Abstract
Background:
Recent evidence suggests that -592 A/C polymorphism in the interleukin-10 (IL-10) gene may influence risk of gastrointestinal tract cancer; however, individual studies have provided conflicting and inconclusive results. Therefore, this meta-analysis was conducted to assess the association between IL-10 -592 A/C polymorphism and gastrointestinal tract cancer susceptibility.
Methods:
EMBASE, PubMed, Web of Science, and China National Knowledge Infrastructure databases were searched for case-control studies published before 1 May 2017. A total of 36 studies involving 8069 cases and 13,089 controls were included in the present meta-analysis according to the inclusion criteria. The random- or fixed-effect model was utilized to calculate pooled odds ratio (OR) with 95% confidence interval (CI), and to survey the association.
Results:
By and large IL-10 -592 A/C (rs1800872) polymorphism was not associated with gastrointestinal cancer risk in five genetic models (A vs. C: OR 1.00; 95% CI 0.93, 1.08; POR = 0.960; AA vs. CC: OR 0.98; 95% CI 0.85, 1.14; POR = 0.835; CA vs. CC: OR 1.01; 95% CI 0.94, 1.08; POR = 0.776; AA+CA vs. CC: OR 1.03; 95% CI 0.94, 1.12; POR = 0.592; AA vs. CA+CC: OR 0.98; 95% CI 0.87, 1.10; POR = 0.666). Similar results were also achieved after stratification by the Hardy–Weinberg equilibrium, ethnicity, source of controls, and cancer type.
Conclusion:
The results of this meta-analysis indicated that there is no association between the IL-10 -592 A/C promoter polymorphism and gastrointestinal tract cancer susceptibility.
Introduction
Cancer remains a main public health challenge and one of the major leading causes of death worldwide. Gastrointestinal tract (GIT) malignancies are one of the most common cancer types, including esophageal, gastric, and colorectal cancers. The latter three cancer types are the eighth, fourth and third most common cancer types worldwide, respectively. 1 During the last two decades, there have been attempts to reduce risk factors, to develop screening tests, and to advance treatment to decrease the incidence of GIT cancer and the resulting death rate.2,3 Many studies reported that cancer is dependent on various factors, and the initiation and development of GIT cancer is a multifactorial and multistep process involving a variety of risk factors, including genetic factors and allelic variation.4-6 Several studies have demonstrated that common polymorphisms in a number of inflammatory genes correlate with the risk of cancer, especially sporadic colorectal cancer susceptibility. 7 The assessment of genetic polymorphism may help to interpret the role of genetic alteration in cancer risk.
Clinical, experimental, and epidemiological evidence have indicated chronic inflammation as a cause of various types of cancer. Inflammation is characterized by a complicated, highly coordinated process, which involves different sorts of cells and molecules. Some of them are responsible for driving the process, others modulate it, and still some others have both these responsibilities at different points. One of the important functions carried out by cytokines is the regulation of inflammatory response. Further concentration on individual cytokines has led to the approval of the possibility that anti-inflammatory cytokines might have a significant role in gastrointestinal carcinogenesis. 8 Interleukin-10 (IL-10) is a principal cytokine during the resolution phase of inflammation and immune-suppressive cytokine. Increasing evidence is positive about the prohibitive effect of IL-10 in tumor carcinogenesis, angiogenesis, and metastasis. 9 On the other hand, the absence of IL-10 can have several consequences, including pro-inflammatory cytokine production, anti-tumor immunity inhibition, and tumor growth advancement. 10
The IL-10 gene is located on chromosome 1q31-q32 in humans, which consists of five exons and four introns. 11 IL-10-592 A/C (rs1800872) polymorphism in the promoter region of the IL-10 gene may affect the activity of transcription factors and its expression level, and thus might modulate the susceptibility of cancer.
Many epidemiological literatures have investigated the association between IL-10-592 A/C polymorphism and esophageal, gastric, and colorectal cancer susceptibility. Previous studies about association between IL-10-592 A/C polymorphism and risk of GIT cancer have provided contradictory results. de Oliveira et al. 12 and Tsilidis et al. 13 reported that IL-10-592 A/C polymorphism is associated with the increased risk of gastric and colorectal cancer, but the studies of Pan et al. 14 and Talseth et al. 15 showed that this polymorphism is not a risk factor. Therefore, in this study, we conducted a meta-analysis to examine the association between the -592 A/C polymorphism in the IL-10 gene and GIT cancer susceptibility.
Materials and Methods
Search strategy
To identify all articles that evaluated the association of IL-10 -592 A/C polymorphism with GIT cancer, literature retrieval was carried out in PubMed and other databases (e.g., EMBASE, Web of Science, and China National Knowledge Infrastructure (CNKI)) using a combination of the following terms: IL-10, polymorphism, esophageal, gastric, colorectal, cancer (last search: May 2017). The references of all identified publications were also checked for additional pertinent studies. It is worth mentioning that there was no language limitation in the literature search.
Inclusion and exclusion criteria
All titles and abstracts of the identified studies were investigated and only the ones meeting these criteria were included in the present study: (a) a case-control study; (b) a study about the association of IL-10 -592 A/C polymorphism with GIT cancer susceptibility; and (c) the provision of required data, including the distribution of genotype frequency, odds ratios (ORs), and 95% confidence intervals (CIs). The major exclusion criteria comprised: (a) not reporting genotype frequency in cases and controls; and (b) case-only studies, case reports, reviews, and duplicated studies.
For each eligible study, the following data were extracted: first author, year of publication, country, ethnicity, cancer type, number of cases and controls, and genotyping methods. We also evaluated whether the distribution of genotype in controls followed the Hardy–Weinberg equilibrium (HWE).
Statistical analysis
ORs with the corresponding 95% CIs were calculated to assess the strength of association between IL-10 -592 A/C polymorphism and GIT cancer risk. A Z-test was performed to evaluate the significance of pooled ORs. Recessive (AA vs. CA+CC), dominant (AA+CA vs. CC), heterozygous (CA vs. CC), homozygous (AA vs. CC), and allelic (A vs. C) genetic models were investigated. Stratified analysis according to HWE, ethnicity, source of controls, and cancer type was carried out to detect the potential relationship for each subgroup.
In order to survey HWE in the control group, the genotype frequency of the IL-10 -592 A/C polymorphism was calculated using the chi-square test, and P< 0.05 was considered as significant deviation from HWE. Heterogeneity was measured by the I-square (I2) test and the Q-test. When I2>40% and the P value of the Q-test was less than 0.1, significant heterogeneity between studies was considered, and a random-effects model was used to calculate the pooled ORs. Otherwise, a fixed-effects model was selected to estimate the pooled ORs. 16 The potential publication bias was measured using Begg’s funnel plots and Egger’s linear regression test. A P value less than 0.05 for Egger’s and Begg’s test suggested significant publication bias. 17 Sensitivity analysis was performed to examine the influence of the individual studies on the overall finding. All the statistical analysis was conducted by Comprehensive Meta-Analysis version 2.2 software (Biostat, Englewood, NJ, USA).
Results
Study characteristics
Relevant articles published before 1 May 2017 were identified through a search in EMBASE, PubMed, Web of Science, and CNKI databases with no language restriction. A flow diagram of the study selection process is displayed in Figure 1. A total of 36 case-control studies on the association of IL-10 -592 A/C polymorphism and susceptibility to GIT cancer were retrieved based on the inclusion criteria.8,12-14,18-47 The distribution of genotypes in the control groups have deviated from HWE in three publications.21,22,30
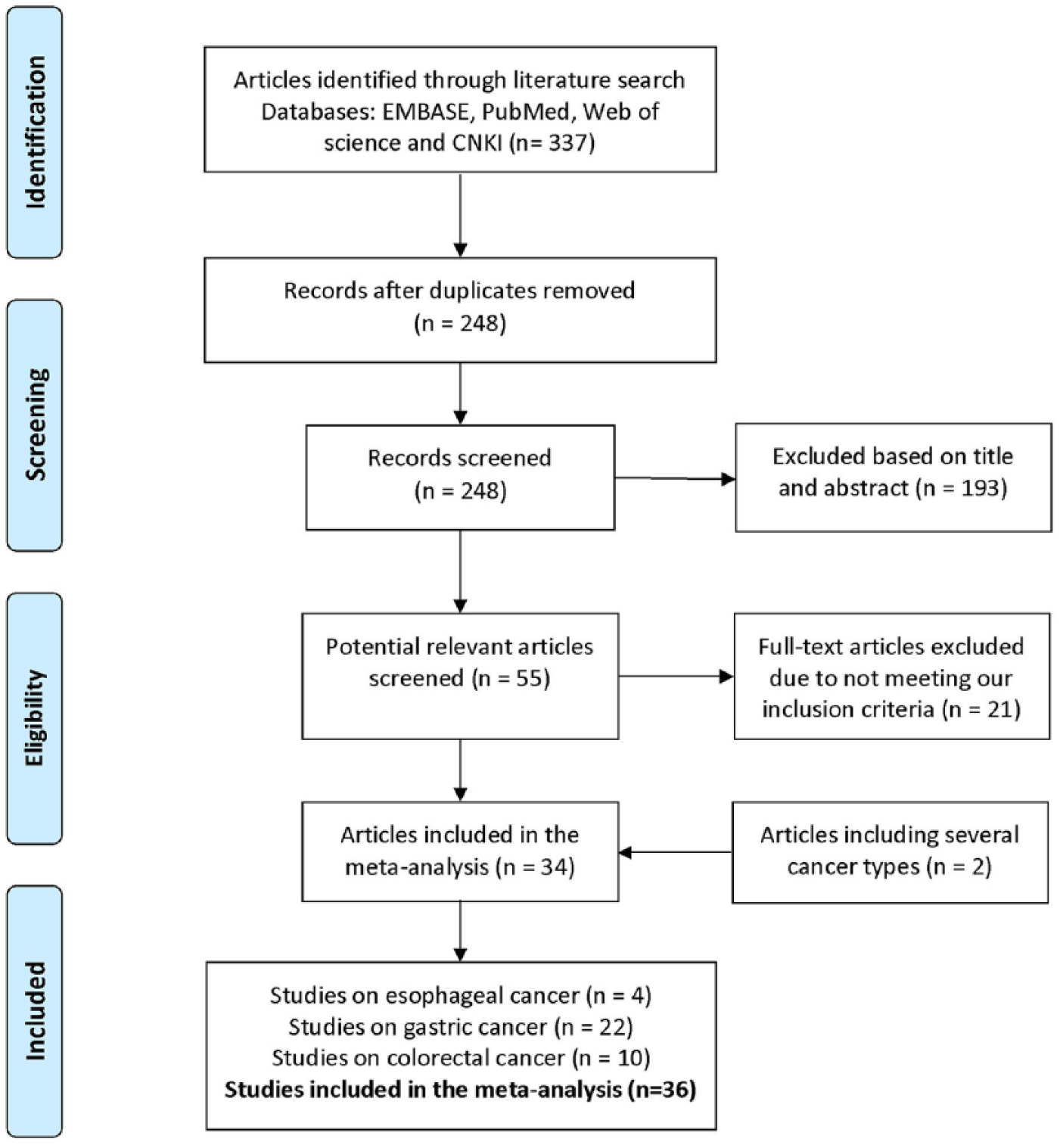
PRISMA flow diagram of included studies in interleukin-10 -592 A/C polymorphism and gastrointestinal tract cancer.
Based on the search criteria, 8069 multiple cancer cases and 13,089 controls from 36 eligible studies were retrieved for this meta-analysis. Four, 22, and 10 studies conducted analysis on esophageal, gastric, and colorectal cancer, respectively. Sixteen studies out of the relevant ones were carried out in the Asian population, 17 in Caucasians, and three Latinos. Regarding the source of controls in the 36 available studies, 17 were based on population controls, and 17 on hospital controls, while two did not refer to their control sources.
Two articles contained different cancer types, so they were treated as two studies in this meta-analysis.25,34 In most of the studies, polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) and TaqMan techniques were used to determine the genotype. The detailed characteristics of the relevant studies are represented in Table 1.
Characteristics of the studies recruited for the meta-analysis.
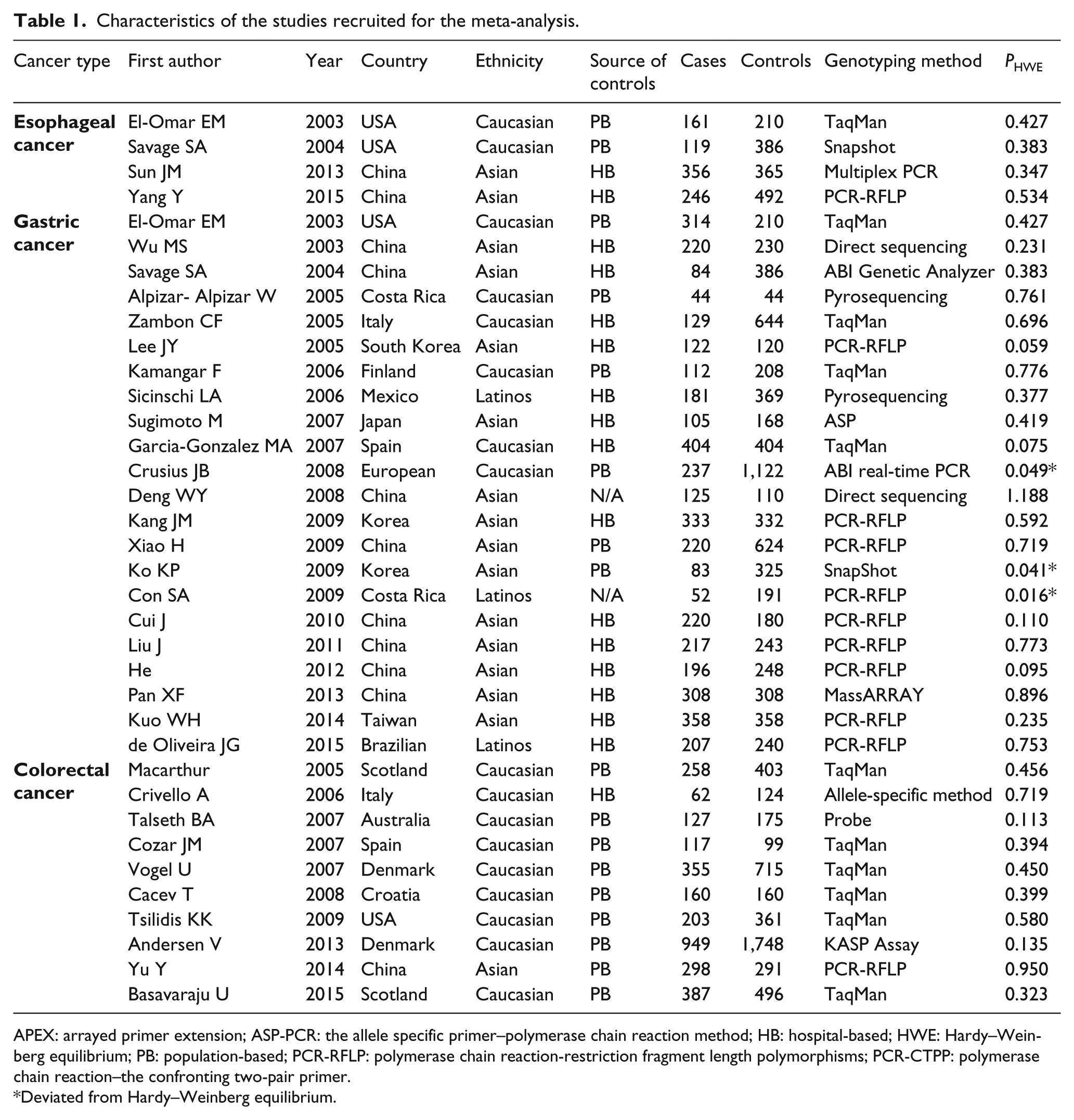
APEX: arrayed primer extension; ASP-PCR: the allele specific primer–polymerase chain reaction method; HB: hospital-based; HWE: Hardy–Weinberg equilibrium; PB: population-based; PCR-RFLP: polymerase chain reaction-restriction fragment length polymorphisms; PCR-CTPP: polymerase chain reaction–the confronting two-pair primer.
Deviated from Hardy–Weinberg equilibrium.
Meta-analysis results
The results of the meta-analysis of IL-10 -592 A/C polymorphism and GIT cancer susceptibility are presented in Table 2 and Figure 2. The pooled ORs of all eligible case-control studies indicated that there were no statistically significant associations between the IL-10 -592 A/C polymorphism and the GIT cancer risk: (A vs. C: OR 1.00; 95% CI 0.93, 1.08; POR = 0.960; AA vs. CC: OR 0.98; 95% CI 0.85, 1.14; POR = 0.835; CA vs. CC: OR 1.01; 95% CI 0.94, 1.08; POR = 0.776; AA+CA vs. CC: OR 1.03; 95% CI 0.94, 1.12; POR = 0.592; AA vs. CA+CC: OR 0.98; 95% CI 0.87, 1.10; POR = 0.666).
Investigating the association between IL-10 -592 A/C polymorphism and gastrointestinal tract cancer.
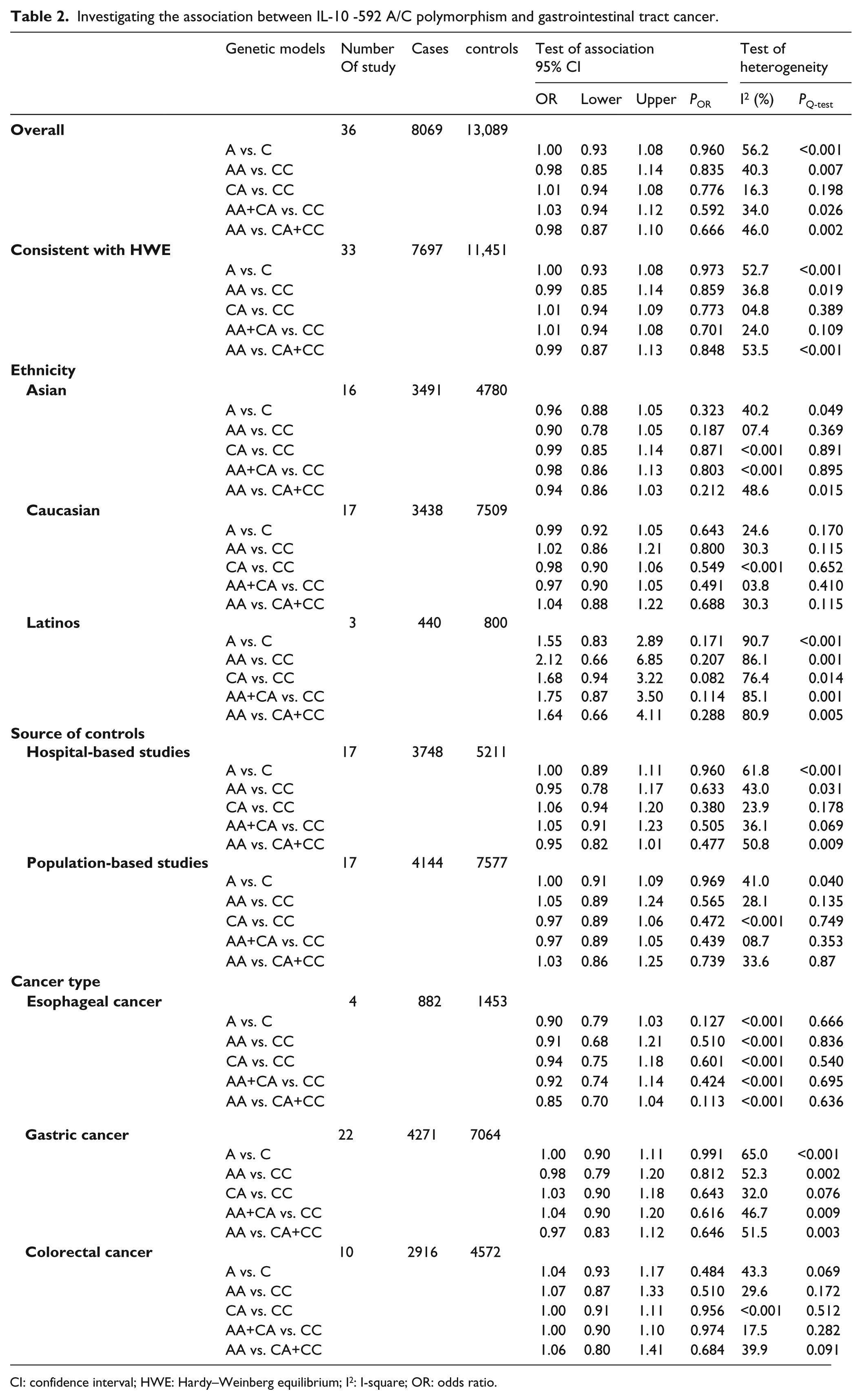
CI: confidence interval; HWE: Hardy–Weinberg equilibrium; I2: I-square; OR: odds ratio.
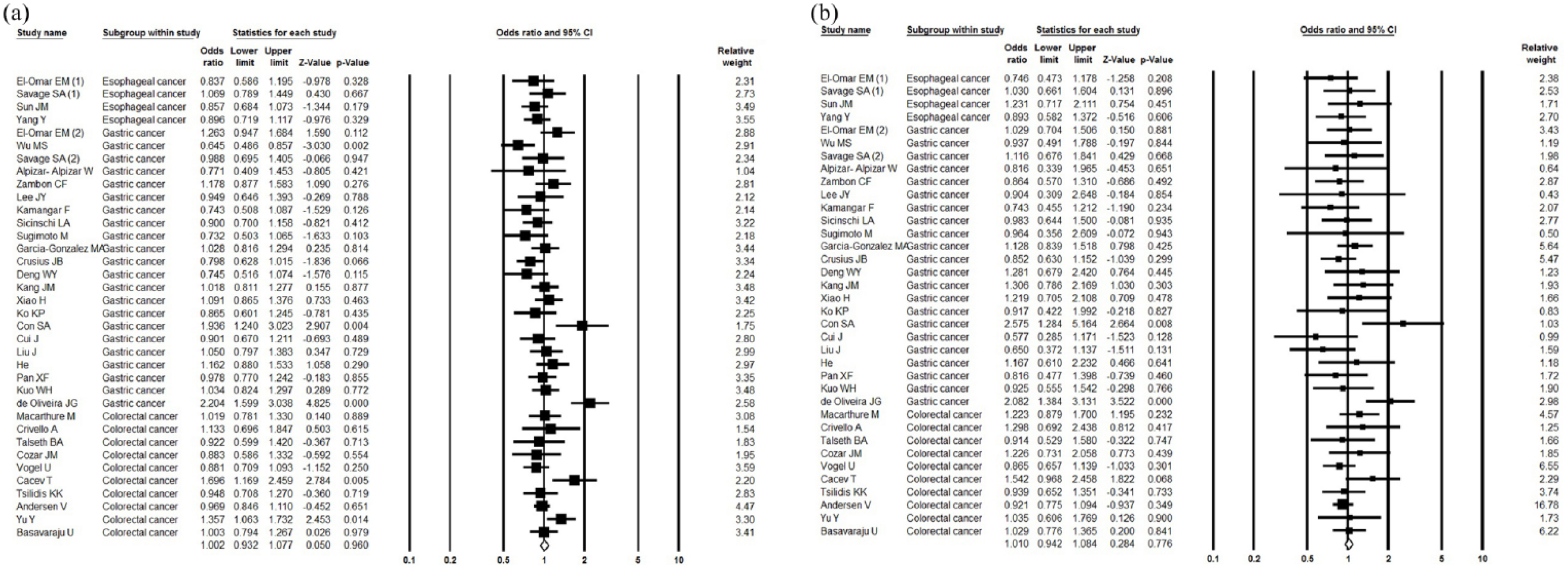
Forest plot of associations between IL-10 -592 A/C polymorphism and GIT cancer risk. (a) Allelic model (A vs. C). (b) Heterozygous model (CA vs. CC).
In stratified analysis by cancer types, no statistically significant association was observed under all genetic models between the IL-10 -592 A/C polymorphism and esophageal, gastric and colorectal susceptibility.
In addition, in a subgroup analysis by HWE, ethnicity (Asian, Caucasian, and Latinos) and source of controls demonstrated no significant associations between IL-10 -592 A/C polymorphism and GIT cancer risk (Table 2 and Figure 3).
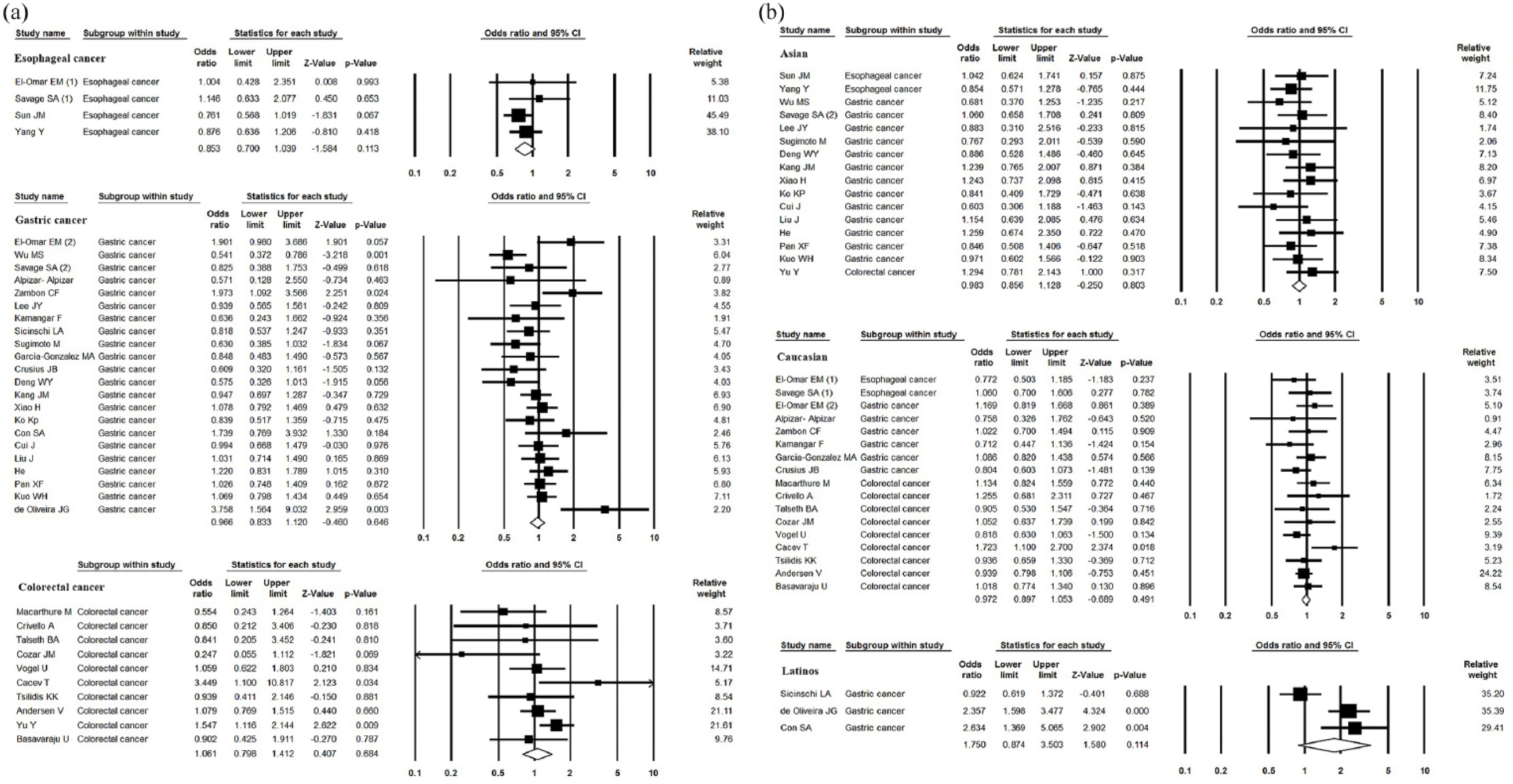
Forest plot of subgroup analysis by ethnicity and cancer type on the association between IL-10 -592 A/C polymorphism and GIT cancer risk. (a) Recessive model of cancer type subgroup (AA vs. CA+CC). (b) Dominant model of ethnicity subgroup (AA+CA vs. CC).
Sensitivity analysis, publication bias and test of heterogeneity
Sensitivity analysis was tested to determine the influence of each single study on the pooled OR by removing individual studies, and the corresponding pooled ORs in all genetic models were not materially altered.
Begg’s funnel plot and Egger’s linear regression test were conducted to evaluate the publication bias in this meta-analysis. The shapes of the funnel plots did not display any evidence of an obvious asymmetry in all genetic models, and the results of Egger’s test showed that there was no publication bias for overall analysis in the current meta-analysis (homozygous model: P = 0.881; heterozygous model: P = 0.396; dominant model: P = 0.501; recessive model: P = 0.986; allelic model: P = 0.627).
Significant heterogeneity existed in the allelic model (PQ< 0.001; I2 = 56.2%), the homozygous model (PQ = 0.007; I2 = 40.3%), the dominant model (PQ = 0.026; I2 = 34.0%), and the recessive model (PQ = 0.002; I2 = 46.0%). Hence, the random-effect model was used to calculate CIs. In contrast, the heterozygous model (PQ = 0.198; I2 = 16.3%) did not present significant heterogeneity, so ORs were pooled according to the fixed-effects model.
Discussion
Cytokine polymorphisms play a key role in genetic functioning of the host. 48 IL-10 is a strong pleiotropic indeed, which plays a significant role in immunity and inflammatory responses. 49 Several studies reported that function of IL-10 might accelerate or prevent tumor growth, but with due consideration to anti-inflammatory and immune regulatory characteristics, it is believed that IL-10 might interfere with the process of tumor cells fleeing from the supervision of the immune system.32,50 Several polymorphisms were identified in the promoter of the IL-10 gene, the most important of which are −592, −819, and −1089. They may influence transcriptional activity of this gene. 51
In the present meta-analysis, the role of IL-10 -592 promoter polymorphism was evaluated in GIT susceptibility. Overall, the results indicated that IL-10 -592 promoter polymorphism may not be correlated with the risk of GIT cancer. Also, no significant association was found when stratified by HWE, ethnicity, source of controls, and cancer types. Publication bias and sensitivity analysis were checked and both of Begg’s funnel plots and Egger’s test results represented no publication bias. In addition, a sensitivity analysis was performed and attested the reliability of the findings.
Just like every other meta-analysis, limitations of the current study should be considered. First, a total of 36 case-control studies with 8069 cases and 13,089 controls were included in this meta-analysis. Only three studies about Latinos were acquired; consequently, an insufficient number of studies may decrease the statistical power to evaluate the ethnicity effect. In the subgroup analysis of cancer type stratified by ethnicity and source of controls, the number of studies and the sample size in esophageal cancer are not large enough, and the studies based on Asian race and hospital-based case-control studies were very limited in colorectal cancer. The number of studies may be insufficient to identify slight associations, and the findings should be interpreted with caution. Second, carcinogenesis is a multi-factorial and complicated process, which encompasses the interaction of environmental and genetic factors. There is a high probability that the role of the polymorphism is covered or diminished due to other gene–gene and gene–environmental interactions. This study has only examined one polymorphism of the IL-10 gene. It needs further mention that it disregards the effect of other genes, environmental factors, and predisposed polymorphisms. Various polymorphism loci in promoter of the same gene may change the OR values. Third, this meta-analysis was according to unadjusted OR estimates because, in most of the included original studies, some factors like age, sex, family background of cancer, smoking, alcohol consumption, etc., have not been taken into account.
Despite its limitations, the present meta-analysis also had its own advantages. This is the first meta-analysis consisting of 36 case-control studies investigating the association of IL-10 -592 A/C polymorphism with GIT cancer susceptibility, and the included studies are exactly related to the association between IL-10 -592 A/C polymorphism and GIT cancer risk.
In conclusion, it can be said that our results found no significant association between IL-10 -592 promoter polymorphism and the risk of GIT cancer. It is recommended that more deliberate and well-designed studies with larger samples be conducted to investigate the relation between this polymorphism and the risk of GIT cancer extensively. Moreover, the potential impact of gene–gene and gene–environment interactions on GIT cancer development needs further assessment in future studies.
Footnotes
Acknowledgements
I would like to express my very great appreciation to Dr. Mojgan Afifezade for her valuable and constructive suggestions during article writing.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
