Abstract
Background
This study aimed to investigate the expression pattern and prognostic significance of HOXB13 in rectal cancer.
Methods
HOXB13 expression in rectal cancer and normal adjacent tissues was detected by immunohistochemistry, and its clinicopathological characteristics and prognosis were statistically tested. Furthermore, we evaluated the association between tumor immune infiltrating cells and HOXB13 using the tumor immune estimation resource (TIMER) database. The potential biological mechanism associated with HOXB13 overexpression was investigated by gene set enrichment analysis (GSEA).
Results
The expression of HOXB13 protein in human rectal cancer tissues were significantly higher than those in the normal adjacent tissues (P < 0.05). HOXB13 expression was significantly correlated with depth of invasion, lymphatic invasion, lymph node metastasis, distant metastasis, and pathological tumor node metastasis stage (P < 0.05). Kaplan–Meier survival curves confirmed that HOXB13 overexpression was correlated negatively with overall survival and disease-free survival in rectal cancer (P < 0.05). Also, multivariate Cox regression analysis demonstrated that HOXB13 expression, age, and lymphatic invasion were independent prognostic factors in rectal cancer (P < 0.05). Plus, the results from the TIMER database indicated that HOXB13 expression has a significant association with several immune cell infiltrates. Finally, the GSEA results indicated that HOXB13 participated in the various immune-associated processes, including natural killer cell-mediated cytotoxicity and the T-cell receptor signaling pathway.
Conclusion
Our study showed an essential role of HOXB13 in rectal cancer immunity and prognosis. Significantly, the overexpression of HOXB13 leads to the worse prognosis for patients with rectal cancer, which will contribute to understanding molecular mechanisms associated with tumor pathogenesis and prognosis in this disease.
Introduction
Colorectal cancer is one of the most typical cancers globally with around 900,000 cancer-related deaths annually, approximately one-fourth of which is rectal cancer (RC). 1 Nowadays, the tumor node metastasis (TNM) stage is the most important method for evaluating the prognosis of RC. However, RC patients with the same TNM stage tend to have a specific outcome, which indicates a single TNM stage is not comprehensive enough to determine the outcome and therapeutic method of some specific RC patients.2,3 Fortunately, in recent decades, biomarkers have been increasingly recognized as the predictive factor in various malignant tumors. Thus, finding new biomarkers to improve the treatment of patients with RC is now urgent.
HOX genes regulate axial regional specification during embryonic development and are often expressed in a tissue-specific and stage-related manner. Previously, many studies have suggested that HOXB13 overexpression in malignant tissues promotes tumor development such as gastric cancer, 4 ovarian cancer, 5 breast cancer, 6 prostate cancer, 7 and hepatocellular carcinoma. 8 Significantly, recent findings have demonstrated the prognostic value of HOXB13 in gastric cancer 9 and prostate cancer. 10 In contrast, some studies believe that HOXB13 exerts a tumor-suppressive effect on renal cancer 11 and melanoma. 12 Interestingly, a study based on tumor gene expression data showed that the HOX genes were more expressed in proximal colon cancer, and their expression decreased along more distal locations in the gastrointestinal tract—yet the expression of HOXB13 showed a reversed pattern. 13
In conclusion, the expression profile and prognostic value of HOXB13 have been confirmed in various malignant tumors. Investigating the overexpression of HOXB13 in RC provides an opportunity to enhance our understanding of RC development and to develop novel therapeutic methods. Herein, we first discovered that the overexpression of HOXB13 leads to a worse prognosis in RC and may be associated with immune cell infiltrates.
Materials and methods
Analysis of clinical data
Formalin-fixed, paraffin-embedded tissues from 103 cases of RC were obtained. The clinicopathological parameters such as age, gender, histological grade, the presence of lymphatic invasion, invasion depth, lymph node, or distant metastasis, and pathological TNM (pTNM) stage were evaluated by reviewing the medical charts and pathological records. The median patient age was 67.0 years, and 82.5% of the patients had undergone curative resection. The clinical outcome was determined from the day of surgery to death or December 31, 2009, which resulted in a follow-up period from 1.9 to 63.5 months (median 53.5 months). The cases that were lost to follow-up and deaths caused by other problems (not RC), were regarded as censored data in the survival analysis. This study was approved by the Institutional Review Boards of The Affiliated Xinhua Hospital Dalian University (Dalian, China). All patients with RC were diagnosed by pathological examination.
Immunohistochemistry
Fresh tissues were fixed in neutral-buffered formalin for 24 h, followed by infiltration with melted Istowax. Then, histologic evaluations were performed to individuate and exclude necrotic and hemorrhagic areas. From paraffin blocks, the most expansive area that could represent the condition of the corresponding rectal tumors was sectioned by 3 μm thickness, and the sections were deparaffinized in xylene and rehydrated with a graded series of ethanol. To augment the expression of antigen in tissues, Trilogy (Cell Marque; CA, USA) solution was added to the samples, which were then boiled in a programmable pressure cooker. To suppress the endogenous peroxidase activity, the samples were treated with peroxidase-blocking solution (DAKO; Mukilteo, Denmark) for 20 min and rinsed with a wash buffer (DAKO; Glostrup, Denmark). These samples were then reacted with 3% normal goat serum for 10 min and rinsed with wash buffer to prevent non-specific immune reactions. Immunohistochemistry (IHC) was used as the antibody to HOXB13 (Santa Cruz; CA, USA). The slides were shaken lightly and then reacted with primary antibody diluted 1:5 at 4°C overnight. After rinsing with the wash buffer, the Polink-1 polymer HRP second antibody (Santa Cruz; CA, USA) was added, reacted for 30 min at room temperature, and rinsed with the wash buffer solution. Subsequently, 1 × DAB + chromogen (DAKO; Glostrup, Denmark) was added, and the mixture was reacted for 7 min at room temperature, and after 5 min, washed with H2O. It was counterstained with hematoxylin and reacted for 1 min at room temperature; subsequently, it was sealed with crystal mount (Sakura; Tokyo, Japan). The negative control was prepared by the same procedure using phosphate-buffered saline (PBS) instead of the primary antibody, and RC tissues were used as the positive control.
Evaluation of IHC
The HOXB13 staining was separately and simultaneously evaluated by two independent observers, and a consensus score was reached for each core. The positive reaction of HOXB13 was scored into four grades according to the intensity of the staining: 0, 1 + , 2 + , and 3 + (Figure 1). The highest intensity of all tissue cores of each tumor was used as its final immunohistochemical result. Scores of 2 + and 3 + were classified as moderate and strong expression, and the score of 1 + was used to define tumors with weak HOXB13 expression.
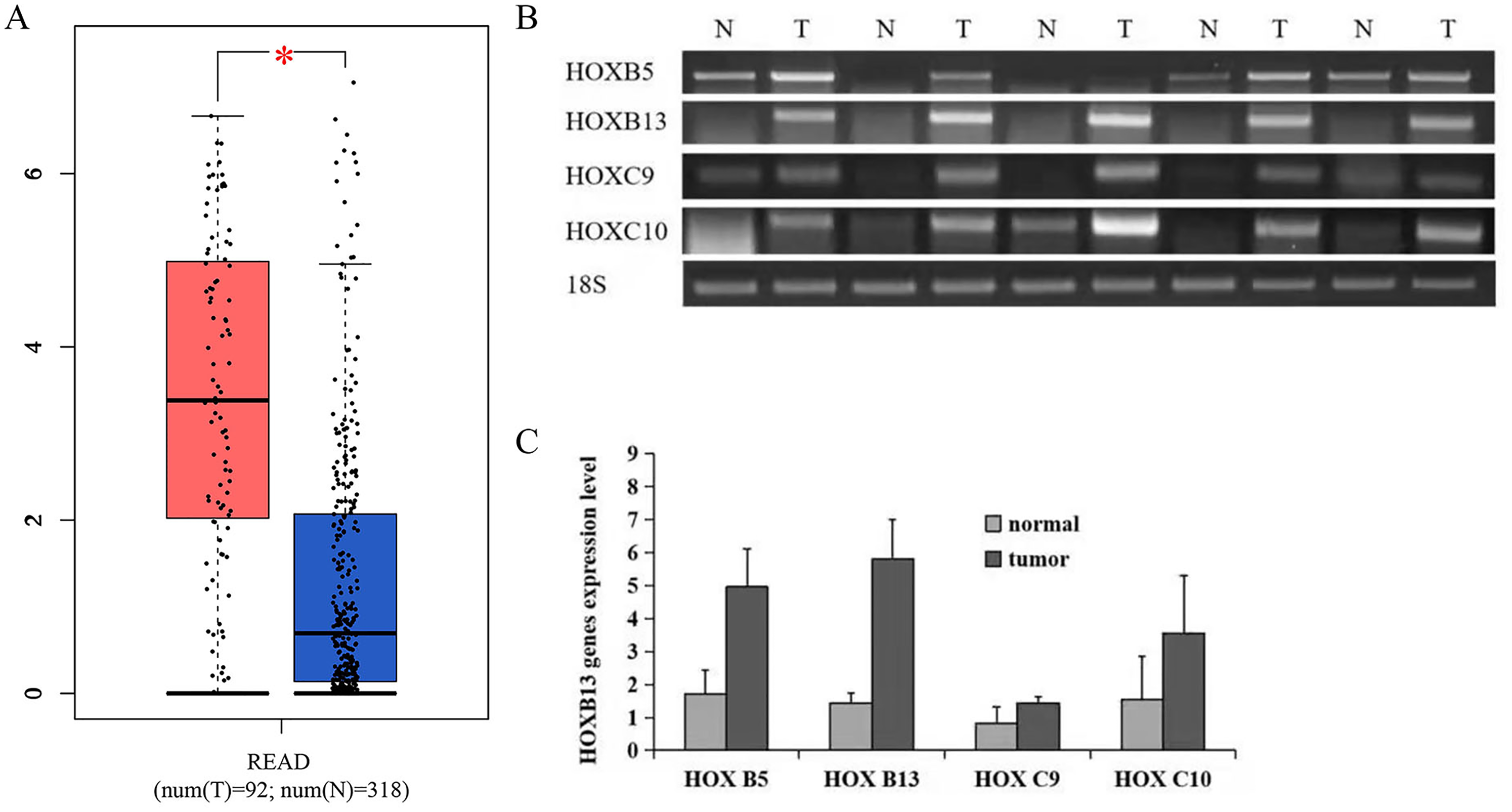
HOXB13 expression in rectal cancer and paired normal tissues using the GEPIA database.
STRING database analysis
The STRING online database (https//string-db.org/) was used to explore the relationship between screen genes correlated with HOXB13.15,16 The confidence score >0.7 was considered significant.
Associations between tumor immune infiltrating cells and HOXB13 using the tumor immune estimation resource and the GEPIA database
Interestingly, genes associated with HOXB13 have been demonstrated to be associated with immune infiltrates. However, the role of HOXB13 in the immune process has not been studied in detail. Thus, we further detected the relationship between HOXB13 and tumor immune infiltrating cells (TIICs), including B-cells, CD4 + T-cells, CD8 + T-cells, dendritic cells, macrophages, and neutrophils. Additionally, the co-expression between HOXB13 and various biomarkers of immune cells was explored using GEPIA and the tumor immune estimation resource (TIMER) database.14,17
Gene set enrichment analysis
To evaluate the potential mechanism underlying the involvement of HOXB13 in the carcinogenesis of RC, we performed gene set enrichment analysis (GSEA) (version 4.0.1; http://software. broadinstitute.org/gsea/index.jsp) to identify the pathways related to the differential HOXB13 expression in rectum cancer tissues according to The Cancer Genome Atlas (TCGA).18,19
Statistical analysis
Differences between and among groups were compared using Pearson's chi-square test for qualitative variables. Survival curves were estimated using the Kaplan–Meier product-limit method, and the significance of differences between survival curves was determined using the log-rank test. Univariate and multivariate analyses were performed using the Cox proportional hazard regression modeling. All statistical tests were two-sided, and statistical significance was defined as P < 0.05.
Results
Expression levels of HOXB13 protein mRNA and protein in RC and noncancerous tissues
GEPIA results showed that the expression of HOXB13 mRNA was higher than in their matched non-malignant tissues (Figure 1). HOXB13 expression in RC and non-tumor tissues were also examined by IHC. RC tissues showed significantly higher levels of the HOXB13 protein than their paired normal mucosa tissues. This result indicated that an increase of HOXB13 protein might facilitate tumorigenesis in RC. HOXB13 was predominantly presented in the cell nucleus in RC cells (Figure 2(a)). Furthermore, the positive rate of HOXB13 expression levels increased as the stage became advanced, with stage IV CRCs being the highest and stage I CRCs the lowest (Figure 2(b)).
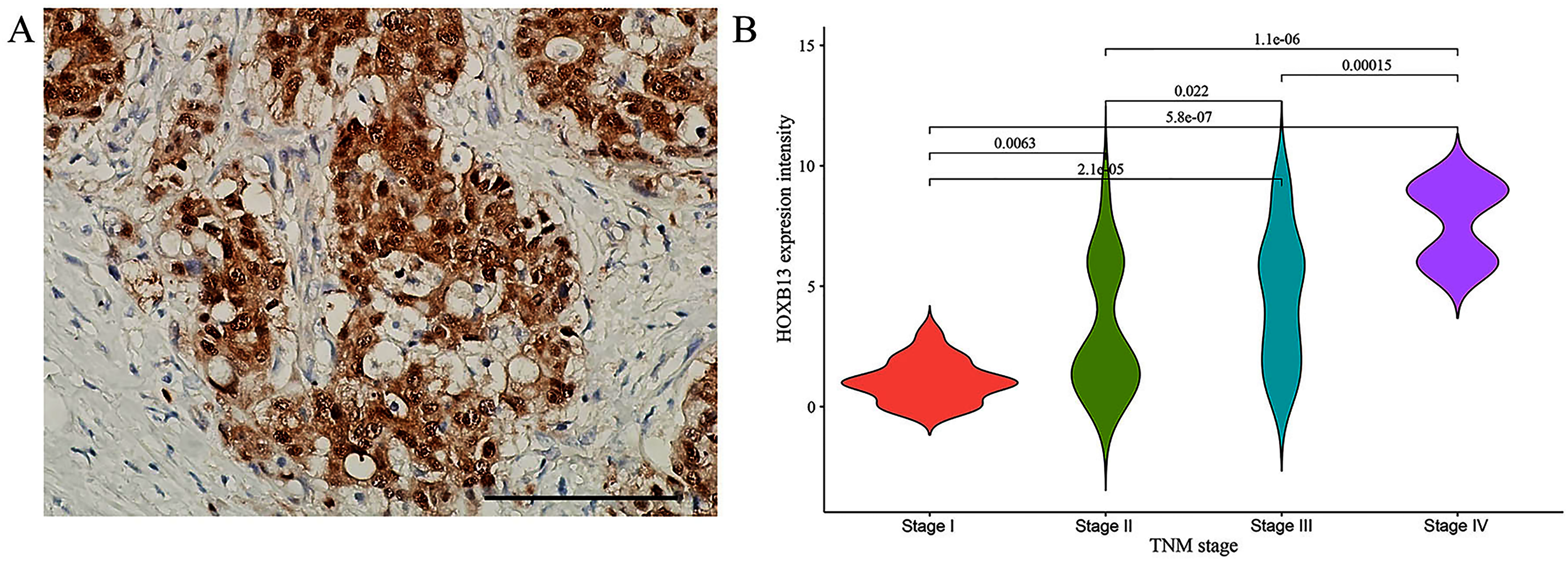
HOXB13 protein expression in rectal cancer tissues. (a) HOXB13 protein overexpression in rectal cancer tissues. Magnifications 40 × . (b) The expression difference of HOXB13 in different stages of rectal cancer. The expression intensity of HOXB13 appear an increasing trend from stages I to IV.
Relationship between HOXB13 expression and clinicopathologic features
The expression status of HOXB13 protein in 103 RC tissues was determined by IHC. Scores of no staining and low expression were regarded as “negative” for HOXB13 protein expression, and moderate and high expression scores as “positive” or “overexpression.” The expression of HOXB13 was significantly correlated with depth of invasion (P < 0.001), lymphatic invasion (P = 0.027), lymph node metastasis (P < 0.001), distant metastasis (P < 0.001), and pTNM stage (P < 0.001) in 103 RC patients. However, the expression of HOXB13 was not correlated with the clinicopathologic features tested, including age, gender, tumor size, and histological grade (Table 1).
Correlation between HOXB13 expression and patient prognosis
In terms of OS, univariate analysis using the Cox proportional hazard revealed that HOXB13 expression (P < 0.001), age (P = 0.020), lymphatic invasion (P = 0.001), lymph node metastasis (P = 0.001), pTNM stage (P < 0.001), and histological grade (P = 0.045) were significantly correlated with prognosis (Figure 4). Multivariate analysis of the Cox proportional hazards model further showed that HOXB13 expression (P = 0.001), age (P = 0.012), and lymphatic invasion (P = 0.022) were significantly correlated with OS (Figure 3(a) and (b)). Therefore, in RC, HOXB13 expression, age, and lymphatic invasion were independent poor prognosis factors. Furthermore, Kaplan–Meier survival curves were constructed using overall survival (OS) and disease-free survival (DFS) to detect whether HOXB13 expression in RC is associated with a poor prognosis. Twenty-nine of the 103 patients (28.2%) died during the follow-up period. The log-rank test showed that the OS and DFS of patients with HOXB13 positive expression were significantly shorter than those without HOXB13 expression. The positive expression of HOXB13 was negatively correlated with the OS and DFS (P< 0.001) (Figure 4(a) and (b)).
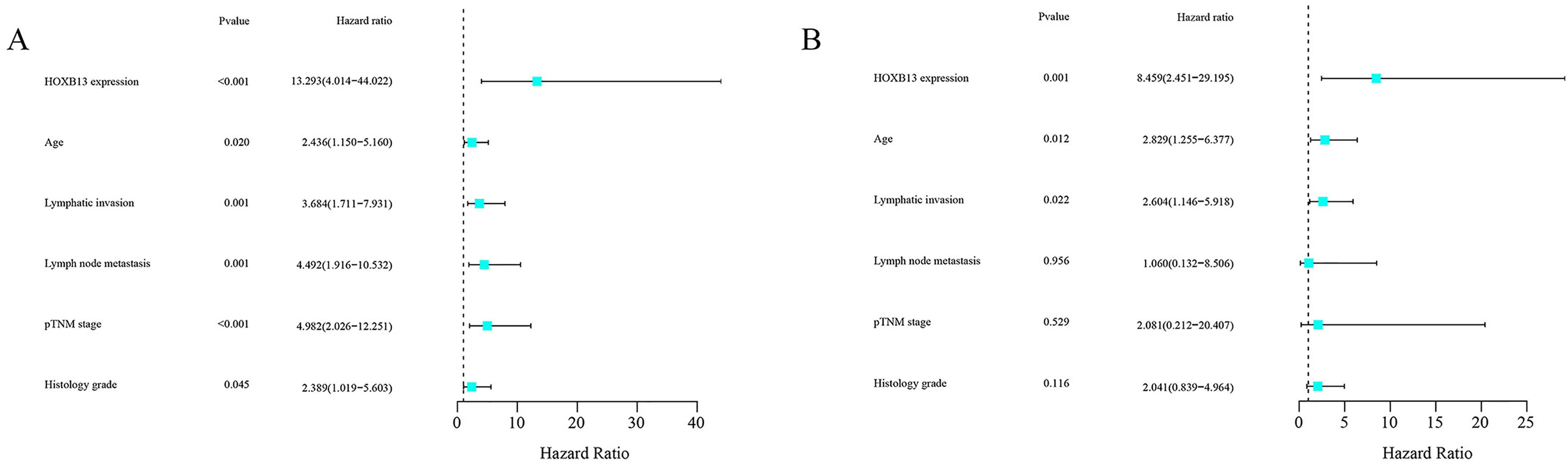
Cox proportional hazards analysis on overall survival in 103 patients with rectal cancer. (a) Univariate Cox proportional hazards regression analyses of HOXB13 expression and clinical features in rectal cancer. (b) Multivariate Cox proportional hazards regression analyses of HOXB13 expression and clinical features in rectal cancer.
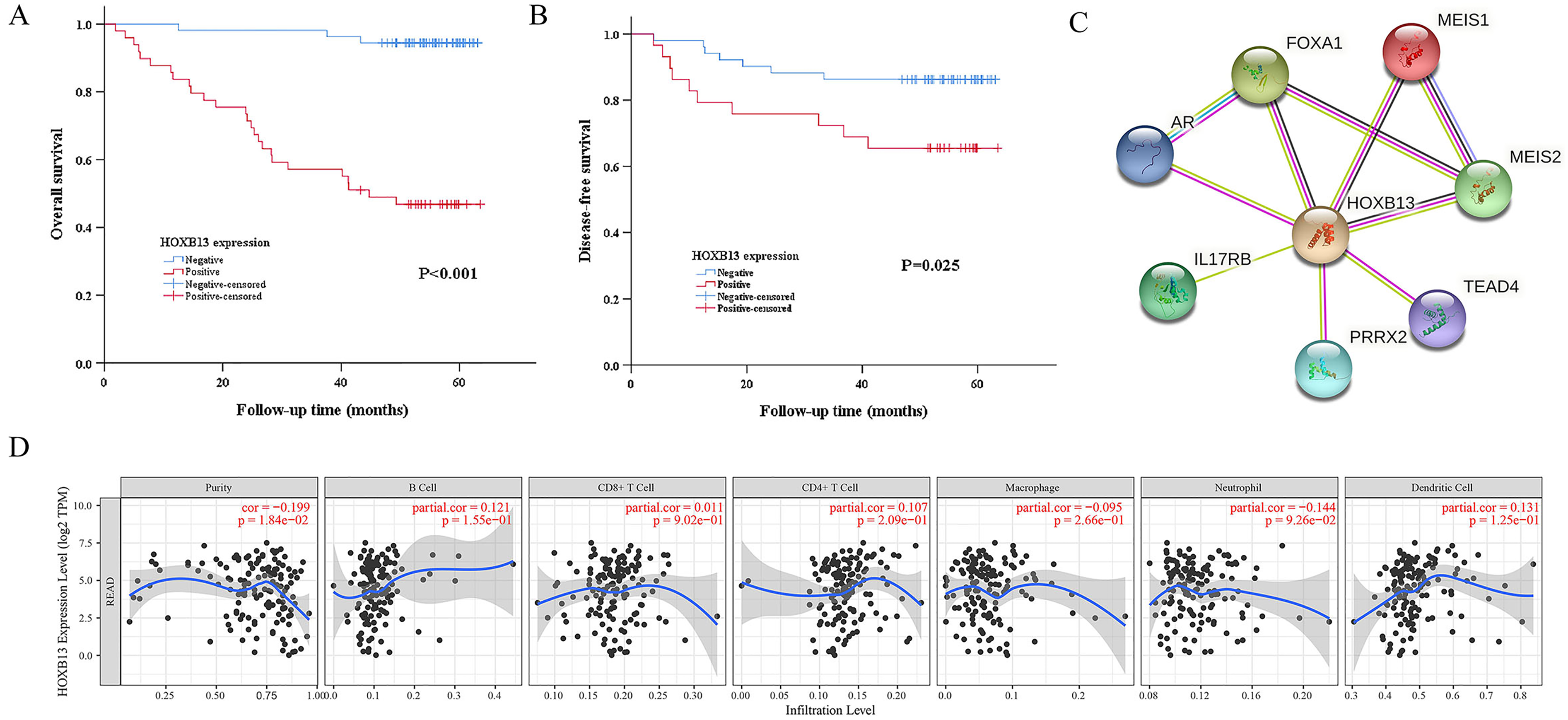
Survival curves using the Kaplan–Meier method by log rank test and associations of co-expression genes of HOXB13, tumor infiltrating immune cells, and HOXB13. (a) Overall survival of RC patients with negative and positive HOXB13 expression. (b) Disease-free survival of RC patients with negative and positive HOXB13 expression. (c) Genes associated with HOXB13 analyzed using the STRING database. (d) Associations between tumor purity, tumor infiltrating immune cells (TIICs; B cells, CD4 + T cells, CD8 + T cells, neutrophils, macrophages, and dendritic cells) and HOXB13 in RC.
Constructing protein interaction networks
The functional interaction between proteins is necessary for a malignancy's molecular mechanism and metabolism. Therefore, we used the STRING tool to analyze the protein–protein interaction (PPI) network of HOXB13 protein to determine their interactions in the progression of RC (Figure 4(c)). The seven proteins and corresponding gene names, annotations, and scores are listed in Table 2. These genes included: androgen receptor (AR), Meis homeobox 1 (MEIS1), Meis homeobox 2 (MEIS2), forkhead box A1 (FOXA1), TEA domain transcription factor 4 (TEAD4), paired related homeobox 2 (PRRX2), and interleukin 17 receptor B (IL17RB).
Correlations between TIICs and HOXB13
Recently, increasing evidence has affirmed the association between immunological features and survival in tumors. Thus, we further investigated the correlation between TIICs and HOXB13. The results were gained from the TIMER database, which confirmed a negative correlation between HOXB13 and tumor purity. In comparison, there were no significant associations between TIICs and HOXB13 (Figure 4(d)). Then, we further found that HOXB13 correlated with various biomarkers of immune cells (Table 3).
Potential mechanisms underlying the effects of HOXB13 on RC
The potential biological mechanism associated with HOXB13 overexpression was investigated by GSEA. The results showed that HOXB13 overexpression was related to natural killer cell-mediated cytotoxicity and the T-cell receptor signaling pathway (Figure 5(a) and (b)).
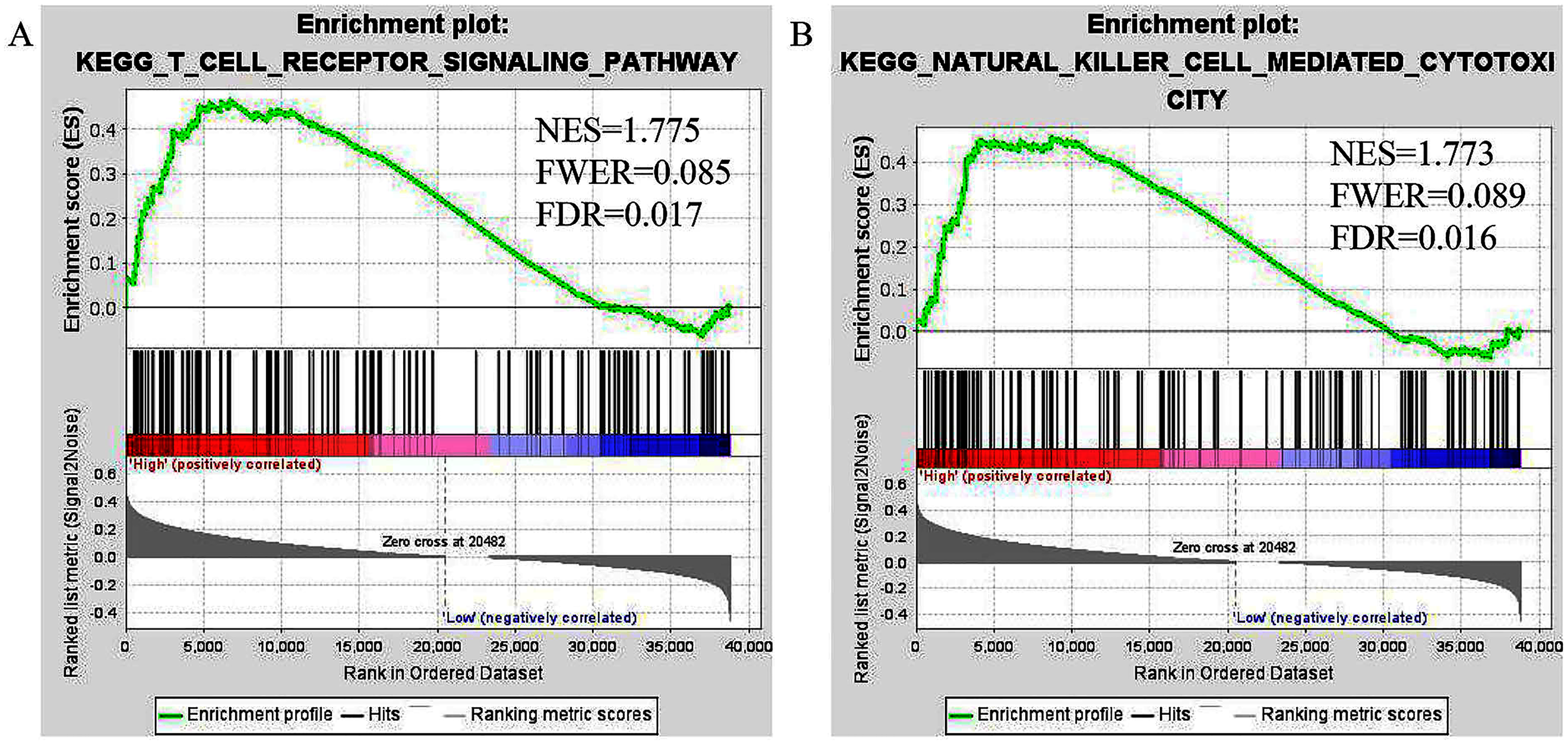
Cancer-related Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways associated with HOXB13 based on a gene set enrichment analysis (GSEA). (a) Natural killer cell-mediated cytotoxicity. (b) T-cell receptor signaling pathway.
Discussion
HOX genes are crucial regulators of normal embryonic development, cell growth and differentiation, and other critical processes of eukaryotic cell life. The tissue-specific HOX genes play essential roles in cancer development. 23 The deregulated expression of HOX genes has been found in esophageal cancer, 24 gastric cancer, 25 breast cancer, 26 prostate cancer, 27 and kidney cancer. 28 Interestingly, some HOX genes control the anteroposterior development of the intestine, 29 and their aberrant expression has already been associated with colorectal carcinogenesis, such as HOXB5 20 HOXB13, 13 HOXC9, 21 and HOXC10, 22 which is consistent with our study. However, little is known about the expression pattern and prognostic role of HOXB13 in RC. Thus, we further made a detailed investigation on the oncological effect and prognostic significance of HOXB13 in RC.
HOXB13, a member of the HOX genes family, is expressed explicitly in the distal colon, rectum, and prostate. Increasing evidence indicates HOXB13 to be associated with various malignant tumors. Interestingly, in recent decades, many studies about HOXB13 have completely contradictory results in several kinds of tumors. In prostate cancer, HOXB13 has been found to significantly inhibit in vitro and in vivo cell growth of PC3 cells with G1 cell cycle arrest mediated by the suppression of cyclin D1 expression 30 ; whereas subsequent data have shown that HOXB13 is overexpressed in prostate cancer and is speculated to lead to the pathogenesis of the neoplasm.8 For gastric cancer, it has been confirmed that decreased expression of HOXB13 mRNA was related to depth of invasion, tumor differentiation, and lymph node metastases, and was regarded as a poor prognostic factor for patients with gastric cancer. 9
In contrast, Guo et al. demonstrated that HOXB13 expression promotes the proliferation, migration, and invasion of gastric cancer cells by PI3 K/AKT/mTOR signaling via IGF-1R2. 4 Such contradictory phenomena were also found in CRC. A study to analyze the expression of the paralogous 13 HOX genes in proximal and distal CRC patients revealed that HOXB13 showed a statistical association with lymph nodes metastasis and was overexpressed in right CRC tissues. 31 Nevertheless, after 1 year, using five datasets from the Gene Expression Omnibus, a team of researchers from China found that HOXB13 was obviously less expressed in right colon cancer than left colon cancer and act as a tumor suppressor in right colon cancer. 32 These results indicate that the oncological effect of HOXB13 is complex and diverse, which may be explained by differences in the study populations and specimens. However, few studies have been conducted to investigate the expression pattern of HOXB13 in RC.
The tumor-suppressive effect of HOXB13 was firstly revealed by Jung et al., 33 who demonstrated that enhanced expression of HOXB13 drove CRC cells into growth suppression. Conversely, our study showed that HOXB13 mRNA and protein expression levels in human RC tissues were significantly higher than that in the normal adjacent tissues and increased as the TNM stage became advanced.
The prognostic significance of HOXB13 has been revealed in many tumors, such as gastric cancer, 9 prostate cancer, 10 and right colon cancer. 34 In our study, to sufficiently attest to the relationship between HOXB13 expression and prognosis in RC, we first analyzed the prognostic significance of HOXB13 in RC from a unitary perspective. Kaplan–Meier survival curves confirmed that HOXB13 overexpression was correlated negatively with OS and DFS in RC. Furthermore, multivariate Cox regression analysis demonstrated that HOXB13 expression, age, and lymphatic invasion were independent prognostic factors in RC. These findings verified that HOXB13 overexpression leads to a worse prognosis of patients with RC, which strengthens our former point of view.
To further detect the mechanism of HOXB13 participation in RC, we further explored seven genes associated with HOXB13: AR, MEIS1, MEIS2, FOXA1, TEAD4, PRRX2, and IL17RB. Interestingly, several studies have confirmed that AR 34 and PRRX2 35 were significantly associated with tumor immunity. Also, Jung et al. confirmed that HOXB13 could suppress the growth of prostate cancer cells by the negative regulation of T-cell factor 4. 33 These findings indicate that HOXB13 may be essential in tumor immunity. Nevertheless, little is known about the association between HOXB13 and the immune process in RC. Thus, we further detected the association between HOXB13 and immune infiltrates in RC. In comparison, the results indicated that HOXB13 has no associations with TIICs, with a negative correlation with tumor purity. More interestingly, we found that HOXB13 was co-expressed with various immune cell biomarkers, including ICOS, IL12RB2, IRF4, STAT3, FOXP3, and CCR8. Finally, the GSEA results indicated that HOXB13 participated in the various immune-associated processes, including natural killer cell-mediated cytotoxicity and the T-cell receptor signaling pathway. Therefore, further efforts are needed to explore the association between HOXB13 and immune cell infiltrates in RC.
In conclusion, this study demonstrates that HOXB13 is associated with the occurrence and development of RC. Overexpression levels of HOXB13 are predictive of a poor prognosis in RC. Our results highlight the significant association between HOXB13 and immune cell infiltrations. In addition, we can speculate that our data may provide an essential element for a more profound understanding of the molecular mechanisms that determine the pathogenesis of a tumor on the rectum.
Comparison of clinicopathological parameters between rectum cancer with HOXB13 negative and positive.
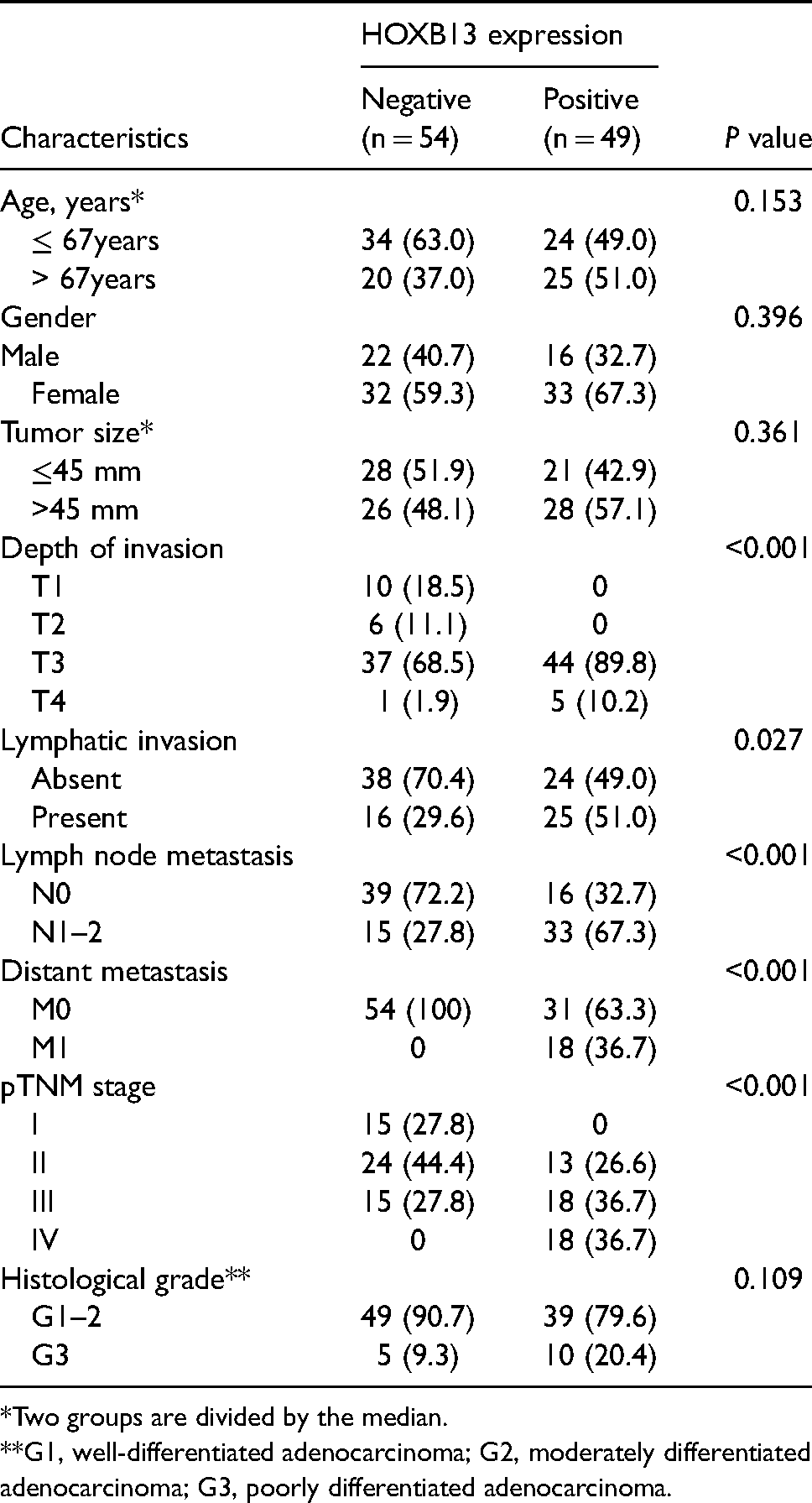
*Two groups are divided by the median.
**G1, well-differentiated adenocarcinoma; G2, moderately differentiated adenocarcinoma; G3, poorly differentiated adenocarcinoma.
Annotation of HOXB13-interacting proteins and their co-expression scores.

Correlations between HOXB13 and gene markers of immune cells in TIMER and GEPIA.
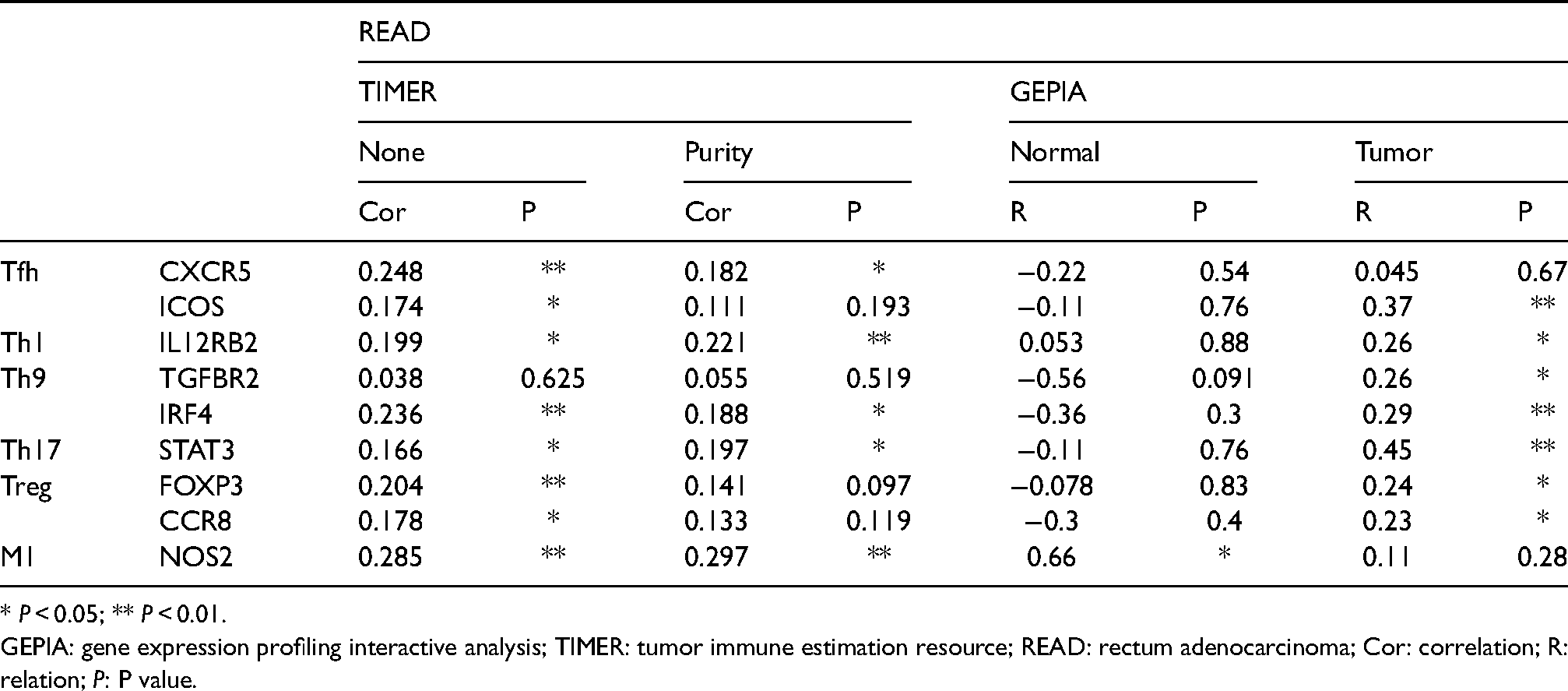
* P < 0.05; ** P < 0.01.
GEPIA: gene expression profiling interactive analysis; TIMER: tumor immune estimation resource; READ: rectum adenocarcinoma; Cor: correlation; R: relation; P: P value.
Footnotes
Ethics approval
The study protocol was authorized by the institutional review board of The Affiliated Xinhua Hospital, Dalian University. Investigators have to obtain informed consent before enrolling participants in clinical trials.
Author contributions
Da-Zhi Gao and Yu-Shen Yang contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Basic Scientific Research Project of Education Department of Liaoning Province (grant number LJKZ1192).
