Abstract
Background:
Cluster of differentiation molecules are markers of immune cells that have been identified as a potential immunotherapeutic target for cancer treatment. MicroRNAs are small non-coding RNAs that act as tumor suppressors or oncogenes whose importance in diagnosis, prognosis, and treatment of gastric and colorectal cancers has been widely reported. However, their association with cluster of differentiation molecules in gastrointestinal cancers has not been well studied. Therefore, our study aimed to analyze the relationship between microRNAs and cluster of differentiation molecules in gastrointestinal cancers, and to identify cluster of differentiation molecule-associated microRNAs as prognostic biomarkers for gastrointestinal cancer patients.
Methods:
Targetscan, Starbase, DIANA microT, and miRDB were used to investigate microRNA profiles that might be correlated with cluster of differentiation molecules in gastrointestinal cancers. Moreover, The Cancer Genome Atlas data analysis was used to investigate the association between cluster of differentiation molecules and microRNA expression in patients with gastric, colon, rectal, pancreatic, and esophageal cancers. The Kaplan–Meier plotter was used to identify the association between overall survival and cluster of differentiation molecule-associated microRNA expression in gastrointestinal cancer patients.
Results:
miR-200a, miR-559, and miR-1236 were negatively associated with CD86, CD81, and CD160, respectively, in almost all types of gastrointestinal cancers, which were further verified in the in vitro studies by transfecting microRNA mimics in gastric cancer, colon cancer, pancreatic, and esophageal cell lines.
Conclusion:
Our study showed that miR-200a, miR-1236, and miR-559 are identified as cluster of differentiation-associated microRNAs in gastrointestinal cancers, providing a novel perspective to identify new therapeutic targets for cancer immunotherapy in gastrointestinal cancer patients.
Introduction
Gastrointestinal (GI) cancer refers to malignant pathological changes in the digestive tract and GI organs, such as the esophagus, stomach, pancreas, colon, rectum, and liver. 1 GI tumors are the most common cancers in both developed and developing countries, 2 which has a poor prognosis and a high mortality rate. It has been shown that GI cancer is closely related to heredity, environment, dietary habits, precancerous lesions, and intestinal (gut) flora. 3
CD molecules play an important role in immunity and are considered as ideal targets for cancer immunotherapy. 4 Many studies have shown that CD73 inhibits the anti-tumor immune response. For example, the silencing of CD73 resulted in the increase of T-cell infiltration into the tumor site. 5 Jin et al. 6 found that CD73 was associated with tumor cell expansion and T-cell suppression. In addition, in vivo experiments have shown that the CD200 signal of tumor cells suppressed the anti-tumor immune response. 7 Accumulating evidence has suggested that CD molecules are associated with prognosis in cancer patients. For example, overexpression of CD274 is associated with poor prognosis and resistance to antitumor therapy in some malignant tumours. 8 In addition, CD9 expression and the number of CD133 (+) cells in gastric cancer were associated with poor prognosis.9,10 Furthermore, CD molecules also act as a biological marker for cancer. Studies have shown that expression of CD274 in tumor cells is a valuable biomarker for cancer prognosis. 4 Expression of CD166 (ALCAM) is a marker of poor survival in patients with esophageal cancer. 11 CD86 expression in lymphoma cells and increased infiltration of CD56 (+) NK cells in tumor tissues are potential markers for identifying a helicobacter pylori-dependent tumor. 12 CD97 promoted proliferation, invasion, and migration of gastric carcinoma cells, and led to the formation of premetastatic niche through exosomes. 13 Thus, it is important to explore novel approaches to target CD molecules in GI cancer, which may help to achieve better outcomes in the clinical application of cancer immunotherapy.
A microRNA (miRNA) is a small non-coding RNA molecule containing 22 nucleotides, and is found in plants, animals, and viruses. Its function is to silence RNA and to regulate gene post transcriptionally through binding to the 3′-untranslated region (3′-UTR) of the target genes. The importance of miRNAs in prognosis, diagnosis, and treatment as well as drug resistance of GI cancer has been widely reported.14-17 Recently, accumulating evidence has indicated that a variety of miRNAs could target CD molecules in GI cancers, thereby modulating the cancer progression. Using the Gene Expression Omnibus database and RNA-Seq colon cancer data from The Cancer Genome Atlas data portal, Gao et al. 18 reported that CD92 was targeted by miR-183, miR-206, and miR-147 in colon cancer, and differentially expressed genes (DEGs) were identified between colon tumor samples and adjacent normal samples. An in vivo experiment confirmed that miR-192 and miR-215 had high expression in gastric cancer and were capable of regulating CD166, and accordingly, the expression of CD166 was reduced in primary gastric cancer. 19 Also, miR-483-5p promotes metastasis of lung adenocarcinoma by acting directly on CD166. 20 However, the CD molecule-associated miRNAs have not received much scholarly attention and are not well studied. Hence, it is important to study the association between CD molecules and miRNA for the treatment of several diseases, including cancer.
This study aimed to: (a) analyze the relationship between miRNA and CD molecules in GI cancers by using RNA-Seq datasets from The Cancer Genome Atlas (TCGA) consortium 21 ; (b) identify miRNAs that may have an effect on particular CD molecules; and (c) validate this hypothesis in the GI cancer cell lines accordingly. The flow chart of this study design as well as all computational and experimental passages are demonstrated in Figure 1. Our study might provide evidence for identifying new miRNA targets or prognostic markers for GI cancer patients who undergo immunotherapy.
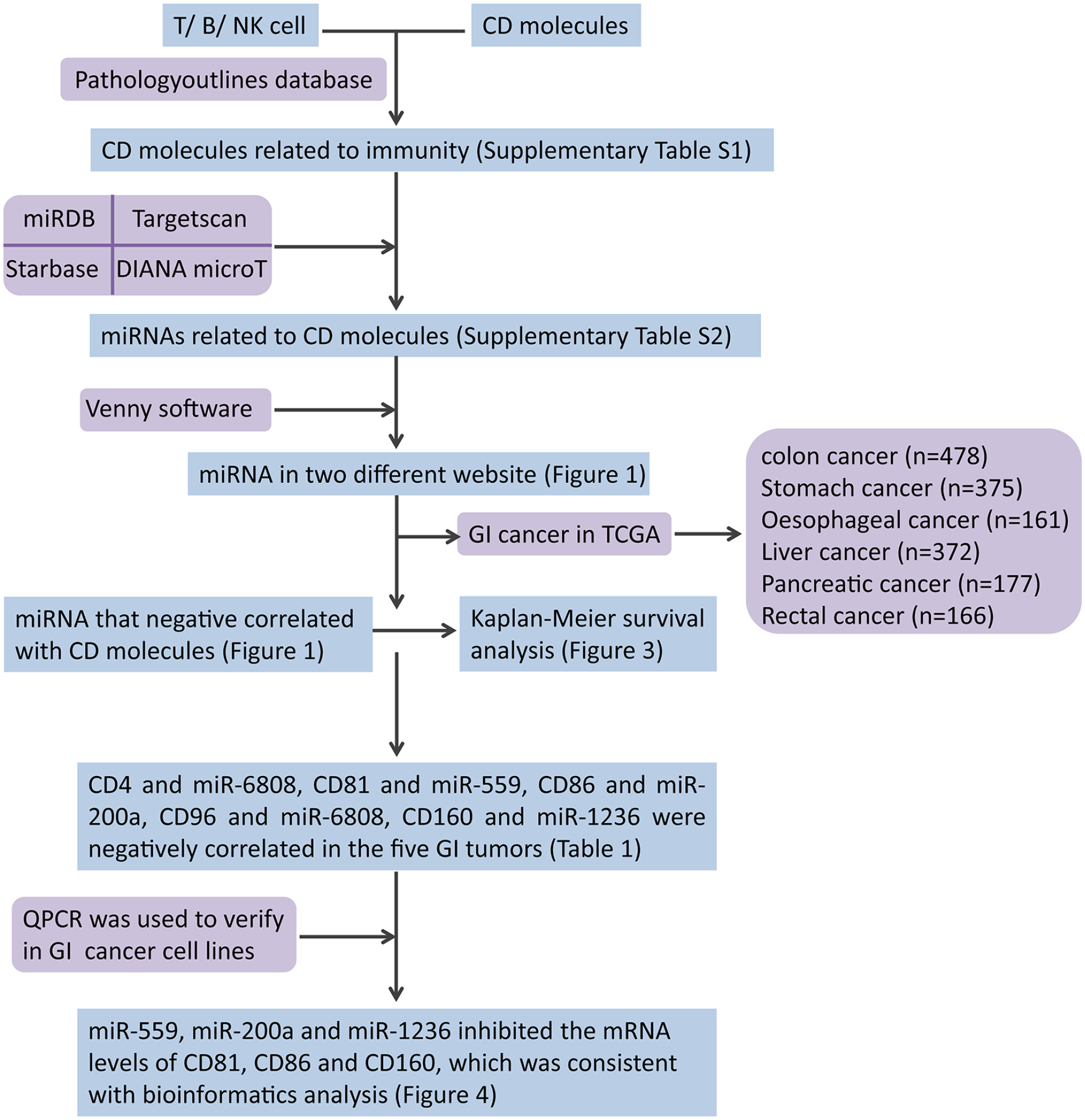
Flow diagram of study design. All computational and experimental passages used have been provided.
Materials and methods
Bioinformatics analysis
CD molecules related to immunity were obtained from Pathology Outlines Database (https://www.pathologyoutlines.com/cdmarkers.html). Additionally, miRNAs related to CD molecules were discovered using Targetscan v7.1 (http://www.targetscan.org/vert_71/), Starbase v2.0 (http://starbase.sysu.edu.cn/), DIANA microT v.5.0 (http://diana.imis.athena-innovation.gr/DianaTools/index.php?r=site/index), and miRDB (https://bioinfogp.cnb.csic.es/tools/venny/index.html). 22 In detail, predicted miRNA targets in the species of human were searched by entering certain CD molecules on the TargetScan or miRDB website. In the website of Starbase, the target information of CD molecules was viewed through the miRNA-messenger RNA (mRNA) in the navigation bar. In the website of DIANA TOOLS, we clicked DIANA microT-CDS and entered the corresponding CD molecules to find the potential miRNA targets. Venny software was used as an interactive application with Venn's diagrams that shows all possible logical relations between a finite collection of different sets. The researchers then used Venny software v2.1.0 (https://bioinfogp.cnb.csic.es/tools/venny/) to find miRNAs in at least two different sites. 23 Cancer datasets from TCGA (https://www.cancer.gov/tcga) incorporating RNA-Seq data files in six different cancer types were also obtained. These datasets contained information on colon adenocarcinoma (478 tissue samples), stomach adenocarcinoma (375), esophageal adenocarcinoma (161), liver adenocarcinoma (372), pancreatic adenocarcinoma (177), and rectal adenocarcinoma (166).
Cell culture and transfection
Gastric cancer cell line TMK1 was a gift from Dr. Eiichi Tahara (University of Hiroshima, Hiroshima, Japan). 24 Gastric cancer cell line BGC823, colorectal cancer cell line SW480, esophageal carcinoma cell line Eca109, and pancreatic cell line Panc-1 were obtained from the American Type Culture Collection (Manassas, VA, USA). All cancer cells were grown in RPMI 1640 (Gibco, Big Cabin, OK, USA), supplemented with 10% fetal bovine serum (FBS) (Gibco) and 1% Penicillin Streptomycin Glutamine (Gibco). MiR-559, miR-200a, and miR-1236 mimics were purchased from GenePharma (Shanghai, PR China). All cancer cell lines were transfected with 40 nM negative control and three kinds of miRNA mimics using 4 μL of jetPRIME reagent (Polyplus transfection, Strasbourg, France) as per manufacturer’s protocol.
Quantitative Real-time PCR
Forty-eight hours after transfection, cells were harvest and total RNA samples were extracted using TRIZOL reagent (Ambion, Life Technologies, Austin, TX, USA) and reverse transcription was performed according to the manufacturer’s protocol of FastKing RT Kit (With gDNase) (TIANGEN, Beijing, China). For detection of gene expression, 2×SYBR Green reagent (FastFire qPCR Premix SYBR green kit, TIANGEN), 0.5 mL cDNA together with primers at the concentration of 0.25 mM were added to a 96-well plate for Quantitative Real-time PCR (qPCR) analysis. qPCR was performed on the Real-time PCR system (CFX96 Touch Real-time PCR detection system, Bio-Rad, Hercules, CA, USA) with GAPDH used as an internal control. The relative expression levels of target genes were validated by the △△Ct calculation method. A Ct > 34 was deemed as transcript not detected. Relative fold changes were determined using the value of 2-△△Ct.
Primer sequences for different CD molecules were list as follows. CD81: Forward 5′-CTGCTTTGACCACCTCAGTGCT-3′, Reverse 5′-TGGCAGCAATGCCGAT GAGGTA-3′; CD86: Forward 5′-CCATCAGCTTGTCTGTTTCATTCC-3′, Reverse 5′-GCTGTAATCCAAGGAATGTGGTC-3′; and CD160: Forward 5′-GACCTACC AGTGTTGTGCCAGA-3′, Reverse 5′-ATCCCGTCACTGTGTAGT TCCC-3′. All the pairs of primers were obtained from ORIGENE (https://www.origene.com/).
Statistical analysis
Statistical analysis was conducted using GraphPad Prism 6 (GraphPad Software Inc., San Diego, CA, USA). The correlation of CD molecules and miRNA was analyzed using the Pearson test and the Spearman test. Survival analysis was represented by Kaplan–Meier curve with P values calculated using the log-rank test. Data were expressed as mean ± SEM. Unpaired student's t-test was used for two groups of comparisons. A P value < 0.05 was considered statistically significant.
Results
Identification of potential miRNAs related to CD molecules in GI cancers
The immune response of CD molecules expressed in immune cells, including T, B, and nature killer (NK) cells, may play an important role in the mucosal innate immune defense and GI cancers. 25 Therefore, CD molecules expressed in T/B/NK cells were summarized (Supplementary Table S1), and miRNAs related to CD molecules obtained from databases including Targetscan, Starbase, DIANA microT, and miRDB were identified. To enhance accuracy, Venny software 23 was used to find predicted miRNAs at least in two different websites (Figure 2) and the overlapping miRNAs were list in Supplementary Table S2.
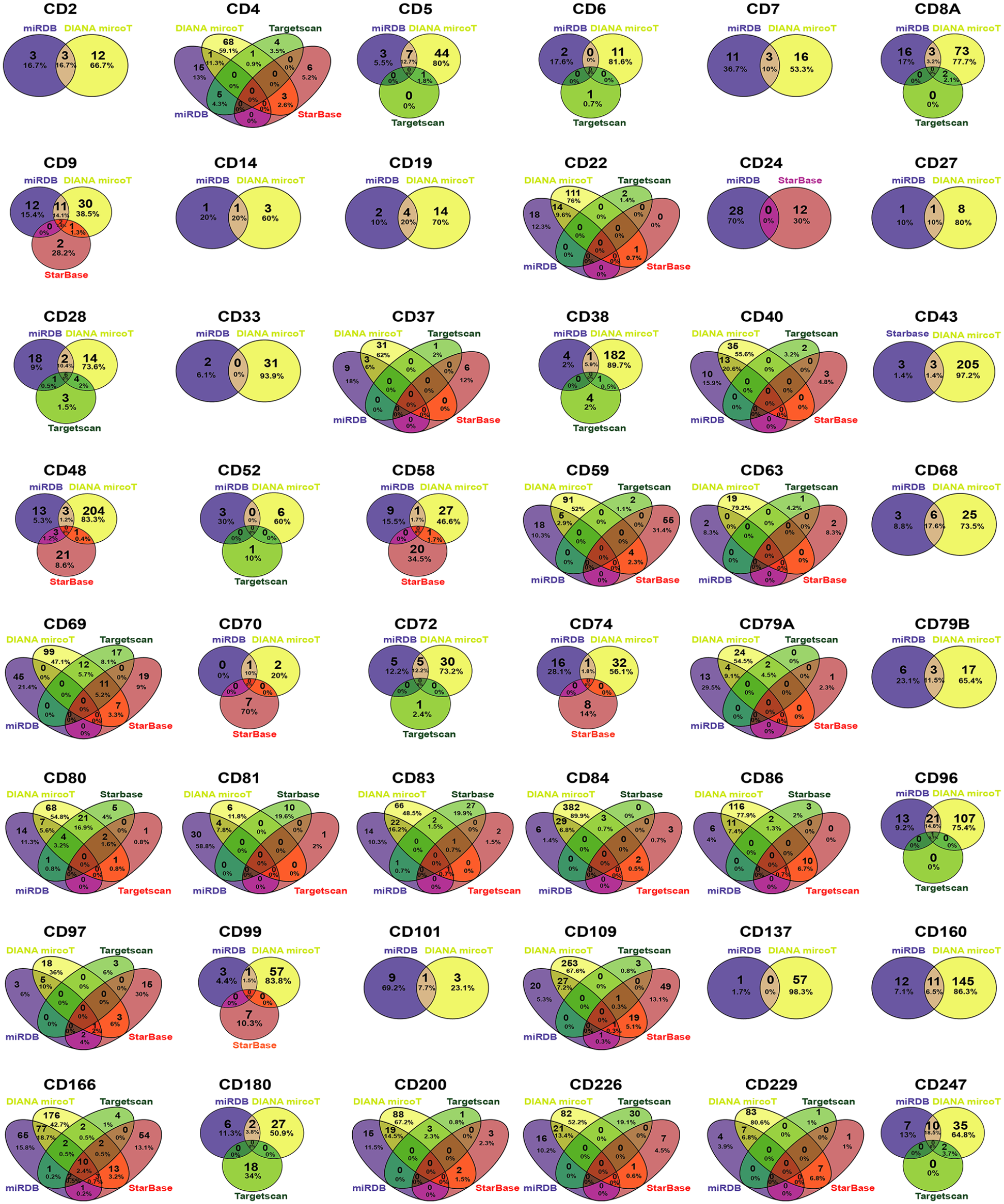
Comparisons of CD-associated miRNAs data collected from different miRNA target predicted tools. Using miRNA datasets obtained from Targetscan, Starbase, DIANA microT, and miRDB, Venny software 2.1.0 was used to find miRNAs that predicted in at least two different datasets.
Negative correlation of miRNA with CD molecules
In order to show accurate correlation between CD molecules and miRNAs, data of patients with GI cancer in TCGA were accessed. The correlation between CD molecules and miRNAs in GI cancers was screened (Figure 3 and Supplementary Figure S1). miRNAs that negatively associated with their corresponding CD molecules in one or two kinds of GI cancers were identified, as displayed in Figure 3. Moreover, we also list these potential CD molecule-miRNA pairs in Table 1. Interestingly, we found that CD4 was negatively correlated with miR-6808, CD81 with miR-559, CD86 with miR-200a, CD96 with miR-6808, and CD160 with miR-1236 in five types of GI cancer. For example, while CD28 was negatively correlated with miR-6838 in rectal cancer, CD86 was negatively correlated with miR-5558 in stomach cancer. Moreover, CD molecules such as CD166 and CD200 were negatively associated with different miRNAs in different cancers. While CD166 was negatively correlated with miR-142, miR-148a, and miR-429 in esophageal, colon, and pancreatic cancer, respectively, CD200 was negatively associated with miR-6884 and miR 6753 in stomach pancreatic cancer, respectively.
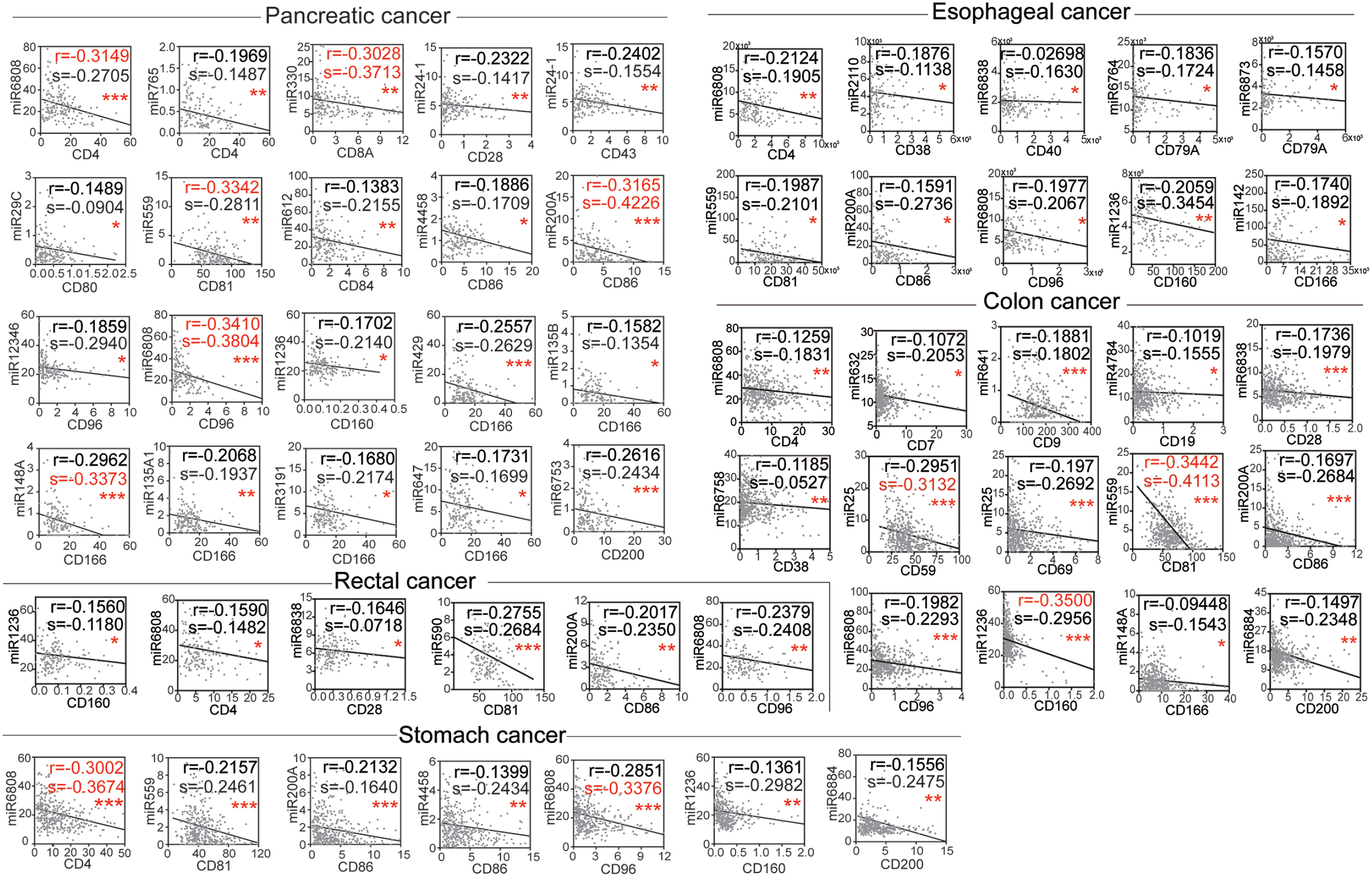
Negative correlations between CD molecules and different miRNAs in GI cancers. Data of GI cancer patients including pancreatic cancer (n=177), stomach cancer (n=375), rectal cancer (n=166), colon cancer (n=478) and esophageal cancer (n=161) in TCGA were downloaded. Prism correlation analysis was used to show the correlation between CD molecules and miRNA. The correlation was verified by the Pearson test and Spearman test. Significant negative correlation (* P<0.05, ** P<0.1, *** P<0.001.) analyzed by the Pearson test (r < −0.3) were highlighted in red.
Negative correlations between certain miRNAs and CD molecules in different kinds of GI cancer.

CD4 was negatively correlated with miR-6808, CD81 with miR-559, CD86 with miR-200a, CD96 with miR-6808, and CD160 with miR-1236 in the five types of GI cancer. Thus, we randomly selected three groups of miRNAs and CD molecules to verify the experiment.
GI: gastrointestinal; miRNA: microRNA.
miRNAs and overall survival of GI cancer patients
We further investigated the prognostic role of these CD molecule-related miRNAs in GI cancer patients based on TCGA datasets. The effect of these selected miRNAs on the survival rate of GI cancer patients is shown in Figure 4. High levels of miR-4458, miR-148a, miR-429, miR-135a1, and miR-647 were all significantly associated with increased overall survival in patients with pancreatic cancer (P<0.05, log-rank test), which might be considered as a potential biomarker for pancreatic cancer prognosis. Patients with stomach cancer with high expression levels of miR-4458 and miR-200a have a preferable prognosis when compared with patients with low levels of miR-4458 and miR-200a. On the other hand, high expression of miR-148a was significantly associated with poor prognosis of patients with colon cancer, while the presence of high miR-6808 might predict decreased overall survival in patients with rectal cancer. miR-148a and miR-4458 exhibited different roles in different GI cancers, indicating the specified miRNA might be specific for one or two cancers. In summary, our study indicated that these CD molecule-associated miRNAs may play an important role in GI cancer prognosis. The bioinformatics analysis of TCGA data provided evidence to find novel prognostic factors for GI cancers.
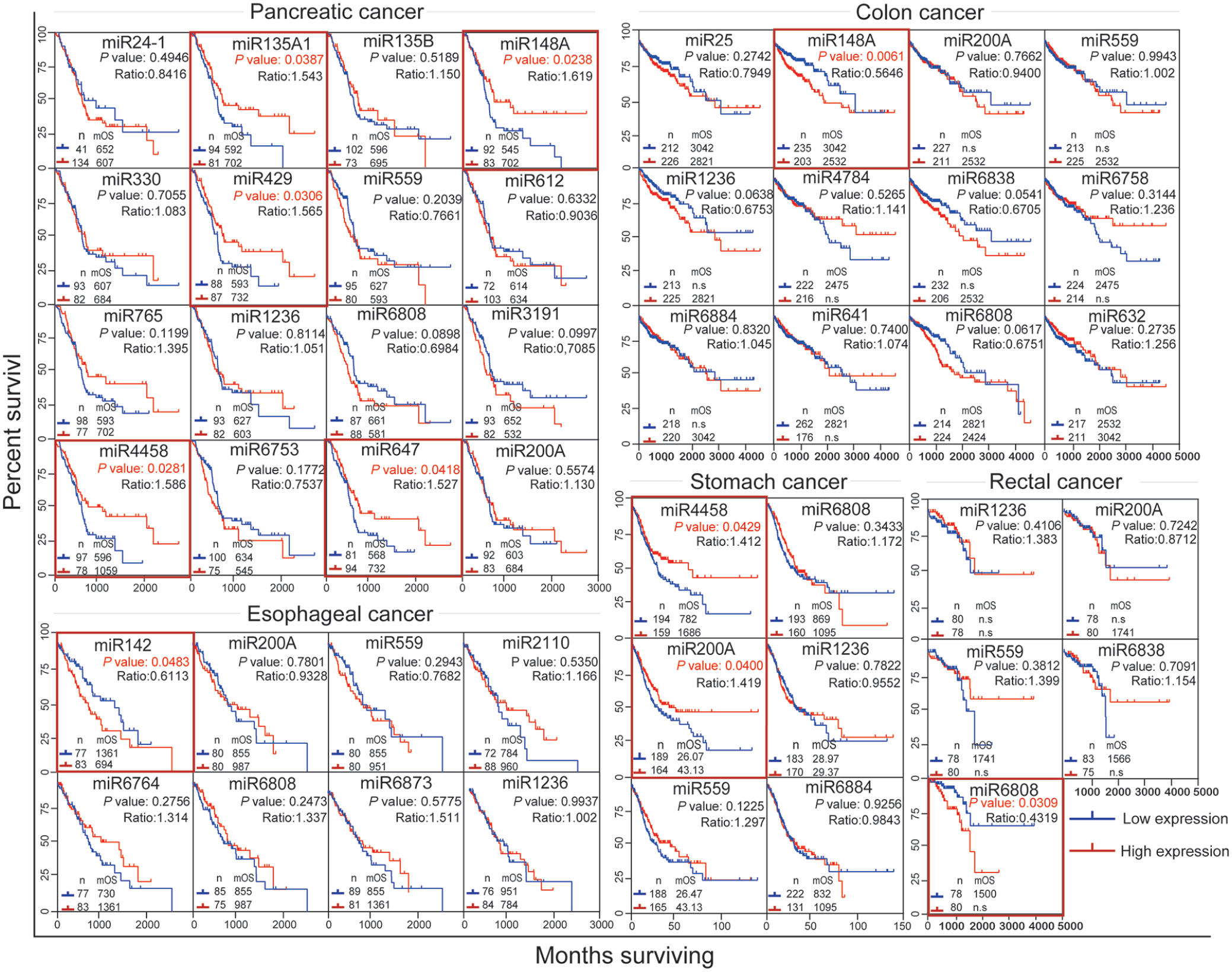
Prognostic role of CD molecules-related miRNAs in GI cancers. We analyzed the correlation of CD-associated miRNAs and overall survival in all kinds of GI cancers, including pancreatic cancer, esophageal cancer, rectal cancer, colon cancer, and stomach cancer. Statistical analysis was carried out using Kaplan–Meier analysis and significant associations were highlighted with red boxes.
Verification of CD-associated miRNAs
In order to verify the relationship between CD molecules and miRNAs, we overexpressed miRNAs in GI cancer cell lines, including two gastric cancer cell lines (TMK1 and BGC823), one colon cancer cell line (SW480), one pancreatic cancer cell line (Panc-1), and one esophageal cancer cell line (Eca109), by using miRNA mimics. Among the five cancer datasets, the negative correlation between miR-559 and CD81, miR-200a and CD86, and miR-1236 and CD160 in five types of cancers was examined. The results showed that increased expression of miR-559, miR-200a, and miR-1236 significantly suppressed the mRNA levels of CD81, CD86, and CD160, respectively, in all of these four cell lines (Figure 5), which were consistent with our previous bioinformatics analysis. These results provided experimental evidence for the identification of CD molecule-associated miRNAs and have laid a good foundation for exploring novel therapeutic targets for GI cancers.
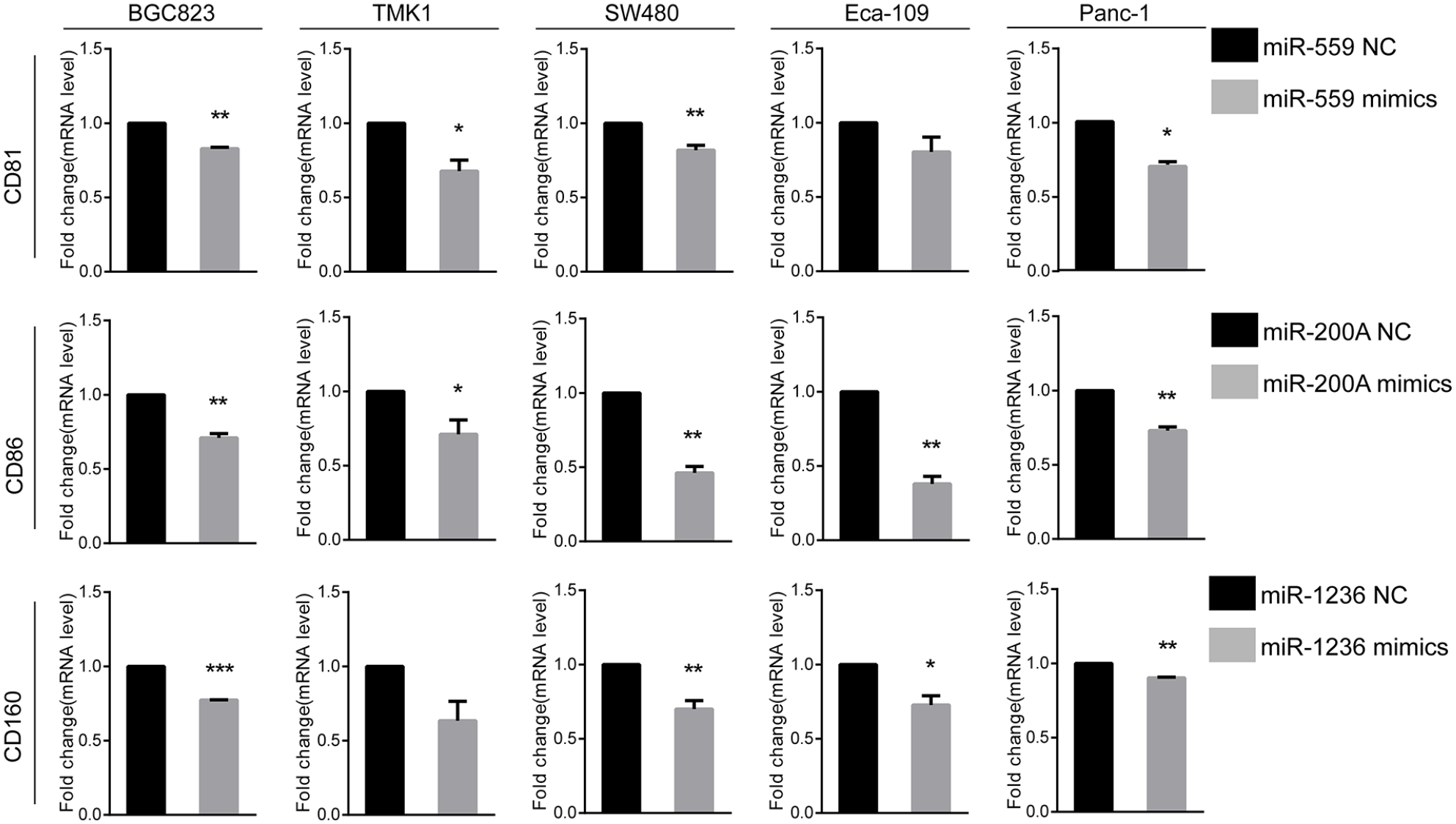
CD81, CD86, and CD160 levels were downregulated by miR-559, miR-200a, and miR-1236, respectively, in five kinds of GI cancer cell lines. The specific miRNA mimics were used to study the effects of miR-559, miR-200a, and miR-1236 on CD81, CD86, and CD160, respectively, in five cancer cell lines including gastric cancer cell lines BGC823 and TMK1, colon cell SW480, esophageal cancer cell Eca-109, and pancreatic cancer cell Panc-1. miRNA mimics and the control were transfected for 48 h in cells, and the effect on mRNA expression of CD molecules after miRNA down regulation was analyzed by qPCR.
Discussion
A large number of studies have shown that miRNAs are involved in the biological processes of cells, such as cell proliferation, differentiation, apoptosis, and migration, by targeting and down-regulating the translation of mRNA.26,27 Several human diseases are connected with the dysregulation of miRNAs, including different types of cancers. 27 CD molecules are closely related with immune cells including lymphocytes, macrophages, etc., and the associated miRNA network in the immune system were also widely studied. 28 It was reported that the immune infiltrates in cancer were associated with cancer genetic alteration, 29 suggesting that both the miRNA and CD expression pattern in TCGA data might predict immune-infiltrate levels. Our study focused on different expression profiles of CD molecule-associated miRNAs in different kinds of GI cancers, which might play important roles in immune response in cancer prevention and treatment.
To date, many studies have shown that miR-200a usually functions as a tumor suppressive gene in a wide range of cancer types including GI cancers, such as gastric cancer and colon cancer,30,31 while in esophageal cancer miR-200a supported cancer progression and invasion, 32 indicating the different role of miR-200a in different kinds of GI cancers. Recently, it was found that overexpression of miR-200a in osteosarcoma cells decreased proliferation and cytotoxicity of CD8 T cells and induced immunosuppression in osteosarcoma through increasing PD-L1 expression. 33 This also suggested that miR-200a might act as a prognostic marker for PD-L1-based cancer immunotherapy. The bioinformatics in the present study showed that miR-200a was negatively associated with CD86. An in vitro experiment also confirmed that miR-200a significantly decreased the levels of CD86 in all transfected GI cancer cell lines. CD86 is a costimulatory molecule expressed on antigen-presenting cells that is responsible for T-cell activation and survival. 34 miR-200a mediated downregulation of CD86 might also result in the deactivation of T cells, and have an immunosuppressive effect on GI cancers, indicating that miR-200a may be considered as a novel target for enhancing the efficacy of cancer immunotherapy. The immune function of miR-200a and CD86 in GI cancer needs further investigation.
CD160 is one of the inhibitory receptors on T cells, 35 which exhibits an immunosuppressive effect in the tumor microenvironment. Negative regulation of CD160 might activate the immune response of T cells, thereby increasing the efficacy of cancer immunotherapy. Our study showed that miR-1236 mimics reduced the levels of CD160 in gastric, colon, and esophageal cancer cell lines, suggesting a potential role of miR-1236 as a therapeutic target for GI cancer immunotherapy. A recent study has also shown that miR-1236-3p, by targeting metastasis-associated protein MTA2, had anti-metastatic effect for gastric cancer, 36 indicating the suppressive effect of the miR-1236 family members on GI cancers.
Our study also showed that CD81 significantly decreased by miR-559 in GI cancer cell lines. Previous study showed that CD81 could promote cancer progression through modulating immune suppressive function of regulatory T cells and myeloid-derived suppressor cells (MDSC) in myeloid tumor model. 37 miR-559 and its association with CD81 in GI cancer cell lines are reported for the first time in our study. Regarding the role of CD81 in immune modulation MDSC, it is anticipated that miR-559-induced CD81 deficiency might be associated with enhanced immune response, thereby exhibiting a tumor suppressive effect in GI cancers. Further research is recommended.
As expected, CD molecules and miRNA expression were closely related in all five GI cancer cell lines. This study suggests that miR-200a, miR-1236, and miR-559 might play an important role in immune modulation through targeting CD86, CD160, and CD81, respectively, and the association between miRNA and CD molecules open novel perspectives for exploring potential immunotherapeutic and prognostic biomarkers for GI cancers.
Conclusions
This study has identified miRNAs that might negatively control CD molecules in GI cancers. Although a few studies have suggested that GI cancers were inhibited by CD-associated miRNAs, direct involvement of CD molecules regulated by miRNA in GI cancers have not been studied until now. In summary, our findings suggested that miR-200, miR-1236, and miR-559 were closely associated with CD86, CD160, and CD81 expressions, respectively. These results advocate novel miRNA/CD molecule-based immunotherapeutic strategies to treat patients with GI cancer, which makes them pathogenic factors of disease and potential immunotherapeutic targets for malignancies in the GI tract.
Supplemental Material
sj-pdf-1-jbm-10.1177_17246008211005473 – Supplemental material for Identification of cluster of differentiation molecule-associated microRNAs as potential therapeutic targets for gastrointestinal cancer immunotherapy
Supplemental material, sj-pdf-1-jbm-10.1177_17246008211005473 for Identification of cluster of differentiation molecule-associated microRNAs as potential therapeutic targets for gastrointestinal cancer immunotherapy by Hanyu Zhang, Mingxing Li, Parham Jabbarzadeh Kaboli, Huijiao Ji, Fukuan Du, Xu Wu, Yueshui Zhao, Jing Shen, Lin Wan, Tao Yi, Qinglian Wen, Xiang Li, Chi Hin Cho, Jing Li and Zhangang Xiao in The International Journal of Biological Markers
Supplemental Material
sj-pdf-2-jbm-10.1177_17246008211005473 – Supplemental material for Identification of cluster of differentiation molecule-associated microRNAs as potential therapeutic targets for gastrointestinal cancer immunotherapy
Supplemental material, sj-pdf-2-jbm-10.1177_17246008211005473 for Identification of cluster of differentiation molecule-associated microRNAs as potential therapeutic targets for gastrointestinal cancer immunotherapy by Hanyu Zhang, Mingxing Li, Parham Jabbarzadeh Kaboli, Huijiao Ji, Fukuan Du, Xu Wu, Yueshui Zhao, Jing Shen, Lin Wan, Tao Yi, Qinglian Wen, Xiang Li, Chi Hin Cho, Jing Li and Zhangang Xiao in The International Journal of Biological Markers
Supplemental Material
sj-tif-3-jbm-10.1177_17246008211005473 – Supplemental material for Identification of cluster of differentiation molecule-associated microRNAs as potential therapeutic targets for gastrointestinal cancer immunotherapy
Supplemental material, sj-tif-3-jbm-10.1177_17246008211005473 for Identification of cluster of differentiation molecule-associated microRNAs as potential therapeutic targets for gastrointestinal cancer immunotherapy by Hanyu Zhang, Mingxing Li, Parham Jabbarzadeh Kaboli, Huijiao Ji, Fukuan Du, Xu Wu, Yueshui Zhao, Jing Shen, Lin Wan, Tao Yi, Qinglian Wen, Xiang Li, Chi Hin Cho, Jing Li and Zhangang Xiao in The International Journal of Biological Markers
Supplemental Material
sj-tif-4-jbm-10.1177_17246008211005473 – Supplemental material for Identification of cluster of differentiation molecule-associated microRNAs as potential therapeutic targets for gastrointestinal cancer immunotherapy
Supplemental material, sj-tif-4-jbm-10.1177_17246008211005473 for Identification of cluster of differentiation molecule-associated microRNAs as potential therapeutic targets for gastrointestinal cancer immunotherapy by Hanyu Zhang, Mingxing Li, Parham Jabbarzadeh Kaboli, Huijiao Ji, Fukuan Du, Xu Wu, Yueshui Zhao, Jing Shen, Lin Wan, Tao Yi, Qinglian Wen, Xiang Li, Chi Hin Cho, Jing Li and Zhangang Xiao in The International Journal of Biological Markers
Footnotes
Acknowledgements
Not applicable
Author contributions
ML, JL, and ZX designed and conceived the study. PJK, HJ, and FD conducted the bioinformatics analysis. HZ and JS performed the experiments. YZ and QW analyzed the data. LW, TY, XL, and CHC provided the resources and experiments for the study. ML and XW wrote the first draft. HZ and ML reviewed and edited the manuscript, and contributed equally to this article. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (Grant nos. 81602166, 81672444, 81770562) and the Joint Funds of the Southwest Medical University & Luzhou (2017LZXNYD-Z05 and 2017LZXNYD-J09).
Ethics approval and consent to participate
Not applicable
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
