Abstract
Background:
Aberrant expression of circular RNAs (circRNAs) is implicated in tumorigenesis and disease progression. However, the underlying molecular mechanisms and physiological functions of hsa_circ_0005075 in human colorectal cancer are still poorly understood.
Methods:
Quantitative real-time PCR (qRT-PCR) was performed to detect the expression level of hsa_circ_0005075 in colorectal cancer. Correlations of hsa_circ_0005075 expression with pathological parameters and overall survival were assessed. CCK-8 and Transwell invasion assays were utilized to determine the effect of hsa_circ_0005075 on proliferation and invasion of colorectal cancer cells.
Results:
Hsa_circ_0005075 was overexpressed in colorectal cancer tissues compared with paracancerous tissues and intestinal polyps. Its expression level was associated with distal metastasis, invasion, tumor node metastasis stage, and tumor diameter in colorectal cancer, and was negatively correlated with overall survival of patients with colorectal cancer. Moreover, enforced expression of hsa_circ_0005075 potentiated the proliferation and invasive behavior of colorectal cancer cells.
Conclusions:
Our findings suggested that hsa_circ_0005075 expression was increased in colorectal cancer and might serve as a promising diagnostic mark and therapy target of colorectal cancer.
Introduction
Colorectal cancer (CRC) is the most common gastrointestinal malignancy, and is the fourth leading cause of cancer-related mortality globally. 1 The symptoms of early CRC are not obvious and are easy for patients to ignore. Symptoms are often accompanied by multiple organ invasion and distant metastasis when diagnosed, resulting in lost opportunity of radical resection and a poor prognosis.2,3 Consequently, endeavoring to explore ideal diagnostic and therapeutic targets will contribute to the amelioration of the outcome for CRC patients.
Circular RNAs (circRNAs) are becoming a new research hotspot in the non-coding RNA world; they have covalently closed loop structures through joining free 3′- to 5′- ends.4-6 Some circRNAs could function as a microRNA sponge to possess microRNA binding sites, affecting the expression of target genes. 7 In contrast to microRNAs and long non-coding RNAs (lncRNAs), circRNAs feature a more stable structure and higher tissue-specific expression. 8 This typical character of circRNAs makes them an emerging choice for acting as molecular markers of CRC.
Emerging evidence indicates that circRNAs are closely related with carcinogenesis and tumor progression. 9 We found that hsa_circ_0005075 might be a CRC-related circRNA by searching from the circRNA database: (circBase: http://www.circbase.org/). Hsa_circ_0005075 is mapped on chromosome 1: 2137735821415706. Its associated gene symbol is a tumor suppressor gene named SHPRH, which could be found in glioblastoma and could affect genomic integrity.10,11 Our data illustrated that high expression hsa_circ_0005075 was observed in CRC samples and cells by quantitative real-time PCR (qRT-PCR), and high expression of hsa_circ_0005075 was correlated with clinical pathological characteristics and survival rate. Moreover, proliferation and invasion of CRC cells were impacted by the expression level of hsa_circ_0005075. Our findings indicated that hsa_circ_0005075 may function as a promising biomarker for CRC diagnosis and treatment.
Materials and methods
Patient samples
Specimens were collected after informed consent from all participants in the Affiliated Hospital of Xuzhou Medical University, China, from April 2013 to June 2015. The paired 93 CRC tissues and adjacent normal tissues were collected from surgical patients. Another 60 intestinal polyp tissues were collected from intestinal mucosa biopsies. All samples were stored immediately at −80°C after acquisition. The tumor stage of CRC tissue was classified according to the eighth American Joint Committee on Cancer (AJCC) classification system. Clinical information of every patient was listed and described in Supplementary Table 1. None of the selected patients underwent any radiotherapy and/or chemotherapy before tissue biopsy and surgical resection.
The protocol was approved by the Ethics Committee of Affiliated Hospital of Xuzhou Medical University.
Cell culture
Human normal colorectal mucosal cells FHC and colorectal cancer cells HCT116, DLD1, LoVo, SW480, and SW620 were obtained from the Cell Bank of Chinese Academy of Sciences (Beijing, China). Cell lines SW620, HCT116, and SW480 were cultured in DMEM medium (Gibco, Carlsbad, CA, USA); LoVo and DLD1 in RPMI 1640 medium (Gibco); and FHC in DMEM/F12 medium (Gibco) was supplemented with 10% fetal bovine serum (Gibco) and 1% penicillin/streptomycin. Cells were incubated in a 37°C humidified atmosphere with 5% CO2.
Cell transfection
HCT116 and SW480 cells (3.0 × 105 per well) plated in a 6-well plate with were transfected. GenePharm (Shanghai, China) designed and synthesized interference sequence of hsa_circ_0005075, LV-hsa_circ_0005075 (LV-HSA) packaged with a lentivirus vector. The sequences for LV-HAS are 5′-GACTCTCTCAAAATTCCTAGA-3′, and sequences for the negative control (LV-NC) are 5′-TTCTCCGAACGTGTCACGT-3′. 12 The hsa_circ_0005075 overexpression plasmid pEX1-hsa_circ_0005075 (HSA-OV) and its negative control (NC) plasmid pEX1-control were purchased from GenePharm.
RNA extraction and qRT-PCR
RNA was isolated using Trizol (Life Technologies, Carlsbad, CA) according to the manufacturer’s protocol. Then cDNA was synthesized with SuperScript IV VILO Master Mix (Waltham, MA, USA). qRT-PCR analysis was performed by SYBR Premix Ex Taq II (Takara, Dalian, China) with 7500 Real-time PCR System (Life Technologies, NY, USA). The sequences of hsa_circ_0005075 primers and GAPDH are as follows (GenePharma): hsa_circ_0005075, forward: 5′-AATGCTGAAAACTGCTGAGAGAA-3′, reverse 5′-TTGAGAAAACGAGTGCTTTGG-3′; GAPDH, forward: 5′-CAAATCTTGCGGCAACGC-3′, reverse 5′-GCGGGAGTGAAGATTCGA-3′. Each reaction was performed in triplicate. The data were normalized to the values of GADPH, and relative expression was calculated by the 2-ΔΔCt method. The ΔCt values of all samples listed and described are in Supplementary Table 1.
Cell proliferation assay
Cell proliferation assays were performed following the manufacturer’s protocol. Briefly, transfected cells (HCT116 and SW480) were resuspended in serum-supplemented medium, plated onto a 96-well plate (3 × 103 cells/well). Then 10 uL of CCK-8 reagent was added to each well and the absorbance (450 nm) was detected at the indicated time. All assays were repeated at least three times.
Transwell invasion assay
Cell suspension with serum-free medium was inoculated in the upper chambers containing 8 μm pores (BD Bioscience, CA, USA) with Matrigel, and the lower chambers were filled with serum-supplemented medium. After 48 h, the invading cells were fixed, stained with 0.5% crystal violet, calculated, and photographed under a microscope in five randomized fields. All assays were repeated at least three times.
Statistical analysis
All statistical analyses were performed using the SPSS 20.0 (IBM, Chicago, IL, USA) and images were graphed using GraphPad Prism 5.0. Two or more group comparisons were performed by the Student’s t-test or one-way analysis of variance (ANOVA), as appropriate. The differences of hsa_circ_0005075 expression in different clinical samples were evaluated by paired t-test. The survival curve was calculated using the Kaplan–Meier method and the log-rank test. All dates were expressed as mean ± SD. P < 0.05 was considered significant.
Results
Hsa_circ_0005075 is abundantly expressed in CRC
Initially, the associated gene (SHPRH) expression of hsa_circ_0005075 was detected. Results from qRT-PCR showed that the SHPRH level was lower in CRC cell lines than in FHC cells (normal colorectal epithelial cells) (Figure 1(a); P < 0.01); it also had lower expression in CRC tissues compared with paracancerous tissues (Figure 1(b); P < 0.05). To verify the bioinformatics analysis, we measured the hsa_circ_0005075 expression in CRC cell lines, CRC tissues, and the plasma of patients with CRC. We found that hsa_circ_0005075 in CRC cell lines were enhanced compared with FHC cells (Figure 2(a); P < 0.05). Moreover, hsa_circ_0005075 was increased in CRC tissue versus intestinal polyp and paracancerous tissues (Figure 2(b); P < 0.05), which was consistent with the result observed in plasma samples (Figure 2(c); P < 0.05). All results illustrated that hsa_circ_0005075 expression was up-regulated in CRC, whereas its expression profile in CRC was different from the circRNA host gene SHPRH.
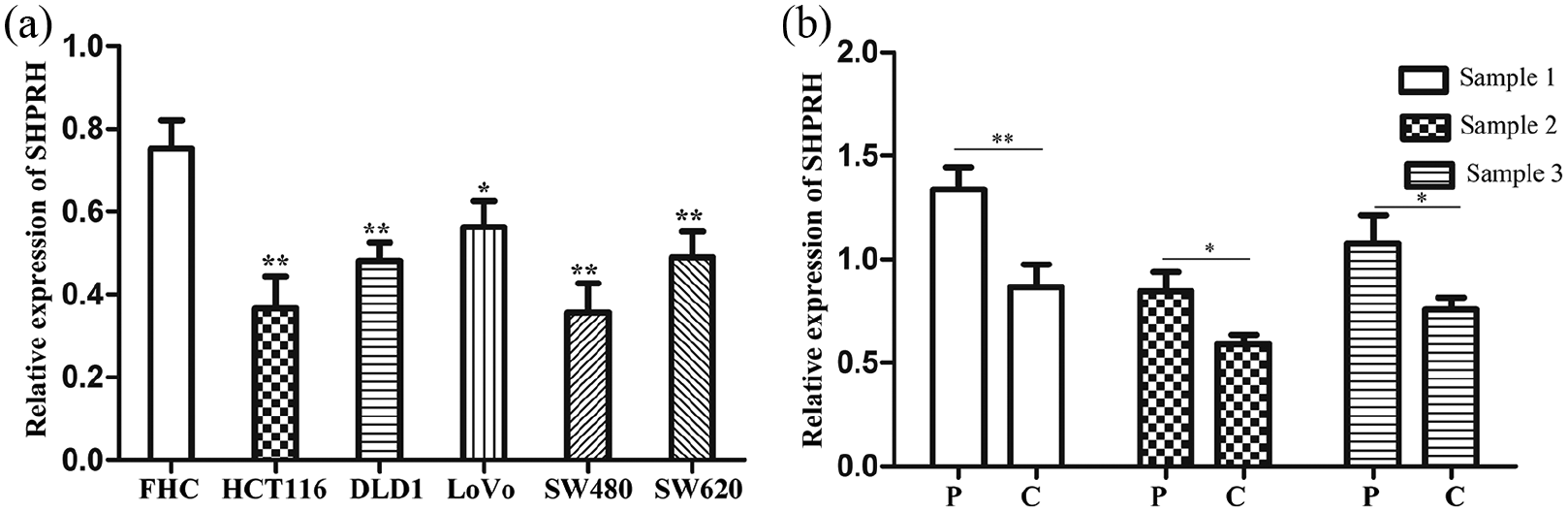
SHPRH is decreased in CRC. (a) qRT-PCR analysis of the relative expression of hsa_circ_0005075 in CRC cells and FHC cells (one-way ANOVA; *P < 0.01). (b) We randomly selectd three pairs CRC tissues and adjacent normal tissues to investigate the SHPRH expression (Student’s t-test; *P < 0.05).
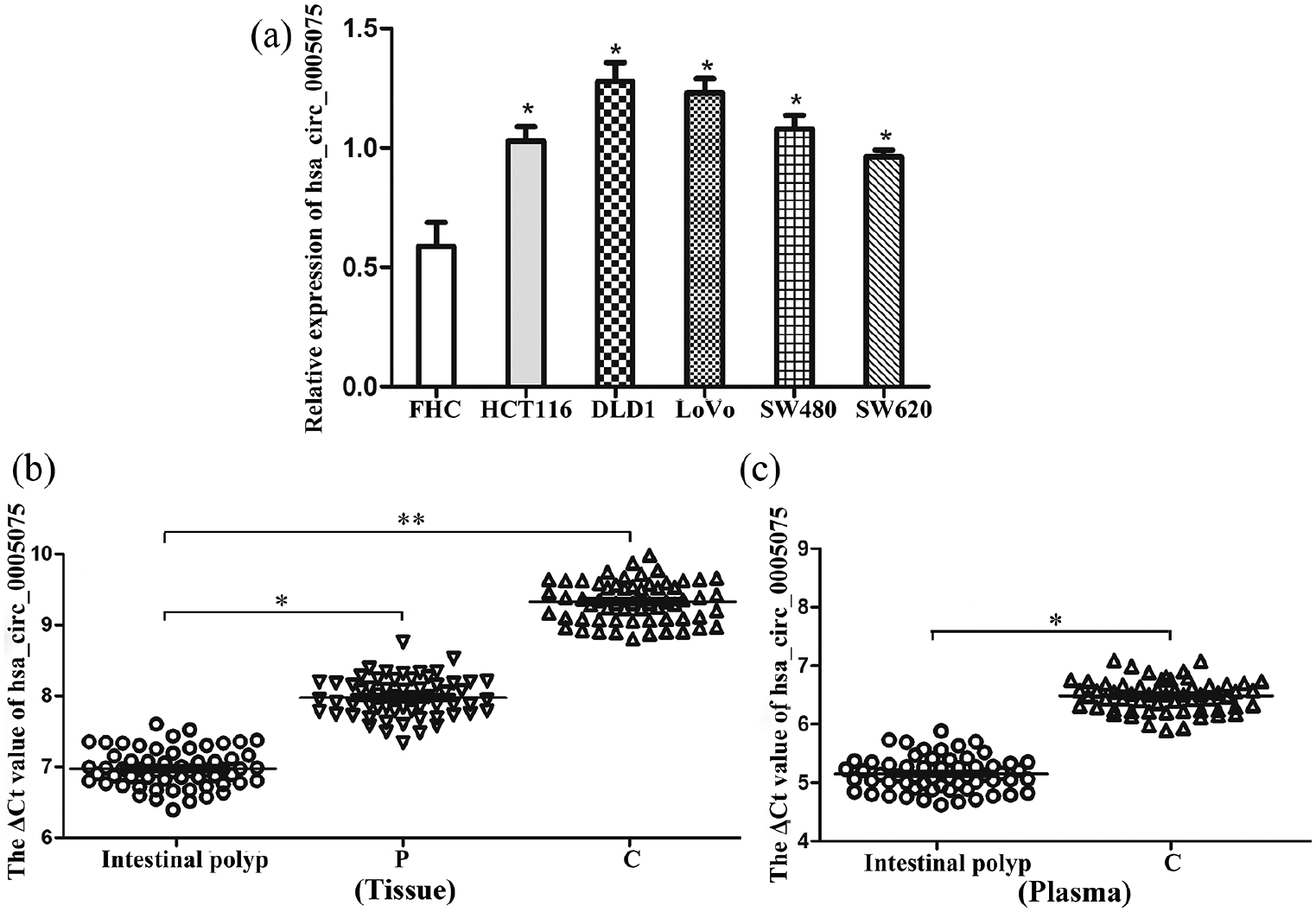
Hsa_circ_0005075 is increased in CRC. (a) qRT-PCR analysis of the relative expression of hsa_circ_0005075 in CRC cells and FHC cells (one-way ANOVA; *P < 0.05). (b) The ΔCt value of hsa_circ_0005075 in CRC tissues compared with intestinal polyp and paracancerous tissues: (n=60, paired t-test; *P < 0.05, **P < 0.01). (c) The ΔCt value of hsa_circ_0005075 in CRC patients’ plasma compared with intestinal polyp patients: (n=60, paired t-test; *P < 0.05).
Increased hsa_circ_0005075 is correlated the clinicopathological parameters in CRC
Results described above showed that hsa_circ_0005075 expression was increased in CRC. To explore the importance and potential function of hsa_circ_0005075, the relationships between its expression and clinicopathological features was analyzed in CRC patients. It was indicated that increased hsa_circ_0005075 was associated with tumor diameter, distal metastasis, invasion, and tumor node metastasis (TNM) stage (Table 1). However, we did not find any association of hsa_circ_0005075 with age, gender, and differentiation (Table 1). Furthermore, the similar association was not found in plasma samples (Supplementary Table 2). Patients were divided into two groups according to the media value of hsa_circ_0005075 expression in CRC tissues (low expression ⩽ media value; high expression > media value). Meanwhile, CRC patients with overexpressed hsa_circ_0005075 possessed poorer survival via Kaplan–Meier survival analysis and log-rank test (Figure 3; P < 0.001). Together, hsa_circ_0005075 might have a critical impact on CRC progression.
The relationship of hsa_circ_0005075 expression levels (ΔCt) in CRC tissues and clinicopathological factors of patients with CRC.
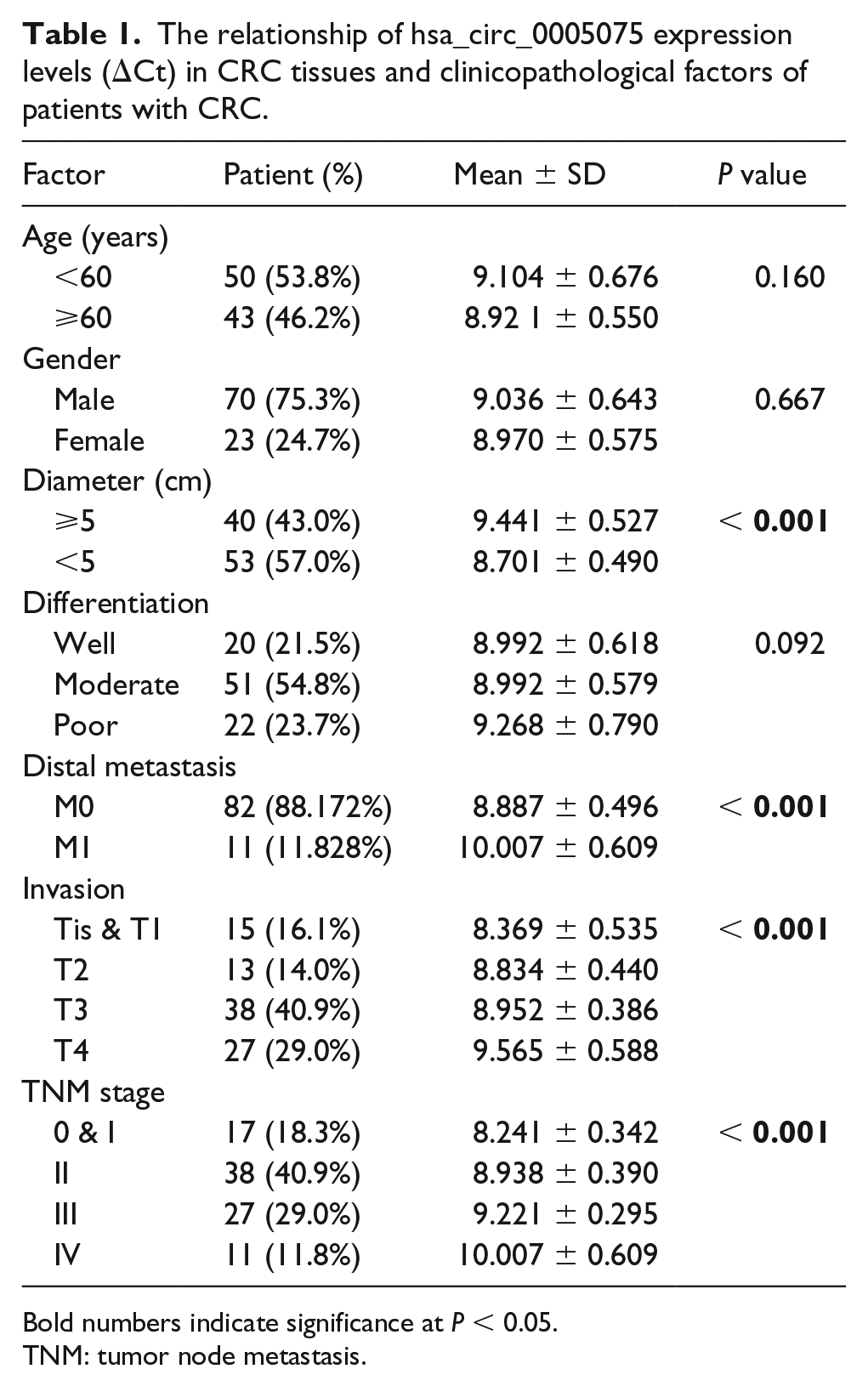
Bold numbers indicate significance at P < 0.05.
TNM: tumor node metastasis.
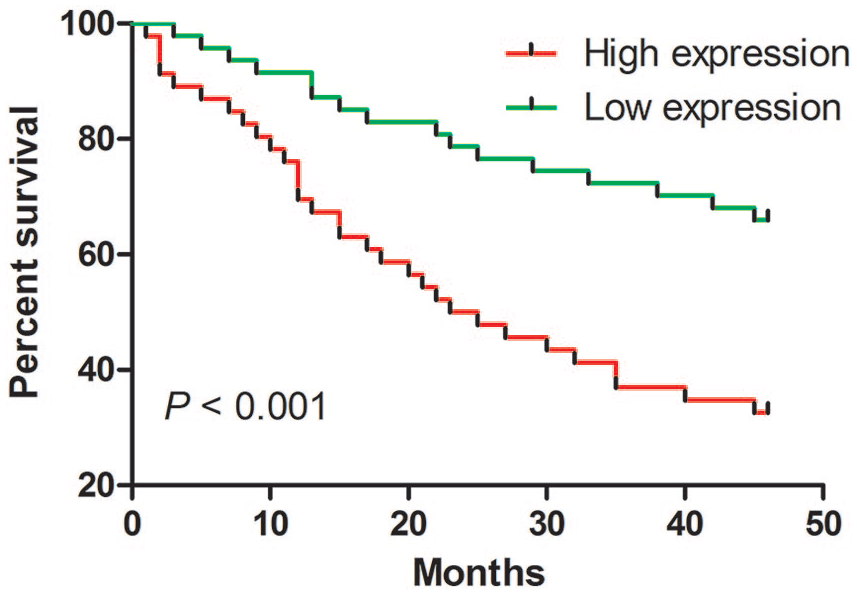
The relationship between hsa_circ_0005075 expression and cumulative survival for CRC. Kaplan–Meier survival analysis for CRC patients with high versus low expression was conducted. (Kaplan–Meier method and log-rank test; P < 0.001).
Hsa_circ_0005075 promotes cell proliferation and invasion
The above findings indicated enforced expression of hsa_circ_0005075 correlated with proliferative and aggressive phenotypes and unfavorable prognosis of CRC. We hypothesized that hsa_circ_0005075 might exert significant influence on the ability of cell proliferation and invasion. We increased and decreased hsa_circ_0005075 expression using overexpression plasmid of hsa_circ_0005075 and interference sequence targeting hsa_circ_0005075 in HCT116 and SW480 cells, respectively. As shown in Figure 4(a), transfection efficiencies of hsa_circ_0005075 overexpression and knockdown in CRC cells were confirmed using qRT-PCR. Subsequently, CCK-8 assay was utilized to determine the effect of hsa_circ_0005075 on cell proliferation. Elevated hsa_circ_0005075 enhanced the proliferation of HCT116 and SW480 cells (Figure 4(b); P < 0.01); conversely, hsa_circ_0005075 silencing markedly attenuated cell proliferation (Figure 4(c); P < 0.01). Furthermore, Transwell assay was used to assess hsa_circ_0005075-associated invasion in CRC cells. As expected, hsa_circ_0005075 could affect cell motility. Hsa_circ_0005075 amplification significantly potentiated invasion of HCT116 and SW480 cells (Figure 5(a); P < 0.01). The ability for invasion was reduced in CRC cells with hsa_circ_0005075 silencing (Figure 5(b); P < 0.001). Collectively, hsa_circ_0005075 could facilitate proliferation and invasion of CRC cells.
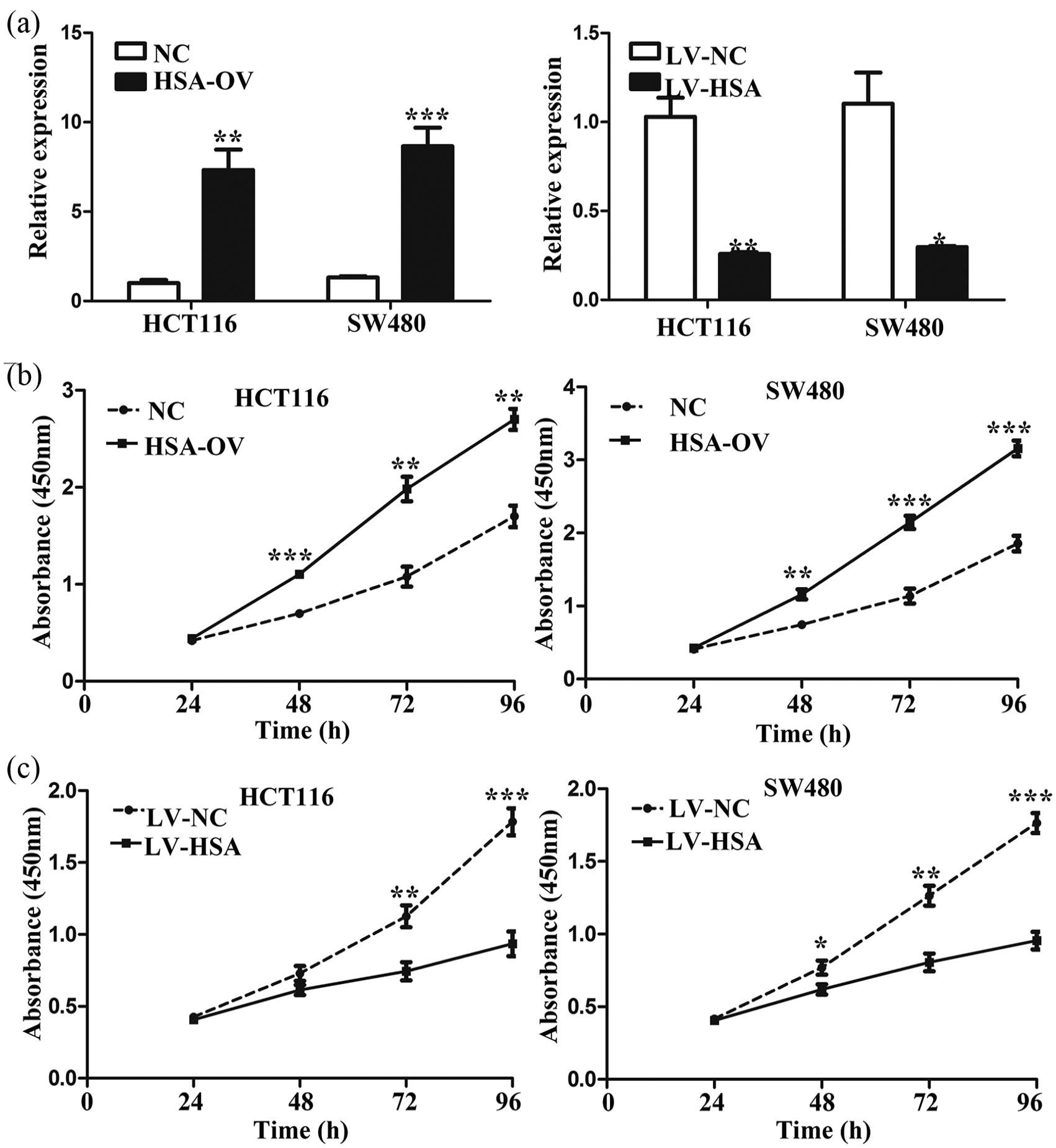
Hsa_circ_0005075 promotes proliferation of CRC cells. (a) qRT-PCR analysis of hsa_circ_0005075 expression in HCT116 and SW480 cells transfected with hsa_circ_0005075 overexpression plasmid, interference sequence targeting hsa_circ_0005075 and respective negative control (Student’s t-test; *P < 0.05, **P < 0.01, ***P < 0.001). (b) and (c) Cell proliferation was analyzed by CCK-8 assays in HCT116 and SW480 cells after hsa_circ_0005075 overexpression and knockdown (Student’s t-test; **P < 0.01, ***P < 0.001).
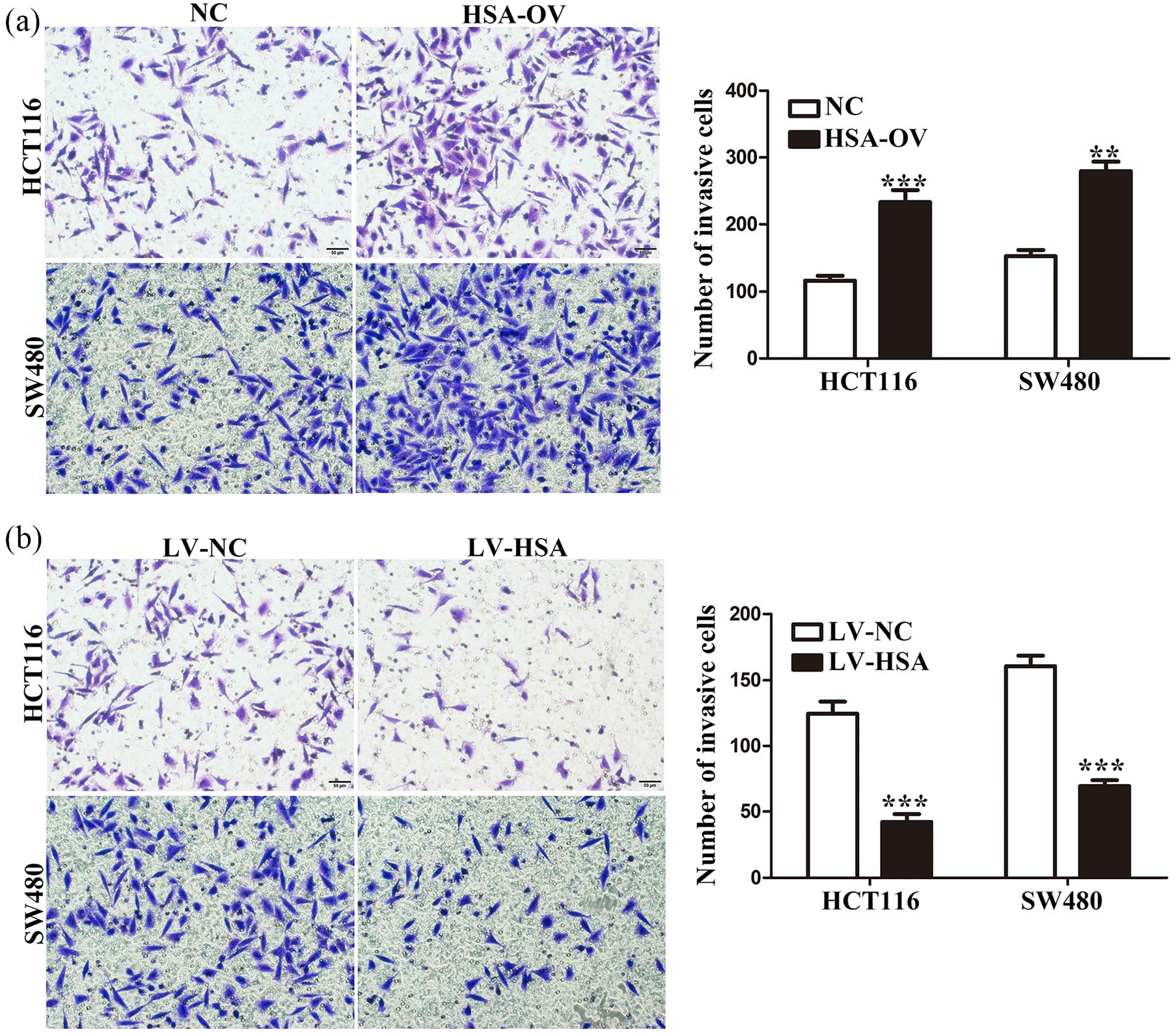
Hsa_circ_0005075 promotes invasion of CRC cells. (a) and (b) Invasive properties of HCT116 and SW480 cells were assessed by Transwell invasion assay after hsa_circ_0005075 overexpression and knockdown (Student’s t-test; **P < 0.01, ***P < 0.001).
Discussion
With the improvement of treatment strategies, clinical efficacy remains suboptimal due to malignant invasion and rapid deterioration of CRC. 13 Through the years, although much progress has been achieved in molecular pathogenesis CRC, intense investigation is still required. 14 Hence, searching for novel biomarkers to ameliorate clinical diagnosis and therapy for CRC seems especially compelling.
The aberrant expression of non-coding RNAs, particularly lncRNAs, microRNAs, and circRNAs, has been confirmed as having a crucial role in tumorigenesis and malignant progression through modulating tumor-related signaling pathways.15-20 Non-coding RNAs were originally considered as the results from transcriptional noise, one of which is circRNAs. 21 Recent reports have demonstrated that circRNAs can act as a microRNA sponge, modulating gene transcription. 22 Emerging evidence has indicated that circRNAs exert critical roles in carcinogenesis, and could serve as new biomarkers. For example, hsa_circ_0006215 expression was increased, and regulated the expression of miR-378a-3p in pancreatic cancer. 23 In lung adenocarcinoma, hsa_circ_0000729 could participate in signal transduction by targeting miR-375. 24 Hsa_circ_0016788 promoted human hepatocellular carcinoma (HCC) tumorigenesis via modulating miR-486/CDK4 signaling, and stimulated cell proliferation and invasion. 25 Hsa_circ_0008039 potentiated the occurrence and aggressiveness of breast cancer by influencing miR-432-5p/E2F3 signaling. 26 Decreased hsa_circ_0137287 expression reversed tumorigenesis and was associated with clinicopathologic characteristics in papillary thyroid carcinoma. 27 Nevertheless, hsa_circ_0005075 function in CRC progression remains limited.
Here, we examined RNA levels of hsa_circ_0005075 in CRC cells, tissue, and plasma using qRT-PCR, and found that hsa_circ_0005075 was elevated in CRC cells compared with FHC cells. A stepwise increased trend in expression of hsa_circ_0005075 was observed as malignant progression of CRC tissues. Hsa_circ_0005075 expression in cancer was also higher than that of intestinal polyps when its level was analyzed from plasma samples. We further evaluated the relationships between hsa_circ_0005075 level, clinicopathologic parameters, and prognostic survival of CRC. Hsa_circ_0005075 expression was positively related to tumor diameter, distal metastasis, invasion, and TNM stage in CRC. Meanwhile, CRC patients with overexpressed hsa_circ_0005075 were involved in a poor prognosis. Subsequently, we investigated hsa_circ_0005075 roles in cell proliferation and invasion via its overexpression and inhibition. Hsa_circ_0005075-transfected CRC cells possessed more potent proliferation and invasion capacity; reverse trends can be found when hsa_circ_0005075 was down-regulated, which evidenced that hsa_circ_0005075 might exert a critical role in CRC occurrence and progression. Previous studies of hsa_circ_0005075 focused on HCC. Hsa_circ_0005075 had been discovered to increase in HCC and could accelerate tumorigenesis by targeting miR-431. 12 Also, hsa_circ_0005075 expression was related to tumor size and potentiated cell adhesion property in malignant progression of HCC. 28 Significantly, Jin et al. 29 recently reported that higher expression of hsa_circ_0005075 in CRC was associated with depth of invasion and advanced TNM stage, and predicted a poor prognosis; knockdown of hsa_circ_0005075 inhibited cells proliferation, migration, and invasion, and expedited apoptosis by activating the Wnt/β‑catenin pathways. Taken together, the hsa_circ_0005075 role may provide a new perspective for CRC diagnosis and treatment.
Conclusions
In summary, hsa_circ_0005075 expression was increased in CRC. Also, there were correlations between hsa_circ_0005075 expression, clinicopathological features, and prognosis survival of CRC. Furthermore, hsa_circ_0005075 promoted proliferation and invasion of CRC cells. Taken together, hsa_circ_0005075 may serve as a novel and potential biomarker for CRC diagnosis and gene therapy.
Supplemental Material
Supplemental_Table_1 – Supplemental material for Hsa_circ_0005075 promotes the proliferation and invasion of colorectal cancer cells
Supplemental material, Supplemental_Table_1 for Hsa_circ_0005075 promotes the proliferation and invasion of colorectal cancer cells by Di Zhong, Peng Li and Pei-yao Gong in The International Journal of Biological Markers
Supplemental Material
Supplemental_Table_2 – Supplemental material for Hsa_circ_0005075 promotes the proliferation and invasion of colorectal cancer cells
Supplemental material, Supplemental_Table_2 for Hsa_circ_0005075 promotes the proliferation and invasion of colorectal cancer cells by Di Zhong, Peng Li and Pei-yao Gong in The International Journal of Biological Markers
Footnotes
Author contributions
Di Zhong and Peng Li contributed equally to this work.
Declaration of conflicting interests
The author(s) declare that there is no conflict of interest.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Science and Technology Bureau of Xuzhou (KC15SH075).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
