Abstract
Background:
Partially clustered trials are trials that, by design, include a mixture of independent and clustered observations. For example, neonatal trials may include infants from a single, twin or triplet birth. The clustering of observations in partially clustered trials should be accounted for when determining the target sample size to avoid treatment arm comparisons being over or under powered. Limited tools are currently available for calculating the sample size for partially clustered trials, particularly when the maximum cluster size is greater than 2. The aim of this article is to introduce a new online application to calculate the target sample size for partially clustered trials covering a broad range of scenarios.
Methods:
The target sample size is calculated using design effects recently derived for two-arm partially clustered trials when the clusters exist prior to randomisation and the outcome of interest is continuous or binary. Both cluster and individual randomisation are considered for the clustered observations (resulting in nested and crossed designs, respectively). The sample size depends on quantities needed for typical sample size calculations, such as the effect size of interest, and the desired significance level and power. In addition, the sample size for partially clustered trials also depends on the range of cluster sizes, the proportion of observations that belong to clusters of each size, the intracluster correlation coefficient, the method of randomisation for the clustered observations, and the model that will be used for analysis. We developed an R Shiny web application that implements these methods in an easy-to-use sample size calculator that is freely available online.
Results:
The sample size calculator is free to access and provides trialists with the ability to determine the target sample size for different types of partially clustered trials. Step-by-step instructions are provided to illustrate the use of the calculator for designing two hypothetical trials. The target sample size that accounts for partial clustering can be quite different to the sample size that is calculated by methods for an independent design that ignore the clustering.
Conclusion:
Partial clustering affects the power and sample size requirements of clinical trials. The calculator presented in this article allows trialists to account for the clustering that occurs in two-arm partially clustered trials for binary and continuous outcomes and ensure their trials are appropriately powered.
Keywords
Introduction
Partially clustered trials are trials that, by design, include a mixture of independent and clustered observations. 1 Often, the clusters exist prior to randomisation. Examples include: neonatal trials, where the cluster is the mother and participants may include infants from singleton and multiple births; orthopaedic trials, where patients are clusters and may have one or more diseased joints eligible for treatment; and trials that use a re-randomisation design, where participants are clusters and may enrol in the trial one or more times. 2 The outcomes of observations from the same cluster are typically more similar than outcomes from independent observations or different clusters, and this can have an impact on the power of the trial. 3 If sample size calculations do not account for the partial clustering, then trials may be over-powered and hence lead to excessive use of resources, or under-powered and hence risk an important intervention effect being missed. 4
Freely available and easy-to-use tools for calculating the target sample size are important for ensuring trials are appropriately powered. Currently, there are limited tools available for partially clustered trials with clusters that exist prior to randomisation (for a discussion of calculating sample size when clusters are created after randomisation, such as in group-based treatment trials, see Roberts and Roberts 5 and the IRGT Calculator 6 ). An online sample size calculator has previously been developed for partially clustered trials with a maximum cluster size of 2; 7 however, larger clusters also occur. Examples include trials in neonatology (e.g. triplets) and dermatology (e.g. skin lesions). We recently developed sample size methods for two-arm, partially clustered trials with pre-randomisation clustering for use in trials where the planned analysis will account for the clustering via generalised estimating equations (GEEs). 8 The methods accommodate continuous and binary outcomes, varying cluster sizes, and different randomisation methods for the clustered observations. In this article we introduce a free, online, R Shiny application that uses these methods to calculate the target sample size for partially clustered trials.
Methods
The web app is available at https://klange.shinyapps.io/partial-clustering-ssizecalc/. R code for the Shiny app can be accessed at https://github.com/klange01/partial-clustering-ssizecalc.
The app uses design effects (DEFFs) to calculate the sample size. The sample size assuming independence is multiplied by the DEFF to produce the target sample size that accounts for the clustering. A DEFF is therefore a multiplication factor that may inflate (if DEFF > 1; typically occurs when all observational units within a cluster receive the same treatment) or decrease (if DEFF < 1; typically occurs when clusters may consist of observational units randomised to different treatment arms) the sample size, depending on the strength of the clustering and features of the trial design. Formulas for DEFFs for partially clustered trials have been published elsewhere, including details of the underlying models and assumptions. 8 To select and calculate the appropriate DEFF, the following information is entered into the app, starting with the ‘User Inputs’ panel (Figure 1):
1. The type of outcome variable, either continuous (e.g. birthweight in grammes) or binary (e.g. mortality).
2. The number of observational units required per trial arm, assuming independence. The sample size assuming independence can be calculated using standard tools for a two-sample t-test or chi-square test for a continuous or binary outcome, respectively, based on the desired power and significance level. 9 For example, the pwr.t.test function from the pwr package in R, or the power twomeans command in Stata, can be used to determine the required sample size for a two-sample t-test. Likewise, pwr.2p.test in R or power twoproportions in Stata can be used for a chi-square test. The sample size calculated by the app is the target sample size needed to achieve the same power and significance level that are used in this calculation but allowing for clustering.
3. The expected prevalence of the outcome in the intervention and control arms (binary outcomes only). Prevalence estimates are likely to have been used in calculating the sample size assuming independence that is entered in step 2. Those same prevalence estimates should be entered here.
4. The intracluster correlation coefficient (ICC). This is the correlation between outcomes of clustered observations. The ICC ranges from −1 to 1, with values closer to −1 or 1 indicating that outcomes from clustered observational units are more strongly negatively or positively correlated, respectively. In partially clustered trials, ICCs will typically be greater than 0. Ideally, estimates of the ICC for the outcome of interest will come from previously published trials or repositories of ICCs that have been collated in specific fields7,10–12 but sometimes an educated guess may be required.
5. The working correlation structure to be used in the planned GEE analysis model. The independence working correlation structure accounts for clustering by applying a robust variance estimate for the treatment effect. The exchangeable working correlation structure estimates a constant correlation between the clustered observations and uses the estimated correlation when accounting for the clustering. More detail on these approaches and their different interpretations can be found elsewhere,13–15 including in the context of partially clustered trials.16,17 If the analysis is planned using mixed effects models, the exchangeable working correlation structure may provide a reasonable estimate of the required sample size. 18 Using the default independence working correlation structure will provide a conservative sample size estimate if an alternate method of analysis is used.
6. The method of randomisation that will be used for the clustered observations. When cluster randomisation is used, all observational units within a cluster are randomised to the same treatment and so clusters are nested within treatment arm. This will be the case when the treatment can only be administered at the cluster level (e.g. treatments given during pregnancy when outcomes will be measured on infants after birth), or it may be chosen for logistical reasons (such as to reduce the risk of contamination of treatment effects between cluster members). When individual randomisation is used, observational units are randomised independently, such that members from the same cluster may be randomised to the same or different treatments (and hence clusters and treatment arms may be ‘crossed’).
7. The analysis model to be used (binary outcomes only; a linear model is assumed for continuous outcomes). The appropriate analysis model depends on the effect measure of interest. A logistic model results in the treatment effect being estimated as an odds ratio (or risk difference when combined with marginal standardisation), 19 and a log binomial model estimates the treatment effect as a relative risk.
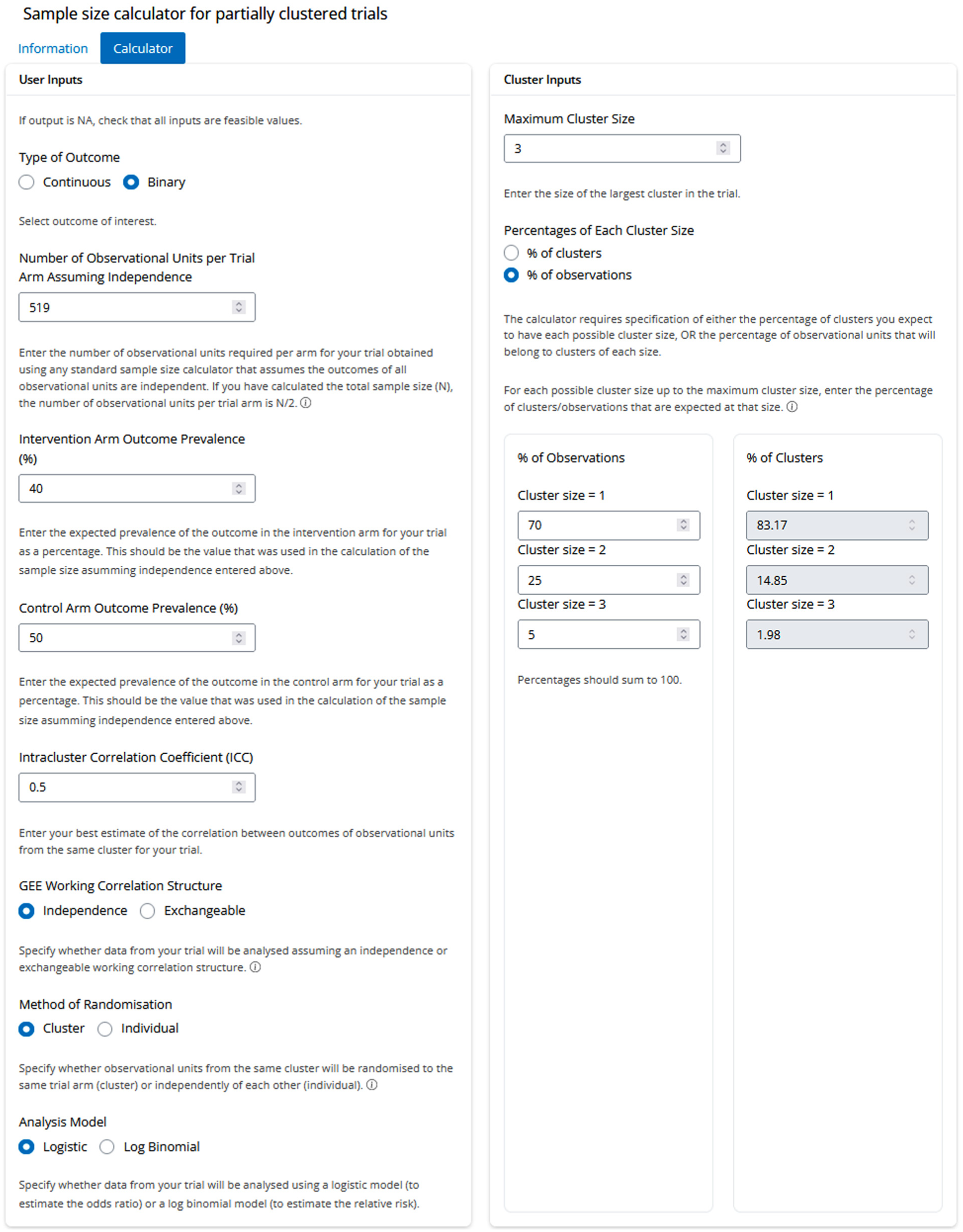
User inputs for calculating the target sample size for the hypothetical neonatal trial.
Next, details of the clustering that is expected in the trial are required. This information is entered into the ‘Cluster Inputs’ panel of the app (Figure 1).
8. The maximum cluster size. This is the size of the largest cluster that is expected in the trial. For example, this will be the largest number of eligible infants per pregnancy in a neonatal trial, or the largest number of skin lesions per participant expected in a dermatology trial.
9. Whether the distribution of cluster sizes will be specified as percentages of clusters or percentages of observations. For example, in a neonatal trial you may have estimates of the percentage of women (clusters) with a twin pregnancy or the percentage of infants (observations) from a twin birth.
10. The percentages (of clusters or observations, depending on the selection at step 9), of each cluster size from 1 (the independent observations) up to the maximum cluster size. This information may be available from existing sources or educated guesses may be required. An error message will be displayed if the entered percentages to do not sum to 100. Equations have been presented elsewhere to convert between the proportions of observations and clusters, 8 and these are used to display the alternative set of percentages within the app.
Once inputs 1–10 are specified, the app selects the appropriate formula for the DEFF and uses it to calculate the target sample size that accounts for the partial clustering, expressed as both the number of observations and the number of clusters required.
Results
To illustrate use of the app, we consider two hypothetical trials. The first is a neonatal trial involving infants from singleton and multiple births. Suppose the aim is to assess the effect of a new infant enteral formula on the development of bronchopulmonary dysplasia among infants born preterm. Infants born before 29 weeks’ gestation will be eligible, with mothers approached for consent to randomise their infants to either a standard formula or the new formula. The sample size calculations will assume a prevalence of bronchopulmonary dysplasia of 50% with the standard infant formula and the minimum clinically important effect is a reduction in prevalence to 40% with the new formula (odds ratio = 0.67). The sample size for a trial in singletons only can be calculated based on a test for two proportions with a two-sided alpha of 0.05 and 90% power. In R, this can be calculated with the command pwr.2p.test(h = ES.h(p1 = 0.5, p2 = 0.4), sig.level = 0.05, power = 0.90), or in Stata, power twoproportions 0.5 0.4, power(0.9) alpha(0.05), and results in a target sample size of 519 infants/mothers per arm (1038 overall). In the planned trial, it is assumed that the ICC for the outcome will be 0.5. The GEE working correlation structure is set to independence, to align with the planned analysis for the primary outcome of an independence GEE with clustering accounted for by robust variance estimation. As the researchers wish to estimate the treatment effect as an odds ratio, the logistic analysis model is selected. Parents of multiples generally prefer that their infants receive the same treatment, 20 and therefore cluster randomisation will be used for the multiples. These settings are entered into the ‘User Inputs’ panel of the app, as shown in Figure 1.
The expected distribution of cluster sizes is entered into the ‘Cluster Inputs’ panel (Figure 1). It is anticipated that in this early preterm population, 70% of infants will be singletons, 25% twins, and 5% triplets. Therefore, the maximum cluster size is entered as 3, ‘% of observations’ is selected, and the estimated percentages are entered into the three displayed input fields. The corresponding percentages of clusters is also displayed (i.e. the percentage of mothers that are estimated to have given birth to singletons, twins and triplets).
The results of the sample size calculation are displayed in the ‘Results’ panel (Figure 2). The use of cluster randomisation results in a DEFF > 1 and therefore the target sample size is increased relative to a trial in singletons only. The value of the appropriate DEFF is 1.175, which when multiplied by the independence sample size, results in a target sample size of 610 infants (observations) or 514 mothers (clusters) per arm for the trial to have the desired power of 90% at a two-sided 5% significance level.
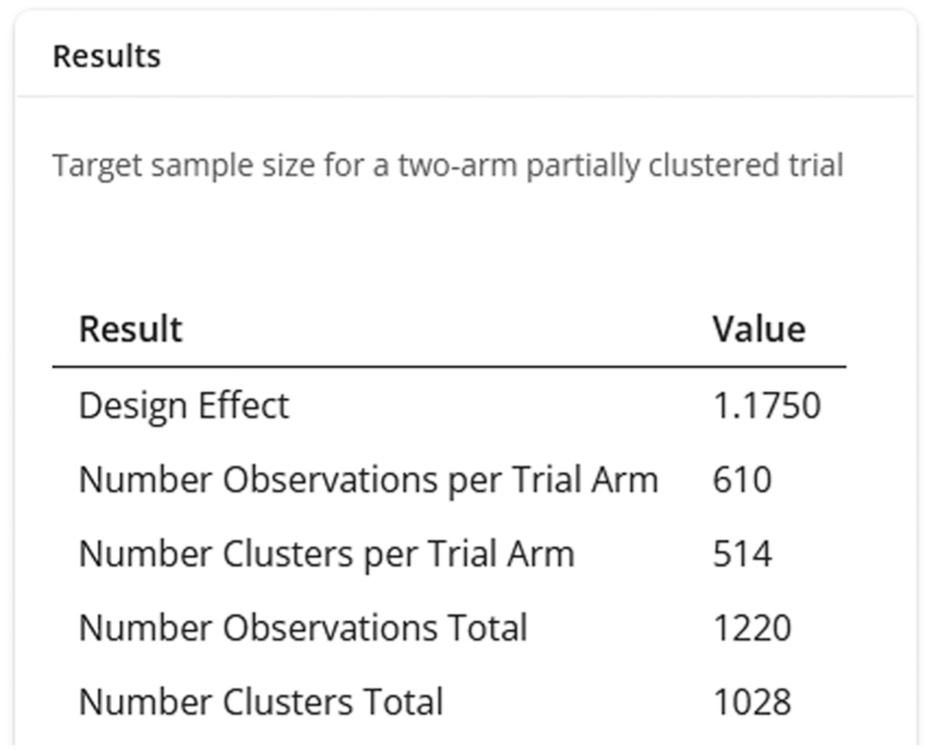
Results of a sample size calculation for the hypothetical neonatal trial.
As a second example, consider an ophthalmology trial to compare two surgical procedures, trabeculectomy and a surgical implant, in patients with uncontrolled glaucoma. The primary outcome is the intraocular pressure and the minimum clinically important effect is a difference of 2.5 mmHg, with an assumed standard deviation of 6 mmHg. The sample size for a trial involving only independent observations can be calculated using the method for a two-sample t-test, with a two-sided significance level of 0.05 and 80% power (in R pwr.t.test(d = 2.5/6, sig.level = 0.05, power = 0.80, type = ‘two.sample’), and in Stata power twomeans 0 2.5, sd(6) power(0.8) alpha(0.05)), and results in a required sample size of 92 patients/eyes per arm (184 total). The planned trial will involve partial clustering as patients with one or two eligible eyes will be enrolled, and outcomes from eyes of the same patient are expected to be correlated. It is estimated that 40% of patients (clusters) will contribute both eyes to the trial and the ICC is assumed to be 0.55. The planned analysis will be via GEE with an exchangeable working correlation structure, and each eye will be randomised independently (individual randomisation). The target sample size for the trial can be calculated using the app as shown in Figure 3. The resulting DEFF is 0.80 (DEFF ≤ 1 due to the use of individual randomisation), which reduces the target total sample size from 184 to 106 patients (clusters) who are expected to contribute 148 eyes, while still achieving 80% power at a two-sided 5% significance level.
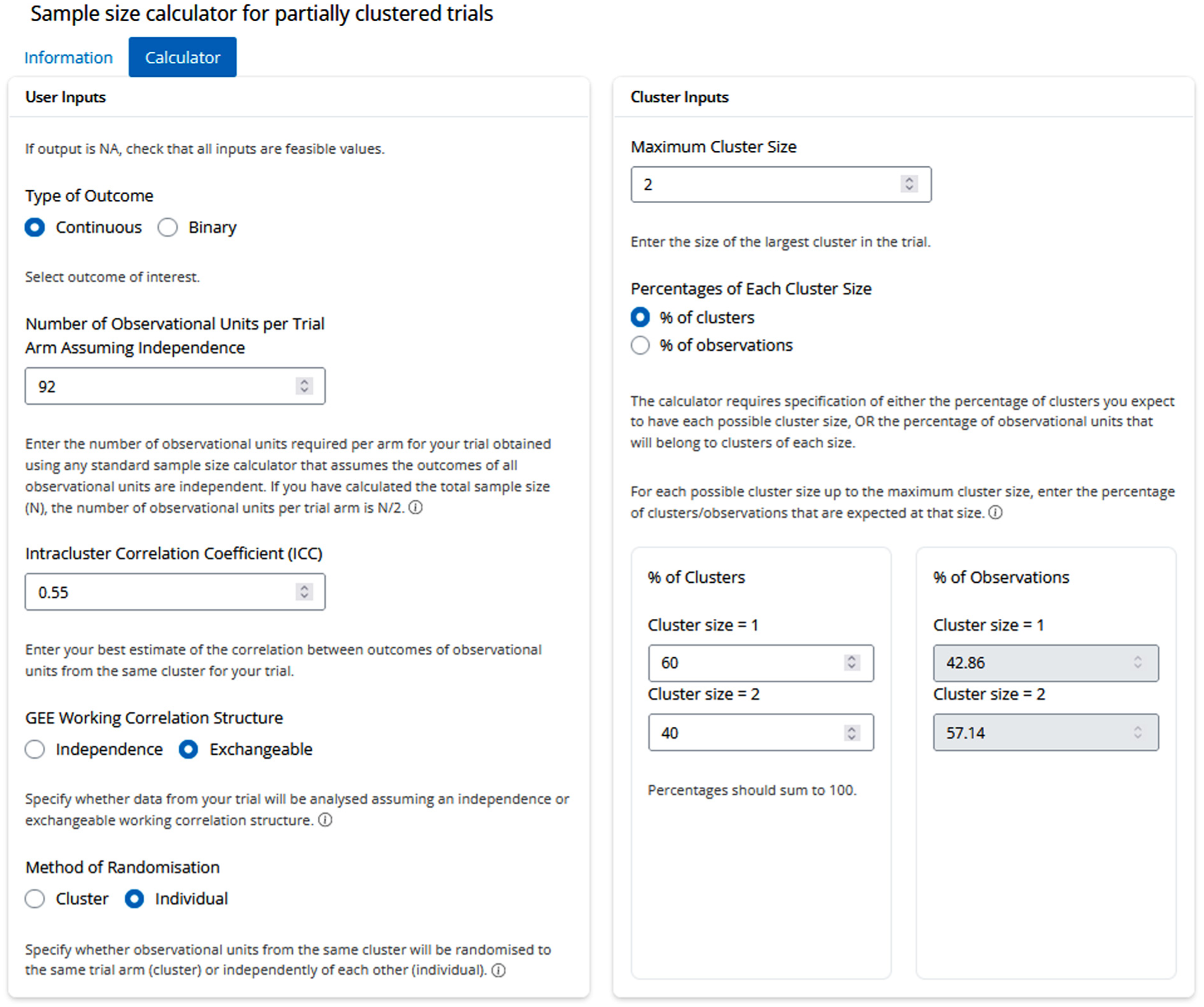
App inputs for the hypothetical ophthalmology trial.
Discussion
Strengths and limitations
The key strength of our work is that we provide a free, user-friendly application for calculating the sample size for partially clustered trials with any distribution of cluster sizes. Users can use the app to assess sensitivity of sample sizes to a range of values for the ICC and percentage of clusters or observations of each cluster size. While designed for trialists, collaboration with a statistician is recommended to ensure that trials are robustly designed. In particular, users are required to perform an initial sample size calculation outside of the app. While this provides the flexibility for researchers to use any appropriate sample size method for independent data, it may require input from a statistician. The choice of working correlation structure for the GEE analysis model may also require advice from a statistician, though the default independence working correlation can be used as a conservative option.
Conclusion
Accounting for partial clustering is important and can have a considerable impact on the sample size. We have presented a web app for calculating the sample size that is required to detect a given effect size for partially clustered trials with pre-existing clusters. The app caters to a variety of scenarios and we have demonstrated its use through two hypothetical examples. By using this app, trialists can easily account for the clustering that occurs in partially clustered trials and ensure their trials are appropriately powered.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship and/or publication of this article: K.M.L. is supported through an Australian Government Training Programme (RTP) Scholarship administered by The University of Adelaide, Australia, and an Australian Trials Methodology (AusTriM) Research Network supplementary scholarship. T.R.S. is supported by a Hospital Research Foundation Group Fellowship 104-83100. J.K. is supported by an NHMRC Leadership Fellowship 2033380. This research was supported by a Centre of Research Excellence grant from the National Health and Medical Research Council (NHMRC), ID 1171422, to the Australian Trials Methodology (AusTriM) Research Network.
