Abstract
Background:
Circulating miRNAs have been proved to be promising biomarkers for disease detection in recent years. The present study aimed at exploring available serum miRNA biomarkers for the detection of colorectal cancer.
Methods:
A three-phase study was performed to select and validate candidate miRNAs with significant dysregulation in colorectal cancer using quantitative reverse transcription-polymerase chain reaction. This study recruited 137 colorectal cancer patients and 145 healthy controls. The diagnostic values of miRNAs were evaluated by receiver operating characteristic analysis. Bioinformatics analyses were utilized to predict target genes of miRNAs, and to conduct functional annotation and enrichment.
Results:
miR-30e-3p, miR-31-5p, miR-34b-3p and miR-146a-5p, miR-148a-3p and miR-192-5p were significantly dysregulated in colorectal cancer serum when compared with healthy controls. The panel composed of miR-30e-3p, miR-146a-5p, and miR-148a-3p exhibited strong diagnostic ability. The area under the receiver operating characteristic curve of the three-miRNA panel was 0.883, with a sensitivity of 0.800 and specificity of 0.787.
Conclusion:
The present study identified a three-miRNA panel in serum with a strong diagnostic ability of colorectal cancer, which may be able to serve as a novel noninvasive biomarker for colorectal cancer detection.
Introduction
Colorectal cancer (CRC) is the third most common cancer worldwide and the second leading cause of cancer-related mortality, with approximately 1.85 million new cases and 880,000 deaths each year. 1 Early diagnosis is the key to a better outcome in CRC patients. The 5-year survival of CRC patients varied depending on the tumor stage at the time of diagnosis. Compared with those who were diagnosed at an early stage, the 5-year survival rate of CRC patients diagnosed at an advanced stage decreased from 90.3% to 12.5%. 2 Currently, colonoscopy is recognized as the gold standard for the detection of CRC. However, its invasiveness, time-consuming nature, high cost and high operator variability have limited its application. 3 The fecal occult blood test (FOBT) is one of the most widely used screening methods for CRC. Although FOBT exhibits high specificity in CRC diagnosis, the sensitivity of it is unsatisfactory. 4 In this situation, identifying novel clinically available biomarkers for CRC detection is of great significance.
microRNA (miRNA) is a type of small non-coding RNA consisting of 19–25 nucleotides, which can regulate the translation of target genes post-transcriptionally. Studies have proved their involvement in CRC cell functions, such as proliferation, migration, invasion, and drug resistance.5-8 As miRNAs are stable in blood and are resistant to exogenous RNase activity and repeated freeze–thaw cycles, 9 circulating miRNAs are potential ideal markers for tumor detection. An increasing number of studies have demonstrated that circulating miRNAs are able to act as diagnostic biomarkers in a variety of cancers.10,11 In fact, several previous studies have identified some circulating miRNA biomarkers for the diagnosis of CRC12-14; however, more research and evidence are required before their clinical use.
This study consisted of three phases: screening, training, and validation. In the screening phase, 25 CRC and 25 healthy control (HC) samples were pooled into 5 CRC samples and 5 HC samples to test the miRNA expression profile. miRNAs selected in the first phase were later evaluated using 32 CRC and 32 HC samples. miRNAs found to be significantly dysregulated in the training phase were further evaluated using 80 CRC and 88 HC samples in the validation phase. Finally, we identified a three-miRNA panel with great diagnostic capability. Furthermore, we conducted a target gene prediction and enrichment analysis of the three miRNAs. In summary, our study identified a three-miRNA panel in serum with the potential to serve as a biomarker for CRC detection.
Materials and methods
Patients and samples
We enrolled 137 patients at the Peking University Shenzhen Hospital during 2017 and 2018 who underwent surgery and were histopathologically confirmed as having CRC. A total of 145 HCs were selected from those who received health examinations in the physical examination center of Peking University Shenzhen Hospital. The inclusion criteria of HCs were: (a) asymptomatic and apparently healthy; and (b) without a previous history of cancer or other benign diseases. The age of the CRC group and the HC group were matched. Our study had gained the permission from the Ethics Committee of Peking University Shenzhen Hospital, and every participant had signed a written informed consent.
Study design
dbDEMC 2.0 is a database containing expression profiles of miRNAs in human cancers based on the Gene Expression Omnibus and The Cancer Genome Atlas database. 15 From this database, we retrieved miRNAs that were dysregulated in the tissue or blood of CRC. Based on the retrieved results and our previous experiment results, we finally selected 40 miRNAs as candidate biomarkers for CRC detection. Later, a three-phase study was carried out to identify novel biomarkers for CRC detection. In the screening phase, 25 serum samples from CRC patients and 25 serum samples from HCs were chosen at random and pooled as 5 CRC samples and 5 HC samples (i.e. 5 samples were pooled as 1 sample). The expression profiles of the 40 candidate miRNAs were tested, and differentially expressed miRNAs were selected for further study. In the training phase, miRNAs selected from the first phase were tested using 32 CRC samples and 32 HC samples. Finally, in the validation phase, another 80 CRC samples and 88 HC samples were utilized to confirm the differential expression of miRNAs. All the procedures above were carried out using quantitative reverse transcription-polymerase chain reaction (qRT-PCR). The diagnostic ability of miRNAs was assessed by receiver operating characteristic (ROC) analysis. miRNA-combined panels were further constructed for the enhancement of diagnostic capability. We performed bioinformatic analyses of the miRNA panel with the aim of understanding its biological mechanism in tumorigenesis and development.
Serum collection and RNA extraction
Before accepting any treatment, 10 mL peripheral serum of each participant was acquired. All serum samples were centrifuged at 1000 g for 10 minutes and 15,000 g for 5 minutes at 4°C within 2 hours from collection. To redress the deviation in the process of extraction and purification, we spiked 2 µL of synthetic Caenorhabditis elegans miR-39 (cel-miR-39) (10 nM, RiboBio, Guangzhou, China) into each serum sample in preparation. According to the manufacturer’s instructions, we used TRIzol LS isolation kit (Thermo Fisher Scientific, Waltham, MA, USA) to extract total RNA from serum. Total RNA were lysed with 30 µL RNase-free water and then stored at −80°C for further experiments. The NanoDrop 2000 spectrophotometer (NanoDrop, Wilmington, DE, USA) was applied to test the concentration and purity of RNA.
qRT-PCR
We conducted the amplification of miRNAs using the specific reverse transcription primers from Bulge-Loop miRNA qRT-PCR Primer Set (RiboBio, Guangzhou, China). The real-time polymerase chain reaction was performed using SYBR Green qPCR kit (SYBR Pre-mix Ex Taq II, TaKaRa) in 384-well plates at 95°C for 30 s, followed by 35 cycles of 95°C for 10 s, 60°C for 20 s, and then 70°C for 10 s on the LightCycler 480 Real-Time PCR System (Roche Diagnostics, Mannheim, Germany). The specificity of the PCR products was evaluated using melting curve analysis. All reactions had been performed in triplicate or more. The relative expression of target miRNAs was calculated using the 2 −△△Cq method 16 and normalized with the spiked-in control cel-miR-39.
Bioinformatics analysis
To better understand the underlying mechanism of the candidate miRNAs in CRC, we used miRWALK 3.0 (http://mirwalk.umm.uni-heidelberg.de)—an online prediction tool—to perform miRNA-mRNA prediction. The predicted target genes were further uploaded to the Enrichr database (https://amp.pharm.mssm.edu/Enrichr/) for functional annotation and enrichment analysis, including Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis.
Statistical analysis
Differences of miRNA expression between CRC patients and HCs was analyzed by the Students’ T-test or Mann–Whitney test. Comparisons of miRNA expression between two groups with different tumor characteristics were conducted using the Students’ T-test or the Mann–Whitney test. The diagnostic value of miRNAs was evaluated by ROC curves and the area under the ROC curve (AUC). The optimal sensitivity and specificity were determined using the Youden index (Youden index = Sensitivity + Specificity − 1). The diagnostic panels were built by the stepwise logistic regression model. All statistical analyses were performed using SPSS software (SPSS 22.0 Inc., Chicago, IL) and GraphPad Prism 7 (GraphPadSoftware Inc., LaJolla, CA). P< 0.05 was considered as statistical significance.
Result
Characteristics of study samples
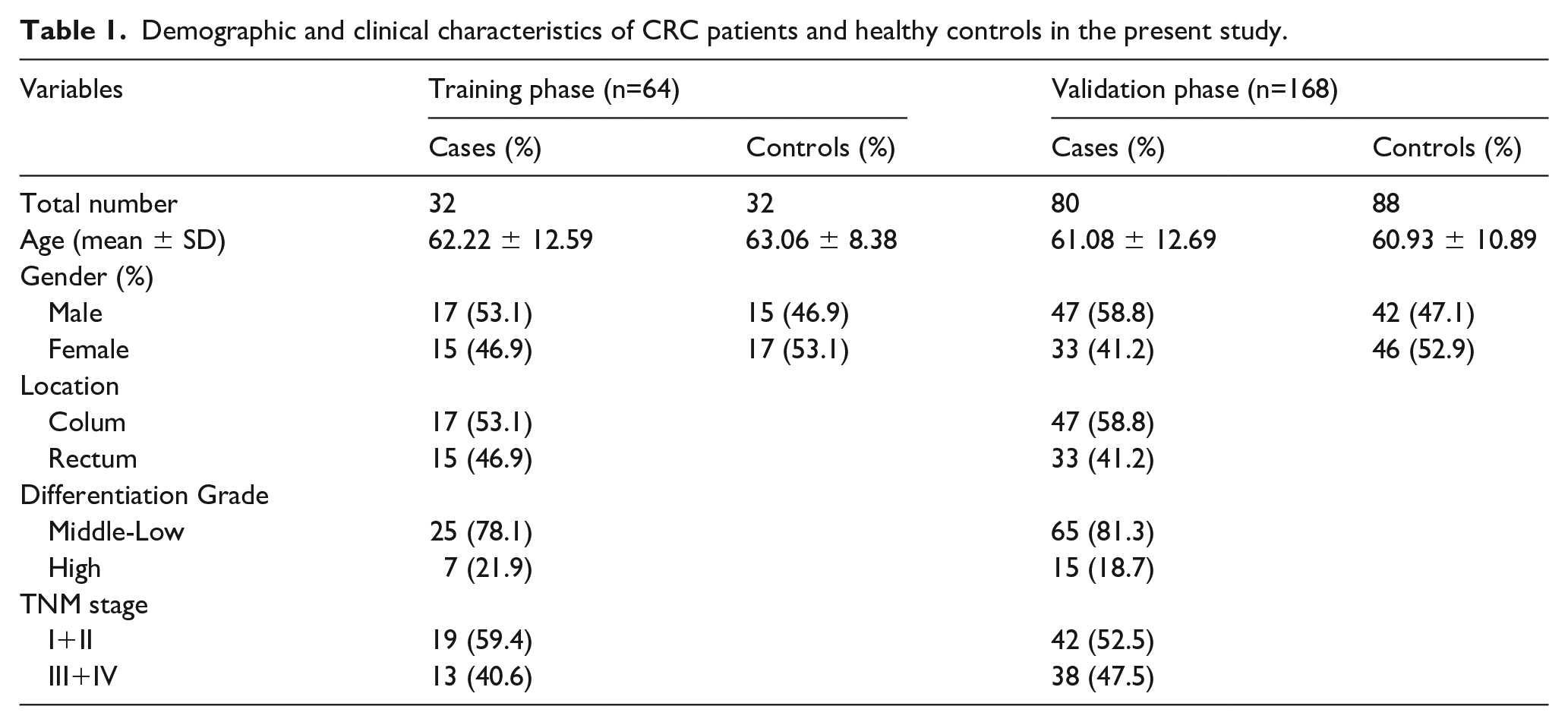
Our study contained 137 CRC and 145 HC samples. The demographics and clinical features of participants are listed in Table 1. The samples were randomly divided into three parts for three study phases: screening, training, and validation. The gender and age between the CRC and HC groups showed no significant difference in this study (P> 0.05).
Demographic and clinical characteristics of CRC patients and healthy controls in the present study.
Candidate miRNA screening
In the screening phase, five CRC pools and five HCs pools were used to test the expression level of 40 candidate miRNAs. A total of 11 significantly altered miRNAs were identified with the criteria of Fold change >2 or <−2. Among them, miR-17-5p, miR-30e-3p, miR-31-5p, miR-34b-3p, and miR-146a-5p were upregulated. miR-142-5p, miR-148a-3p, miR-181a-5p, miR-192-5p, miR-215-5p, and miR-409-3p were downregulated (Figure 1). We selected these 11 miRNAs for further validation.
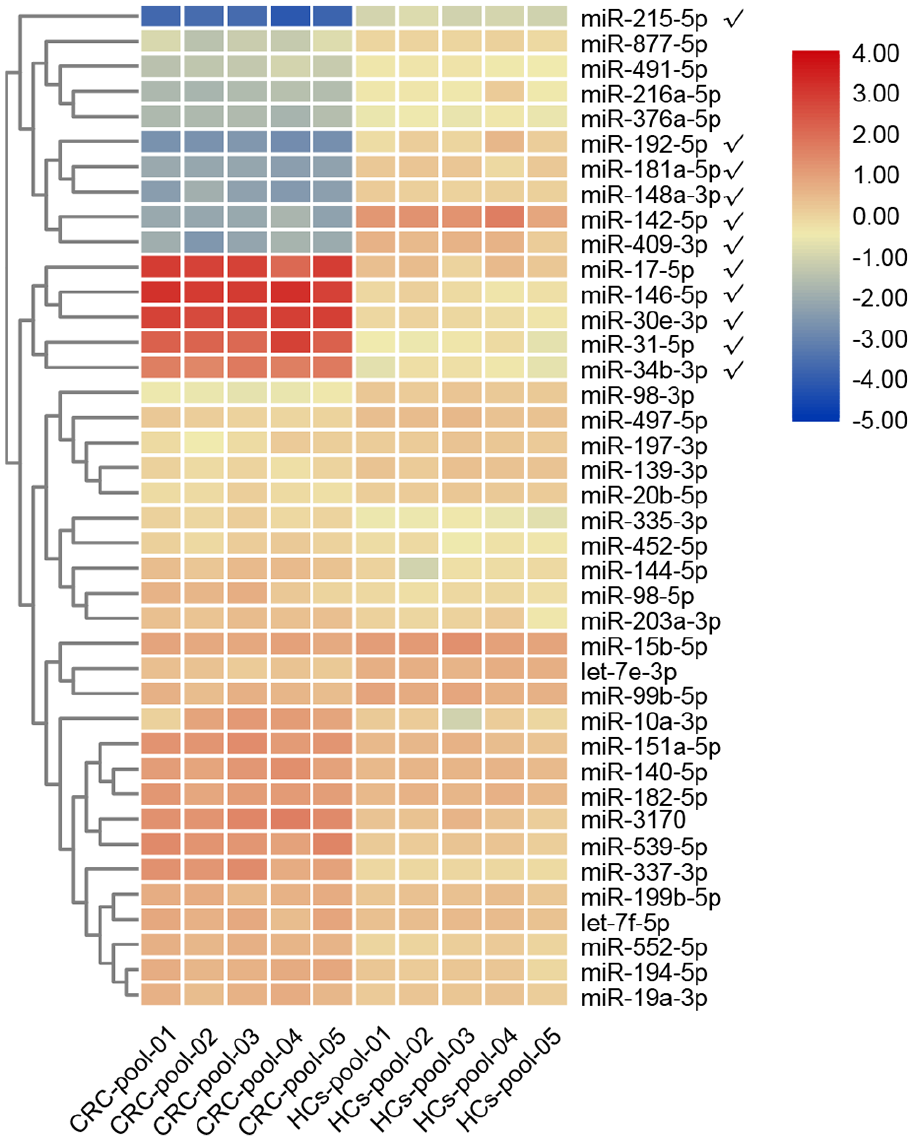
Heatmap of the expression profile of 40 candidate miRNAs in the screening phase. Eleven miRNAs marked with ticks were differentially expressed between CRC pools and HCs pools and were selected for further study (Fold change >2 or <−2).
Candidate miRNA training and validation
In the training phase, 32 CRC and 32 HC samples were used to confirm the differential expression of 11 candidate miRNAs. As a result, 6 miRNAs were verified to be significantly dysregulated (Figure 2). miR-30e-3p (P < 0.0001), miR-31-5p (P = 0.0054), miR-34b-3p (P = 0.0026), and miR-146a-5p (P = 0.0004) were upregulated. miR-148a-3p (P = 0.0266) and miR-192-5p (P = 0.0014) were downregulated. miR-17-5p (P = 0.0608), miR-142-5p (P = 0.1876), miR-181a-5p (P = 0.0996), miR-215-5p (P = 0.1295), and miR-409-3p (P = 0.1553) showed no significant differential expression.
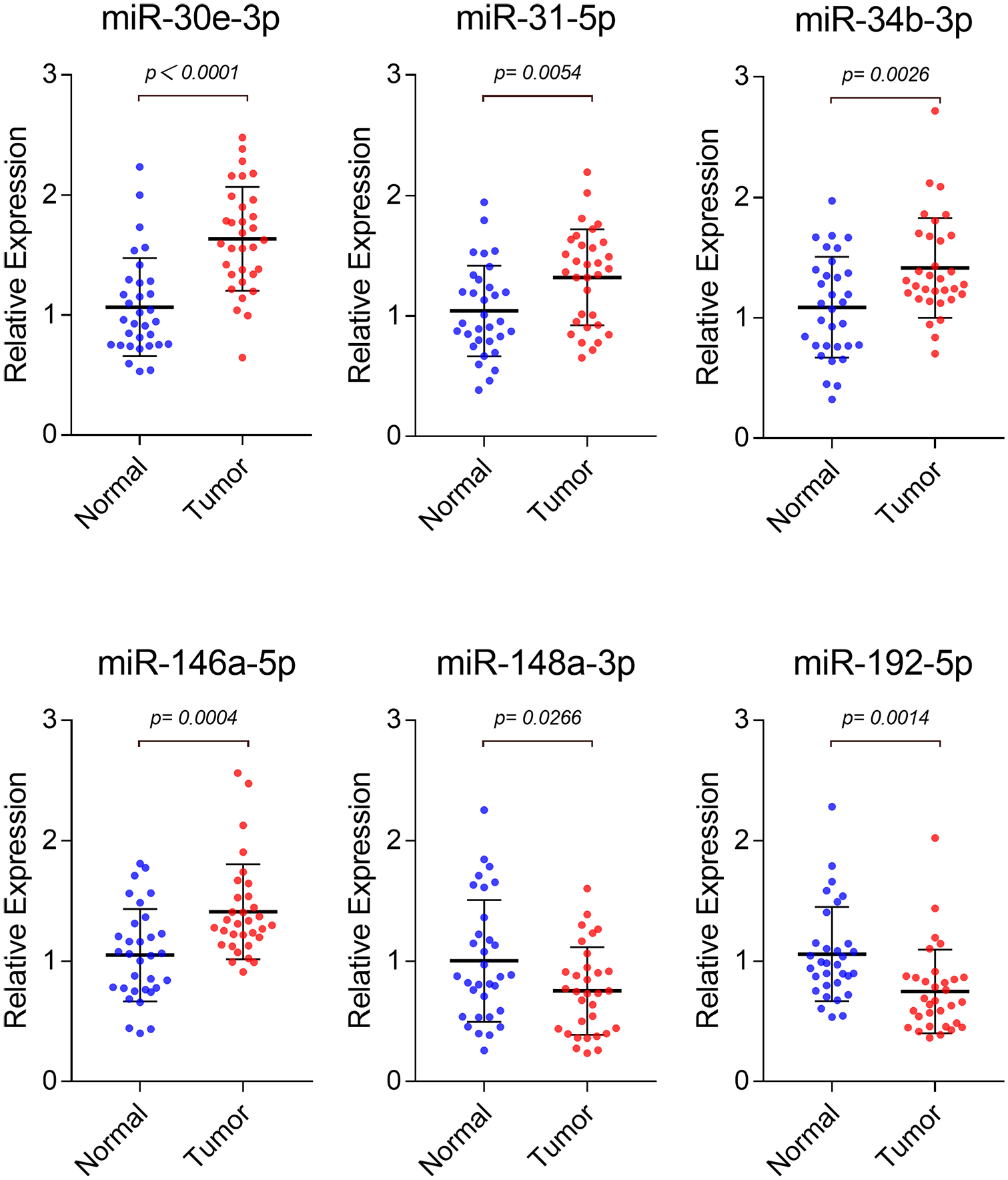
Expression levels of candidate miRNAs in 32 CRC patients compared with 32 HCs in the training phase. Six miRNAs were found to be significantly dysregulated in the training phase (P < 0.05). Horizontal line: mean with 95% CI.
Larger sample verification with 80 CRC and 88 HC samples was later conducted for further confirmation. The results were consistent with the training phase. Expression of miR-30e-3p (P = 0.0019), miR-31-5p, miR-34b-3p (P < 0.0001), and miR-146a-5p (P < 0.0001) were significantly increased while miR-148a-3p and miR-192-5p were decreased when compared with HCs (Figure 3).
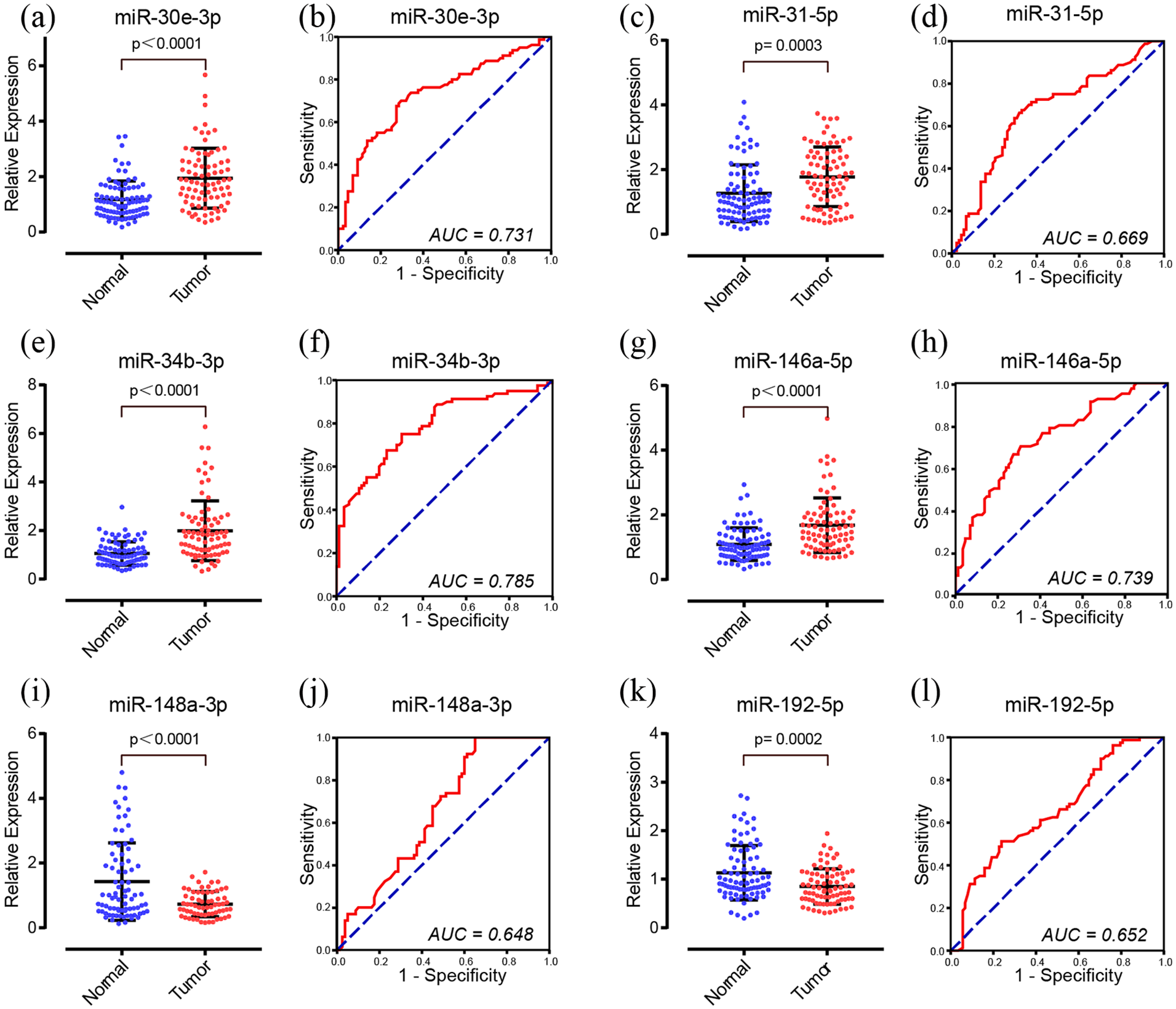
Expression levels and ROC curves of the six candidate miRNAs in 80 CRC patients compared with 88 HCs in the validation phase. (a) and (b) miR-30e-3p; (c) and (d) miR-31-5p; (e) and (f) miR-34b-3p; (g) and (h) miR-146a-5p; (i) and (j) miR-148a-3p; (k) and (l) miR-192-5p.
Diagnostic value of serum miRNA for CRC
We applied ROC analysis to evaluate the diagnostic capability of the six miRNAs. The AUC of the six miRNAs (miR-30e-3p, miR-31-5p, miR-34b-3p, miR-146a-5p, miR-148a-3p, miR-192-5p) were 0.731 (95% confidence interval (CI) 0.654, 0.808), 0.669 (95% CI 0.586, 0.751), 0.785 (95% CI 0.715, 0.855), 0.739 (95% CI 0.665, 0.813), 0.648 (95% CI 0.559, 0.737), and 0.652 (95% CI 0.569, 0.735), respectively (Figure 3).
Diagnostic value of serum miRNA panel for CRC detection
We combined the six proved miRNAs into a panel by stepwise logistic regression model to enhance the diagnostic capability. The model was calculated by the equation:
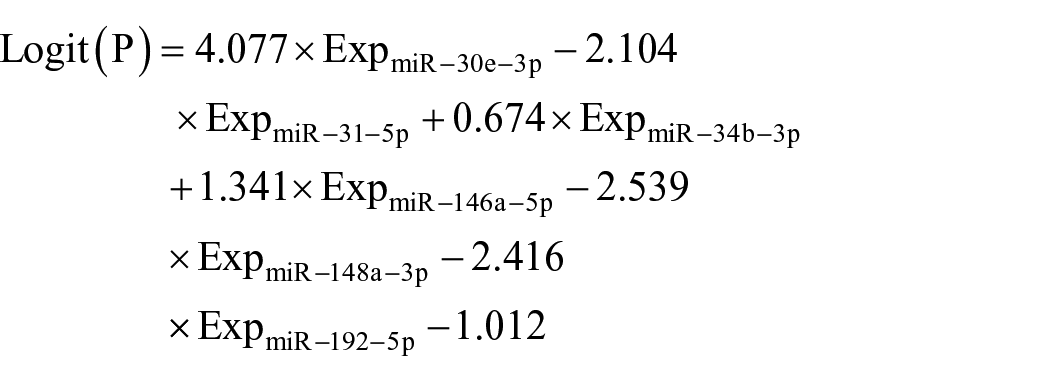
As a result, the six-miRNA panel showed excellent capability to distinguish CRC from HC patients with an AUC of 0.932 in ROC analysis (95% CI 0.895, 0.970; sensitivity, 0.846; specificity, 0.861) (Figure 4(a)).
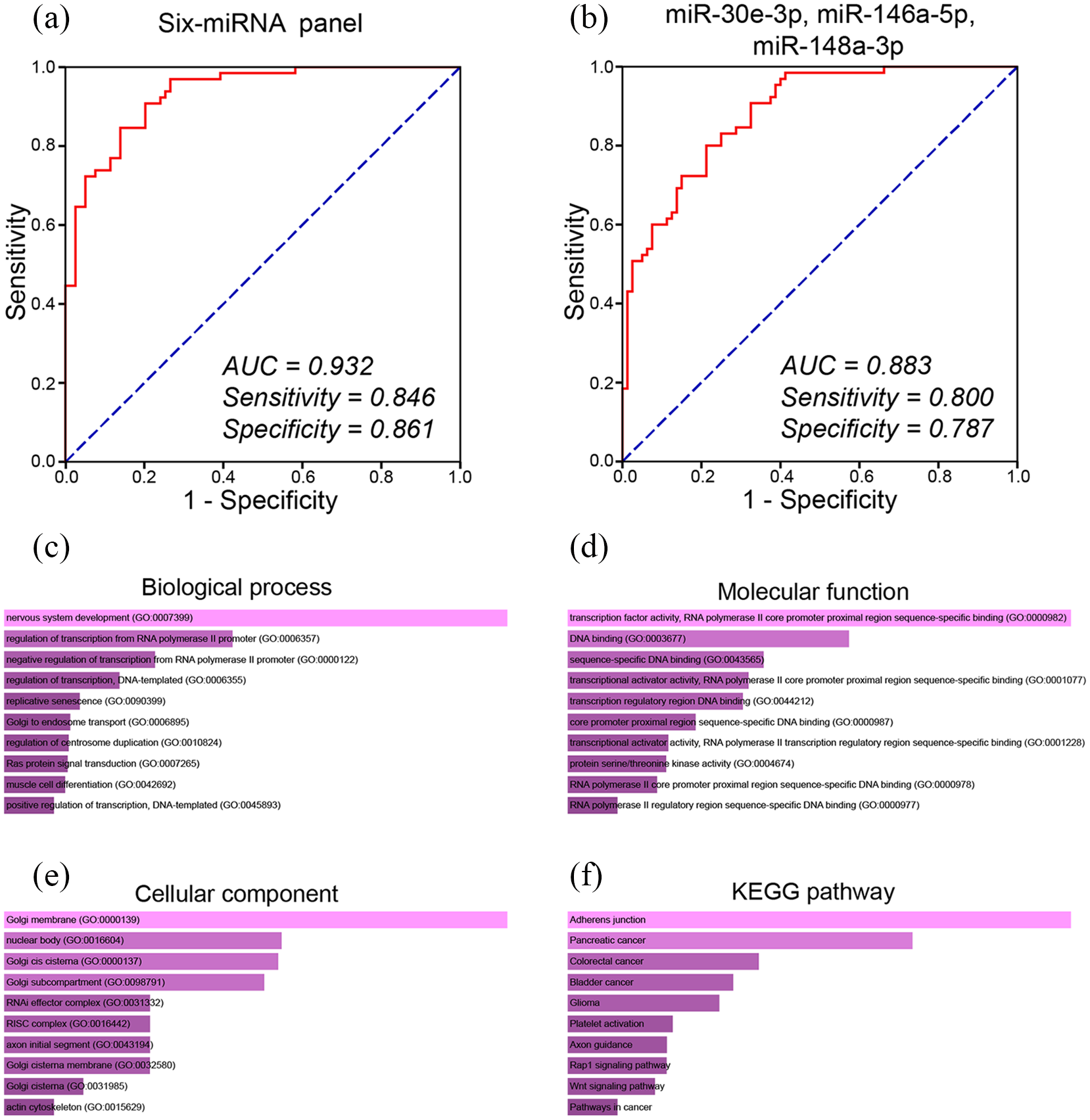
ROC curves of miRNA panels for diagnosis of CRC. (a) miRNA panel composed of miR-30e-3p, miR-31-5p, miR-34b-3p, miR-146a-5p, miR-148a-3p, and miR-192-5p. (b) The refined three-miRNA panel composed of miR-30e-3p, miR-146a-5p, and miR-148a-3p. The optimal sensitivity and specificity were determined using the Youden index. Functional enrichment analysis of the predicted target genes of miR-30e-3p, miR-146a-5p, and miR-148a-3p. (c) Biological process (BP) analysis. (d) Molecular function (MF) analysis. (e) Cellular component (CC) analysis. (f) Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis.
However, if too many biomarkers appear in the panel, the detection process becomes complicated. Therefore, it is important to balance the relationship between the number of biomarkers in the combination and the diagnostic ability of the panel. Our strategy is to remove miRNAs that contribute little to the combined diagnostic capability from the six-miRNA panel. We simplified the panel by conducting multiple logistic regression analysis. The result showed that a three-miRNA panel composed of miR-30e-3p, miR-146a-5p, and miR-148a-3p achieved a similar performance compared with the six-miRNA panel (Figure 4(b)). The model was calculated by the equation:
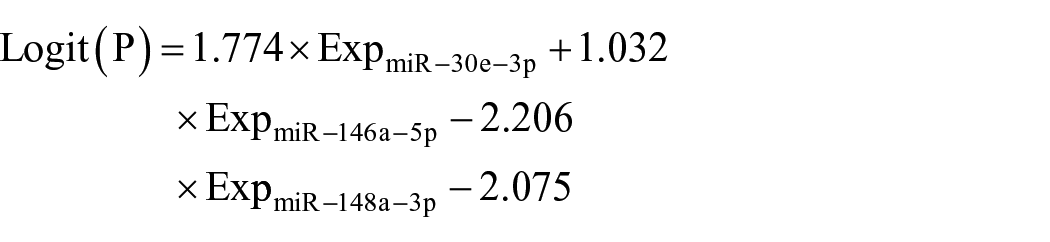
The AUC of the three-miRNA panel was 0.883 (95% CI 0.831, 0.35; sensitivity 0.800; specificity 0.787). These results had revealed the potential of the three-miRNA panel to be a clinically available biomarker for CRC detection.
Relationship between candidate miRNAs and tumor characteristics
To explore more clinical value of these three miRNAs, we evaluated the relationship between the miRNA expression level and several tumor characteristics, including tumor stage, grade, and location. The results showed no significant difference between the three miRNAs and the tumor stage and location (P> 0.05). The expression level of miR-30e-3p and miR-34b-3p seemed to be higher in the high-grade group than the middle-low group (P< 0.05). However, most of our CRC samples were in the middle-low-grade group, and the sample size of the high-grade group was relatively small. More high-grade samples are needed to further prove the correctness of this result.
Target gene prediction and enrichment analysis of the three-miRNA panel
Using the miRWALK website, we predicted the target genes of miR-30e-3p, miR-146a-5p, and miR-148a-3p. The results contained 1806 predicted target genes. Later, we uploaded these target genes on the Enrichr database for enrichment analysis. GO analysis contained biological process (BP), molecular function (MF), and cellular component (CC). The top 10 enriched GO terms of BP, MF, and CC are shown in Figure 4(c) to (e), respectively. The most enriched BP terms were nervous system development (GO:0007399), regulation of transcription from RNA polymerase II promoter (GO:0006357), negative regulation of transcription from RNA polymerase II promoter (GO:0000122), regulation of transcription, DNA-templated (GO:0006355), and replicative senescence (GO:0090399). The most significantly enriched MF terms were transcription factor activity, RNA polymerase II core promoter proximal region sequence-specific binding (GO:0000982), DNA binding (GO:0003677), sequence-specific DNA binding (GO:0043565), transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding (GO:0001077), and transcription regulatory region DNA binding (GO:0044212). The result of CC revealed that the three-miRNA panel was significantly enriched in Golgi membrane (GO:0000139), nuclear body (GO:0016604), Golgi cis cisterna (GO:0000137), Golgi subcompartment (GO:0098791), and RISC complex (GO:0016442). As for KEGG analysis, the top 10 most enriched terms are shown in Figure 4(f), among which the colorectal cancer pathway was one of the most significantly enriched terms.
Discussion
At present, colonoscopy is still the gold standard for CRC detection. However, limited by its invasiveness, high cost, discomfort, and high operator variability, it is unsuitable to be a widely used detection measure. 17 Ideal biomarkers should conform with criteria of simplicity, high reliability, and inexpensiveness. Recently, increasing evidence has demonstrated that the availability of circulating miRNAs are diagnostic biomarkers for malignant tumors with their unique superiority.18,19
In the present study, a three-phase study was carried out to identify potential miRNA biomarkers for CRC detection. In the screening phase, we identified 11 miRNAs from 40 candidate miRNAs, which showed prominent dysregulation in CRC patients using serum sample pools. In the training phase, 6 candidate miRNAs were proved. Further confirmation with a large sample size in the validation phase demonstrated that the expression of miR-30e-3p, miR-31-5p, miR-34b-3p, and miR-146a-5p were significantly increased while miR-148a-3p and miR-192-5p were decreased in CRC serum samples. Usually, single miRNA as a diagnostic biomarker cannot reach strong diagnostic ability, sensitivity, and specificity at the same time while the combination of several miRNAs may be able to achieve this. The three-miRNA panel constructed by miR-30e-3p, miR-146a-5p, and miR-148a-3p showed an AUC of 0.883 in ROC analysis (95% CI 0.831, 0.935) with a sensitivity of 0.800 and a specificity of 0.787, revealing its strong ability for CRC detection. The results of the KEGG pathway analysis showed that their target genes were enriched in the pathways of colorectal cancer, the Rap1 signaling pathway, and the Wnt signaling pathway, revealing their potential role in CRC tumorigenesis and development. BP analysis revealed their involvement in the regulation of transcription from RNA polymerase II promoter, negative regulation of transcription from RNA polymerase II promoter, regulation of transcription, DNA-templated, and replicative senescence. MF analysis indicated that they were enriched in transcription factor activity, RNA polymerase II core promoter proximal region sequence-specific binding, DNA binding, sequence-specific DNA binding, transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding, and transcription regulatory region DNA binding. Golgi membrane, nuclear body, Golgi cis cisterna, and Golgi subcompartment were the most enriched cellular components. The results of GO and KEGG analyses have provided the direction for future mechanism research. The serum miRNA test is both convenient and low cost. This panel, along with colonoscopy, could likely become a powerful weapon for CRC detection. To gain a better understanding of these three miRNAs, a literature review was carried out.
Previous studies have shown the relationship between miR-30e-3p and several types of cancers. Plasma miR-30e-3p was downregulated in non-small-cell lung cancer patients and was associated with short disease-free survival. 20 Moreover, miR-30e-3p was identified as a diagnostic biomarker for non-small-cell lung cancer in the early stage. 21 In ovarian carcinoma, miR-30e-3p was significantly dysregulated and was correlated with prognosis. 22 The study of Balci et al. 23 revealed that miR-30e-3p could be one of the novel biomarkers for non-melanoma skin cancer. Additionally, Wang et al. 24 demonstrated that miR-30e-3p could inhibit the cell invasion and migration function of clear cell renal cell carcinoma by targeting Snail1. No study of miR-30e-3p in CRC had been reported before; therefore, our study has been the first to explore their relationship.
The role of miR-146a-5p in different kinds of cancers was complicated and was identified as a potential biomarker and therapeutic target in various cancers.25,26 For example, miR-146a-5p was proved to function as a tumor suppressor in oesophageal squamous cell carcinoma, breast cancer, prostate cancer, and pancreatic cancer. However, it was upregulated in squamous cell carcinoma of the oral cavity, 27 head and neck, 28 cervix, 29 and lung. 30 In oral squamous cell carcinoma, it was also upregulated and was involved in oncogenicity.27,31 The research of Lu et al. 32 confirmed that miR-146a-5p was an oncogene in CRC with promotion of migration and invasion functions in CRC cells via the carboxypeptidase M/src-FAK pathway, which provided evidence for our study results.
miR-148a-3p has been proved to have important roles in various tumors, including esophageal squamous cell carcinoma, cervical cancer, ovarian cancer, lung cancer, and more.33-36 Moreover, several studies have shown the potential of circulating miR-148a-3p to be a noninvasive biomarker in tumor diagnosis and surveillance. For instance, Abdollahi et al. 37 identified a panel containing miR-148a-3p, which could be a diagnostic biomarker of non-small cell lung cancer. The study of Bellissimo et al. 38 demonstrated that miR-148a-3p could act as a non-invasive biomarker to evaluate the therapeutic efficacy and the prognosis of thymic epithelial tumors. In DNA mismatch repair-deficient colorectal cancer, decreased miR-148a-3p contributes to the immunosuppressive tumor microenvironment by targeting PD-L1. 39 A previous study proved that the expression level of miR-148a-3p was higher in CRC tissues compared with normal tissues. 40 Our study was the first to evaluate the expression level of serum miR-148a-3p in CRC patients.
Some limitations of our study must be mentioned. First, the candidate miRNAs in this study was relatively small; there should be a more extensive screening phase. Second, the sample size of our study was relatively small and there was a lack of external validation. The small sample size in the screening phase and training phase may cause biases, resulting in the neglect of some other useful miRNAs. The present study was in the discovery phase. We need more serum samples of early stage CRC, benign gastrointestinal diseases, and other types of cancers, to further assess the potential role of the three-miRNA panel in CRC screening. Third, only the preoperative serum was tested and the postoperative comparison was deficient. Moreover, The exploration of the potential mechanism that the three-miRNA panel particapates in CRC was only based on bioinformatics analysis. Our future studies will continue to explore the exact biological effect of the three-miRNA panel in CRC and validate their clinical value.
In conclusion, our study has proved the significant dysregulation of serum miR-30e-3p, miR-146a-5p, and miR-148a-3p in CRC patients compared with HCs. The three-miRNA panel comprised of them has shown strong diagnostic ability with high sensitivity and specificity (AUC, 0.883; sensitivity, 0.800; specificity, 0.787). This panel has the potential to be a clinically available biomarker for CRC detection.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Basic Research Project of Peking University Shenzhen Hospital (JCYJ2017001, JCYJ2017004, JCYJ2017005, JCYJ2017006, JCYJ2017007, JCYJ2017012), Clinical Research Project of Peking University Shenzhen Hospital (LCYJ2017001), Science and Technology Development Fund Project of Shenzhen (no. JCYJ20180507183102747), and Clinical Research Project of Shenzhen Health Commission (no. SZLY2018023).
