Abstract
The manuscript analyzes potential pre-mRNA biomarkers of Myasthenia gravis (MG) in thymoma in silico. GSE11967 data set and platform5188 from the Gene Expression Omnibus database apply for data preprocessing, normalization, and quality control. Quality metrics indicated high overall data integrity, with no significant outliers or batch effects detected. Differential expression analysis (DEG.) uses the limma package in R. We compared thymoma samples to normal thymus tissue to identify DEGs. The significance criteria are adjusted p-value <0.05 and a |log2 fold change| > 1. Functional enrichment and pathway analysis, ontology analysis, and KEGG pathway analysis further investigate the potential underlying biological processes. Despite the extensive use of gene expression profiling for identifying potential biomarkers and therapeutic targets, this study identifies DEGs ENSG00000112345 and ENSG00000234567 and pathways like hsa04110, hsa03013 and hsa04115 in thymoma with MG compared to normal thymus tissue using the GSE11967 dataset Plots like UMAP, Boxplot, Expression density and Mean variance demonstrate differential expression in disease and control group. The GSE11967 data set shows the presence of significant DEG and pathways in thymoma-associated MG tissue, compared to healthy tissue. A broader and integrative approach is needed to understand the complex expression biomarkers in thymoma in MG patients and other regulatory mechanisms that may contribute to the disease by multi-omic approaches.
Introduction
The landscape of biomedical research is affected by diverse factors ranging from literature review, theoretical frameworks, technological advancements, and organizational dynamics.1,2 Developing technology and artificial intelligence facilitate gathering and understanding information, enabling better clinical decision-making processes where societal norms sharp attitudes towards innovation and advancing biomedical knowledge.1,2 Moreover, organizational hierarchies and power structures influence the implementation and adoption of novel molecular signaling insights. 3 Long-chain non-coding RNA contributes to complex regulatory networks driving cancer tumorigenesis and disease progression like PI3K/AKT (Phosphoinositide 3-Kinases/Protein Kinase B), Wnt/β-Catenin (Wingless-Related Integration Site Beta-Catenin), TGF-β (Transforming Growth Factor Beta), and NF-κB (Nuclear Factor Kappa B) signaling.4,5 The research landscape is influenced by additional factors such as education, skills training, knowledge sharing, and policy frameworks, affecting the applicability of scientific knowledge.6,7 Understanding patient outcomes in technical and organizational contexts is a must, while navigating through the complex interplay of risk, uncertainty, and change. A culture of enthusiasm and motivation requires a concrete effort aligned with organizational structures to foster embracing new knowledge and technologies in clinical and scientific settings, enhancing business operations.8,9 In silico methodology enables fast data analysis, leveraging bioinformatic knowledge, extracting meaningful insights from various data sets, accessing literature, and facilitating informed decisions. 10 Appropriate utilization of hardware, knowledge, attitudes, and skills, supported by effective management and IT (Informational Technology) competence, depends on the successful implementation into scientific and clinical organizations, harnessing the power of data-driven insights to drive innovation and improve patient outcomes. 11 For example, MALAT1 (Metastasis-Associated Lung Adenocarcinoma Transcript 1) expression influences cancer development and progression biology, whereas therapeutic targets address lncRNA (Long Non-Coding RNA) to improve the disease outcome. 12 32 factorial design evaluated the effect of independent variables over particle size and the entrapment efficiency of nanoparticles, providing an idea that use of simvastatin nanoparticles with an anticancer agent can be used for patients who have cancer and hyperlipidemic conditions. 13 Computer simulations and models apply for in silico analysis, which represents analog to silicon material used to make computer chips. 14 In silico analysis is applied in diverse fields of biology, chemistry, and medicine to evaluate and forecast multiplex biological actions within the organism, chemical interactions, and drug responses without the necessity of physical experiments. 15 In silico analysis includes several processes like data analysis, modeling and simulation, drug discovery and development, genomics and proteomics, and system biology. 15 In order to identify patterns and make predictions, large data sets of genetic sequence and clinical trial results are processed and analyzed to enable stimulation of physiological processes of biological systems of specific organisms and chemical reactions with mathematical models. 15 Potential drug candidates, interaction with targets, and structure optimization are identified by in silico methodology. 15 Complex interactions within biological systems are predictable by integrating various biological data like genomic and proteomic to understand gene function, protein interaction, and regulatory networks within diverse signaling pathways. 15 For example, modification of melatonin receptors can have an anticancer effect affecting cell proliferation, epigenetics, metastasis, angiogenesis, cell energy, and immune invasion. 16
Biological markers are measurable indicators of an organism’s biological state that assess the disease progression, monitor the effect of a treatment, and understand the biological processes and signaling network within the organism and specific cells.17–20 Biomarkers can be any molecule, gene, protein, cell, or entire system, like temperature and blood pressure, depending on the researcher and the level of exploration.17–20 Molecular, imaging, physiological, and pathological markers divide based on characteristics and type, for example, molecular biomarkers like DNA (Deoxyribonucleic Acid), RNA (Ribonucleic Acid), protein, lipids, metabolites, and specific gene mutations.17,20 Other biomarkers divide based on imaging technologies, and they include MRI (Magnetic Resonance Imaging), CT (Computed Tomography), and PET (Positron Emission Tomography) scans where the size and shape of a tumor represent a marker for cancer progression, measurable changes of physical state like blood pressure, heart rate, and body temperature; tissue samples and biopsies to analyze specific metabolite od different level, like amyloid plaques in the brain of Alzheimer disease.17,19,20 Depending on the purpose, biomarkers can be diagnostic, prognostic, predictive, pharmacodynamic, and safety biomarkers.17,18 Some will enable us to identify the presence and course of a disease, predict how the patient will respond to treatment, and indicate the positive effect or potential adverse effect.17–20 Precursor messenger RNA (pre-mRNA) is a significant intermediate molecule in the messenger RNA (mRNA) biogenesis.21–23 The biological significance of pre-mRNA molecules lies in gene expression regulation, pre-mRNA processing, alternative splicing, quality control, regulation of mRNA stability and translation, disease association, and expression of therapeutic potential.21–23 Integrative multi-omics approaches are necessary to understand the complex gene expression in tumor microenvironment. The tumor microenvironment contains diverse cell types, signaling pathways affecting disease progression, and therapy outcomes. A holistic understanding of proteins, metabolites, and epigenetics in tumor microenvironment will help unravel genetic modifications lying between.21–28 For example, pre-mRNA and mRNA play a role in the development, differentiation, cell proliferation, apoptosis, and metabolism of health and disease.22,23 Diseases in which they involve represent cancer, cardiovascular disease, and neurological disorders.21–23 Understanding pre-mRNA processing and regulation enables insights into disease mechanisms and potential therapeutic targets.21–24 Moreover, mRNA-based therapies include mRNA mimicking to restore function, inhibit overactive mRNA, or even modulate activity in the therapeutic context.21,22 An emerging area of research includes the involvement of mRNA and pre-mRNA in Myasthenia gravis (MG) associated with thymoma in the regulation of immune response.25–28 Aberrant alternative splicing in the Acetylcholinesterase (AChE) gene is present in patients with MG-thymoma, along with dysregulation of splicing factors SF2/ASF (Splicing Factor 2/Alternative Splicing Factor) and CS35 (Monoclonal antibody) can lead to immune dysregulation. 26 mRNA are key molecules in regulating the function of immune cells like T and B lymphocytes and AChRs and other neuromuscular junction components. 26 Several specific mRNAs involved in disease mechanism and progression in MG include acetylcholine receptor (AChR) subunits encoding alpha (α), beta (β), delta (δ), and epsilon (ε) subunits of the AChR affecting neuromuscular transmission, muscle-specific kinase (MuSK) causing severe muscle weakness, Low-density lipoprotein receptor-related protein 4 (LRP4) affecting neuromuscular transmission and AChE impacting neuromuscular signaling.26,27 In thymoma, the most significant mRNA are autoimmune regulator (AIRE) promoting central tolerance, thymic stromal lymphopoietin (TLSP) affecting T cell development, forkhead box protein p3 (FOXP3) impairing Treg function, cytokines, and chemokines like interleukine (IL) IL-25 and IL-33. 28 Another study involves the synthesis of Ch-TGP (chitosan-Tamarind Gum polysaccharide) polyelectrolyte complex stabilized nanoparticles of simvastatin against MCF-7 cells, evaluating antitumor efficacy with further theoretical modeling for the perfect designing system. 13 CXCL12 (C-X-C Motif Chemokine Ligand 12) influences the tumor microenvironment, immune cell recruitment, and immune responses, affecting tumor progression and autoimmunity, as all the rest are representative. 28 mRNA influence on the development of MG and thymoma by autoimmunity and immune cell regulation attacking neuromuscular junction, abnormal microenvironment affecting mRNA expression, regulation of inflammation, and cytokine production.25–28 Based on the proposed functions and influence in the disease development, pre-mRNA and mRNA serve as a diagnostic and therapeutic implication like potential biomarkers, therapeutic targets, and personalized medicine following disease monitoring and progress, restoring normal mRNA function and reducing autoimmunity and tumor growth leading to personalized treatment plans, optimizing therapeutic efficacy and minimizing side effects.25–28 These molecules are crucial targets for further research with potential applications in diagnosis and therapy.25–28
Fatigue of skeletal muscles and weakness represent the main characteristics of autoimmune neuromuscular disorders such as MG.13,25–28 Usually, MG is associated with thymoma, a tumor originating from epithelial cells of the thymus gland. Thymoma and MG connect with autoimmune mechanisms and immune dysregulation.13,25–28 Autoantibodies are produced against AChR at the neuromuscular junction, leading to a damaged transmission of nerve impulses to muscles.13,25–28 When the thymus is abnormal, an autoimmune response (autoantibodies among which target AChRs) appears in MG because the thymus gland plays a crucial role in immune system cell development like T-cells.13,25–28 Patients with MG have thymoma in 10%–15%, while 30%–50% of patients with thymoma develop MG as a disease.13,29 Clinical symptoms are different for MG and thymoma.13,25–31 The muscle weakness in MG worsens with physical activity and improves with rest.13,25–31 Depending on the disease severity, the most commonly affected muscles include those controlling eye and eye movement (ptosis and diplopia), facial expression, chewing, swallowing, and speaking, but in severity cases affects respiratory muscles, leading to myasthenic crisis.13,25–31 On the other hand, thymomas can be asymptomatic and symptomatic. The diagnosis of asymptomatic cases is confirmed by accident with imaging techniques in most cases, while symptomatic cause pain in the chest, cough, breath shortening, or superior vena cava syndrome (swelling of face, neck, and upper chest) due to obstruction. 32 Diagnosis of these two interconnected diseases is different. MG includes several steps like clinical evaluation and history, electromyography (EMG) and nerve conduction studies, serum assay for anti-AChR antibodies or anti-MuSK antibodies as well ass endophonium test in the diagnostic procedures of MG.13,25–32 The biopsy, and surgical resection for histopathological confirmation with imaging studies such as chest X-ray, CT scan, and MRI apply to identify the presence of thymic mass.13,25–32 Treatment in MG gravis includes medications, plasmapheresis, and intravenous immunoglobulin, while for thymoma includes surgical resection, radiation therapy, and chemotherapy.13,25–32 AChE inhibitors like pyridostigmine improve neuromuscular transmission, and immunosuppressive therapies like corticosteroids and azathioprine reduce autoimmune activity in MG.13,25–33 The exacerbation pharmapheresis and intravenous immunoglobulin apply as a treatment in MG disease to reduce fast-circulating antibodies in myasthenic crisis.13,25–33 Mangifera indica (Mango) extracts from diverse plant parts represent a promising breast cancer therapy affecting diverse signaling pathways, molecules, and cell cycles along with the fabrication of nanofiber matrices. 34 Removal of the thymus (thymectomy) is the primary treatment of thymoma despite the presence or absence of MG disease since it represents a high risk of malignancy where postoperatively in the case of disease invasiveness or not of the thymoma radiation therapy applies. In the cases of advanced or metastatic thymomas, chemotherapy is in place.13,26–34 The prognosis of diseases like MG and thymoma is different. The prognosis is significantly improved in patients by controlling the symptoms, leading to normal lives, with modern treatments, while thymectomy in MG can lead to remission or improvement in MG symptoms.35,36 Depending on the stage and invasiveness of the tumor, the prognosis of thymoma depends.35,36 In the early stage of the disease, patients with surgical resection have a good prognosis, and in the advanced stage, the prognosis requires additional treatment.35,36
Population-based epidemiological cohort studies determine the incidence and prevalence of thymoma-associated MG in diverse geographic regions, revealing variability and focusing on the general incidence and prevalence. 37 The overall incidence, prevalence, geographic variations, and study examples for MG and thymoma vary depending on diverse factors like study quality, geographic location, and diagnostic criteria, where high heterogeneity limits precise interpretation and disease management. 37 The overall incidence of MG range from 1.7 to 21.3 cases per million persons per year, where the prevalence of MG ranges from 15 to 179 cases per million persons, with poll estimates like 77.7 cases per million with substantial heterogeneity of precise interpretations.38,39 Incidence rates of MG and thymoma-associated cases present literature geographic variability. 38 For example, incidence rates in Europe range from 4.3 to 18.0 permillion persons per year, whereas the data show less comprehensiveness for Asia, Africa, and South America regions due to fewer studies. 38 For example, studies indicate a higher prevalence of childhood MG onset in the Asian population compared to Western, suggesting regional differences in genetic susceptibility and influence of environmental factors affecting disease onset. 38 Northern and Southern Europe, Northern America, and Canada have excellent incidence and prevalence rates. 38 Northern Europe shows a smaller incidence rate of 4.6 million persons per year, while Southern Europe has a greater incidence of 4.9 million persons per year, whereas data from the Asian population indicate a higher proportion of childhood-onset MG. 38 Moreover, the Swedish study showed 55 studies from the literature between 1950 and 2007 years where they included 1.7 billion population-years estimating incidence rate (IR), prevalence rate (PR), and mortality rate (MR) for all MG estimated pooled IR: 5.3 per million person-years, range: 1.7 to 21.3; estimated pooled PR: 77.7 per million persons, range 15 to 179; MR range 0.1 to 0.9 per millions person-years. AChR MG eIR: 7.3, range: 4.3 to 18.0; MuSK MG IR range: 0.1 to 0.32. Marked variation persisted between populations studied with similar methodology and in similar areas. 38 Thymoma-associated MG tends to be more frequent in the German population, where these patients have more severe symptoms and higher antibody titters like titin, which is more prevalent in thymoma-associated cases. 40 Denmark study reported the incidence and prevalence of MG, providing detailed data on thymoma-associated cases like the significant association between thymoma and more severe MG symptoms. 39 Thymoma-associated MG occurs at any age, but it is usually present in late-onset, >50 years of age. 40 Thymoma-associated MG affects both genders and presents variation in gender distribution depending on a specific population. Male predominance, in some females, and some equal distribution between male and female participants of the research population group depending on the year of research.40–42 However, the newest study confirms that more women than men develop the disease, the ratio being three to two. 43 Saudi population showed that MG occurrence in thymoma patients is more likely to occur at a younger age, higher TNM (A cancer staging system that describes the extent of cancer’s spread) classification, and advanced MASAOKA (A staging system for thymomas based on tumor size and spread) stage. Although no significant association exists between MG and complications and mortality, MG exhibited a protective role in thymoma by providing a lower recurrence rate and longer survival duration. 44 Moreover, thymoma-associated MG may be associated with various comorbidities, including autoimmune diseases such as autoimmune thyroid disease, rheumatoid arthritis, systemic lupus erythematosus or paraneoplastic syndromes beyond MG like pure red cell aplasia or hypogammaglobulinemia. 45 Patients with thymoma-associated MG may have severe symptoms at diagnosis compared to those without thymoma in MG. 45 Compared with non-thymoma-associated MG, Thymoma-associated MG patients have a significantly shorter disease duration from symptom onset to the crisis (17.95 ± 40.9 vs 51.31 ± 60.61 months, p < 0.0001) and a higher proportion of MG Foundation of America Classification IVa as the initial onset clinical classification (6.67% vs 0, p = 0.0205), and a prolonged hospital stay (39.24 ± 22.09 [6–111] vs 33.2 ± 23.42 days [7–120]; p = 0.0317) during the crisis. Incidence rates were lower in the first years of the COVID-19 period. The previous trend was confirmed when the pandemic ended. 46 Patients with thymoma but without MG had a high probability of developing PMG. Although the incidence of Pure Red Cell Aplasia (PMG) was very low, thymectomy could not prevent the occurrence of MG. Preoperative seropositive AChR-Ab level, open thymectomy, non-R0 resection (surgical term indicating that no cancer cells are seen at the margins of the removed tissue), WHO type B (classification of thymomas based on histological features by the World Health Organization), and postoperative inflammation were risk factors for PMG. 47 10 years follow-up study in the Beijing population revealed a higher prevalence rate of autoimmune diseases, especially rheumatoid arthritis, in MG patients after thymectomy. 48 In a Spanish study, Thymoma-associated MG patients had more severe myasthenic symptoms and worse prognosis. 49 Thymoma recurrence was frequently associated with transient worsening of MG, but the long-term prognosis did not differ from nonrecurrent thymoma. 49 Patients with nonresectable thymoma tended to present severe forms of MG. 49 Reactive carbonyl species are associated with cancer and neurodegenerative diseases. 50 Rubiadin is a promising natural anthraquinone for new drug discovery and development in preclinical, and clinical trials, bioavailability, permeability, and administration of safe doses. 51
Few recent expression profiling studies of genome-wide mRNA expression profiling in thymoma tissues with or without MG identified dysregulated mRNA associated with the disease. 52 The Long Non-Coding RNA 00452/MicroRNA/Carbohydrate Sulfotransferase 4 (LINC00452/miR-204/CHST4) axis is involved in the progression of thymoma-associated MG, highlighting the role of specific mRNAs in the disease’s pathogenesis regulating thymic regulatory T cells in the progression of MG, confirming the competitive endogenous hypothesis proposing a crosstalk between mRNA and non-coding RNA (ncRNA) by competitive binding to the microRNA. 52 Other studies identified dysregulated miRNA-mRNA like miR-548k, which plays a crucial role in MG by regulating CXCL13 (C-X-C Motif Chemokine Ligand 13) expression included in the immune response. 53 Dysregulated miRNAs such as miR-30 family members and miR-188-5p have been reported, showing altered expression patterns in thymoma tissues from MG patients. 53 RNA sequencing (RNA-seq) is a powerful tool for revealing pre-mRNA expression patterns and identifying aberrant spliced transcripts in thymoma tissues. RNA sequencing techniques include sample preparation, RNA extraction, library preparation, and sequencing.52,53 Tissues from patients with or without MG and total RNA extraction include pre-mRNA with standard RNA extraction kits, enabling high-quality RNA for sequencing technique.52,53 Library preparation usually includes mRNA enrichment or rRNA (Ribosomal RNA) depletion, cDNA (Complementary DNA) synthesis, fragmentation and adapter ligation, and PCR (Polymerase Chain Reaction) amplification.52,53 mRNA molecules using Poly A selection or rRNA depletion method enriches focusing on coding RNA to avoid abundant ribosomal RNA for comprehensive analysis to gain cDNA, which is fragmented and sequenced.52,53 High throughput sequencing like Illumina HiSeq or NovaSeq enables detailed data analysis dependent on quality control, alignment and mapping, transcript assembly and quantification, and splicing analysis.52,53 Read quality assessment, trimming, and filtering enable further alignment to the reference genome, mapping reads to transcript along with transcript assembly, quantification, identification of splicing events, and data information visualization. Tools like FastQC, Trimmomatic, Cutadapt, STAR, HISAT2, StringTie, Cufflinks, HTSeq, and featureCounts along with transcript level quantification, software DEXSeq, MATS, SUPPA2, and IGV enable in-detail analysis, assessing to quality of raw sequencing reads to identify and quantify novel splicing events in MG and thymoma associated and non associated with MG.52,53 Examples of studies and findings include miR-548k impacting the CXCL13 gene in MG and other aberrant splicing genes involved in immune regulation and cellular signaling.52,53 The aberrant activation of the JAK-STAT (Janus Kinase/Signal Transducers and Activators of Transcription) pathway reports in various disease states, including inflammatory conditions, hematologic malignancies, and cancer, indicating the overall significance in pathogenesis and therapy of thymoma in MG affecting miRNA interaction leading to changes in interaction to mRNA molecules. 54
Functional studies enable validation of dysregulated mRNA and pre-mRNA using in vitro and in vivo models of MG-associated thymoma, focusing on understanding molecular interaction of disease pathogenesis and discovering therapeutic targets.55–57 Dysregulated mRNA is usually validated by in vitro and in vivo studies.55–57 In vitro studies mRNA knockdown and overexpression in cell lines with transfection with mRNA mimicking inhibitors as a technique elucidate roles in cell proliferation, apoptosis, and other cell functions as well as luciferase reported assay that confirms direct interaction between mRNA and target mRNA validating miRNA binding and regulation activities.55–57 In vivo studies are validated with xenograft models and genetically modified mice.55–57 Usually, dysregulated mRNA expression in human thymoma cells is injected into immunodeficient mice to form a tumor-like mRNA for gene MYC, TP53 (Tumor Protein P53), BCL2 (B-Cell Lymphoma 2), EGFR (Epidermal Growth Factor Receptor), KRAS (Kirsten Rat Sarcoma Viral Oncogene Homolog), VEGF (Vascular Endothelial Growth Factor), and AKT1 (AKT Serine/Threonine Kinase 1).55–57 The comprehensive genomic and transcriptomic analysis identifies dysregulated mRNA before studying them in xenograft modes.55–57 Focused therapy of thymoma in MG disease includes siRNA (Small Interfering RNA) and antisense oligonucleotides, gene editing with CRISPR-Cas9 (Clustered Regularly Interspaced Short Palindromic Repeats associated protein 9) of AIRE or tumor-specific genes, mRNA vaccines and immune modulations encoding autoantigen peptides and regulatory cytokines, direct mRNA-based therapeutics encoding tumor suppressor and immune-modulating proteins, combination therapies with immune checkpoint inhibitors and thymectomy as well as specific targets for thymoma and MG.55–57 Clinical trials and experimental study research are still ongoing for specific mRNA-based therapies.55–57 Targeting specific mRNA reduces the expression of proteins involved in tumor growth and autoimmune response.55–57 Knock-out mRNA determines relevance to thymoma development and MG progression.55–57 On the other hand, aberrant spliced transcripts are validated in vitro and in vivo studies.55–57 In vitro studies include splicing reported assays, RNA Sequencing (RNA-Seq), and quantitative PCR (qPCR).55–57 Exon-intron regions of interest are transfected into thymoma cell lines to identify aberrant splicing events.55–57 RNA-Seq identifies deferentially spliced transcripts, whereas qPCR validates the findings by specific spliced isoforms.55–57 In vivo studies include mouse models, splicing modulators along functional assay in mice assessing the impact of splicing changes in thymoma and MG to identify potential therapeutic interventions that correct aberrant splicing and investigate the physiological effect of specific splicing variants involved in cell populations, neuromuscular junction, integrity and overall muscle function in mouse model organism.55–57 Molecular mechanisms underlining the effect of dysregulated mRNA or pre-mRNA on thymic epithelial cell function, immune dysregulation, and neuromuscular junction pathology are present in the literature.55,57 In cell proliferation, survival, and apoptosis, miRNA like mir-21 targeting Phosphatase and tensine homolog (PTEN) to activate PI3K/AKT pathway and miR-125a-5p affecting p53 (protein encoded bt TP53) and BAK1 (BRI1-associated kinase 1) as well as FOXN1 (Forkhead Box N1) leading to abnormal cell growth thymic architecture disruption leading to thymoma development.55,57 Thymoma tissue associated with MG usually includes miR-155 enhancing pro-inflammatory cytokines, disrupting the balance of regulatory T cells and effector T cells contributing to autoimmunity, miR-146a tasting ass negative regulator of the NK-kB signaling leading to chronic inflammation and autoimmunity interacting with TRAF6 (TNF Receptor-Associated Factor 6) and IPAK1 (Ixodes p21-activated kinase).55,56 Mutations in the splicing factor SF3B1 (Splicing Factor 3b Subunit 1) cause aberrant splicing of immune regulatory genes like T cell receptor signaling, leading to an increased risk of autoimmunity. 57 In neuromuscular junction pathology, miRNAs involved in synaptic function and alternative splicing is miR-132 targeting AChE involved in Acetylcholine breakdown at the synaptic cleft where disruption leads to impaired synaptic transmission and muscle weakness, miR-206 involved in muscle regeneration and repair where down-regulation leads to progressive muscle weakness as well as Connexin 43 and Utrophin involved in neuromuscular maintenance.55,57 Moreover, aberrant splicing of agrin and LRP4 (Low-Density Lipoprotein Receptor-Related Protein 4) leads to disruption of synaptic stability and signaling. 56 The scientific literature confirms pre-mRNAs and dysregulated mRNAs as diagnostic biomarkers and therapeutic targets. For example, regulation of HMGB1 (High Mobility Group Box 1) in thymoma cells affects T cell proliferation, leading to a change in the occurrence of immune abnormalities in MG-associated thymoma. 58 So far no new driver gene was identified in advanced thymomas, but arm-level chromosomal copy number variations were detected in patients with MG in Tokyo. 59 This information indicates the influence of genetic and environmental factors contributing to disease initiation, progression, and development. In silico analysis represents a promising tool in biomedical research, allowing the unraveling of intricate molecular mechanisms underlining the disease using analysis of complex data like pre-mRNA as potential biomarkers in disease pathology. Pre-miRNA is crucial in gene expression regulation between DNA transcription and protein synthesis, which dysregulation leads to disease. Aberrant gene splicing of AChR, MuSK, Titin, LRP4, RAPSIN, and RYR1 genes lead to the development of thymoma in MG disease. These examples illustrate the complexity of pre-mRNA processing and its role in disease development and progression. Additionally, alterations in pre-mRNA processing of genes involved in cell proliferation and apoptosis influence disease progression and clinical outcomes. The intricate interplay between molecular pathways involved in thymoma development and the pathogenesis of MG remains an area of active investigation.
In this study, we aim to leverage in silico analysis techniques to explore the potential of pre-mRNA molecules as biomarkers for MG in thymoma. By interrogating publicly available datasets and employing advanced computational methods, we seek to identify dysregulated pre-mRNA signatures associated with disease progression and outcome. Understanding the molecular landscape of thymoma-associated MG could provide valuable insights into disease mechanisms and facilitate the development of novel diagnostic and therapeutic strategies. This can establish a significant relationship among individual reliability, internal consistency, and discriminant validity of analyzed data, demonstrating satisfactory validity and reliability before examining the relationship within the clinical model studies.
Methodology
Description of the tissue with specific accession number.

Transcript group of analyzed samples with accession number.

Results
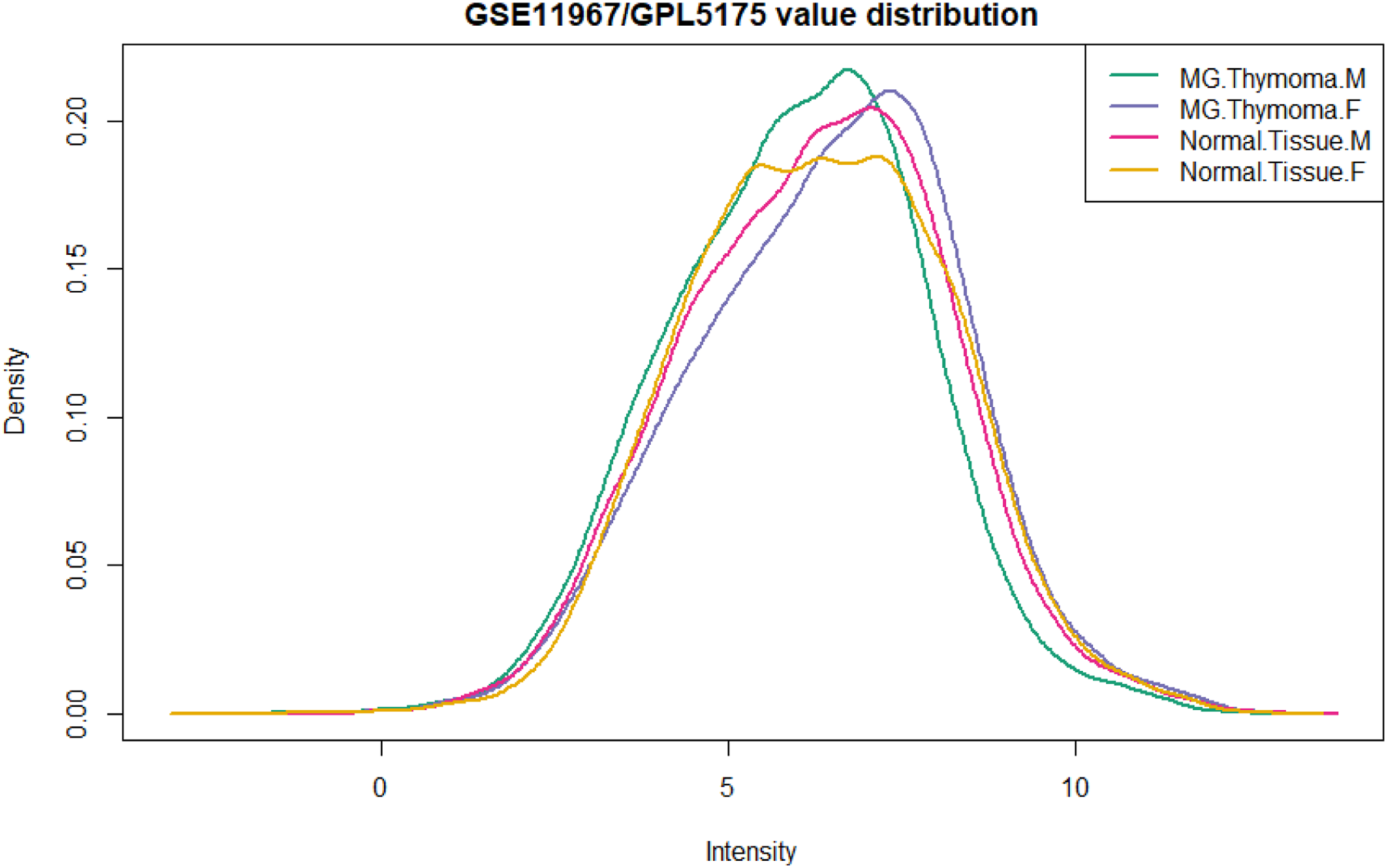
Results ensure that the findings are logically structured and easy to follow for the readers. Results directly address the research objectives outlined in the introduction section. Findings contribute to answering the research hypothesis. Tables, Figures, and Graphs effectively illustrate key findings appropriately labeled and explained in the text. The depth of interpretation is present for each result. The significance and implications of the findings are adequately discussed, including unexpected or contradictory results. Results are consistent with the methodology described in the previous section. Any potential deviations or unexpected outcomes are addressed, explained, and ensured. Limitations of the study are appropriately acknowledged and discussed. Future directions are suggested based on the obtained findings. GEO2R analysis compares the data of the control group and the group with the disease to identify differentially expressed genes between the experimental conditions. In this case, the results are not present. R programming language is the analysis basis with the limma R package (Linear Models for Microarray Analysis) to evaluate differentially expressed genes. We obtain results in a fast and efficient manner. Four summary datasets for each sample are available in Boxplot data, presented in Graph 1. From the analysis, UMAP (Uniform Manifold Approximation and Projection) applies as a dimensional reduction technique for visualizing that the samples are interconnected. Graph 2 shows the results indicating the presence of a control group and a group with the disease and their relationships. We generated a box plot by R to show the distribution of sample values and assess whether the data are for differential expression analysis. Based on these results, they apply for differential expression analysis. We perform expression density analysis using limma in R to see the distribution values of the analyzed samples, Graph 3. Moderate t statistic quantile-quantile (q-q plot) generates using the limma package. Points are present in the amount of sample data relative to the theoretical model of the t test distribution. The ideal case is a straight line, which indicates that the values of the moderated t-statistic test follow the theoretical prediction of the distribution. Analysis of the q-q plot reveals a robust statistical framework for identifying deferential expressed RNA molecules. Summary set of data for each sample in Boxplot data. Graph 1. Enables an understanding of the distribution, central tendency, and variability of the summarizing data in the boxplot for each sample. Representation of the connection between samples. Graph 2. Enables and understands the structure and meaning of the connection between samples, highlighting relationships, similarities, and features within the data. Expression density plot distribution values of analyzed samples. Graph 3. Present and understanding of the distribution and density of expression values across samples.

Discussion
The fast-paced development of electronic communication and the widespread availability of public databases such as GEO led to a revolution in biomedical research by providing vast data for analysis. However, there is a need for careful consideration of the regulatory framework to ensure data integrity and reliability. Previous studies established concrete standards for data analysis and interpretation with GEO2R databases, highlighting the importance of standardization, but challenges persist in data quality, consistency, and ethics. 63 Data regarding thymoma associated with MG underlining molecular mechanism of the disease is paramount, correlating with the same biomarkers in diverse diseases, making it interconnected with others and indicating the influence of molecular mechanisms in cancer like pre-miRNA splicing in the process, modulating gene expression patterns influencing cell process, signaling pathways and development processes. The study aimed to utilize in silico method with a publicly available database of thymoma associated with MG to asses pre-mRNA interaction for pathological implication. This study shed light on pre-miRNA relevance to disease pathology, the development of novel diagnostic tools, therapeutic targets, and personalized treatment approaches for patients with this disease condition, enabling more focused and concise information regarding the proposed topic with a clear rationale for the study. Accessing publicly available data involves more than just accessing and analyzing the data. Data privacy, consent process, adequate citing of sources, and applying ethical guidelines must be present in the researcher doing data analysis of this type. Cost efficiency, speed, and the ability to test hypotheses that are not experimentally represent the advantages of in silico studies. Researchers can conduct preliminary studies before moving into in vitro or in vivo studies, enabling faster research and development of their hypothesis with specific biomarkers and model system organisms.
Personalized medicine treatment will use biomarkers to tailor treatment plans based on individual differences. 64 Personalized medicine can lead to the development of complex regulations due to malpractice claims and deviations in treatment. 64 Because of that, there is a need to establish standards for evidence-based guidelines of molecular diagnostic and target therapies. 64 Liability risk can arise from failure to incorporate molecular data appropriately into the treatment decision-making process, indicating the significance of robust risk management strategies for the best practice in personalized medicine and patient care. 64 In personalized medicine, legal elements have national and international level significance. 64 Healthcare regulatory framework at the national level can vary depending on the adoption and implementation of personalized cancer care strategies like data privacy, informed consent, liability standards, and reimbursement of policies for molecular testing and target therapies. 64 Common guidelines and standards exist at the international level to harmonize ethical and legal efforts of personalized medicine through global collaboration, sharing knowledge, and meeting cross-border regulation challenges that can arise. 64 Individual molecular profiles enable tailored treatment strategies in personalized medicine, reshaping the landscape of cancer care and significantly impacting healthcare policy policy-making processes and evidence-based guidelines. 64 Issues that can arise in policymakers are equable access to molecular profiling technologies, reimbursements, and integration of molecular data for decision-making algorithm processes. Biomarkers will be valuable prognostic and predictive indicators for treatment selection and patient care. 64 Molecular profiling and targeting therapies meet the complexity of informed consent in personalized medicine. 64 The patients taking part in the study must receive comprehensive information regarding the potential risks, benefits, and uncertainties associated with the targeted treatment and molecular testing. 64 Ongoing consent processes are necessary because of the dynamic nature of molecular data. Patient autonomy is significant throughout the consent process for clinicians. 64 The most significant is fertility preservation for patients undergoing personalized medicine approaches because they represent a high risk. 65 Oocyte or sperm cryopreservation, as advancements in reproductive technologies, enable the preservation of fertility in patients undergoing cancer treatment. 65 However, it can represent a barrier for some patients due to service access and insurance coverage. 65 This lines the need for enabling equal access to fertility preservation resources. 65 Improved diagnostic and therapeutics for cancer treatment lead to fertility preservation as a moral obligation in a free society. 65
Benefits and challenges exist for in silico analysis, enabling the identification of effective drugs without ethical or financial barriers, leading to faster detection and eradication of disease. In the case of the chosen data set in this manuscript, GSE111967 applies to analyze significant pri-miRNA and miRNA biomarkers. Data integrity enables secure data storage and transfer with the possibility of electronic signatures supported by non-coding RNA in personalized medicine of cancer research. Broaden implications are emphasized by recognizing public interest, morality, national and international laws, secure data management, fostering innovation, and improved public communications during the COVID-19 pandemic, making it indispensable in clinical practice. The significance of the study findings in personalized/precision medicine, molecular classification, and non-coding RNAs depends on the comparative analysis of current research data. Malpractice litigation and medical missus of legal practice exist, although different federal, state, and professional society limitations can lead to ramifications. 66 New drugs and therapies development enables a more efficient and targeted approach to disease management by utilizing biomarkers. Pre-mRNA represents a crucial intermediate in the mRNA biogenesis pathway, making it essential for functional mRNA that regulates gene expression, having a significant implication in biological science and the development of specific mRNA-based therapeutic strategies. MG and thymoma represent interrelated conditions where proper diagnosis and treatment can improve patients’ disease outcomes. In thymoma-associated MGs, dysregulation of various non-coding RNAs, including lncRNAs and miRNAs, has been reported. These molecules can interact with mRNA transcripts and influence gene expression, contributing to the pathogenesis of the disease. Studying the interactions between lncRNAs, miRNAs, and mRNAs can provide insights into the regulatory networks underlying thymoma-associated MG and may lead to potential biomarkers or therapeutic targets. ncRNA is a tool in cancer diagnosis and treatment in personalized medicine to improve patients’ health, creating an intracellular network of crossed interactions that fine-tune the gene expression in both health and disease. The present knowledge of these mechanisms illustrates disease treatment, the current challenges and pitfalls, the roles of environmental and lifestyle-related contributing factors, and ethical, legal, and social issues arising from their (improper) use. 67 Genetic and epigenetic changes show that ncRNA are player in endometrium cancer. 68 An effective bioindicator of anti-cancer drug effectiveness is RPL28 (ribosomal protein L28) used as an epigenetic therapeutic in breast cancer. 69 Potential therapeutics for triple negative breast cancer with worst prognosis include miRNA and lncRNA accompanying chemo/radio therapy with surgery. 70 Recent publications from January 2024 indicate a new approach for managing cancer by effectively targeting and reducing the expression of molecules like proteins—osteopontin (OPN), EpCAM, estrogen receptor-alpha (ER-α), and carbonic anhydrase IX (CA IX) involved in tumor progression with advanced drug delivery system of Berberine liquid crystalline nanoparticles to overcome the limitations of conventional therapies increasing the effect of natural compounds. 71 A study from October 2023 revealed a complete exosome-based theragnostic signature in Ovarian Cancer Metastasis and the power of exosome research in cancer progression, metastasis, and drug resistance for next-generation cancer therapy. 72 Matrix Metalloproteinase enzymes have a significant role in chemoresistance and can be used for furture therapy enhancing chemosensitivityvia interaction with different molecules and signaling pathways. 73 Epidermal growth factor receptors have associated gene expression profiling between mitochondrial genes and carcinoma, as well as in vitro and in vivo models for screening synergistic effects in various combinational treatments, however, there is still a gap between experimental and clinical studies. 74 State-of-the-art advances and future perspectives in cancer therapy represent nanomaterials-assisted photo-thermal therapy for best performance in clinical settings. 75 Since ROS mediates tissue homeostasis, cellular signaling, differentiation, and survival, comprehensive methodology improvement for detection establishes its relationship with antioxidants and cancer, bringing more clarity to signaling pathways using omics technology and understanding antioxidant pathways and ROS complexities. 76 This highlights the importance of studying tumor-specific molecular profiles to tailor treatment strategies effectively. The population-based cohort studies show the significance of regional studies in understanding the epidemiology of thymoma-associated MG.37–49 Differences between incidence and prevalence rates within diverse geographic areas enable informative targeted healthcare strategies and improve patient outcomes. Expression profiling studies enable significant implications in differential miRNA expression, comprehensive profiling with understanding the molecular mechanisms underlining thymoma-associated MG and highlighting the potential therapeutic targets for disease management. Genome-wide miRNA profiling using high-throughput sequencing techniques has revealed numerous miRNAs differentially expressed in thymoma tissues associated with MG implicated in various cellular processes, including immune regulation and tumor biology, which are critical in understanding the disease mechanism.52–54 Translating molecular findings into potential therapeutic strategies of thymoma-associated MG is significant in functional studies, enabling an understanding of how dysregulated miRNA and splicing affect disease mechanisms and therapeutic targets.56–60
Presented findings indicate the complex interplay between dysregulated miRNA splicing events and the effect on the cell function of the thymus, immune cells, and neuromuscular junction integrity in MG associated with thymoma. Scientists need to fill the gaps in the existing literature regarding the understanding of thymoma associated with MG that will enhance diagnostic and therapeutic strategies for patients worldwide. This manuscript provides an overview of the analysis of pre-mRNA as a potential biomarker for cancer, specifically focusing on MG in thymoma, utilizing in silico methods. The strengths of the research include clear objectives (examining the molecular-biological characteristics and pathological implications of pre-mRNA interactions), comprehensive methodology, data presentation, and discussion. The aim is clearly defined, whereas the methodology section enables a detailed description of the data collection process, analysis, and statistical methodology, enabling easy reproducibility of the results. Findings are present with various visualization plots (UMPA, box plot, and expression density), enabling an easier understanding of the analyzed data. The study addresses the implications of the findings, including challenges with publicly available data and in silico analysis, along with guidance for further research in this area. The study is clear and concise, with an overview of the background literature and research gaps. The study findings are limited in interpretation due to results, but detailed analysis and discussion strengthen the manuscript, providing a more comprehensive summary of the findings and their implication in further research and clinical practice. The manuscript represents valuable insight into the analysis of pre-miRNA in thymoma with MG, enhancing the quality and impact of the manuscript. Clearly stating the intentions to conduct these analyses in the methods section and discussing their potential implications in the results section, the researcher provided transparency about the conducted research plan and paved the way for future analyses and interpretations. The analysis identified PRPF19 pre-mRNA as a potential biomarker for thymoma based on its significant differential expression and functional relevance in RNA processing and cell cycle regulation. Manuscript results are consistent with previous studies reporting similar gene expression changes, providing validation and reliability. Uncontrolled proliferation and tumor growth raise dysregulation of cell cycle-related genes, leading to aberrant activation of the p53 pathway, impairing DNA repair, and promoting genomic instability, leading to tumor progression and metastases. We gain insight into molecular mechanisms driving disease development and progression that can guide future research utilizing the GSE11967 dataset. Results from this study indicate counterintuitive data regarding the lack of significant findings in protein-protein interaction network analysis alongside results in other analyses due to different sensitivity and specificity, data characteristics, biological content, sample size and power, and bioinformatic tools and methods. Every analysis type targets different data aspects, where the sensitivity and specificity of methodology vary depending on whether it is a protein or gene analysis. There is an existence of dataset and methodologies differences where DEA (differentially expressed analysis) uses high-thorough sequencing data with a broad view of changes in gene expression, and PPI (protein-protein interaction) relies on known interactions from databases that can be incomplete or not covered in the thymoma associated MG disease. There is the possibility that the biological process of thymoma associated with MG is easier captured with gene expression changes and pathway enrichment in comparison to protein interactions. In the complexity of this disease, the interaction between miRNA and mRNA expression could be easily detectable through RNA-based methodologies. Since we used only 8. Available samples, the statistical power might be a lover for PPI analysis or the data quality is not subtle since DEA data pathway analysis is more robust in detecting significant changes based on the data nature and applicable statistics. Used methods and tools have differences in the refined models and binary interactions, making PPI analysis not sensitive to expression level changes as DEA and enrichment analysis. Due to relevance to the research objective, the quality of data and its alignment with the specificity of thymoma-associated MG guided the researcher to select the GSE11967 dataset with a small sample size (8 in total). This limited, unique available specificity dataset with only thymoma tissue with and without MG is significant for analyzing the molecular mechanism of the disease, making it highly relevant regardless of the small sample size. High-quality and reliable data are enabled by the robust detection capabilities of exon-level comprehensive human genome expression with analyzing splicing events, ensuring experimental rigor through the use of the Affymetrix Human Exon 1.0 ST Array (GPL188) platform. Scientific and practical considerations include the feasibility of in-depth analysis and resource constraints. Using smaller datasets allows for more reliable conclusions and deeper insights into the molecular underpinnings of thymoma-associated MG. Detailed and exhaustive analysis of each sample makes the resource-efficient in analysis and ensures high quality. The selected dataset is the perfect foundation for informing future studies after testing generated hypotheses in exploratory studies to justify further investigation, funding, and support. Limitations of smaller sample size are mitigated by the use of advanced statistical methods and bioinformatic tools like DESeq2 and edgeR, along with combining GSE11967 with other datasets, increasing the sample size, improving statistical power analysis, confirming robust methodology, and enabling reliable differential expression results. Preliminary findings of the study emphasize further investigation and acknowledge study limitations by transparent reporting and contextual relevance. Since the results showed differentially expressed genes in our analysis of the GSE11967 dataset, we can understand the complexity of the disease at the transcriptomic level, subtle transcriptomic changes, alternative molecular mechanism, pathway level insights, methodological considerations with recommendations for further research, clinical validation, and experimental integration. The detection of significant gene expression changes depends on the cell population. In this case, the tumor microenvironment is complex and contains epithelial cells, lymphocytes, and stromal components that can impact the visibility and detection of genetic changes. Results indicate that tumor complexity is not visible on transcriptome analysis alone. Potential subtle transcriptomic changes are due to low-frequency genetic alterations, small sample sizes, or sensitivity of detection methods. Apart from different expression genes, other mechanisms like post-transcription modifications, epigenetic changes, or non-coding RNAs could play a significant role in disease pathogenesis. RNA splicing, stability, and translation efficiency changes could influence protein function without reciprocity changes in mRNA level. Moreover, tumor progression and development depend on chromatin remodeling, histone modification, or DNA methylation. Gene expression modulates post-transcriptional level with microRNA, long non-coding RNA, and other regulatory RNA in thymoma with MG disease. Paper results observed trends in pathways related to cell cycle regulation and immune response with known biological characteristics of thymoma and enabled a focused hypothesis-driven approach. Despite the insights gained from our study, several limitations warrant consideration. When considering methodology, sample size, age, typographical errors, and data accuracy influence final result interpretation. In this case, the sample size of the GSE11967 dataset is potentially insufficient to detect significant differential expressed genes. Future studies should increase the sample size to enhance statistical power if that kind of data exists freely available in databases for this disease. Age differences can confound the results. Further studies should control or stratify samples by age to minimize the finding impact. The correct data entry in the GEO database is crucial for appropriate conclusions. Rigorous data validations and quality control procedures maintain data integrity. Multi-omic approaches like genomics, proteomics, metabolomics, and epigenomics could overcome the limitations observed in this study for a comprehensive understanding of the disease in the future. Understanding the differences in the tumor microenvironment leads to potential therapeutic targets. It is crucial to conduct further in vitro and in vivo studies to validate these findings since our in silico analysis has provided valuable insights into potential biomarkers for thymoma associated with MG. Experimental validation of the identified biomarkers will help confirm their clinical application and biological relevance. Further studies should include experiment validation with patient samples or animal models after in silico studies as a necessity for enhancing the reliability and applicability of the results in clinical practice with translational relevance. Scientific rigor increases the reliability of the results and improve patient outcomes. Validation of trends observed in pathways or biomarkers could include qPCR, immunohistochemistry, and another molecular assay to confirm protein and gene expression levels in patients’ samples. Patient data with age, gender, and disease stage variables can provide direct comparison and verification of the GSE11967 dataset, along with RNA sequencing and proteomic analysis. Functional implications of the pathways and genes identified enable the identification of potential therapeutic targets for tumor growth, progression, and response to therapies. Experimental studies and independent data sets are necessary for potential biomarkers in thymoma diagnosis and prognosis, demonstrating the strength of gene expression analysis with functional enrichment and pathway analysis in clinical studies. This study shed light on gene expression analysis limitation in thymoma associated with MG disease, suggesting the applicability of an integrative multi-omic approach. Future research should unravel novel therapeutic targets and biomolecules by better understanding disease complexity. Overall, the manuscript provides a comprehensive analysis of thymoma associated with MG, an understanding of the molecular basis of pathogenesis, guiding future research and clinical practice with in silico analysis, enhancing clarity and in-depth analysis impact.
Conclusion
We can conclude that the manuscript provides a clear overview of the research approach, including data collection, ethical considerations, and analysis plan. It demonstrates a thoughtful and transparent approach to research using publicly available data and computational methods and provides valuable insight into the molecular landscape of thymoma associated with MG disease. Despite limited sample size, dataset GSE11967 justifies high relevance to research objectives, the quality of data, and practical consideration associated with exploratory studies, providing a foundation for generating initial hypothesis and insight into thymoma-associated MG molecular mechanism by additional validation in future studies, making informative guidance for extensive research effort. The results of a comprehensive analysis of the GSE11967 data set indicate the complexity of thymoma in MG disease, revealing the presence of differentially expressed genes, indicating that genetic changes alone may not fully enable revealing the complexity of this disease. Researching additional approaches in integrative and multi-dimensional studies will probably reveal insights into pathway levels, transcriptomic changes, and alternative molecular mechanisms leading to this disease. Broaden analysis in the segment of omic research and tumor microenvironment will uncover potential biomarkers and therapeutic targets for thymoma in MG disease. A valuable field of research is in silico analysis, enabling advances and challenges and providing better detection and eradication of disease. In silico methodology removes ethical and financial barriers, providing research of potential therapeutic strategies. The COVID-19 pandemic underlines the significance of technology application and public communication in research. Although in silico analysis enables valuable insight into disease mechanisms and treatment strategies, potential clarity of publicly available data sets must address ethical considerations, data privacy, security, and misuse. Data sets are anonymized and securely stored to protect the confidentiality of patient identity and personal health information to minimize the risk of data breaches. Ethical and regulatory documents such as the Health Insurance Portability and Accountability Act (HIPAA) and the General Data Protection Regulation (GDPR) are significant to consider promoting the responsible use of in silico methodology in the research scientific fields. Genomic patient data should be protected from unauthorized access and potential cyber threats by encryption, secure data storage, and controlled access to keep sensitive information safe. Misuse of data information can lead to genetic discrimination and unauthorized genetic modifications, representing challenges in ethical discussions. Clear guidelines and regulations prevent the exploitation of genomic information and enable public awareness, where ethical training mitigates the potential risks that can arise. Results should be applicable in clinical practice by integrating clinical validation and addressing ethical considerations as a comprehensive approach, ensuring data privacy and security in silico research and contributing to functionally significant biomarkers leading to appropriate therapeutic strategies for thymoma associated with MG disease research. Detailed analysis of legal implications in personalized medicine with broader relevance and scope enables implementation into patient care, clinical practice, and health care policy. Policy-making and evidence-based guidelines implicate the complexity of informed consent for fertility preservation and provide a more comprehensive analysis of the precision medicine implication approaches discussed in society and ethical considerations in implementation and personalized medicine.
The article effectively communicates findings, relevance, and impact in the field of thymoma associated with MG research and health care, appealing to a broader readership and making a valuable contribution to the scientific literature. The manuscript effectively conveys its findings, significance, and implications within cancer research and health care.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
