Abstract
Covariate adjustment is an approach to improve the precision of trial analyses by adjusting for baseline variables that are prognostic of the primary endpoint. Motivated by the SEARCH Universal HIV Test-and-Treat Trial (2013–2017), we tell our story of developing, evaluating, and implementing a machine learning-based approach for covariate adjustment. We provide the rationale for as well as the practical concerns with such an approach for estimating marginal effects. Using schematics, we illustrate our procedure: targeted machine learning estimation (TMLE) with Adaptive Pre-specification. Briefly, sample-splitting is used to data-adaptively select the combination of estimators of the outcome regression (i.e. the conditional expectation of the outcome given the trial arm and covariates) and known propensity score (i.e. the conditional probability of being randomized to the intervention given the covariates) that minimizes the cross-validated variance estimate and, thereby, maximizes empirical efficiency. We discuss our approach for evaluating finite sample performance with parametric and plasmode simulations, pre-specifying the Statistical Analysis Plan, and unblinding in real-time on video conference with our colleagues from around the world. We present the results from applying our approach in the primary, pre-specified analysis of eight recently published trials (2022–2024). We conclude with practical recommendations and an invitation to implement our approach in the primary analysis of your next trial.
Keywords
Imagine you are tasked with pre-specifying and implementing the primary analysis of a population-level randomized trial. Concretely, you are responsible for evaluating effectiveness in the SEARCH Universal HIV Test-and-Treat Trial, which was conducted from 2013 to 2017 in Kenya and Uganda (NCT01864603). 1 Based on standard sample size calculations for cluster-randomized controlled trials, 2 you anticipate that 32 communities (each with ∼10,000 persons) will provide sufficient power for the primary endpoint of HIV incidence, a rare binary outcome. Given the randomization and follow-up procedures, you know the unadjusted effect estimator will be unbiased. However, you have also heard promises about the potential gains in precision and power with covariate adjustment: an analytic approach to adjust for baseline variables that are prognostic of the outcome.3,4 You may have seen promising results from simulation studies for a wide range of causal estimands and outcomes.5–12 You may be particularly intrigued by targeted machine learning estimation (TMLE), which is a model-robust, locally efficient approach and illustrated in Figure 1 (eAppendices A-B in Supplementary Material).13,14
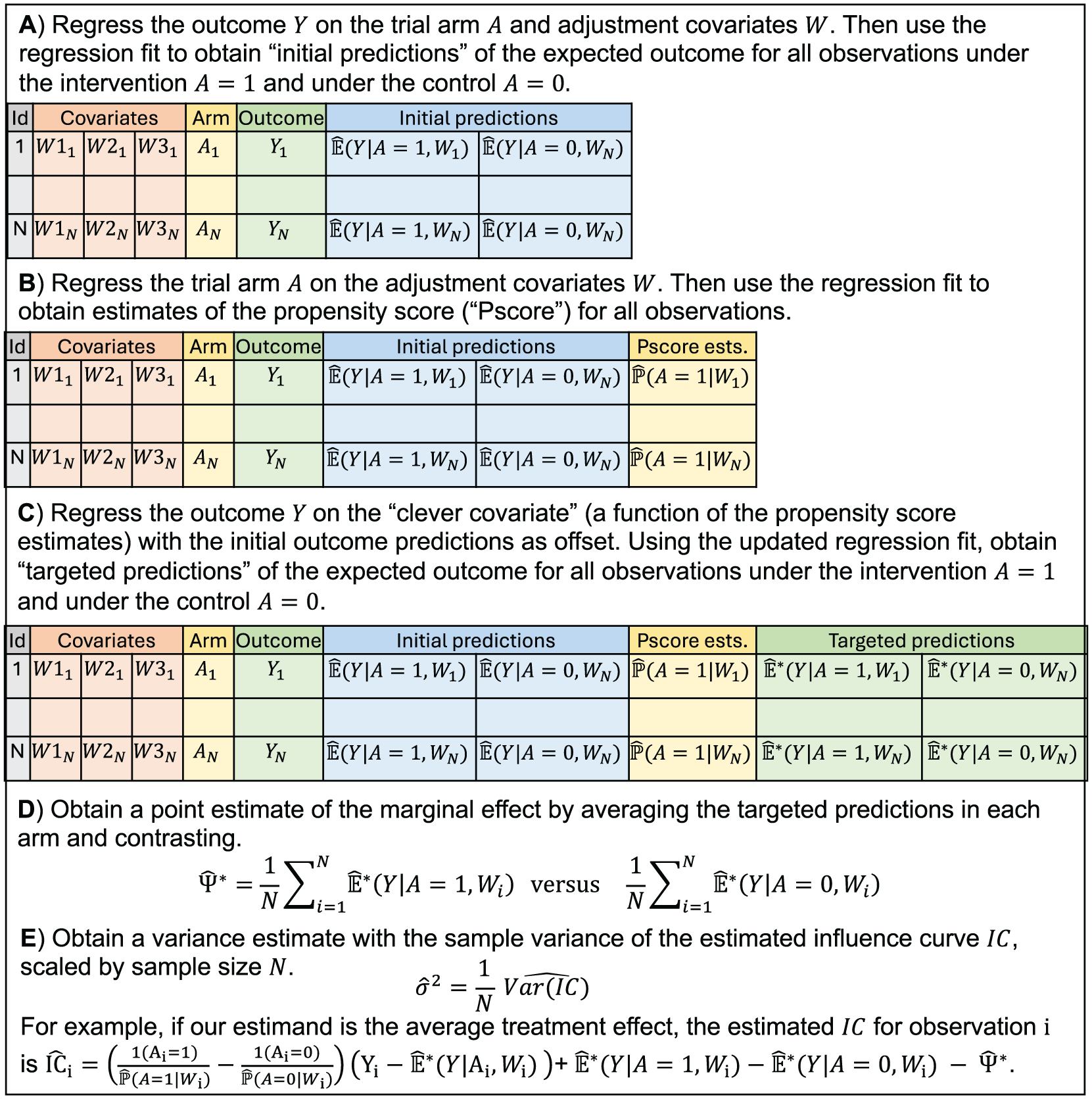
Schematic of steps to obtain a point and variance estimate with TMLE for the marginal effect defined as the contrast of the expected counterfactual outcomes.The targeting procedure and the form of the influence curve will depend on the estimand. See eAppendix A in Supplementary Material for further details.
However, you may also be troubled by practical concerns—at least we were … How do we a priori specify the optimal adjustment approach? What if we bet incorrectly and implement an adjusted analysis that actually harms precision? What if the adjusted analysis gives wildly different results from the unadjusted one? How will the procedure perform in small sample settings—we only have 32 independent units? How will the procedure perform with a rare binary outcome? Is there a risk of underestimating the variance and inflated Type-I error rates? Will stakeholders understand and trust the results? Overall, we needed a method that was data-adaptive, yet fully pre-specified and maintained excellent performance.
Due to treatment randomization, trials are highly robust for conventional intention-to-treat analyses with minimal outcome missingness. This setting allows us to focus our analytic efforts on minimizing variance. Therefore, building on the theory of Empirical Efficiency Maximization,15,16 we proposed a practical and automated procedure using machine learning to select the optimal adjustment approach in randomized trials. Given the procedure is both pre-specified and adaptive, we called it “Adaptive Pre-specification.” 7
For the implementation of this procedure with TMLE, we must pre-specify:
Candidate estimators of the “outcome regression,” which is the expected outcome given the trial arm and covariate(s).
Candidate estimators of the “propensity score,” which is the conditional probability of being randomized to the intervention given the covariates(s). Although this probability is known in trials, estimating it can improve precision. 17
Cross-validation procedure, including the process for splitting the data, evaluating the candidates, and obtaining final point and variance estimates.
Loss function to quantify the performance of the candidates. We use the estimated influence curve (function)-squared, which provides a variance estimate for each candidate. 19
Altogether, the “best” estimator is the candidate minimizing the cross-validated variance estimate and, thereby, maximizing empirical efficiency. The unadjusted estimator must be included as a candidate for the outcome regression and as a candidate for the propensity score so that it can be selected if none of the candidates using covariate adjustment improves precision.
As illustrated in Figure 2, we use sample-splitting to first select the outcome regression estimator that minimizes the cross-validated variance estimate. Then, using the selected outcome regression estimator, we select the propensity score estimator that further minimizes the cross-validated variance estimate. Together, the selected outcome regression estimator and propensity score estimator form the optimal TMLE, which is guaranteed to be at least as precise as the unadjusted effect estimator.
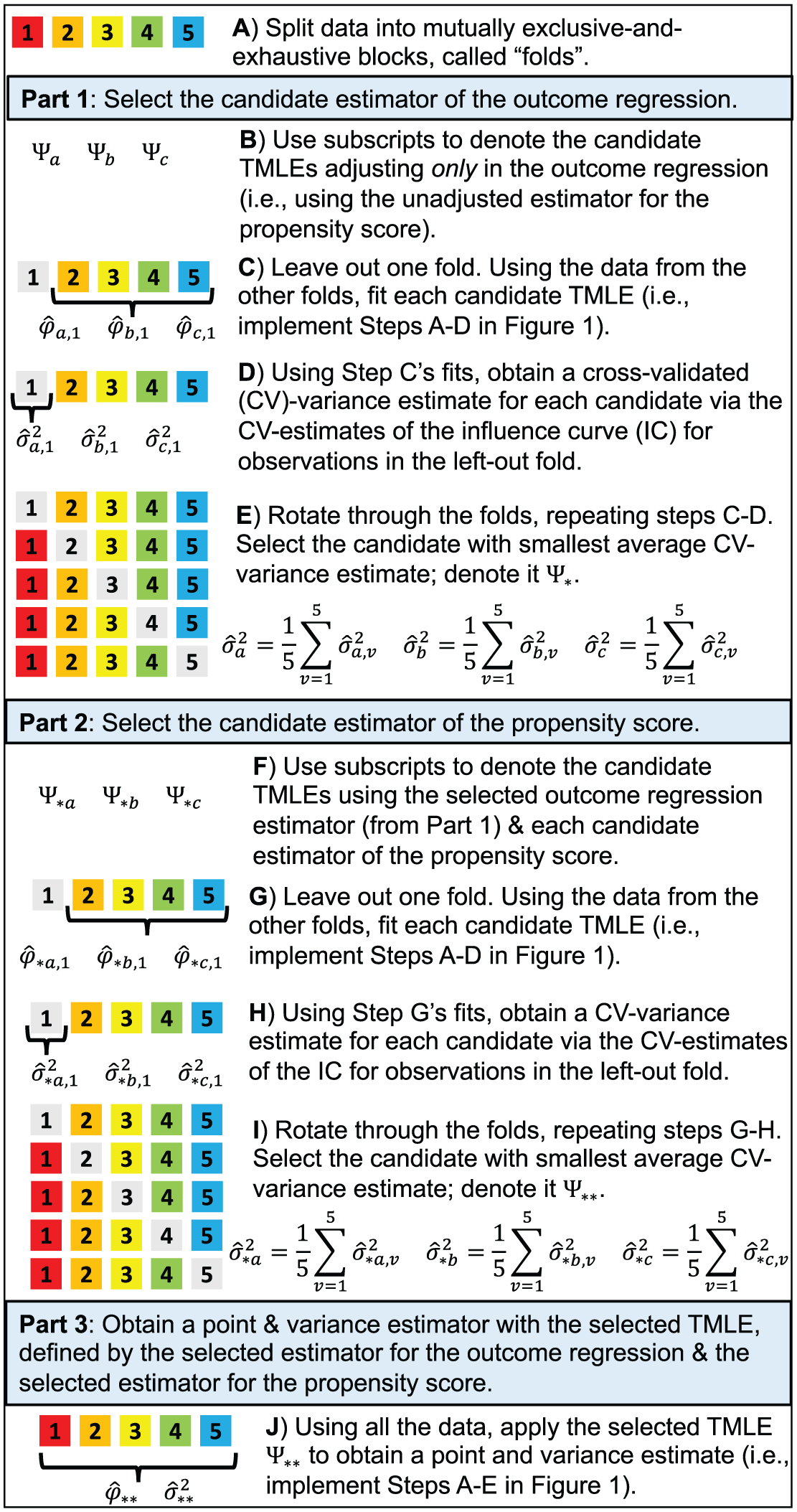
Schematic of TMLE with Adaptive Pre-specification—using an example with three candidates for the outcome regression, three candidates for the propensity score, and 5-fold cross-validation. In Step J, we can alternatively use a cross-validated variance estimate, obtained as the sample variance of the cross-validated influence curve (IC)-estimates scaled by sample size. The procedure should only be implemented after fully pre-specifying the estimand (implying the loss function), the candidate estimators of the outcome regression, the candidate estimators of the propensity score, and the cross-validation procedure (including the approach for variance estimation).
Then, the question was how does TMLE with Adaptive Pre-specification perform in practice and, most importantly, in settings mirroring our trial? To assess performance across a variety of scenarios, we conducted parametric simulations, which demonstrated that our procedure offered meaningful gains in power while maintaining nominal confidence coverage. 7 To assess expected performance for our trial, we conducted plasmode simulations, which sample partially from the empirical distribution.9,18–20 Specifically, we sampled baseline covariates from the SEARCH data, randomized the trial arms according to the study design, and simulated the outcome using mathematical models of the HIV epidemic. 21 These simulations demonstrated that our adaptive procedure improved power while preserving confidence interval coverage when there was an effect and protected Type-I error control under the null.
Finally, we were ready to pre-specify the Statistical Analysis Plan and analytic code to implement TMLE with Adaptive Pre-specification in the primary analysis of SEARCH.1,21,22 The Statistical Analysis Plan and code were locked prior to outcome measurement and while the primary trial statistician (LBB) remained blinded. As a final check, we conducted treatment-blind plasmode simulations to confirm Type-I error control. We loaded in the actual SEARCH data, permuted the arm labels between clusters (the randomized unit), implemented our primary analytic approach, and repeated many times—verifying the proportion of times that the true null hypothesis was rejected was ≤5%.
Then on April 6, 2018, we unblinded. Live on video conference with our colleagues from around the world, the primary statistician (LBB) shared her screen, loaded the actual data into RStudio, and executed the code. Together and in real-time, we learned not only the effectiveness of the intervention, but also which variables were selected for covariate adjustment (if any). The primary analyses, using this machine learning approach, were published in the New England Journal of Medicine. 1
The efficiency gains achieved in SEARCH have previously been published. 11 For HIV incidence (primary endpoint), the primary analysis—TMLE with Adaptive Pre-specification—was nearly 5-times more precise than the unadjusted effect estimator (the simple contrast in arm-specific average outcomes). Specifically, the estimated variance of TMLE was nearly one-fifth that of the unadjusted approach. For the secondary endpoints of tuberculosis incidence and hypertension control, TMLE was 2.6 and 1.8 times more precise, respectively. For HIV viral suppression, another secondary endpoint, we saw no difference in precision. This range of precision gains—from substantial to none, but with no instances of harm—is a key characteristic of our approach. Our procedure defaults to the unadjusted estimator when none of the pre-specified candidates improves empirical efficiency. We are protected from forced adjustment to the detriment of precision.
Given our motivation to maximize power in a trial with only 32 randomized units and a rare outcome, we originally limited the candidate estimators to “working” generalized linear models (GLMs) adjusting for at most one covariate. 6 Recently, we extended the procedure for larger trials and to include “well-behaved” machine learning algorithms as candidates: stepwise regression, penalized regression, and multivariate adaptive regression splines with and without screening. 12 These approaches respond flexibly to the data, but are less likely to overfit (i.e. they satisfy the “Donsker” conditions).13,23 Across a variety of settings, our simulations demonstrated meaningful precision improvements—translating into 20% to 43% reductions in sample size for the same statistical power. 12 (These sample size savings were calculated as 1 minus the mean squared error [MSE] of a covariate-adjusted estimator divided by the MSE of the unadjusted estimator.) 9 Importantly, precision gains were achieved for both binary and continuous endpoints and without sacrificing 95% confidence interval coverage. These simulations also highlighted the precision gains with data-adaptive adjustment versus forced adjustment for the variables used in stratified randomization. As a real-data demonstration, we re-analyzed ACTG Study 175 and found the selected TMLE varied by pre-specified subgroups—highlighting the importance of avoiding a “one-size-fits-all” approach. 12
Since its publication in 2016, we have applied TMLE with Adaptive Pre-specification in the primary analyses of over a dozen trials.1,24–35 In Table 1, we present the precision gains and optimal adjustment approach for eight recently published trials (NCT04810650; NCT05549726). As before, there was a range of efficiency gains, but crucially, we never lost precision. Notably, within and across trials, the selected TMLE varied—highlighting the flexibility of our pre-specified, yet data-adaptive approach. In all cases and as expected, the point estimates from the TMLE and the unadjusted estimator were very similar (eAppendix C in Supplementary Material).
Showcase of recently published trials where TMLE with Adaptive Pre-specification was used in the primary analysis (NCT04810650; NCT05549726).
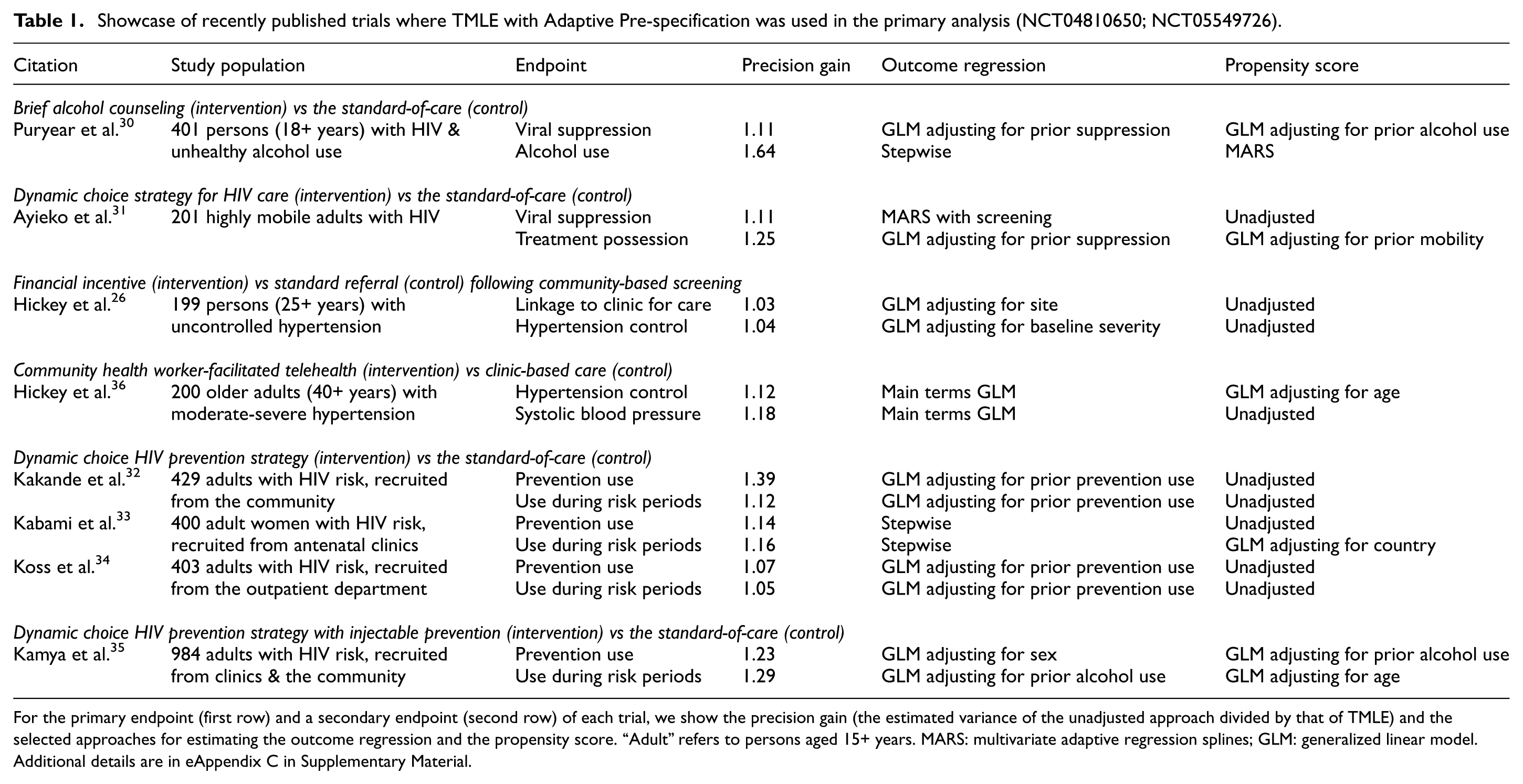
For the primary endpoint (first row) and a secondary endpoint (second row) of each trial, we show the precision gain (the estimated variance of the unadjusted approach divided by that of TMLE) and the selected approaches for estimating the outcome regression and the propensity score. “Adult” refers to persons aged 15+ years. MARS: multivariate adaptive regression splines; GLM: generalized linear model. Additional details are in eAppendix C in Supplementary Material.
Reflecting the motivating studies (Table 1), we have focused on trials with intent-to-treat estimands and ignorable missingness. Other trials may be subject to differential missingness or censoring, which can cause meaningful bias. For individually randomized trials, various TMLEs have been developed and applied to address these common complications.8–10,37,38 For cluster randomized trials, we developed and applied Two-Stage TMLE to first account for missingness and other potentially biasing factors at the individual-level and then to evaluate effectiveness with maximum precision by applying a cluster-level TMLE with Adaptive Pre-specification.1,11,24,25,36,39–41
While our real-data analyses demonstrate the value of our approach, skepticism remains. We are often asked about the rationale for using an adaptive approach when the team “knows” the most prognostic covariate. As illustrated by our prior analyses, it is difficult to bet a priori on the best adjustment covariate(s) and the form of the working regressions. Furthermore, the optimal approach may vary by subgroup or endpoint. Therefore, we recommend letting the data decide through our fully pre-specified, yet adaptive procedure. We offer similar advice to teams who have built a prognostic score based on historical data and plan to adjust for that score in their analysis (a.k.a. “PROCOVA”).42,43 While this score may effectively summarize the covariate set into a single measure, it may not be the optimal adjustment covariate. Therefore, we recommend including it as another candidate in our procedure.
We emphasize that our algorithm requires detailed pre-specification of the primary estimand (implying a specific loss function—the estimated influence curve-squared), the candidate approaches for estimating the outcome regression and propensity score, and the cross-validation procedure (eAppendix C in Supplementary Material). Conducting simulations, imitating the key features of the trial, can help inform these choices and confirm implementation of the computing code.20,21 While recognizing each trial offers unique opportunities and challenges, we offer the following recommendations. First, if you have concerns about data sparsity (e.g. due to rare outcomes or few independent units), limit the candidate adjustment algorithms to a handful of working GLMs adjusting for at most 1 candidate. Second, if you have concerns about perceived or actual overfitting, use the cross-validated variance estimate, which is a natural by-product of the algorithm (Figure 2). In our experience, the cross-validated variance estimate has been overly conservative, but this may not always be the case, especially when using more aggressive learners. Finally, we reiterate that the unadjusted estimator must always be included as a candidate so that it can be selected if none of the candidate approaches using covariate adjustment improves precision. (See eAppendix D in Supplementary Material for implications for power calculations.)
We also emphasize the importance of reproducibility and transparency. First, in line with best practices, the analytic code must be pre-specified. When using data-adaptive methods, such as our approach, we must include additional safeguards, including setting-the-seed and documenting the software version. Second, we have thoroughly appreciated the process of unblinding and running the primary analysis in real-time on video conference. However, we recognize this might not be possible for all studies. At minimum, we recommend using RMarkdown (or a similar markup language) to generate a user-friendly document with time-stamped results for sharing immediately after unblinding. Finally, we recommend using TMLE with Adaptive Pre-specification in the primary analysis and including the details of the selected approach and comparisons to the unadjusted analysis in Supplementary Materials.
In conclusion, TMLE with Adaptive Pre-specification is a fully pre-specified, model-robust, and automated procedure to data-adaptively select the adjustment approach that maximizes empirical efficiency in randomized trials. It is theoretically supported, guaranteed not to harm precision, and widely applicable. Prior use meaningfully improved precision in several high-profile trials, which were published in top-tier journals. Covariate adjustment is endorsed by both the U.S. Food and Drug Administration as well as the European Medicines Agency.3,44,45 Computing code to implement TMLE with Adaptive Pre-specification is publicly available. 46 More precise analyses translate to reduced uncertainty, improved power, and fewer Type-II errors. So perhaps you will consider applying our approach in your next analysis?
Supplemental Material
sj-docx-1-ctj-10.1177_17407745261417227 – Supplemental material for Machine learning to optimize precision in the analysis of randomized trials: A journey in pre-specified, yet data-adaptive learning
Supplemental material, sj-docx-1-ctj-10.1177_17407745261417227 for Machine learning to optimize precision in the analysis of randomized trials: A journey in pre-specified, yet data-adaptive learning by Laura B Balzer, Mark J van der Laan and Maya L Petersen in Clinical Trials
Footnotes
Acknowledgements
We gratefully thank the leaders of and our collaborators on the Sustainable East Africa Research in Community Health (SEARCH) Consortium (https://www.searchendaids.com/): Diane V. Havlir, Moses R. Kamya, James Ayieko, Jane Kabami, Elijah Kakande, Gabriel Chamie, Matthew Hickey, and many others who contributed to the trials motivating this work. The trials reported in ![]() were supported by the NIH (U01AI150510), and ViiV Healthcare provided cabotegravir long-acting in one trial. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH or ViiV. On behalf of the SEARCH team, we also thank the Ministry of Health of Uganda and of Kenya; our research teams and administrative teams in San Francisco, Berkeley, Uganda, and Kenya; our advisory boards, and especially all communities and participants involved.
were supported by the NIH (U01AI150510), and ViiV Healthcare provided cabotegravir long-acting in one trial. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH or ViiV. On behalf of the SEARCH team, we also thank the Ministry of Health of Uganda and of Kenya; our research teams and administrative teams in San Francisco, Berkeley, Uganda, and Kenya; our advisory boards, and especially all communities and participants involved.
Correction (March,2026):
Format errors in text corrected in pages [Page 2 and 4].
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported, in part, by the National Institutes of Health (NIH) under award number U01AI150510 and a philanthropic gift from the Novo Nordisk corporation to the University of California, Berkeley for the Joint Initiative for Causal Inference (JICI). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH or Novo Nordisk.
Trial registrations
NCT01864603, NCT04810650, NCT05549726 at ClinicalTrials.gov.
Supplemental material
Supplemental Material for this article is available online.
