Abstract
Introduction:
Prostate cancer has a high incidence in men and is the second cause of cancer death among americans male. microRNA (miR) is becoming a potential new prognostic factor for prostate cancer. Single nucleotide polymorphisms (SNPs) are common polymorphisms, characterized by a single exchange of nitrogen based in the DNA. This polymorphism is present in the miRs, altering their function.
Objective:
To evaluate the role of SNP rs1834306 of miR100 and rs2910164 of miR146a in the development and prognosis of prostate cancer.
Methods:
One hundred patients diagnosed with prostate cancer and 68 controls were selected. The identification of SNP was rated by quantitative polymerase chain reaction from blood samples, and the analysis was performed within the presence of SNP and the prognostic variables.
Results:
In the SNP rs1834306 (miR100), a smaller presence of the polymorphic homozygous genotype was identified in patients with PSA >10 ng/mL, (
Conclusions:
SNP of miR146a acts as a poor prognostic factor (Gleason ⩾7), and the SNP of miR100 is linked to better prognostic data (PSA <10). MiR100 was overexpressed in prostate cancer with worse prognostic factors.
Keywords
Introduction
Prostate cancer (PCa) is the most common non-cutaneous tumor in men. Approximately 191,930 newly diagnosed cases were estimated in the United States in 2020, and 33,330 deaths from this pathology will appear. 1
microRNA (miR) is a short non-coding RNA sequence with approximately 18–22 nucleotides whose main function is to act as a silencer for target genes. The dysregulation of miR is related to carcinogenesis, due to the alteration of key biological pathways. 2 By targeting tumor suppressors and oncogenes, miRs can be a cellular regulator key for processes that explain the cancer cell phenotype, and could be promising therapeutic targets. 3
miR-146a is defined in the regulation of inflammation and its expression appears to be regulated by inflammatory factors such as IL-1 and TNF-alpha. miR146a blocks the expression of some target genes that are involved in the TLR pathways. Sun et al. 4 describe that miR146a plays a role in PCa through negative Rac1 reduction. Lin et al. 5 reported that miR146a can function as a tumor suppressor miR during a modulation of the HA/ROCK1.
Many studies described that miR100 could be important in carcinogenesis, with a role to target key genes in cellular progression. It is possible for the function in tumor promotion and tumor suppression. miR-100 might have importance in the prognostic and/or diagnosis of PCa. The dysregulation and aberrant expression could be present in the carcinogenesis and tumor progression. On the other hand, miR100 re-sensitizes these carcinogenic cells to chemotherapeutic drugs. Thus, these findings suggested that miR100 could be a new target in treatments and a new molecular marker for PCa. 6
Single nucleotide polymorphisms (SNP) can be identified in approximately 100 independent variants, and have been associated with gene expression, changes in normal prostate tissue, PCa, and prostatic secretions.7-9
Some studies have shown the presence of SNPs not only in DNA but also in miRs, modifying their biological behavior and effect. 10 In a systematic review it was observed that SNP (rs2910164)-C in miR146a is implicated in the susceptibility to gynecological cancer. The SNP miR100 (rs1834306)-G was not associated with risk of female neoplasms. 11
Methods
Patients
This study included 100 patients diagnosed with clinically localized PCa, submitted to radical prostatectomy and 68 patients without PCa in a control group.
We assessed the distributions of polymorphisms related to level of preoperative PSA <10ng/mL and ⩾10ng/mL, Gleason score (GS) subdivided in ⩽6 and 7–10, pathological staging, divided into T2 (organ-confined) and T3 tumors (extra-prostatic disease) and biochemical recurrence, which were considered positive when the post-treatment PSA was higher than 0.2 ng/mL. In the control group, we included 68 healthy patients that were randomly selected. The inclusion criteria for the control group were PSA below 2.5 ng/mL, no family history of PCa, and over 60 years old. The same surgeon performed all surgeries. Blood samples were collected and stored at −20°C. All patients agreed with the study and signed an informed consent form authorizing their participation in the study and allowed their biological samples to be genetically analyzed. This study was approved by the Ethics Committee of University of São Paulo Medical School no. 1.912.899.
The patients’ samples were analyzed by polymerase chain reaction (PCR) for frequency of 2 SNPs:
Analysis of SNPs
Genomic DNA was extracted from patients’ blood samples using QIamp DNA mini kit (QIAGEN). The SNPs were genotyped using probes labeled with fluorophores (Taqman®-SNP Genotyping Assay Kit) and an ABI 7500 fast system for amplification by PCR.
The PCR was performed using these volumes: 2.0 μL of DNA, 2.75 μL distilled water for PCR, 0.25 μL primer specific to each polymorphism and 5.0 μL of Master Mix TaqMan, totalizing 10 μL for each reaction. The PCR template was amplified by 40 cycles at 95°C for 15 seconds and at 60°C for 1 minute, proceeded by 2 minute at 50°C, and 10 minute at 95°C.
MiR isolation and real time PCR
For an analysis of the expression of in vitro assays, microRNA was extracted using the mirVana kit (Ambion, Austin, TX, USA), according to the manufacturer’s instructions. How agents were quantified in a spectrophotometer (Nanodrop DN-1000, Wilmington, USA) (260/280 nM) and stored at −20ºC. These samples were transformed in complementary DNA (cDNA) from total RNA extracted, using High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, CA, USA) and RT primer for miR100.
The miR expression reactions were performed according to the kit protocol (Applied Biosystems® TaqMan® Universal PCR Master Mix). The thermal cycler used was ABI 7500 fast (Applied Biosystems, CA, USA) in standard mode. To quantify the samples, TaqMan® chemistry (Applied Biosystems) was used. Reagents were used in the following concentrations: 0.5 µL of the specific primer TM for miR100; 5 µL of TaqMan® master mix; 2.5 µL of nuclease-free water and 2.0 µL of cDNA (concentrated in 15 ng). The level of gene expression was obtained by the 2-ΔCT method, acquired from Applied Biosystems DataAssist Software. All reactions were performed in duplicate and the RNU48 was used as an endogenous control for miR (Applied Biosystems).
Statistical analysis
We analyzed the allele frequency in control group in comparison to patients with PC to verify the degree of significance, using chi-square test (X2). To verify the association of each polymorphism and prognostic factors—such as preoperative PSA, GS, and pathological staging—we assessed by calculating the odds ratio (OR) and its respective 95% confidence interval (95% CIs). Statistical analyses were performed in SPSS 19.0 software.
For expression and correlation analysis, a Kolmogorov-Smirnov normality test was performed, and subsequently the Mann–Whitney statistical test, using the GraphPad Prism 8.0.1 software.
Results
The rs1834306 (miR100) polymorphism was identified in 70.6% and 75% of the PCa and control groups, respectively. SNP ID rs2910164 (miR146a) was identified in 52% of the PCa group and in 57.4% of the control group.
We performed an analysis using a Chi-square test to elucidate whether any of the genotypes (wild homozygous, heterozygous, or polymorphic homozygous) of the studied polymorphisms have a higher prevalence in the group of patients with PCa or in the control group.
The Chi-square test was performed with SNPs rs1834306 (miR100) and rs2910164 (miR146a), respectively (Table 1).
Distribution of the genotypes of the two SNPs in patients diagnosed with PCa and in individuals in the control group.
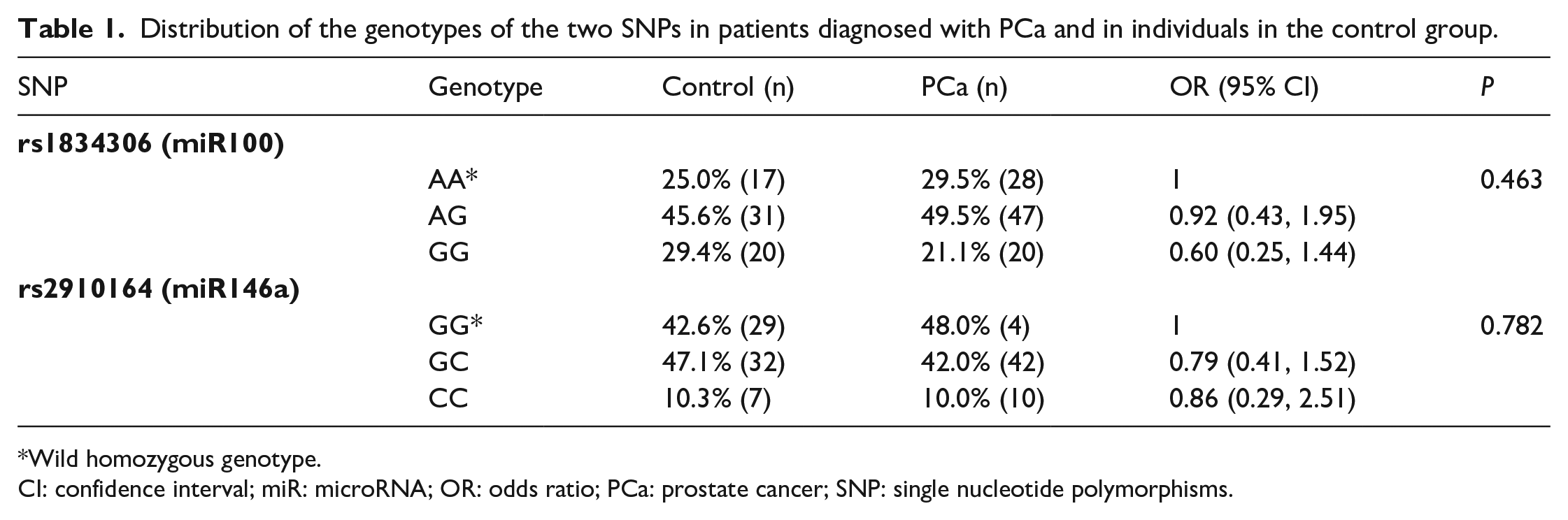
Wild homozygous genotype.
CI: confidence interval; miR: microRNA; OR: odds ratio; PCa: prostate cancer; SNP: single nucleotide polymorphisms.
We did not find statistical differences in miR100 and miR146a, as shown by the
When evaluating only the presence of the polymorphic allele in our two groups, the results were maintained without the statistical difference as determined in Table 2.
Allele distribution of the two polymorphisms between patients with PCa and control group.
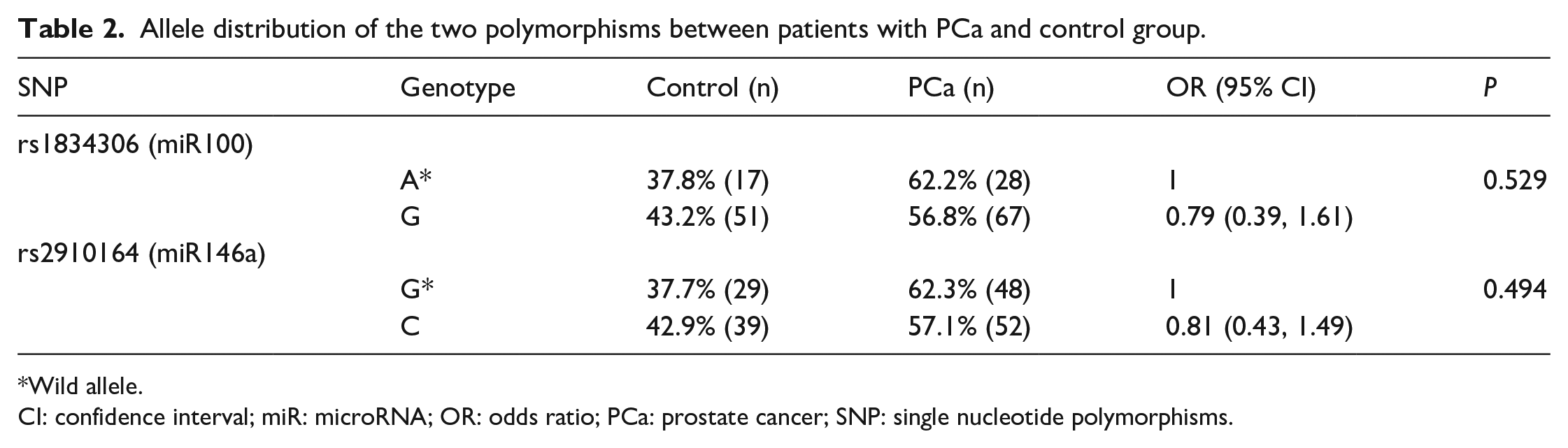
Wild allele.
CI: confidence interval; miR: microRNA; OR: odds ratio; PCa: prostate cancer; SNP: single nucleotide polymorphisms.
When we analyzed the patients according to the classic prognostic factors of PCa, some findings were identified. First, we subdivided patients according to preoperative PSA levels into two groups: PSA <10 ng/mL and patients with PSA ⩾ 10 ng/mL. In this comparison, we found statistical significance for the SNP rs1834306 (miR100). The polymorphic homozygous genotype was less present in the group of patients with PSA ⩾10 ng/mL, (OR 0.12; 95% CI 0.01, 1.04;
SNP genotypes and allele distribution in relation to the preoperative PSA level.
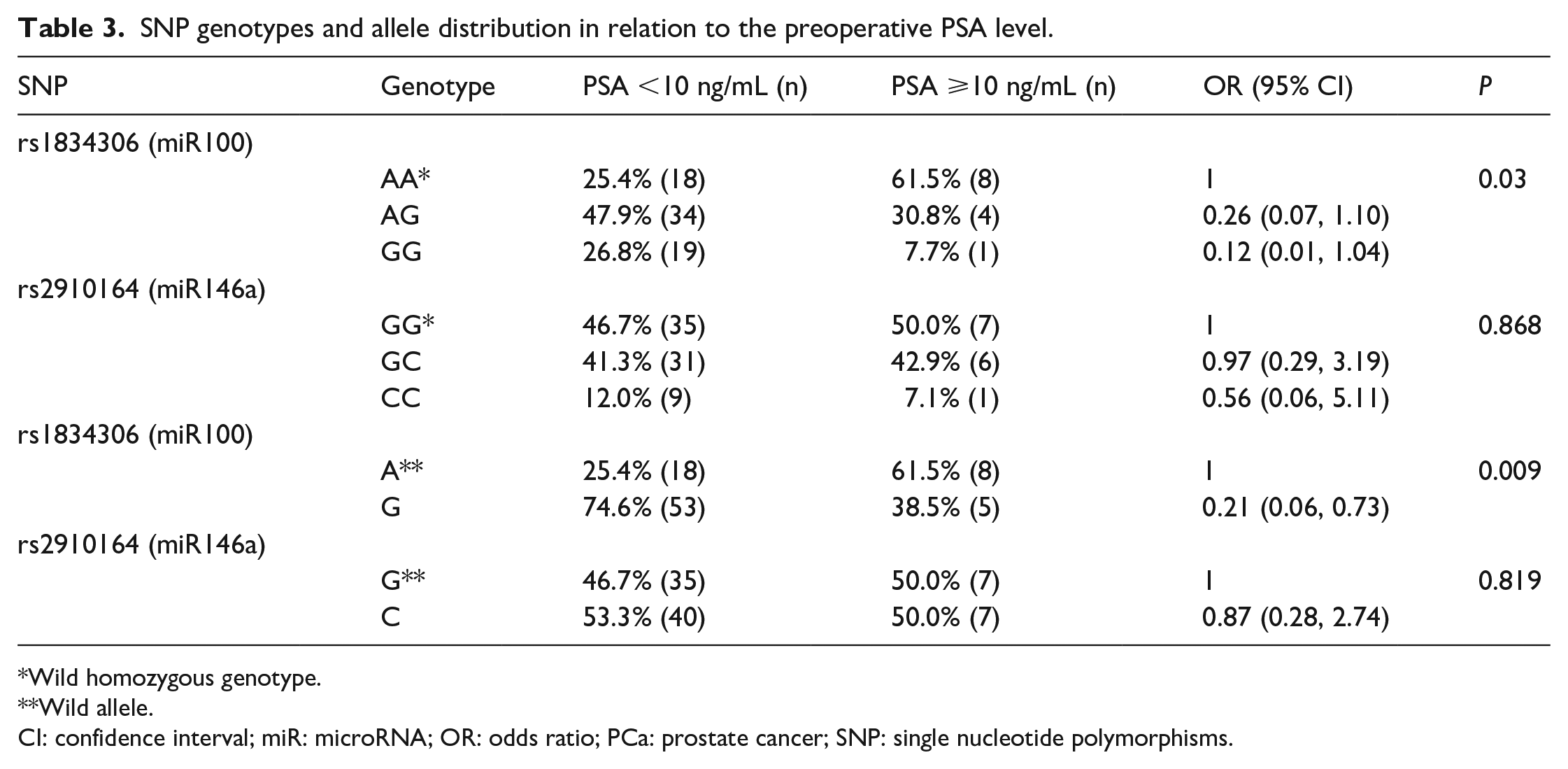
Wild homozygous genotype.
Wild allele.
CI: confidence interval; miR: microRNA; OR: odds ratio; PCa: prostate cancer; SNP: single nucleotide polymorphisms.
We analyzed patients diagnosed with PCa and categorized in GS <7 and ⩾7, comparing their genotypes in both SNPs. No statistical significance was identified between these variables. However, when we consider only the presence of the polymorphic allele in the SNPs, we observed that the C polymorphic allele in the SNP rs2910164 G>C, located in miR146A, is more frequent in patients with GS ⩾7 compared to patients with GS <7, shown to be more prevalent in this group (OR 3.19; 95% CI 1.00, 10.15;
Genotypes and allele distribution in relation to the Gleason score.
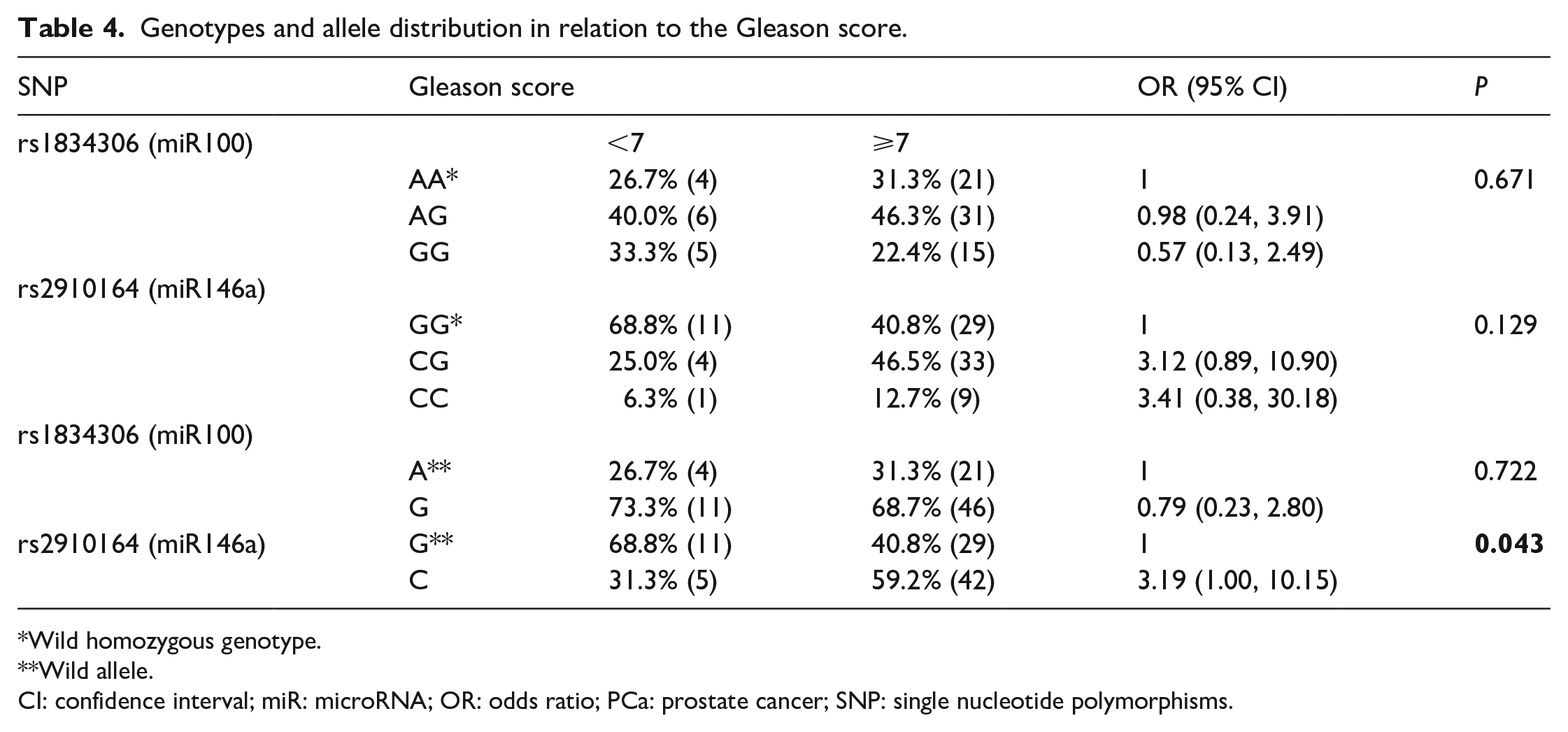
Wild homozygous genotype.
Wild allele.
CI: confidence interval; miR: microRNA; OR: odds ratio; PCa: prostate cancer; SNP: single nucleotide polymorphisms.
When evaluating the expression of miR100 in patients with PCa, we found no significant difference in relation to the control group. However, among patients with PCa, those with the worst prognostic factors, such as pathological staging and biochemical recurrence, (
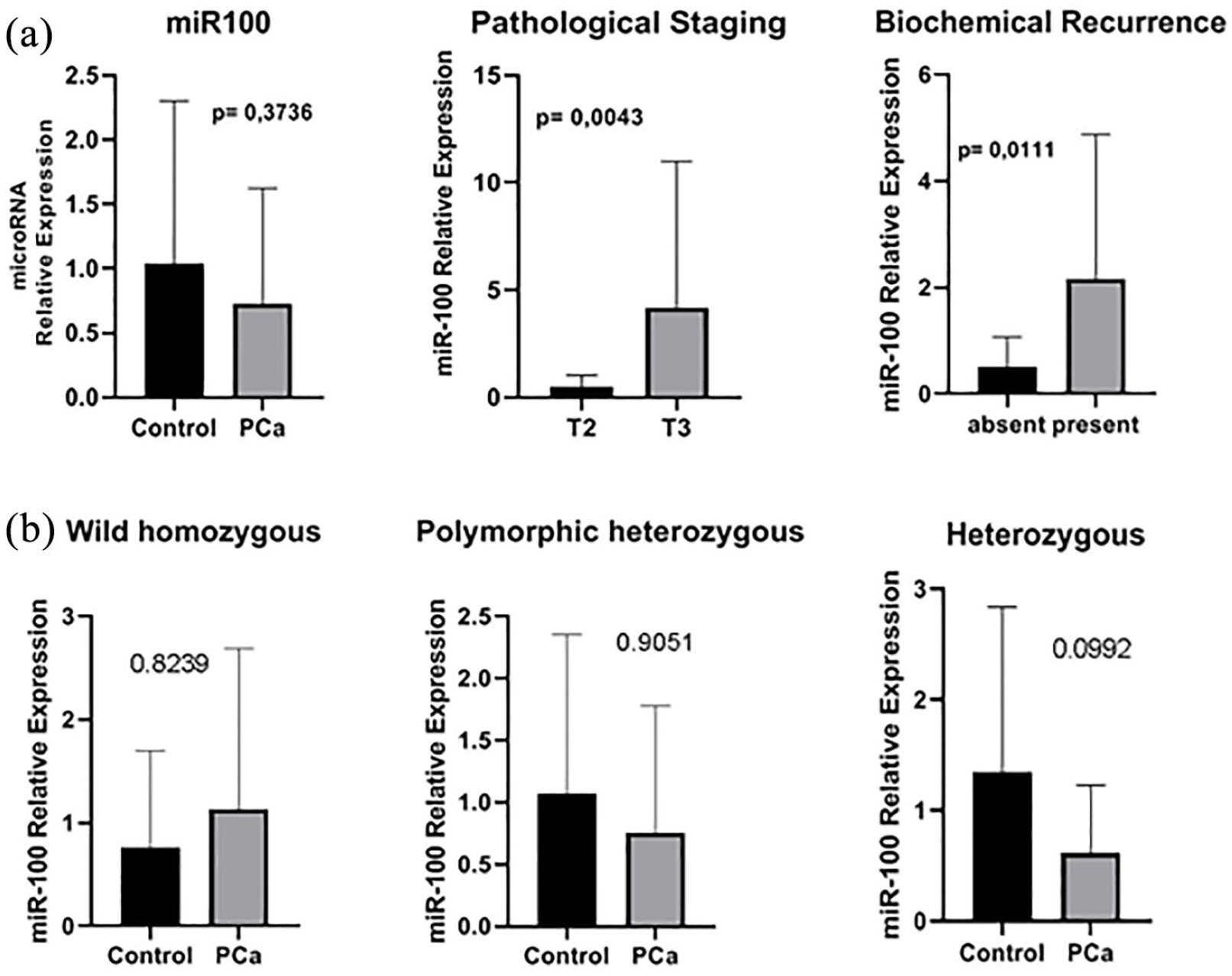
miR100 expression in patients with PCa. (a) Correlation with prognostic factors, with pathological staging, comparing the expression of miR100 in a localized tumor (T2) with locally advanced tumor (T3) and biochemical recurrence, comparing patients who relapsed with those who did not. (b) Expression of miR100 in patients with PCa compared to controls categorized by wild-type homozygous genotypes; polymorphic homozygous and heterozygous.
When we compared the expression of miR100 in relation to SNP genotyping rs1834306, we found no statistical difference in the expression of the miR100 wild homozygote in men with PCa compared to the control group. However, we found a possible tendency of increasing the expression of miR100 in the heterozygous genotype of the control group in patients with PCa (
Discussion
In our study, we sought to assess the prevalence of SNPs rs2910164 and rs1834306 in miR146a and miR100, respectively, and their relationship with prognostic variables through patients’ blood analyses with PCa and controls.
We did not find statistically significant association between the SNPs tested in miRs and the presence of PCa. No difference was observed in the frequency of possible genotypes between patients with PCa and the control group. Our findings are in line with those described in the literature for SNP rs2910164 (miR146a), 12 and for SNP rs1834306 (miR100). 10 However, we observed relevant findings associated with the genotypes of patients with PCa and the variables of prognostic determination.
Chen et al. 13 showed that miR146a is a suppressor among several types of cancers. In a recent review, miR146a was shown to have an important role in the regulation of the T lymphocyte, where its absence results in cell proliferation and activation of the immune system. 14 Consequently, animals with a genetic deletion of miR146a developed chronic inflammatory diseases, with an increased production of myeloid cells, resulting in the development of sarcomas and lymphoma. Thus, miR146a has been characterized as a suppressive miR at least in the context of the immune system.15,16
In our study, we observed an association between the SNP rs2910164 in miR146a and patients with a GS ⩾7, which is known to be a factor of worse prognosis. This finding allows us to suggest that this SNP may affect the binding of this miR with the target gene or inactivate it, inhibiting the function as a suppressor miR. In a study published in 2018, Damodaran et al. 11 investigated the presence of SNP rs2910164 among 100 patients with PCa and 100 controls. When assessing its association with Gleason ⩾7 and PSA at diagnosis, no statistical significance was identified. We believe that the difference in findings could be related to a higher prevalence of the C (polymorphic) allele in the Indian population compared to others, such as American, African, and European. In another similar study, in a Chinese population, a 0.65-fold decreased chance of the occurrence of PCa was identified in those with a CC (polymorphic homozygote) genotype. 17
miR100 is an important diagnostic marker 18 and prognosis estimator 19 of several tumors resulting from the dysregulation or aberrant expression. In addition, new studies demonstrate its ability to re-sensitize tumors to chemotherapy,20,21 which opens the possibility of becoming a new target against tumors.
Nabavi et al. 22 demonstrated that the inhibition of miR100 in metastatic cell lines (LNCaP and DU145) significantly reduced cell proliferation. Therefore, it is concluded that miR100 is necessary for the survival of dormant PCa cells exposed to androgen deprivation.
In our study, miR100 was overexpressed in patients with worse prognostic factors, with statistical significance for pathological staging and biochemical recurrence (
In previous studies, we demonstrated an increase in the expression of miR100 three times in patients who had biochemical recurrence after prostatectomy versus those who did not present recurrence, which would position miR100 as oncomir. 25 In this study, we identified the tendency to earlier biochemical recurrence in patients with polymorphic homozygous alleles in both SNPs for miR100 and miR146a, despite the lack of statistical significance. Chen et al. 26 analyzed the SNP rs2910164 (miR146a) relationship and biochemical recurrence after radical prostatectomy in 72 men in southern China with an average follow-up of 2 years; 33% (n = 24) of the patients had biochemical recurrence.
The SNP rs1834306 is located in the pri-miR100 region and our findings suggest an association with PSA <10, especially when the presence of the isolated polymorphic allele was evaluated (OR 0.21; 95% CI 0.60, 0.73;
The data in the literature are still scarce regarding the polymorphisms in miRs. This is the first SNP study rs1834306 on miR100 in PCa. Our data showed the impact of SNPs on prognostic factors of the disease, but further studies are needed to clarify the prevalence of these SNPs in different populations as well as the influence of these polymorphisms on target genes, and consequently, the possible impact of this mutation on protein expression.
In conclusion, our data reinforce this hypothesis by the association of SNP rs1834306 with PSA <10, especially when the presence of the isolated polymorphic allele was evaluated.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The author Gabriel Fernandes Dias Ferreira received financial support grant #2015/23248-3, Sao Paulo Research Foundation (FAPESP) for this research.
