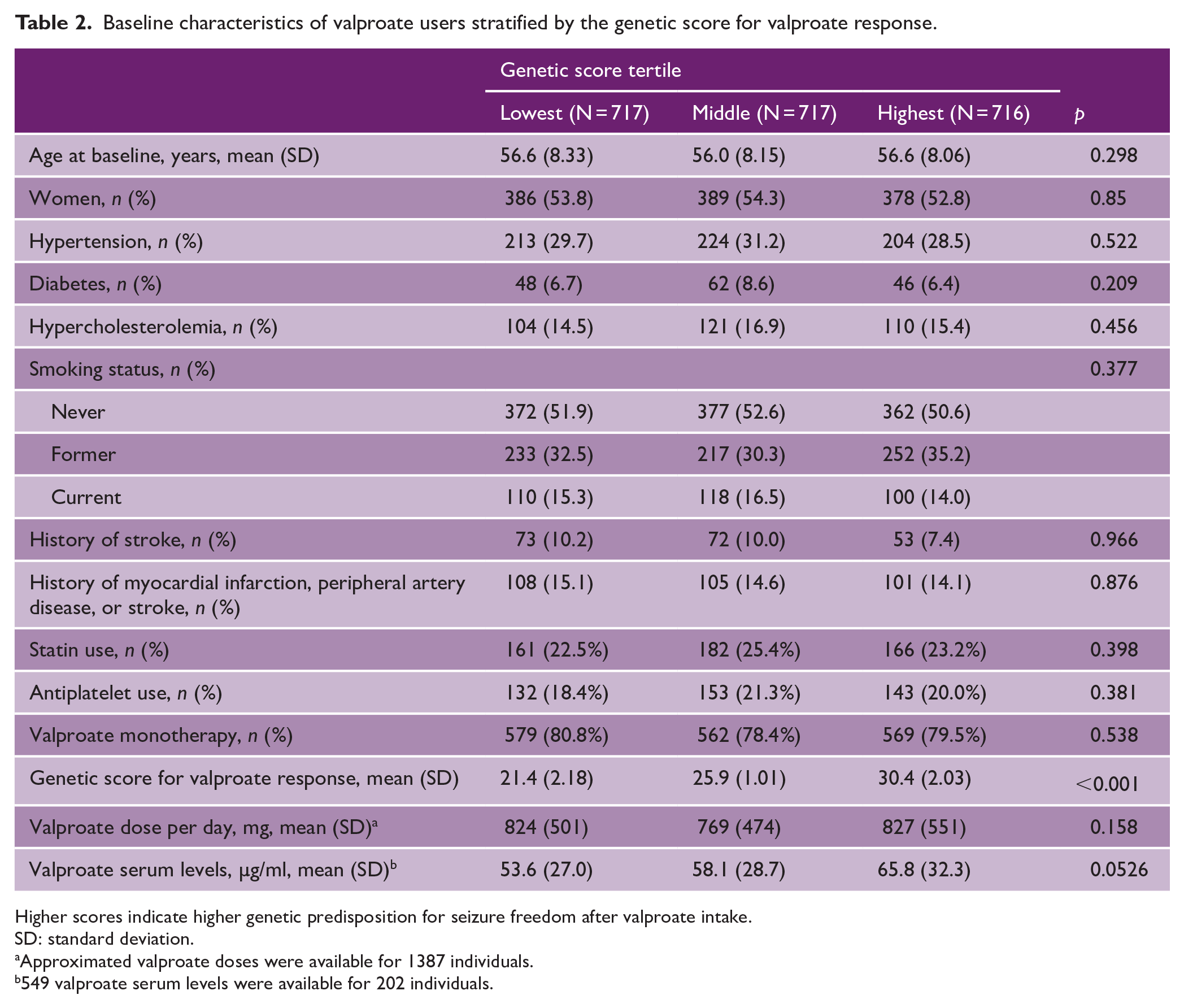
Higher scores indicate higher genetic predisposition for seizure freedom after valproate intake.
SD: standard deviation.
Approximated valproate doses were available for 1387 individuals.
549 valproate serum levels were available for 202 individuals.
Validation of the genetic score on serum valproate response to normalized valproate dosing
A total of 549 valproate serum level values were available for 202 valproate users after exclusion of 178 invalid values (Figure 2(b)). The approximated daily valproate dose was significantly associated with valproate serum levels (1.25 μg/ml per 100 mg/day, 95% CI (0.85, 1.66), Figure 2(c)). We found a significant association of the genetic score with valproate serum levels, indicating higher average serum levels in participants with higher genetic scores (+5.78 µg/ml per 1 SD, 95% CI (3.45, 8.11), Figure 2(c)). These associations were replicated with the alternative genetic score (Supplemental Table S7).
Association of the valproate genetic instrument with incident ischemic stroke
Among the 2150 valproate users, 82 ischemic strokes occurred over a mean follow-up of 11.6 years. A higher genetic score for valproate response was associated with a lower risk of incident ischemic stroke (hazard ratio (HR) 0.73, 95% CI (0.58, 0.91) per 1 SD increase, Figure 3(a)). Individuals in the lowest tertile of the genetic score had an almost two-fold increased absolute risk for ischemic stroke compared to those in the highest tertile (4.8% vs 2.5%, p trend = 0.027, Figure 3(b)). Sensitivity analyses confirmed robustness of the findings among the 1967 unrelated individuals (HR 0.75, 95% CI (0.59, 0.95), Supplemental Table S8) as well as with the alternative genetic score for valproate response using a clumping and thresholding approach (HR 0.75, 95% CI (0.59, 0.95), Supplemental Table S9). Restricting our cohort to the patients on valproate monotherapy (n = 1675), the associations were replicated but with wider confidence intervals due to lower power (Supplemental Table S10). We found no associations between the genetic scores for lamotrigine and levetiracetam response and ischemic stroke in the 1108 lamotrigine and 789 levetiracetam users, respectively (p = 0.76 and p = 0.24, Supplemental Tables S11 and S12).
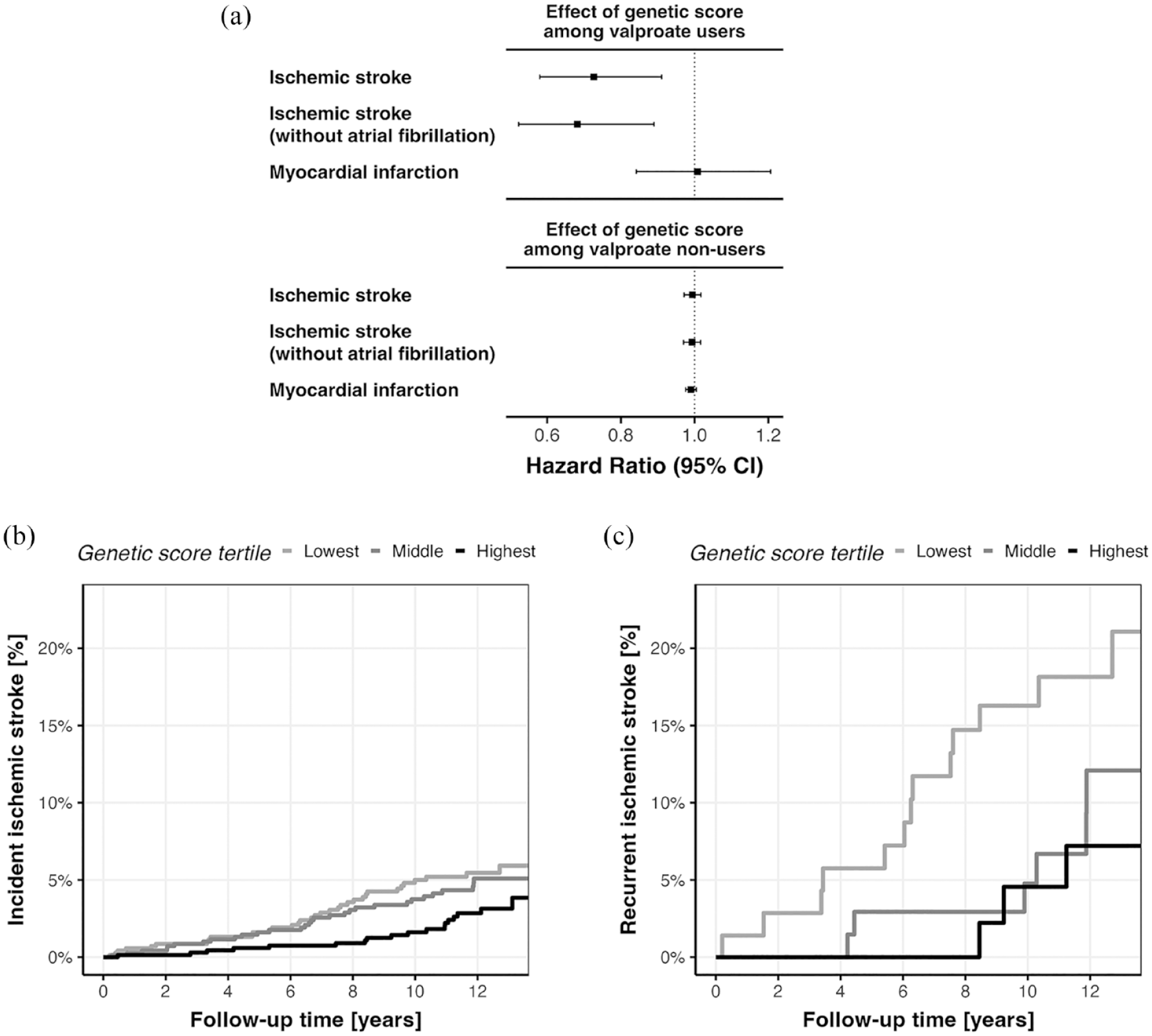
Associations between the genetic score for valproate response with outcomes in the UK Biobank: (a) hazard ratios of valproate on ischemic stroke, ischemic stroke without concurrent diagnosis of atrial fibrillation, and myocardial infarction. (b) and (c) Kaplan–Meier plots for incidence of (a) 82 ischemic strokes among the 2150 valproate users and (b) 22 recurrent ischemic strokes among the 194 valproate users with prevalent stroke at baseline, stratified by genetic score tertiles.
In the cohort of valproate users, 67 individuals had a concurrent diagnosis of atrial fibrillation, and 24 ischemic strokes occurred among them. No interaction between prevalent atrial fibrillation and the genetic score was found (p = 0.40). No association between the genetic score and ischemic stroke was found in this subgroup (p = 0.51); however, when removing the 67 individuals with atrial fibrillation from the cohort of valproate users, the association of the genetic score with incident ischemic stroke was more pronounced (HR 0.68, 95% CI (0.52, 0.89), Figure 3(a)).
Association of the genetic score with recurrent ischemic stroke
Among the 194 individuals with prevalent stroke at baseline, 22 recurrent ischemic strokes occurred over a mean follow-up of 11.2 years. A higher genetic score was associated with decreased risk for recurrent ischemic stroke (HR 0.53, 95% CI (0.32, 0.86) per one SD increase). Although only few cases in total, individuals in the lowest tertile of the genetic score had a three-fold higher absolute risk for ischemic stroke compared to those in the highest tertile (13/71, 18.3% vs 3/51, 5.9%, p trend = 0.026, Figure 3(c)).
Associations of the genetic score with myocardial infarction
No associations of the genetic score for valproate response were found for myocardial infarction (133 events, p = 0.93) among valproate users. Also, no associations were found among valproate non-users, reassuring that our genetic instruments were not affected by horizontal pleiotropy (p = 0.18, Figure 3(a)).
Replication in the MGBB
We identified 1241 valproate users with available genetic data in the MGBB, among which 99 ischemic stroke events and 126 myocardial infarction events occurred over a median follow-up of 6.7 years. We found 839 patients with 6353 valproate serum-level measurements. The associations between prescribed valproate dose and the genetic score with valproate serum levels replicated with almost identical effect estimates (Supplemental Table S13). We also found significant association between the genetic score for valproate response and ischemic stroke (HR 0.77, 95% CI (0.61, 0.97)), and no association between the genetic score and myocardial infarction (p = 0.68).
Discussion
We leveraged data from a GWAS of seizure control after valproate intake from a cohort of patients with epilepsy and applied it to medication prescription and intake data of 502,000 individuals from a population-based observational cohort study. We found that a higher genetically predicted seizure response to valproate was associated with higher valproate serum levels. In turn, this higher genetically predicted response to valproate was associated with a lower risk of ischemic stroke, among valproate users only. There was no such association among valproate non-users, and no association among lamotrigine and levetiracetam users, ruling out independent pleiotropic effects of the genetic variants or an antiepileptic drug effect. The robustness of our results is supported by replication in the MGBB, an independent electronic health record database from a different continent. Our results support a causal effect of valproate on ischemic stroke risk and demonstrate the utility of leveraging genetic data in observational cohorts to model drug response in silico for drug repurposing.
Valproate is a nonspecific HDAC inhibitor 16 and with the detection of HDAC9 as genetic risk locus for large-artery stroke,6,9,10 the role of valproate for stroke prevention has been postulated. Various studies have investigated the association of valproate with vascular outcomes in observational studies with conflicting findings.1 –3,17 –20 Our study confirmed the presumed effect of valproate on ischemic stroke, but failed to confirm previously hypothesized associations with myocardial infarction despite better statistical power, suggesting confounding by indication or attrition bias in previous analyses. To test its clinical value, valproate is currently under investigation in the Sodium Valproate to Prevent Stroke (SOLVE) trial for the prevention of atherosclerosis progression in patients with large-artery stroke (ISRCTN12685153).
We were unable to investigate valproate’s effect on specific stroke subtypes because this information is not available in the UKB, limiting our insight into a specific mechanism. However, we found a stronger association in individuals without atrial fibrillation despite reduced statistical power, suggesting that valproate is not preventing cardioembolism. The lack of association with myocardial infarction could suggest that valproate’s stroke prevention mechanism acts through other pathways than slowing of atherosclerosis, such as protection of the blood brain barrier, 21 increase in ischemic tolerability,22,23 inhibition of platelet aggregation, 24 or increase in tissue plasminogen activator. 25
We found the strongest effect of valproate on ischemic stroke among individuals with prior stroke, despite a small cohort size. Existing data shows that post-stroke epilepsy is common, with an overall incidence of up to 7%. 26 If future studies confirm our results, future trials could evaluate its utility as dual-use secondary prevention for post-stroke epilepsy.
The most likely reason for the conflicting evidence for the effect of valproate from observational studies is confounding by indication. In our cohort, valproate users had a higher number of vascular risk factors compared to the general population. Thus, association of valproate in solely epidemiological models would yield an increased risk for vascular outcomes due to confounding. Our analyses show that in these cases, Mendelian randomization can overcome this challenge by randomizing individuals to similar baseline characteristics, providing an unbiased methodology similar to a clinical trial. This intriguing approach is enabled through the increasing availability of genetic markers for drug response. 27 While valproate’s adverse and teratogenic effects limit its overall attractiveness for drug repurposing, our findings potentially apply to other HDAC inhibitors that have been tested in vitro and in vivo, 28 providing further justification for stroke prevention trials employing HDAC inhibitors.
Our study has limitations. First, the SNPs used to construct the genetic score for valproate response were discovered in individuals of European ancestry 12 and applied to a predominantly European UKB population. However, because the leveraged genetic variants only act as instruments for randomization in our study, it is likely that our results will hold in non-European populations. Second, we could not investigate valproate’s effect on specific stroke subtypes because they are not available in the UKB. Third, since we gathered data on valproate prescriptions, and not on valproate use, we cannot be certain that our whole cohort was in fact using valproate, but non-compliance would only have diluted, not increased our effect estimates. Finally, our study cannot answer which patients would benefit most from valproate; however, we found the largest effect for prevention of recurrent ischemic stroke.
In conclusion, our Mendelian randomization–based study supports a causal role for valproate in the prevention of ischemic stroke, with the largest risk decrease for recurrent ischemic stroke. Our results provide actionable evidence for the performance of clinical trials with valproate and other HDAC inhibitors for the prevention of ischemic stroke.
Supplemental Material
sj-docx-1-wso-10.1177_17474930231190259 – Supplemental material for Genetic variation supports a causal role for valproate in prevention of ischemic stroke
Supplemental material, sj-docx-1-wso-10.1177_17474930231190259 for Genetic variation supports a causal role for valproate in prevention of ischemic stroke by Ernst Mayerhofer, Livia Parodi, Kaavya Narasimhalu, Stefan Wolking, Andreas Harloff, Marios K Georgakis, Jonathan Rosand and Christopher D Anderson in International Journal of Stroke
Supplemental Material
sj-docx-2-wso-10.1177_17474930231190259 – Supplemental material for Genetic variation supports a causal role for valproate in prevention of ischemic stroke
Supplemental material, sj-docx-2-wso-10.1177_17474930231190259 for Genetic variation supports a causal role for valproate in prevention of ischemic stroke by Ernst Mayerhofer, Livia Parodi, Kaavya Narasimhalu, Stefan Wolking, Andreas Harloff, Marios K Georgakis, Jonathan Rosand and Christopher D Anderson in International Journal of Stroke
Supplemental Material
sj-xlsx-3-wso-10.1177_17474930231190259 – Supplemental material for Genetic variation supports a causal role for valproate in prevention of ischemic stroke
Supplemental material, sj-xlsx-3-wso-10.1177_17474930231190259 for Genetic variation supports a causal role for valproate in prevention of ischemic stroke by Ernst Mayerhofer, Livia Parodi, Kaavya Narasimhalu, Stefan Wolking, Andreas Harloff, Marios K Georgakis, Jonathan Rosand and Christopher D Anderson in International Journal of Stroke
Footnotes
Acknowledgements
Declaration of conflicting interest
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: C.D.A. has received sponsored research support from Bayer and has consulted for ApoPharma unrelated to this work. J.R. reports compensation from the National Football League and Takeda Development Center Americas for consultant services unrelated to this work.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: C.D.A. is supported by NIH R01NS103924, U01NS069673, AHA 18SFRN34250007 and 21SFRN812095, and the MGH McCance Center for Brain Health. M.K.G. is supported by the FöFoLe program of LMU Munich (Reg.-Nr. 1120), the DFG Germany’s Excellence Strategy within the framework of the Munich Cluster for Systems Neurology (EXC 2145 SyNergy-ID 390857198), and the Fritz-Thyssen Foundation (10.22.2.024MN). J.R. receives research grants from NIH and the AHA-Bugher Foundation. A.H. received funding from the the Berta-Ottenstein program for Advanced Clinician Scientists, University of Freiburg. S.W. receives funding from the German Research Foundation (WO 2385/2-1).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
