Abstract
Phantom tooth pain (PTP) is a rare and specific neuropathic pain that occurs after pulpectomy and tooth extraction, but its cause is not understood. We hypothesized that there is a genetic contribution to PTP. We focused on solute carrier family 17 member 9 (SLC17A9)/vesicular nucleotide transporter (VNUT) and purinergic receptor P2Y12 (P2RY12), both of which have been associated with neuropathic pain and pain transduction signaling in the trigeminal ganglion in rodents. We sought to corroborate these associations in humans. We investigated gene polymorphisms that contribute to PTP. We statistically examined the association between genetic polymorphisms and PTP vulnerability in 150 patients with orofacial pain, including PTP, and 500 healthy subjects. We found that the rs735055 polymorphism of the SLC17A9 gene and rs3732759 polymorphism of the P2RY12 gene were associated with the development of PTP. Carriers of the minor allele of rs735055 and individuals who were homozygous for the major allele of rs3732759 had a higher rate of PTP. Carriers of the minor allele of rs735055 reportedly had high SLC17A9 mRNA expression in the spinal cord, which may increase the storage and release of adenosine triphosphate. Individuals who were homozygous for the major allele of rs3732759 may have higher P2RY12 expression that is more active in microglia. Therefore, these carriers may be more susceptible to PTP. These results suggest that specific genetic polymorphisms of the SLC17A9 and P2RY12 genes are involved in PTP. This is the first report on genes that are associated with PTP in humans.
Keywords
Introduction
The dental pulp nerve is connected to the tooth at the peripheral end of the trigeminal nerve. This nerve is severed during pulpectomy in routine dental practice. Approximately one million pulpectomies are performed annually in Japan. However, neuropathic pain is not usually remained after pulpectomy despite nerve damage. Moreover, tooth extraction usually does not cause persistent pain. In most cases, the procedure is performed on teeth with infected pulp, with the objective of removing pain. The pain rarely remains for a long period of time. However, there are cases of persistent pain after pulpectomy, although such cases are extremely rare. Marbach et al. referred to this condition as phantom tooth pain (PTP), similar to phantom limb pain after finger amputation,1,2 with a reported incidence of 3–6%.1,3 The actual incidence is likely far less because there are cases of prolonged root cracking, periodontitis, and residual pulpitis after pulpectomy. In the most recent classification by the International Classification of Orofacial Pain, first edition (ICOP), 4 PTP is referred to as “6.3.3. Persistent idiopathic dentoalveolar pain with somatosensory changes.” PTP is still not widely recognized, and it is often mistakenly diagnosed as temporomandibular joint disorder, trigeminal neuralgia, sinusitis, denture incompatibility, typical neuralgia, or myofascial pain. 5 The presentation of PTP is often clinically troublesome because repeated endodontic treatment, apicoectomy, or extractions do not eliminate the pain and may instead worsen pain severity or increase pain distribution. 5
Most patients with PTP meet the Diagnostic and Statistical Manual of Mental Disorders, fifth edition, criteria for somatoform pain disorder, often referred to as psychogenic pain, but there is no evidence that it derives from a pre-morbid personality.5,6 Melzack’s neuromatrix theory may be relevant because of its essentially similar characteristics to phantom limb pain after limb amputation, 5 but the cause of PTP remains unclear.
Recent studies have shown that solute carrier family 17 member 9 (SLC17A9)/vesicular nucleotide transporter (VNUT)-mediated adenosine triphosphate (ATP) release activates satellite glial cells and microglia/macrophage-like cells (MLCs) to release brain-derived neurotrophic factor and nerve growth factor as pain transduction signals in the trigeminal ganglion. 7 Purinergic receptor P2Y12 (P2RY12) is expressed on satellite glial cells, and the purinergic receptor P2X4 is expressed on MLCs. 7 In the spinal cord, SLC17A9-dependent ATP release from dorsal horn neurons8,9 and high P2RY12 expression in microglia10,11 have been reported to be involved in neuropathic pain.
SLC17A9 is a member of the SLC17 family of phosphate transporters. It is a VNUT of ATP8,12–14 and involved in the uptake and release of ATP into vesicles.8,12–15 In the nervous system, SLC17A9 is expressed in neurons of the dorsal root ganglia, astrocytes, microglia, trigeminal ganglion neurons, and satellite glial cells in the central nervous system.7–9,15,16 The SLC17A9 gene is located on chromosome 20, consists of 14 exons and 13 introns9,14 (Figure 1), and encodes SLC17A9 protein.13,14,17 Previous studies reported that the rs548728088 single-nucleotide polymorphism (SNP) of the SLC17A9 gene is associated with disseminated superficial actinic porokeratosis.
18
However, no studies have reported associations between SNPs of the SLC17A9 gene and pain. State of LD among SNPs in the SLC17A9 gene region, including 10 kbp upstream and downstream (LD Plot-r2). Numbers in squares in which two SNPs face represent the percentage of r
2
values that were calculated from genotype data of the SNPs. The white boxes represent D’ <1 and log of the likelihood odds ratio (LOD) <2. The shades of pink or red boxes represent D’ < 1 and LOD ≥2. The blue boxes represent D’ = 1 and LOD <2. The bright red boxes represent D’ = 1 and LOD ≥2. The solid horizontal line above the LD plot represents the SLC17A9 gene, including 10 kbp upstream and downstream of the gene. The orange boxes represent exons, and solid lines represent untranslated region or introns in the SLC17A9 gene structure. The gray arrows represent the direction of transcription. The yellow square represents the SNP on which we focused in this study. LD, linkage disequilibrium; SNP, single-nucleotide polymorphism; LOD, likelihood odds ratio.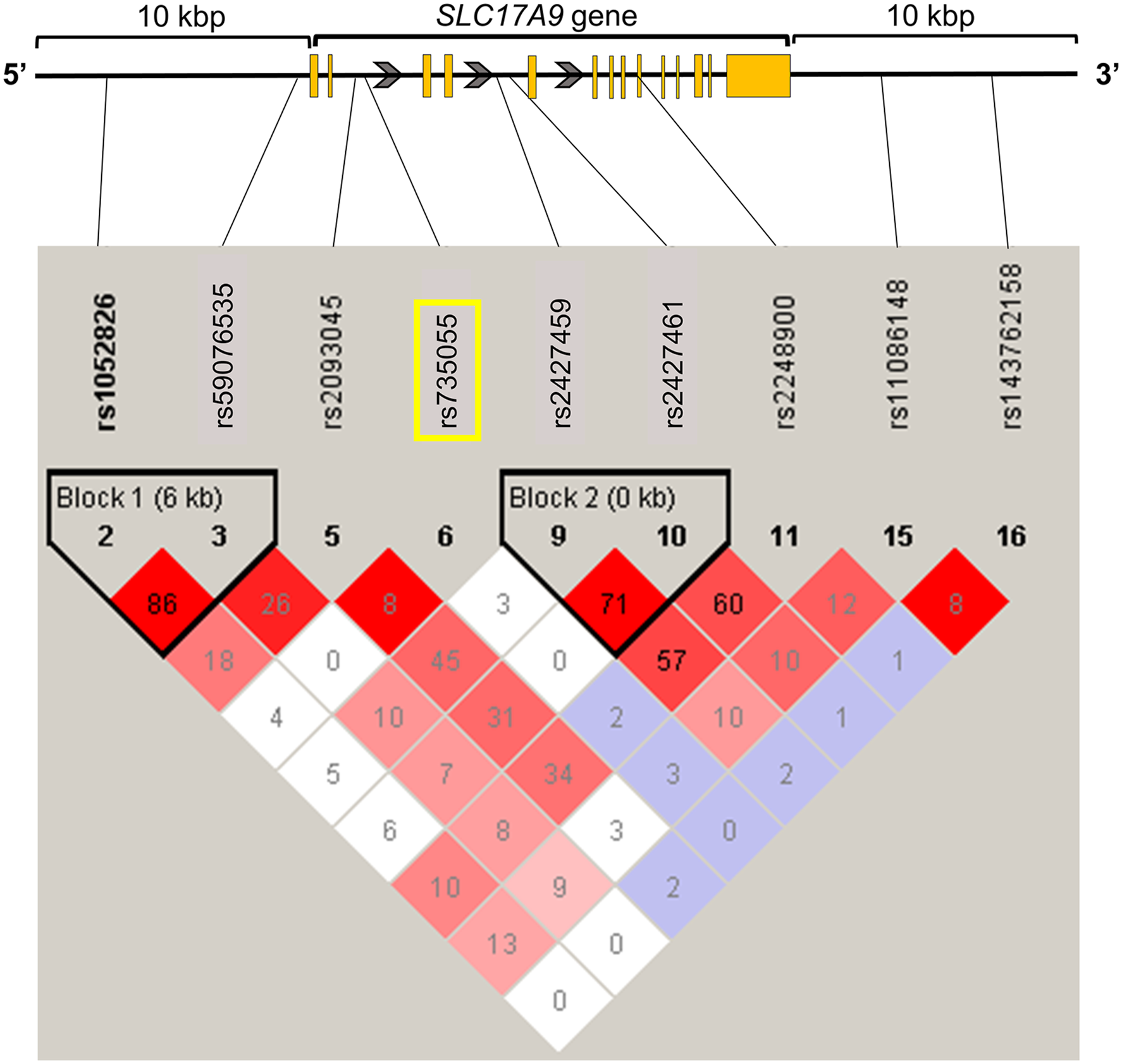
P2RY12 has seven hydrophobic transmembrane regions that are linked by extracellular/intracellular loops. It is activated by adenine and uracil nucleotides (ADP, UTP, UDP, and UDP-glucose), binds to the Giα subunit of G proteins, and decreases intracellular cyclic adenosine monophosphate production.
19
P2RY12 is reported to preferentially bind to ADP.
20
In the nervous system, P2RY12 is expressed in dorsal root ganglia, microglia, and satellite glial cells of the trigeminal ganglion.19–21 The P2RY12 gene is located on chromosome three and consists of three exons and two introns
22
(Figure 2). Previous studies reported that minor alleles of P2RY12 SNPs (rs3732765, rs9859538, rs17283010, rs11713504, and rs10935840) are associated with the severity of cancer pain.
23
State of LD among SNPs in the P2RY12 gene region, including 10 kbp upstream and downstream (LD Plot-r
2
). Numbers in squares in which two SNPs face represent the percentage of r
2
values that were calculated from genotype data of the SNPs. The white boxes represent D’ <1 and log of the likelihood odds ratio (LOD) <2. The shades of pink or red boxes represent D’ <1 and LOD ≥2. The blue boxes represent D’ = 1 and LOD <2. The bright red boxes represent D’ = 1 and LOD ≥2. The solid horizontal line above the LD plot represents the P2RY12 gene, including 10 kbp upstream and downstream of the gene. The orange boxes represent exons, and solid lines represent untranslated region or introns in the P2RY12 gene structure. The gray arrows represent the direction of transcription. The yellow square represents the SNP on which we focused in this study. LD, linkage disequilibrium; SNP, single-nucleotide polymorphism; LOD, likelihood odds ratio.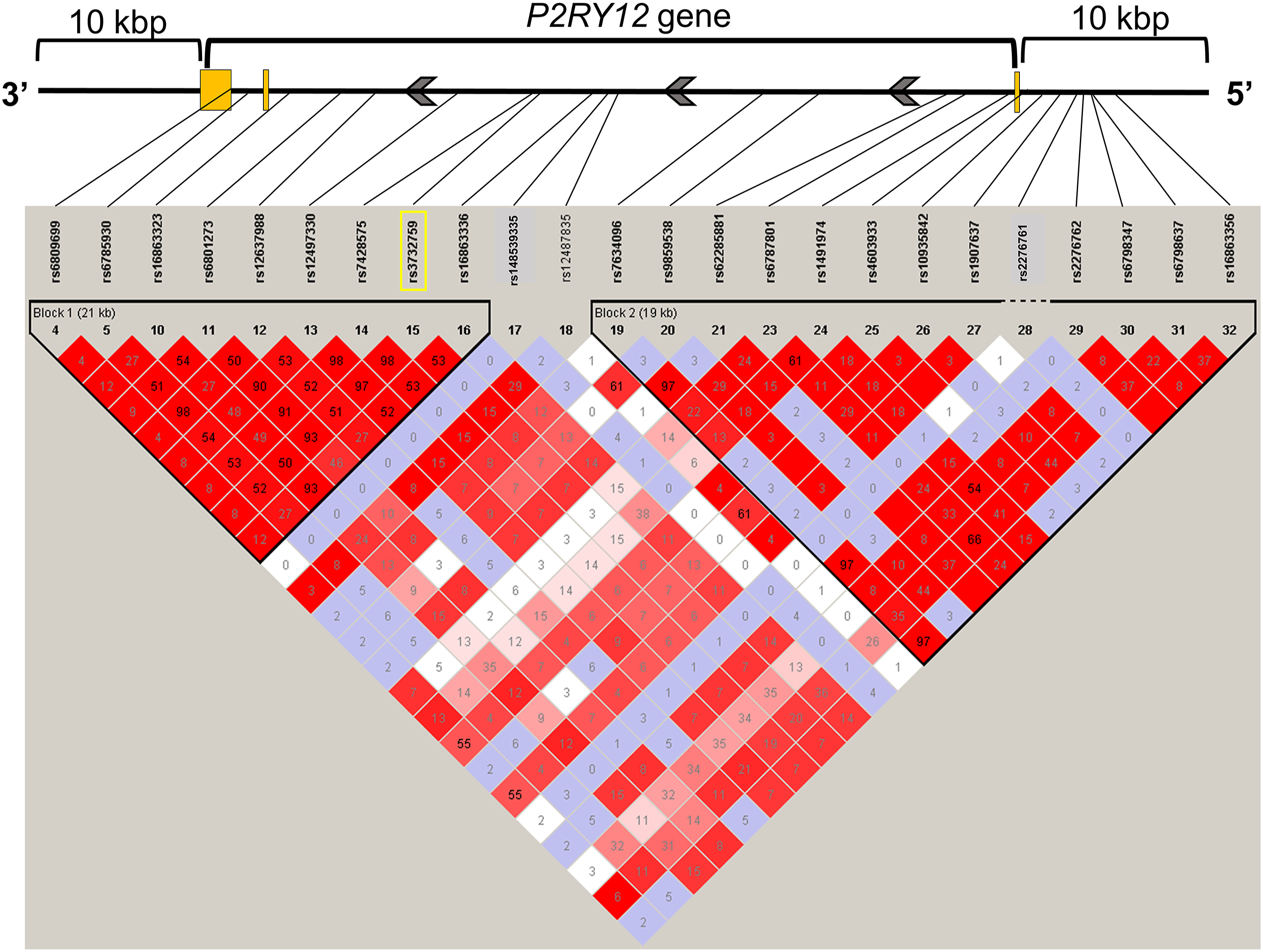
Speculating on the cause of the rare PTP condition is difficult when based solely on phenotype, such as surgical technique and environment. Some reports suggest that genetic polymorphisms are involved in pain sensitivity and resistance to the development of chronic pain.24,25 Therefore, we hypothesized that there is a genetic contribution to the vulnerability to PTP, a neuropathic pain condition in the trigeminal nerve region. In the present study, we focused on SLC17A9 and P2RY12 to investigate the cause of PTP and analyzed gene polymorphisms, particularly SNPs. The rs735055 SNP of the SLC17A9 gene and rs3732759 SNP of the P2RY12 gene were shown to be associated with the development of PTP.
Materials and methods
Patients, healthy subjects, and diagnosis
The protocol for this study was approved by the Ethics Committees of Tokyo Dental College and Tokyo Metropolitan Institute of Medical Science (approval no. 810 and 17–37, respectively) and conformed with the provisions of the Declaration of Helsinki. All of the subjects provided informed, written consent for the genetics studies.
Diagnostic criteria for phantom tooth pain that were used in the present study.
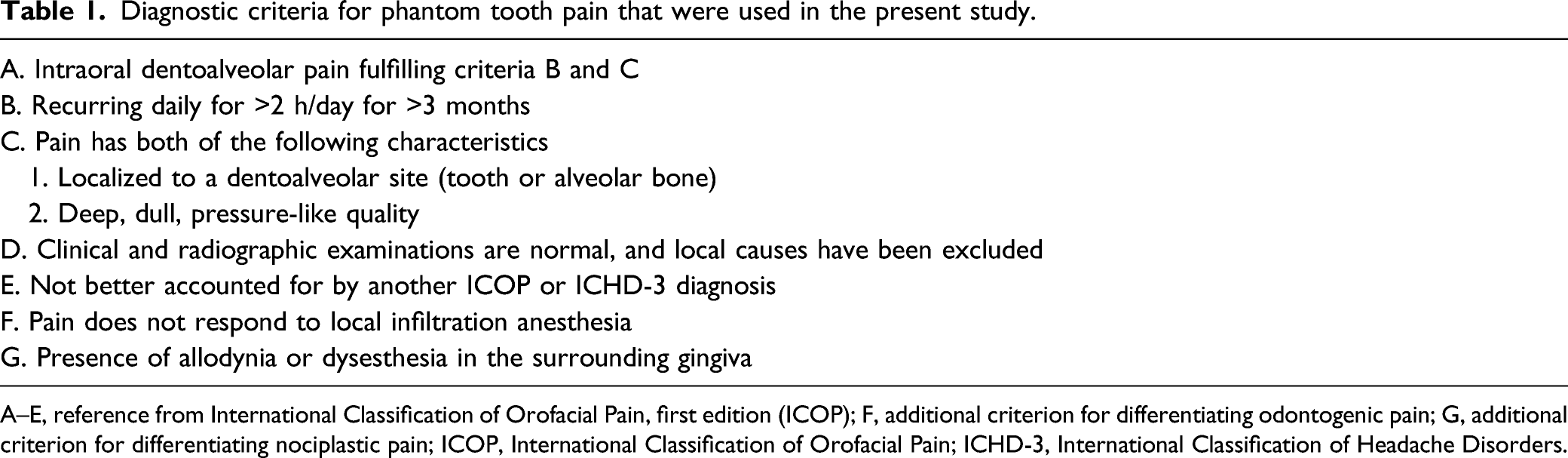

A–E, reference from International Classification of Orofacial Pain, first edition (ICOP); F, additional criterion for differentiating odontogenic pain; G, additional criterion for differentiating nociplastic pain; ICOP, International Classification of Orofacial Pain; ICHD-3, International Classification of Headache Disorders.
We also enrolled 117 patients with pain or sensory disturbance in the orofacial area, excluding PTP (i.e., orofacial pain [OFP]; 23–89 years old), who visited Tokyo Dental College Suidobashi Hospital between May 2012 and December 2019. They were classified according to the ICOP and International Classification of Diseases, 11th revision (ICD-11), as traumatic trigeminal neuropathy, trigeminal neuralgia, postherpetic neuralgia, neuralgia-inducing cavitational osteonecrosis (NICO), 26 and nociplastic pain.
Finally, we enrolled 500 healthy adult volunteers who lived in the Kanto area of Japan and agreed to participate in the study (American Society of Anesthesiologists Physical Status Ⅰ, 20–72 years old, 253 males, 242 females, and five gender-unknown subjects). They were recruited between December 2004 and April 2005 as the control group.27,28
Genotyping and linkage disequilibrium analysis
We examined SNPs of the SLC17A9 and P2RY12 genes. We analyzed 16 and 32 SNPs around the SLC17A9 and P2RY12 gene regions (including 10 kilobase pair [kbp] upstream and downstream), respectively, using genotype data from whole-genome genotyping in 150 patients who had PTP or OFP. Genomic DNA was extracted from whole-blood samples using standard procedures. The extracted DNA was dissolved in TE buffer (10 m
Among the SNPs within SLC17A9 and P2RY12 gene regions, including 10 kbp upstream and downstream, SNPs were selected for the association studies. To identify relationships of the SNPs that were used in the study, linkage disequilibrium (LD) analysis was performed using Haploview v. 4.1. 29 To estimate the LD strength between SNPs, the commonly used D’ and r 2 values were pairwise calculated using the genotype dataset of each SNP. LD blocks were defined among the SNPs that showed “strong LD,” based on the default algorithm of Gabriel et al., in which the upper and lower 95% confidence limits on D’ for strong LD were set at 0.98 and 0.7, respectively. TagSNPs in the LD block were then determined using the Tagger software package, which is incorporated in Haploview and was detailed in a previous report. 30
In 500 healthy subjects, genotype data from whole-genome genotyping were used for the rs735055 SNP. The TaqMan® assay was performed on 500 samples of the rs3732759 SNP because whole-genome genotyping data were unavailable for the rs3732759 SNP in healthy subjects. The DNA concentration was adjusted to 5–50 ng/μL for genotyping the rs3732759 SNP. To perform the TaqMan® assay with a LightCycler® 480 (Roche Diagnostics, Tokyo, Japan), TaqMan® SNP Genotyping Assays (Thermo Fisher Scientific) were used that included sequence-specific forward and reverse primers to amplify the polymorphic sequence and two probes that were labeled with VIC® and FAM™ dye to detect both alleles of the P2RY12 SNPs. The sequences of the primers for rs3732759 were not disclosed. Real-time polymerase chain reaction was performed in a final volume of 10 μL that contained 2 × LightCycler® 480 Probes Master (Roche Diagnostics), 40 × TaqMan® SNP Genotyping Assays, 5–50 ng genomic DNA as the template, and up to 10 μL H2O equipped with 2 × LightCycler® 480 Probes Master. The thermal conditions were the following: 95°C for 10 min, followed by 45 cycles of 95°C for 10 s and 60°C for 60 s, with final cooling at 50°C for 30 s. Afterward, endpoint fluorescence was measured for each sample well, and the G/G, A/G, and A/A genotypes of rs3732759 were determined based on the presence or absence of each type of fluorescence.
Statistical analysis
The patients’ demographic and clinical data are expressed as mean ± SD. Student’s t-test was performed for age and gender. The χ 2 test was performed for all genotype frequency data to investigate deviations of the distributions from those in theoretical Hardy–Weinberg equilibrium and for the association analysis with the clinical data on PTP. We performed a statistical analysis of 500 cases in the control group. Seven cases of SLC17A9 and 21 cases of P2RY12 were excluded because genotyping data were indeterminate. Student’s t-test and the χ 2 test were performed using SPSS 28 software (IBM Japan, Tokyo, Japan). The Cochran–Armitage test was performed using PLINK v1.07 (http://zzz.bwh.harvard.edu/plink/). In all of the statistical tests, the criterion for significance was set at p <.05. A trend toward a positive correlation was set at .05 ≤ p < .1.
Results
Demographic and genotype distributions
Patients’ demographic and diagnosis data.
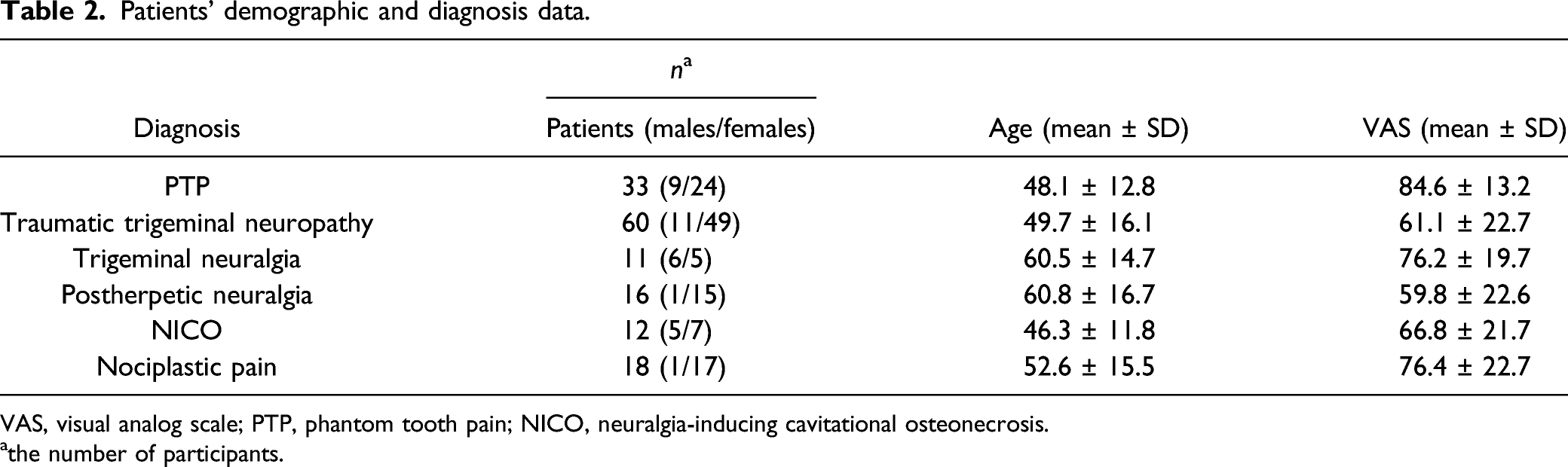
VAS, visual analog scale; PTP, phantom tooth pain; NICO, neuralgia-inducing cavitational osteonecrosis.
athe number of participants.
rs735055 single-nucleotide polymorphism of the SLC17A9 gene was associated with phantom tooth pain
Genotype distributions of SLC17A9 rs735055 single-nucleotide polymorphism.

PTP, phantom tooth pain; OFP, orofacial pain.
asubjects (males/females).
bthe number of participants.
Comparisons of genotype data between phantom tooth pain, orofacial pain and healthy subjects of SLC17A9 rs735055 single-nucleotide polymorphism.
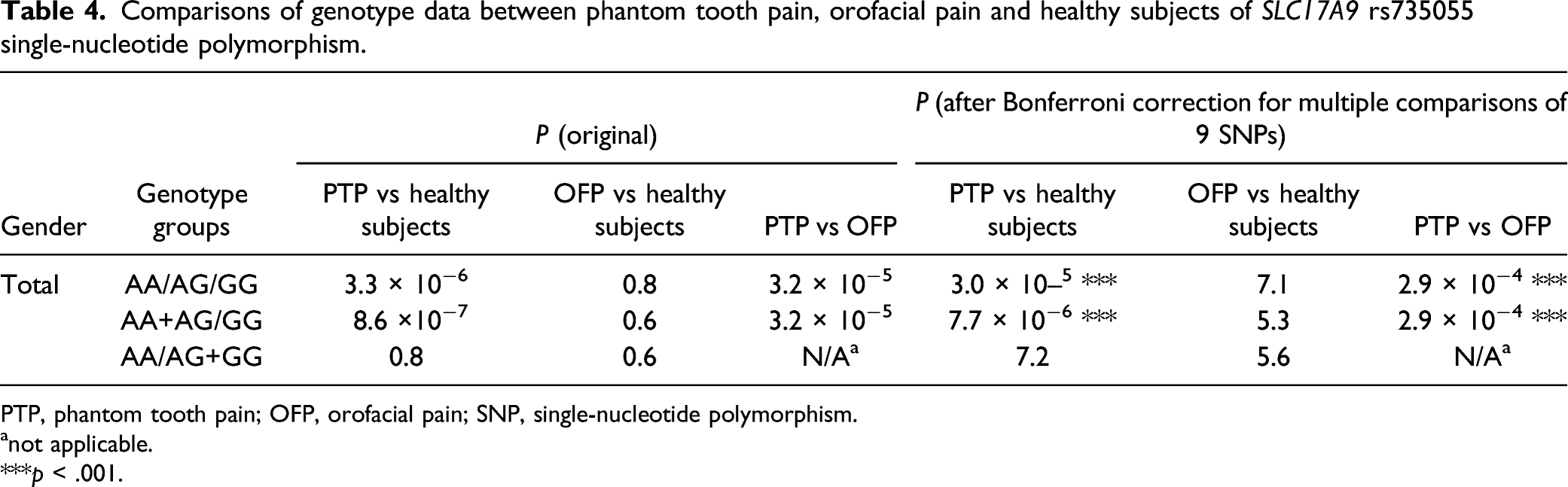
PTP, phantom tooth pain; OFP, orofacial pain; SNP, single-nucleotide polymorphism.
anot applicable.
***p < .001.
rs3732759 single-nucleotide polymorphism of the P2RY12 gene was associated with phantom tooth pain
Genotype distributions of P2RY12 rs3732759 single-nucleotide polymorphism.
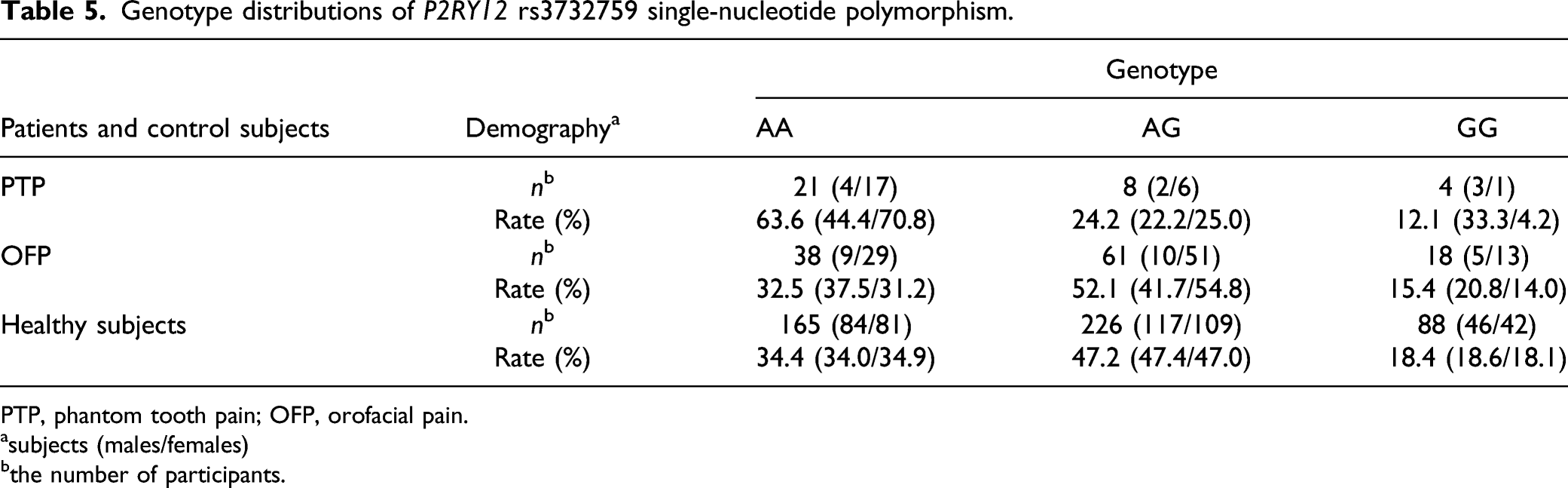
PTP, phantom tooth pain; OFP, orofacial pain.
asubjects (males/females)
bthe number of participants.
Comparisons of genotype data between phantom tooth pain, orofacial pain and healthy subjects of P2RY12 rs3732759 single-nucleotide polymorphism.
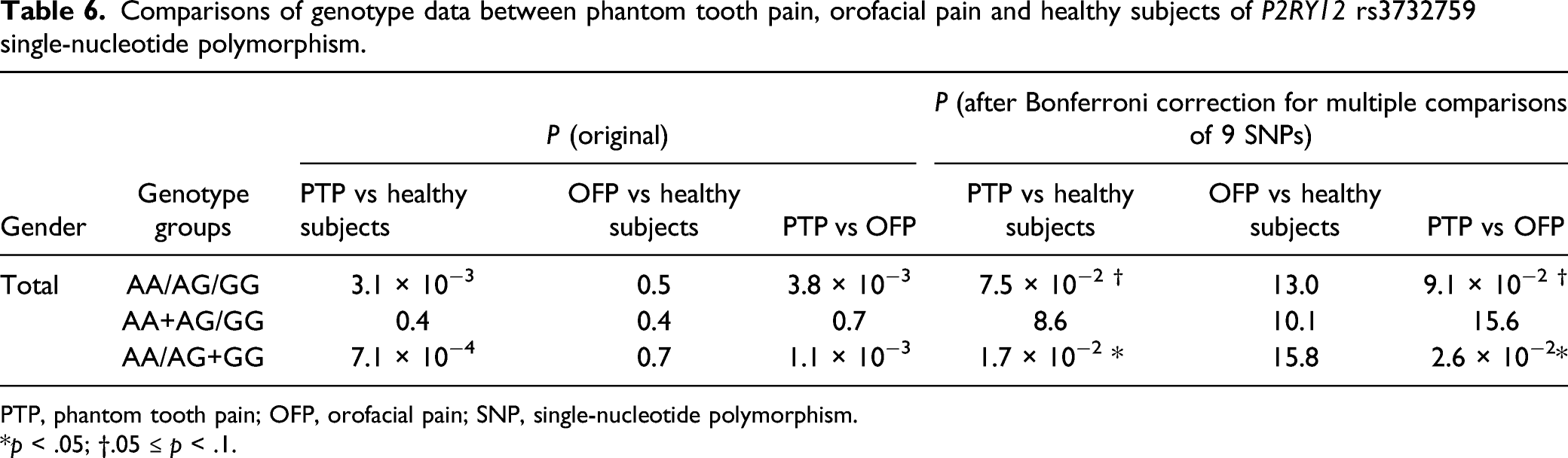
PTP, phantom tooth pain; OFP, orofacial pain; SNP, single-nucleotide polymorphism.
*p < .05; †.05 ≤ p < .1.
Discussion
The present findings suggest that the rs735055 SNP of the SLC17A9 gene and rs3732759 SNP of the P2RY12 gene are associated with the development of PTP. In males and females combined and in females alone, carriers of the minor allele of the rs735055 SNP of the SLC17A9 gene and individuals who were homozygous for the major allele of the rs3732759 SNP of the P2RY12 gene were significantly more likely to be affected by PTP. The rs735055 SNP was positively correlated and the rs3732759 SNP was negatively correlated with the copy number of the minor allele based on the linearity analysis, which indicated that they were dependent on the copy number of the minor allele. To our knowledge, this is the first report on genes that are associated with PTP in humans.
SLC17A9/VNUT-dependent ATP release from dorsal horn neurons is an essential mechanism of tactile allodynia in neuropathic pain after peripheral nerve injury,8,9 and peripheral nerve injury is reported to increase SLC17A9 expression in the spinal cord in rodents.15,17 In humans, carriers of the minor allele of the rs735055 SNP have high mRNA expression in the spinal cord, 31 which may cause high SLC17A9 expression and facilitate the efficient storage and release of ATP. Therefore, they may be more susceptible to PTP, which is a neuropathic pain condition. The rs735055 SNP of the SLC17A9 gene is located in an intron and isolated outside the LD block (Figure 1), but it is located in the promoter region, 32 which may affect transcription. The rs735055 SNP of the SLC17A9 gene has an A allele frequency of 50% and G allele frequency of 50% in African populations. 32 The A allele frequency is higher than in other populations (control group in this study: A allele 3%, G allele 97%; American populations: A allele 18%, G allele 84%; European populations: A allele 17%, G allele 83%; East Asian populations: A allele 5%, G allele 95%; South Asian populations: A allele 15%, G allele 85%). 32 The University of Benin in Africa reported a higher prevalence of orofacial pain in Benin City than in the rest of the world. 33 Although detailed information is lacking, it is highly likely that such orofacial pain also includes PTP. In the present study, A allele carriers had a higher rate of PTP, suggesting that the rs735055 SNP of the SLC17A9 gene is associated with PTP.
In rodents, nerve injury increases the expression of P2RY12 in microglia in the spinal cord and trigeminal spinal subnucleus caudalis.11,34 Nerve injury activates the guanosine triphosphate—Ras homolog gene family, member A (RhoA)—Rho-associated protein kinase 2 (ROCK2) signaling pathway and increases excitatory synaptic transmission in the dorsal horn, leading to mechanical and thermal hyperalgesia.10,35 P2RY12 microglial receptors in the trigeminal nucleus caudalis play an important role in the pathogenesis of chronic migraine by regulating microglial activation in the trigeminal nucleus caudalis via the RhoA/ROCK pathway. 36 In humans, PTP has been reported to be involved in migraine. 37 The rs3732759 SNP of the P2RY12 gene has been reported to be associated with clopidogrel sensitivity in patients with coronary artery disease. 38 The rs6809699 SNP and rs16863323 SNP in the LD block that also contains the rs3732759 SNP of the P2RY12 gene have been reported to be associated with intravenous immunoglobulin resistance in Kawasaki disease patients 39 and early neurological deterioration in acute ischemic stroke. 40 However, no previous studies have shown that the rs3732759 SNP of the P2RY12 gene and SNPs in the same LD block are associated with pain. Individuals who are homozygous for the major allele of the rs3732759 SNP may have higher P2RY12 expression, which may be more active in microglia and predispose these individuals to neuropathic pain, including PTP. However, the effect of the rs3732759 SNP on P2RY12 expression levels is only speculative because no data on such an effect have been reported. SNPs of the P2RY12 gene may affect the expression of genes in the vicinity of the P2RY12 gene. We cannot exclude the possibility that proteins that are encoded by these genes are also associated with the development of PTP. The rs3732759 SNP of the P2RY12 gene is located in the intron region (Figure 2) where the promoter flanking region exists. 32 When an allele of one SNP in the LD block changes, the alleles of other polymorphisms that are in strong LD with the SNP also change. This may have a relatively large impact on gene expression and other factors.
SLC17A9 and P2RY12 are reported to be involved in nucleotide release in platelets in humans 41 and may act together in the spinal cord. After lingual nerve injury, ATP is released from somatic cells of trigeminal ganglion neurons and activates P2RY12 in satellite glial cells, which plays a role in enhancing trigeminal ganglion neural activity and nociceptive reflex behavior, causing neuropathic pain in the tongue in rodents.7,42,43 SLC17A9-dependent ATP release in the trigeminal ganglion (but not in the spinal cord) and trigeminal ganglia-specific signaling via P2RY12 on satellite glial cells may also be involved in PTP (Supplementary Figure S2). This possibility is based on studies in rodents. Further studies are needed to reveal the detailed mechanisms of PTP signal transduction in humans.
In the present study, to better exclude differential diagnoses, PTP was diagnosed according to ICOP criteria and additional information about local anesthesia and allodynia (Table 1, Supplementary Figure S1). The presence or absence of the disappearance of pain with local anesthesia or presence of allodynia would have been useful for differentiating odontogenic pain or nociplastic pain. The OFP group comprised a heterogeneous group of patients with pain or sensory disturbance in the orofacial area and included various pathological conditions, excluding PTP. Because OFP was heterogeneous, it was further classified into three groups (neuropathic pain, trigeminal neuralgia, and nociplastic pain) and compared with the PTP group and healthy subjects, although the number of trigeminal neuralgia and nociplastic pain patients was too small for accurate statistical analysis (Supplementary Tables S5 and S6). Traumatic trigeminal neuropathy, postherpetic neuralgia, and NICO were analyzed together as the neuropathic pain subgroup for correctness because of the small number of patients in each group. The neuropathic pain subgroup was significantly different from the PTP group (SLC17A9 rs735055, p = 2.2 × 10−3, genotypic model and dominant model, total of males and females; P2RY12 rs3732759, p = .02, dominant model, total of males and females) and not significantly different from healthy subjects. Trigeminal neuralgia and nociplastic pain were not significantly different from PTP as well as healthy subjects, likely because of the insufficient number of cases that resulted in an insufficiently powered analysis. We analyzed trends of genotype ratios among each genotype. The genotype ratio for trigeminal neuralgia and nociplastic pain had similar trends as the genotype ratios for neuropathic pain and healthy subjects (in trigeminal neuralgia, nociplastic pain, neuropathic pain and healthy subjects: over 90% of carriers of the SLC17A9 rs735055 SNP were homozygous for the major allele; about 30% of carriers of the P2RY12 rs3732759 SNP were homozygous for the major allele). On the other hand, the genotype ratios for trigeminal neuralgia and nociplastic pain had different trends as the genotype ratio for PTP (in PTP: 70% of carriers of the SLC17A9 rs735055 SNP were homozygous for the major allele; 64% of carriers of the P2RY12 rs3732759 SNP were homozygous for the major allele). These results suggest that the genotype ratio for trigeminal neuralgia and nociplastic pain had similar trends as the genotype ratios for neuropathic pain and healthy subjects. We infer that significant results could be obtained if the number of cases for trigeminal neuralgia and nociplastic pain increased. These results suggest that the OFP group was an appropriate control group for the statistical analysis in the present study.
No significant results were found for OFP, despite the inclusion of neuropathic pain other than PTP. Pulpectomy and tooth extraction that can lead to PTP are frequent treatment procedures in clinical dentistry. Despite being procedures that damage nerves, these treatments usually leave no residual symptoms and are less susceptible to such environmental factors as the surgical technique. On the other hand, neuropathic pain, such as traumatic trigeminal neuropathy, was included in the OFP group in the present study. Neuropathic pain often occurs after the extraction of a buried wisdom tooth or after implant placement. This is often influenced by environmental factors, such as the level of nerve damage and pattern of damage. We may have observed significant results only for PTP because PTP is less influenced by environmental factors and thus more likely to reflect the influence of genetic factors. Additionally, the comparisons between the OFP group and healthy subjects showed no significant results, whereas the comparisons between the PTP group and healthy subjects and between the PTP group and OFP group showed significant results. These findings suggest that PTP is different from other pain and sensory disorders in the orofacial area, including neuropathic pain.
In the present study, the results were significant in females but not in males. Previous studies have shown no significant difference in the incidence of PTP between genders.2,5 Most mouse studies of neuropathic pain have used only male mice. Microglia were reported to be involved in hyperalgesia in chronic pain in males, 44 and high P2RY12 expression in microglia in male mice was shown to result in neuropathic pain. 35 These previous findings do not contradict the present results. The same results could be found for males alone as for males and females combined. One reason why we found no significant difference in males may be that the sample size was relatively small, in which only nine patients with PTP were included in the study. The total sample size was not sufficiently large, and further analysis with a larger sample size is needed.
Supplemental Material
sj-xlsx-11-mpx-10.1177_17448069221089592 – Supplemental Material for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain
Supplementary Material, sj-xlsx-11-mpx-10.1177_17448069221089592 for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain by Moe Soeda, Seii Ohka, Daisuke Nishizawa, Junko Hasegawa, Kyoko Nakayama, Yuko Ebata, Ken-ichi Fukuda, and Kazutaka Ikeda in Molecular Pain.
Supplemental Material
sj-xlsx-11-mpx-10.1177_17448069221089592 – Supplemental Material for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain
Supplementary Material, sj-xlsx-11-mpx-10.1177_17448069221089592 for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain by Moe Soeda, Seii Ohka, Daisuke Nishizawa, Junko Hasegawa, Kyoko Nakayama, Yuko Ebata, Ken-ichi Fukuda, and Kazutaka Ikeda in Molecular Pain.
Supplemental Material
sj-xlsx-11-mpx-10.1177_17448069221089592 – Supplemental Material for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain
Supplementary Material, sj-xlsx-11-mpx-10.1177_17448069221089592 for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain by Moe Soeda, Seii Ohka, Daisuke Nishizawa, Junko Hasegawa, Kyoko Nakayama, Yuko Ebata, Ken-ichi Fukuda, and Kazutaka Ikeda in Molecular Pain.
Supplemental Material
sj-xlsx-11-mpx-10.1177_17448069221089592 – Supplemental Material for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain
Supplementary Material, sj-xlsx-11-mpx-10.1177_17448069221089592 for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain by Moe Soeda, Seii Ohka, Daisuke Nishizawa, Junko Hasegawa, Kyoko Nakayama, Yuko Ebata, Ken-ichi Fukuda, and Kazutaka Ikeda in Molecular Pain.
Supplemental Material
sj-xlsx-11-mpx-10.1177_17448069221089592 – Supplemental Material for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain
Supplementary Material, sj-xlsx-11-mpx-10.1177_17448069221089592 for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain by Moe Soeda, Seii Ohka, Daisuke Nishizawa, Junko Hasegawa, Kyoko Nakayama, Yuko Ebata, Ken-ichi Fukuda, and Kazutaka Ikeda in Molecular Pain.
Supplemental Material
sj-xlsx-11-mpx-10.1177_17448069221089592 – Supplemental Material for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain
Supplementary Material, sj-xlsx-11-mpx-10.1177_17448069221089592 for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain by Moe Soeda, Seii Ohka, Daisuke Nishizawa, Junko Hasegawa, Kyoko Nakayama, Yuko Ebata, Ken-ichi Fukuda, and Kazutaka Ikeda in Molecular Pain.
Supplemental Material
sj-xlsx-11-mpx-10.1177_17448069221089592 – Supplemental Material for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain
Supplementary Material, sj-xlsx-11-mpx-10.1177_17448069221089592 for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain by Moe Soeda, Seii Ohka, Daisuke Nishizawa, Junko Hasegawa, Kyoko Nakayama, Yuko Ebata, Ken-ichi Fukuda, and Kazutaka Ikeda in Molecular Pain.
Supplemental Material
sj-xlsx-11-mpx-10.1177_17448069221089592 – Supplemental Material for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain
Supplementary Material, sj-xlsx-11-mpx-10.1177_17448069221089592 for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain by Moe Soeda, Seii Ohka, Daisuke Nishizawa, Junko Hasegawa, Kyoko Nakayama, Yuko Ebata, Ken-ichi Fukuda, and Kazutaka Ikeda in Molecular Pain.
Supplemental Material
sj-xlsx-11-mpx-10.1177_17448069221089592 – Supplemental Material for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain
Supplementary Material, sj-xlsx-11-mpx-10.1177_17448069221089592 for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain by Moe Soeda, Seii Ohka, Daisuke Nishizawa, Junko Hasegawa, Kyoko Nakayama, Yuko Ebata, Ken-ichi Fukuda, and Kazutaka Ikeda in Molecular Pain.
Supplemental Material
sj-xlsx-11-mpx-10.1177_17448069221089592 – Supplemental Material for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain
Supplementary Material, sj-xlsx-11-mpx-10.1177_17448069221089592 for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain by Moe Soeda, Seii Ohka, Daisuke Nishizawa, Junko Hasegawa, Kyoko Nakayama, Yuko Ebata, Ken-ichi Fukuda, and Kazutaka Ikeda in Molecular Pain.
Supplemental Material
sj-xlsx-11-mpx-10.1177_17448069221089592 – Supplemental Material for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain
Supplementary Material, sj-xlsx-11-mpx-10.1177_17448069221089592 for Single-nucleotide polymorphisms of the SLC17A9 and P2RY12 genes are significantly associated with phantom tooth pain by Moe Soeda, Seii Ohka, Daisuke Nishizawa, Junko Hasegawa, Kyoko Nakayama, Yuko Ebata, Ken-ichi Fukuda, and Kazutaka Ikeda in Molecular Pain.
Footnotes
Acknowledgments
We thank Mr Michael Arends for his assistance with editing the manuscript. We are grateful to the volunteers for their participation in the study and anesthesiologists and surgeons for collecting the clinical data.
Author contributions
MS, SO, DN, KF, and KI conceived the study and designed the experiments. MS, SO, KN, YE, and DN performed the statistical analyses. MS wrote the manuscript. MS and KF collected clinical samples and data. MS and JH performed the genotyping procedures. SO, DN, KF, and KI supervised the experiments and finalized the manuscript. All of the authors contributed to writing the manuscript, and all authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the Japan Society for the Promotion of Science (JSPS) KAKENHI (no. 21H03028, JP16H06276 [AdAMS], 20K07774, 17K09052, 20K09259, 17K08970, and 17H04324) and Japan Agency for Medical Research and Development (AMED; no. JP19ek0610011).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
