Abstract
Although the molecular mechanisms of chronic pain have been extensively studied, a global picture of alternatively spliced genes and events in the peripheral and central nervous systems of chronic pain is poorly understood. The current study analyzed the changing pattern of alternative splicing (AS) in mouse brain, dorsal root ganglion, and spinal cord tissue under inflammatory and neuropathic pain. In total, we identified 6495 differentially alternatively spliced (DAS) genes. The molecular functions of shared DAS genes between these two models are mainly enriched in calcium signaling pathways, synapse organization, axon regeneration, and neurodegeneration disease. Additionally, we identified 509 DAS in differentially expressed genes (DEGs) shared by these two models, accounting for a small proportion of total DEGs. Our findings supported the hypothesis that the AS has an independent regulation pattern different from transcriptional regulation. Taken together, these findings indicate that AS is one of the important molecular mechanisms of chronic pain in mammals. This study presents a global description of AS profile changes in the full path of neuropathic and inflammatory pain models, providing new insights into the underlying mechanisms of chronic pain and guiding genomic clinical diagnosis methods and rational medication.
Keywords
Introduction
Chronic pain is a major source of distress, imposing a substantial personal and economic burden. 1 Over 30% of the global population suffers from chronic pain, which interferes with daily functioning and stands as the leading cause of chronic disabilities. 2 Inflammatory and neuropathic pain are two primary types of chronic pain, sharing several neural mechanisms, including heightened primary afferent excitability, reduced spinal cord inhibitory tone, and involvement of higher brain centers. The development of therapeutic drugs for chronic pain is challenging on account of multiple interdependent factors involved, including pathological stimuli, neuronal sensitization and remodeling, inflammation, etc. Despite progress in uncovering neuronal alterations in pain pathways, the genetic programs and molecular mechanisms underpinning these alterations remain unclear.
Clinically, pain symptoms are primarily assessed based on the patient’s subjective statement. There is a lack of biomarkers to predict and classify the type of chronic pain at the individual level, which may be largely due to the molecular mechanism of pain generation that remains unclear. With the advancement of RNA sequencing technology, numerous studies, including our previous works,3–7 have identified differentially expressed genes (DEGs) related to chronic pain through transcriptomic analysis. However, the transcriptome information remains incomplete, lacking a comprehensive analysis of alternative splicing (AS) patterns of transcripts.
AS is a ubiquitous transcriptional regulatory mechanism in higher eukaryotes, contributing to transcriptome and proteome diversity.8–10 Extensive AS is a major source of protein diversity in over 90% of human genes, serving as important molecular markers of various human diseases and potential targets for novel therapeutics. 11 Aberrated AS of genes has been implicated in vital roles in cancer, autoimmune, and neurodegenerative diseases.12–15 Growing evidence indicates that translational regulation of mRNAs in the peripheral and central nervous systems plays a crucial role in the regulation of chronic pain.16–19 A systematic analysis of pain-related AS can aid in understanding the biological process of transcript isoform diversity under inflammatory neuropathic conditions and provide new targets for pain treatment.
In the current study, we conducted a comprehensive and systematic analysis of AS patterns in two common types of chronic pain. We identified differentially alternatively spliced (DAS) genes in the mouse brain, dorsal root ganglion (DRG), and spinal cord in inflammatory and neuropathic pain induced by plantar injection of complete Freund’s adjuvant (CFA) and spared nerve injury (SNI), respectively. In the brain, DRG, and spinal cord of the CFA model, 1100, 955, and 1245 DAS genes were identified, while 1080, 1014, and 1101 DAS genes were identified in the corresponding tissues of the SNI model. Furthermore, we identified the DAS genes shared between neuropathic and inflammatory pain and demonstrated the potential role of these genes through Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis. The brain, DRG, and spinal cord in both chronic pain models shared 478, 428, and 512 DAS genes, respectively. Additionally, we extracted all DEGs in DAS genes and further analyzed the expression patterns of these DEGs in different tissues of the pain conduction pathway in neuropathic and inflammatory models. To the best of our knowledge, the present study is the first to provide a comprehensive view of AS in chronic pain on a genome-wide scale. It offers a global description of AS profile changes in the full path of neuropathic and inflammatory pain models, providing new insight into the underlying mechanisms of chronic pain and guiding genomic clinical diagnosis and rational medication.
Methods
Data collection
We obtained a dataset of 27 samples sequenced based on the Illumina HiSeq 2000 platform, paired-end RNA-seq data with 100 nt read length, from NCBI’s Gene Expression Omnibus 20 with accession number GSE111216, including inflammatory and neuropathic pain models. 21 A total of nine 18-week-old BALB/c female mice were randomly assigned into 3 groups: control (CTR, naïve), treated with complete Freund’s adjuvant (CFA), and underwent spared nerve injury (SNI). To reduce the overall number of mice required, the CFA injection or SNI surgery was performed on both the left and right sides. Bilateral tissues and biological information were collected from L3-L5 DRG, the dorsal horn of the SC, and the whole brain. Since CFA injection–induced hypersensitivity peaks on day 3 and SNI-induced hypersensitivity develops to its maximal extent on day 7, tissues were extracted 3 days after CFA injection and 7 days after SNI. The Summary of RNA-Seq data from each group with 3 biological replicates is shown in Supplemental Table S1.
Screening of differentially alternative splicing genes
The twopassMode of STAR (version 2.7.10a) was used to map reads to the mouse reference genome (mm10), 22 and samtools (version 1.3.1) was used to sort the bam file after mapping and to establish the index. 23 rMATS (version 4.1.2) and rmats2sashimiplot (version 2.0.4) are used to identify and visualize the alternative splicings. 24
GO and KEGG analysis
Gene ontology (GO) analysis and significantly enriched Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways of gene functional annotation clustering were performed by ClusterProfiler package (version 4.10.0) of R with mus musculus as background, default options, and annotation categories were used. 25 Statistics were performed using a modified Fisher’s exact test followed by Benjamini-Hochberg multiple hypothesis testing corrections.
Screening the intersection genes between DAS genes and DEGs
The clustering analysis of DAS genes and DEGs was performed using EVenn. 26 (https://www.ehbio.com/test/venn/#/ (accessed on 8 September 2023)). The list of DEGs was downloaded from the previously published work. 21 (https://links.lww.com/PAIN/A722). DAS genes from each group screened by the current work can be found in Supplemental Tables S2–7.
Statistical analysis
All statistical analyses were conducted using R language (version 4.3.2). Statistical tests used are indicated where the tests are performed. Correction for multiple testing is applied whenever multiple tests are involved, along with a statement indicating which method of correction is used. p values less than 0.05 were considered to be significantly different for each test unless specifically indicated at the location of the test.
Results
Summary of alternative splicing events in brain, DRG, and spinal cord of CFA and SNI models
Eukaryotic genomes contain mutually spaced exons and introns, and the transcribed mRNA precursors can produce different mRNA splicing isoforms through alternative splicing (AS), potentially leading to structurally and functionally diverse proteins.9,27 AS is a major source of complexity for eukaryotic transcripts, occurring in approximately 95% of multi-exonic genes in eukaryotic organisms, and can be divided into 5 basic forms (Figure 1(a)): skipped exon (SE), retained intron (RI), Alternative 5′ splice site (A5SS), Alternative 3′ splice site (A3SS), and mutually exclusive (MXE).28,29 To investigate the distribution of AS events in inflammatory and neuropathic pain models, we conducted a statistical analysis of the frequency of AS events in brain, DRG, and spinal cord tissues, respectively. We identified 53,446, 50,299, and 54,020 AS events in the brain, DRG, and spinal cord of the CFA model, and 52,300, 49,936, and 53,008 in the SNI model, respectively (Figure 1(b)). Exon skipping events predominated in all three tissues of both chronic pain models. Classification and statistics of alternatively splicing (AS) events. (a) Illustrations for five types of AS events, including alternative 3′ splice site (A3SS), alternative 3′ splice site (A5SS), mutually exclusive (MXE), skipped exon (SE), and retained intron (RI). (b) The number of AS events in the brain, DRG, and spinal cord from the CFA and SNI pain models, respectively. The color bar represents the number of different types of preliminarily detected AS events.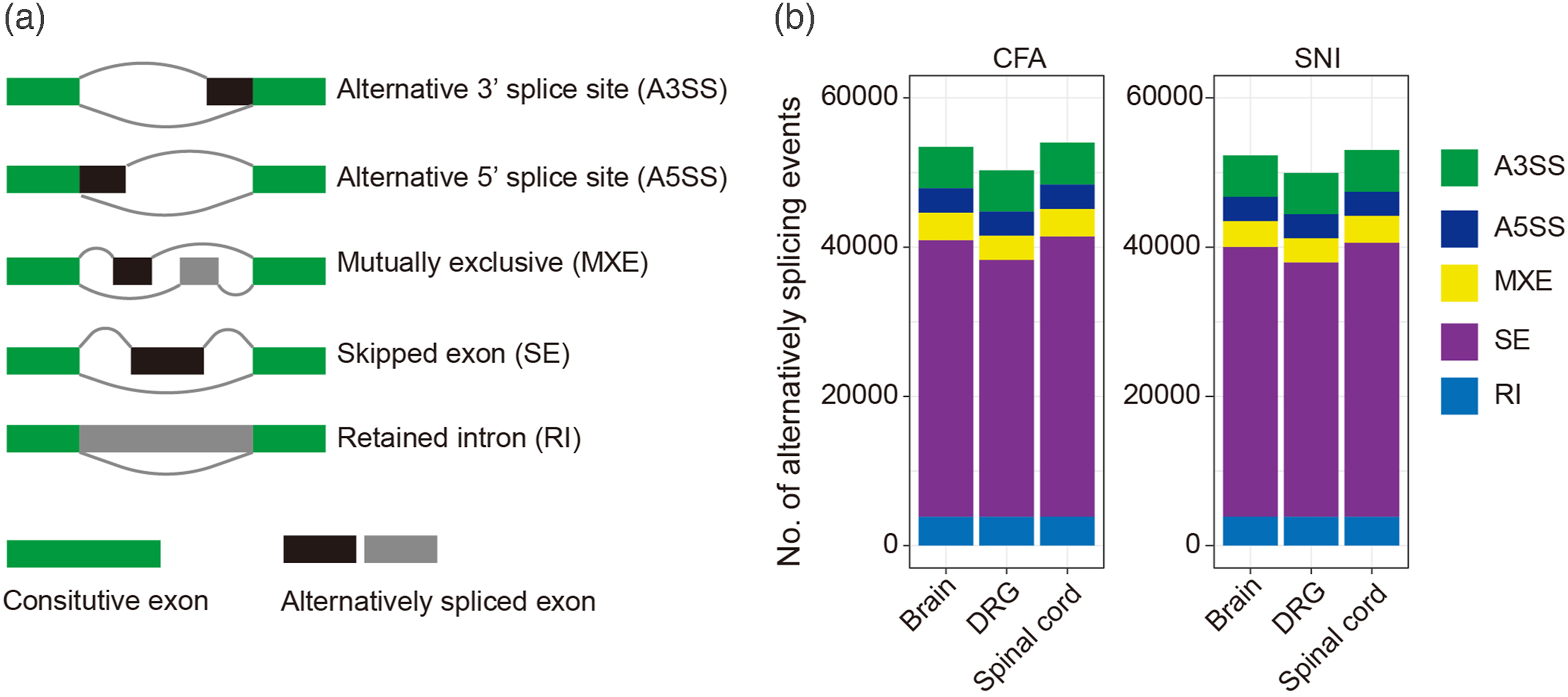
Screening and identification of differential alternative splicing events and genes
We identified DAS events with a p-value less than 0.05 by comparing them to naïve mice as the control group for both CFA and SNI models. A total of 1384, 1187, and 1674 DAS events were identified in the brain, DRG, and spinal cord of the CFA model, and 1341, 1289, and 1422 DAS events were identified in the SNI model, respectively (Figure 2(a)). Next, we identified DAS genes according to the screening results. There were 1100, 955, and 1245 DAS genes identified in the brain, DRG, and spinal cord of the CFA model, and 1080, 1014, and 1101 DAS genes identified in the SNI model, respectively (Figure 2(b)). The two chronic pain models shared 478, 428, and 512 DAS genes in the brain, DRG, and spinal cord, respectively. Examples of DAS genes shared by the two chronic pain models in different tissues are shown in Figure 2(c). Screening and comparison of differentially alternatively spliced (DAS) genes in the brain, DRG, and spinal cord of two chronic pain models. (a) The distribution of DAS events in different tissues of CFA and SNI pain models. (b) Venn diagram shows the number of DAS genes shared by two chronic pain models in the brain, DRG, and spinal cord. (c) Examples of DAS genes shared by two chronic pain models in different tissues.
Functional enrichment analysis of differentially alternative spliced genes
To gain deeper insights into the biological processes, cellular components, and molecular functions of the DAS genes shared between neuropathic and inflammatory pain, we used the clusterProfiler package for GO analysis to conduct enrichment analysis and classifications. The GO analysis identified that in the brain of both chronic pain models, the biological processes of “cell junction assembly,” the cellular component term of “actin binding,” and the molecular function of “cell cortex” were significantly enriched (Figure 3(a)). In the DRG, the enriched biological processes were mainly for “axonogenesis” and “synapse organization,” and the regulation of “cell morphogenesis,” “synapse organization,” and “synapse structure or activity.” The enriched cellular components were the “asymmetric synapse,” and the molecular function terms were “actin binding” and “transcription coregulator activity” (Figure 3(b)). In the spinal cord, biological processes of “synapse organization,” the cellular component term of “neuron to neuron synapse,” and the molecular function of “Ras and small GTPase binding” were significantly enriched in both chronic pain models (Figure 3(c)). These results suggest that DAS genes are mainly involved in synapse organization in the peripheral nervous system and spinal cord of neuropathic and inflammatory pain. Taken together, these findings suggest different mechanisms underlying chronic pain and that different AS profiles are involved in the peripheral and central nervous system in neuropathic and inflammatory pain. Functional analysis of DAS genes in the brain, DRG, and the spinal cord from shared genes between neuropathic and inflammatory pain. (a)–(c) GO analysis showing the enriched terms of DAS genes in biological process (purple font), cellular component (blue font), and molecular function (green font) from pairs of comparable groups. The significantly enriched GO terms determined by –log10 (corrected p-value) are plotted as the ordinate and the enriched gene number is plotted as the abscissa.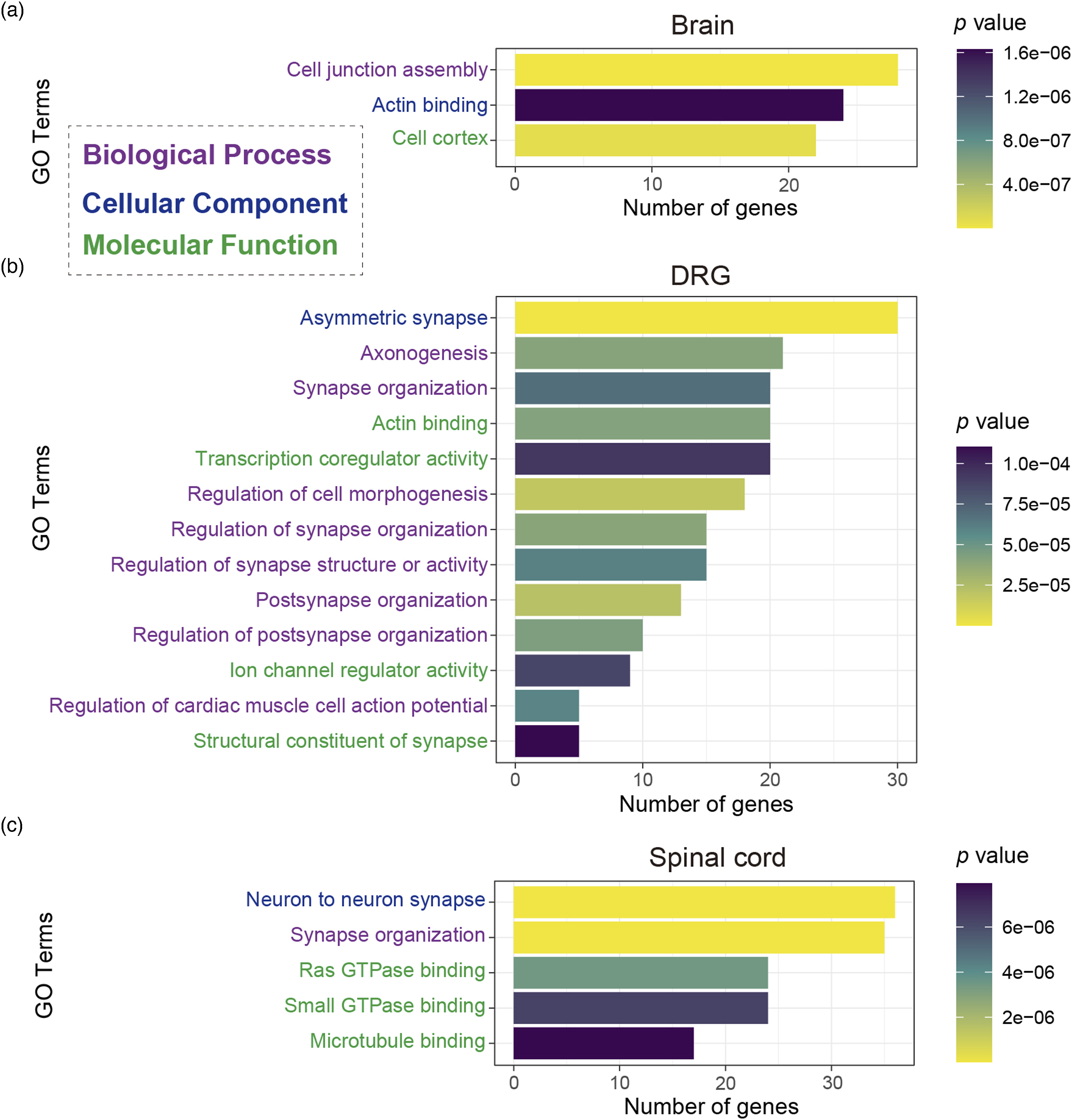
KEGG analysis of differentially alternatively spliced genes
To investigate the major signaling pathways involved in DAS genes shared between neuropathic and inflammatory pain, KEGG pathway enrichment was performed using the clusterProfiler package. The analysis showed that the DAS genes were significantly enriched in the classifications of the “MAPK signaling pathway,” “endocytosis,” and “JAK-STAT signaling pathway” in the brains of both chronic pain models (Figure 4(a)). Consistent with previous findings, the DAS genes were mainly enriched in “pathways of neurodegeneration,” “Parkinson’s disease,” “calcium signaling pathway,” and “axon guidance” in the DRG and spinal cord, respectively (Figure 4(b) and (c)). Notably, the “calcium signaling pathway” and “axon guidance” were enriched in both the DRG and spinal cord of both chronic pain models. These results suggest that neurodegeneration, axon guidance, and the involvement of calcium signaling pathways are the major signaling pathways in the peripheral nerves and spinal cord levels, whereas pathways associated with cell signal transduction are the main processes of chronic pain in the higher central nervous system. KEGG classifications of DAS genes in the brain, DRG, and the spinal cord from shared genes between neuropathic and inflammatory pain. (a)–(c) The comparison of pathway enrichment from pairs of comparable groups. The top 20 KEGG terms determined by the rich factor are plotted as the ordinate. The rich factor is plotted as the abscissa. The gene number involved in each pathway is represented by the size of the dots.
Identification of the intersection genes between DAS genes and DEGs
To further investigate the landscape and expression patterns of DAS genes involved in inflammatory and neuropathic pain, we overlapped DEGs and DAS genes from the brain, DRG, and spinal cord of CFA and SNI models, respectively. The list of the intersection of DAS genes and DEGs, referred to as DAS DEGs, in different tissues of CFA and SNI models can be found in Table S8. We found that a small portion of DAS genes overlapped with DEGs, ranging from 2.7% to 17.7% in the brain, DRG, and spinal cord of CFA and SNI models, respectively (Figure 5(a) and (b)). Notably, the proportion of DAS DEGs in the spinal cord of CFA (12.9%) and DRG of the SNI model (17.7%) was quite high compared to other subgroups (less than 5%, Figure 5(a) and (b). These results indicate that AS events of DEGs often occur in the spinal cord of the CFA model and the DRG of the SNI model, suggesting that these AS events may be linked to distinct pathological mechanisms underlying inflammatory and neuropathic pain, respectively. Identification of the intersection genes between DAS genes and DEGs. (a) and (b) Venn diagram shows the number of intersection genes between DAS genes and DEGs in the brain, DRG, and spinal cord of CFA and SNI pain models, respectively. (c) Venn diagram and Bar plots show the statistics of all intersection genes from each comparable group. (d) The list of intersection genes shared by different comparable groups. The red font highlights the upregulated DAS genes and the blue font indicates the downregulated DAS genes in the left corresponding groups. Up and down arrows indicate the same DAS genes were up-regulated and down-regulated respectively in different situations.
We then analyzed the distribution and expression levels of DAS DEGs in CFA and SNI models. Our study revealed that a small subset of DAS DEGs is shared in different tissues of CFA and SNI models. Specifically, we found 132 DAS DEGs that only occur in the spinal cord of the CFA model and 153 of that only occur in the DRG of the SNI model, accounting for the majority of pain-related AS events (Figure 5(c)). These results suggest that DAS DEGs-driven AS events lead to different pathological mechanisms of inflammatory and neuropathic pain. To further explore the similarities of DAS DEGs involved in the mechanisms of inflammatory and neuropathic pain, we analyzed the gene expression patterns of DAS DEGs shared by CFA and SNI models (Figure 5(d)). We identified 3 DAS DEGs, including Apc, Ly6e, and Atrx, that were shared by both CFA and SNI models across more than two tissues. Sp100 was the only DAS DEGs shared in all three types of tissue in the CFA model. Additionally, we found 3, 7, and 8 DAS DEGs shared in the brain, DRG, and spinal cord between the CFA and SNI models, respectively. Interestingly, all the DAS DEGs shared by the DRG and spinal cord showed the same trend of regulation, while DAS DEGs shared by the brain showed the opposite trend of regulation. Notably, there are only 10 DAS DEGs shared between the spinal cord of the CFA model and the DRG of the SNI model, with 9 of them showing the same trend of regulation and 1 showing the opposite regulation trend (Figure 5(d)). These results suggest that the AS mechanisms of a few DAS DEGs are shared between inflammatory and neuropathic pain.
Discussion
The conduction and transformation pathways of pain in mammals are highly complex. Peripheral signals indicative of inflammation or nerve injury are detected by sensory neurons. The input information from DRG neurons is processed at multiple levels of the central nervous system, from the spinal cord to the brain. 30 Numerous cellular molecules and processes can affect neuronal properties, thereby altering the information encoded within the pathway. Many of these molecules can be alternatively spliced and targeted to control pain information transmission. 31 While the molecular mechanisms of chronic pain have been extensively studied, a global understanding of alternatively spliced genes and events in the peripheral and central nervous systems of classic chronic pain, including inflammatory and neuropathic pain, is still lacking. In this study, we conducted a transcriptomic analysis of the brain, DRG, and spinal cord selected from inflammatory and neuropathic pain models to investigate the similarities and differences in AS events induced by inflammation and nerve injury, respectively. Therefore, this study contributes to a comprehensive understanding of the role of gene alternative splicing in different pathogenesis of inflammatory and neuropathic pain.
Our study revealed that the total number and subtypes of AS events that occurred in the brain, DRG, and spinal cord tissues of mice with inflammatory and neuropathic pain were similar. The total number of AS events in DRG was slightly less than that in the spinal cord and brain (Figure 1). Consistent with these findings, nociception-associated AS events have also been observed in humans, 32 horses, 33 and Drosophila 34 through experimental studies. However, a systematic investigation of the AS events and genes associated with pain or nociception in other species has not been reported. It has been demonstrated that AS can increase proteome diversity and regulate pain sensitivity and morphine tolerance in humans and rodents.32,35–37 Most splicing errors are eliminated by mRNA monitoring mechanisms, such as nonsense-mediated decay. 38
By conducting DAS genes screening, we identified 1187 to 1674 DAS events involving 955 to 1245 DAS genes in the CFA and SNI models. Notably, there were 478, 428, and 512 DAS genes shared by CFA and SNI models in the brain, DRG, and spinal cord, respectively (Figure 2). Our GO and KEGG analyses revealed significant enrichment of DAS genes in GO terms such as “cell cortex” and “cell junction assembly,” as well as KEGG pathways including the “MAPK signaling pathway,” “endocytosis,” and “JAK-STAT signaling pathway” in the brain of both CFA and SNI models (Figures 3 and 4). While few studies have reported associations between AS occurring in the brain and pain sensation or analgesia, evidence suggests that AS of the Mu opioid receptor is required for analgesia in rodents and the human brain.32,36,37 Additionally, AS of NMDA receptor NR1 subunit induced by morphine administration is involved in morphine tolerance. 35 Future studies may focus on the AS events that occurred in the “MAPK signaling pathway” and “JAK-STAT signaling pathway” in the brain associated with chronic pain.
In the DRG and spinal cord, DAS genes were significantly enriched in GO and KEGG terms related to synapse organization, axon regeneration, the calcium signaling pathway, and neurodegeneration disease (Figures 3 and 4). To date, limited information is available about the functional implications and control of the expression of splice variants associated with pain sensation and analgesia. However, existing knowledge primarily centers on the functional implications of expression of different splice variants of voltage-gated calcium channels16,39,40 and sodium channels,41–43 particularly Cav2.2.18,44,45 The alternative splice variant of Cav2.2 is enriched in DRG and the spinal cord, and its presence increases sensitivity to neuronal inhibition through opioid receptors.24,46–48 Similarly, VEGFA splice variants have been widely studied in peripheral neuropathy,49,50 diabetic peripheral neuropathy, 17 and pelvic pain. 24
Furthermore, splice variants of several neurotransmitters, growth factors, and ion channels have also been reported, including acetylcholine, 51 nestin, 52 and TRPA1 channel.53,54 Emerging evidence suggests that secreted phosphoprotein 1, 55 carbonic anhydrase-8, 56 and pituitary adenylate-cyclase activating polypeptide 57 are associated with the generation of pain and analgesia. Our findings align with previous research that has reported on splice variants of neuregulin 1, which regulate neuronal cell adhesion, and the neuronal cell adhesion molecule (NrCAM), which links neuronal cells, playing critical roles in neuropathic pain.58–60 While splicing control in pain pathways has been a well-established area of research for chronic pain interventions, the majority of previous studies have concentrated on splicing variants in various receptors and channels crucial to pain pathways. Consequently, the potential implications of many new splicing events identified in the current study necessitate further investigation.
By intersecting the DEGs and DAS genes, we identified 30 to 179 DAS DEGs associated with inflammatory and neuropathic pain, accounting for 2.3% to 6.1% of DEGs in the brain, DRG, and spinal cord, respectively (Figure 5(a) and (b)). These results suggest that AS may not be the primary cause of gene expression changes for most DEGs associated with inflammatory and neuropathic pain. In other words, changes in AS of genes do not necessarily result in changes in gene expression. This conclusion is supported by previous research on heat/cold stress in rats and plants, which indicated that AS has an independent regulatory pattern distinct from transcriptional regulation.61,62
While most of the DAS DEGs exhibited consistent expression levels, several genes, including Apc, Atrx, Col11a1, Dgkh, Homer1, Fbln2, TRP53i11, and Enah, showed controversial expression levels across different tissues or models (Figure 5(d)). Despite the alternative splicing (AS) of Nrcam, which has been implicated in neuropathic pain,58,59 and the involvement of Nrxn2 in spinal muscular atrophy has recently been reported, 63 no other splicing variants of the DAS genes shared by CFA and SNI models have been reported to be involved in pain or analgesia. Some of the splicing variants of these DAS genes have been implicated in cancer and other diseases. Many DEGs related to chronic pain are accompanied by alternative splicing, while the potential functions and underlying regulatory mechanisms are still not well understood and require further exploration in future studies.
It is worth noting that in the DRG and spinal cord, the DAS genes shared by the CFA and SNI models are involved in similar biological processes and signaling pathways, distinct from those in the brain according to the GO and KEGG analysis (Figures 3 and 4). Subsequent analyses revealed an uneven distribution of DAS DEGs in the brain, spinal cord, and DRG, with a predominant presence in the spinal cord of the CFA model and the DRG of the SNI model, and only a small overlap of DAS DEGs shared between both models. Interestingly, the shared DAS DEGs exhibited a consistent expression trend in the DRG and spinal cord but a reversed trend in the brain (Figure 5). These findings suggest that the functions of DAS genes in the DRG and spinal cord in inflammatory pain and neuropathic pain are similar, while the involvement of DAS DEGs and underlying mechanisms may differ between inflammatory and neuropathic pain. Consequently, further research is warranted to investigate the specific DAS genes and pathways contributing to these distinctions.
Recent studies have demonstrated that SNI would induce the formation of new neuronal clusters in the DRG, which also undergo dynamic changes in their molecular characteristics at various time points following SNI. 64 Further investigation is required to understand the dynamic AS alteration in transcriptome associated with the development of chronic pain in SNI and CFA pain models. While data from animal models can provide valuable insights, further studies are needed to explore the role and mechanism of AS events in the development of chronic pain in higher animals and humans.
Conclusions
This study investigated the changing patterns of AS in mice brain, DRG, and spinal cord tissue under inflammatory and neuropathic pain. A total of 6495 DAS genes were identified, with the molecular functions of shared DAS genes enriched in calcium signaling pathways, synapse organization, axon regeneration, and neurodegeneration disease. Additionally, 509 DAS DEGs were identified and found to be shared by inflammatory and neuropathic pain. However, the potential function and underlying regulatory mechanisms of these genes are not yet fully understood. Overall, these findings suggest that AS plays a crucial role as a molecular mechanism in chronic pain in mammals. The DAS DEGs identified in this study could potentially serve as candidates for classifying chronic pain at the gene level or as targets for further research into analgesia.
Supplemental Material
Supplemental Material - Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain
Supplemental Material for Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain by Mingzhu Zhai, Jiabin Huang, Shaomin Yang, Na Li, Jun Zeng, Yi Zheng, Wuping Sun and Benqing Wu in Molecular Pain.
Supplemental Material
Supplemental Material - Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain
Supplemental Material for Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain by Mingzhu Zhai, Jiabin Huang, Shaomin Yang, Na Li, Jun Zeng, Yi Zheng, Wuping Sun and Benqing Wu in Molecular Pain.
Supplemental Material
Supplemental Material - Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain
Supplemental Material for Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain by Mingzhu Zhai, Jiabin Huang, Shaomin Yang, Na Li, Jun Zeng, Yi Zheng, Wuping Sun and Benqing Wu in Molecular Pain.
Supplemental Material
Supplemental Material - Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain
Supplemental Material for Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain by Mingzhu Zhai, Jiabin Huang, Shaomin Yang, Na Li, Jun Zeng, Yi Zheng, Wuping Sun and Benqing Wu in Molecular Pain.
Supplemental Material
Supplemental Material - Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain
Supplemental Material for Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain by Mingzhu Zhai, Jiabin Huang, Shaomin Yang, Na Li, Jun Zeng, Yi Zheng, Wuping Sun and Benqing Wu in Molecular Pain.
Supplemental Material
Supplemental Material - Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain
Supplemental Material for Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain by Mingzhu Zhai, Jiabin Huang, Shaomin Yang, Na Li, Jun Zeng, Yi Zheng, Wuping Sun and Benqing Wu in Molecular Pain.
Supplemental Material
Supplemental Material - Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain
Supplemental Material for Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain by Mingzhu Zhai, Jiabin Huang, Shaomin Yang, Na Li, Jun Zeng, Yi Zheng, Wuping Sun and Benqing Wu in Molecular Pain.
Supplemental Material
Supplemental Material - Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain
Supplemental Material for Transcriptomic analysis of differentially alternative splicing patterns in mice with inflammatory and neuropathic pain by Mingzhu Zhai, Jiabin Huang, Shaomin Yang, Na Li, Jun Zeng, Yi Zheng, Wuping Sun and Benqing Wu in Molecular Pain.
Footnotes
Author contributions
Mingzhu Zhai: Conceptualization; Data curation; Funding acquisition; Writing – original draft; Writing – review & editing. Jiabin Huang: Conceptualization; Data curation; Writing – original draft; Writing – review & editing. Shaomin Yang: Data curation; Writing – review & editing. Na Li: Project administration; Data curation; Writing – review & editing. Jun Zeng: Project administration; Data curation; Writing – review & editing. Yi Zheng: Data curation; Writing – review & editing. Wuping Sun: Conceptualization; Supervision; Writing – original draft; Writing – review & editing. Benqing Wu: Conceptualization; Funding acquisition, Supervision; Writing – original draft; Writing – review & editing.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by The Medical and Health Science and Technology Plan Project of Shenzhen Yantian District Science and Technology Innovation Bureau (Number: YTWS20220208), The Shenzhen Municipal Science, Technology, and Innovation Commission (Number: JCYJ20220531094815034), and The Guangming District special fund for economic development (2021R01055).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
