Abstract
Objectives:
To evaluate the correlation of miRNA-related single nucleotide polymorphisms (miR-SNPs) with the risk of dermatomyositis (DM) development.
Introduction:
MicroRNAs (MiRNAs) are involved in a variety of activities such as cell differentiation, proliferation, apoptosis, tumorigenesis, and immunological response. MiR-SNPs alter the expression levels of miRNAs, leading to increased susceptibility to DM.
Methods:
We genotyped six miR-SNPs for miRNA processing machinery genes, including XPO5 (rs11077), RAN (rs14035), Dicer (rs3742330), TNRC6B (rs9623117), GEMIN3 (rs197412), and GEMIN4 (rs2740348), and two miR-SNPs for microRNA binding site, including SET8 (rs16917496), and KRT81 (rs3660), in a case-control study to assess the impact of these miR-SNPs on DM risk. Then we assessed cytokine expression and ROS levels in DM to determine the relationship between risk-related miR-SNPs and cytokines.
Results:
We discovered that Dicer’s (rs3742330) AA genotype had a decreased chance of developing DM than the AG + GG type (odds ratio, 0.527; 95% confidence interval: 0.281–0.987; p = 0.045). The subsequent analysis showed that the AA genotype carrier had greater levels of IL-4 (p = 0.034).
Conclusion:
The SNP of Dicer (rs3742330) maybe an attractive predictor of DM, moreover the cytokine of IL-4 may act as the factor that distinguishes SNP of Dicer (rs3742330) into AA and AG + GG.
Introduction
Dermatomyositis (DM) is an idiopathic inflammatory myopathy (IIM) defined by gradual symmetrical muscle weakening and characteristic skin lesions. It may also be a manifestation of interstitial lung disease, cardiac injury, esophageal dysfunction, and an underlying malignancy. Approximately, the prevalence of DM ranges from 9.54 to 32.74 per 100,000 people and the incidence is between 2.47 and 7.8 per 1,000,000 person-years.1,2 Based on individual genetic predisposition, it is commonly believed that environmental exposures cause the development of autoimmune diseases. 3 Changes in environmental factors partially influence the susceptibility of autoimmune diseases through epigenetic modifications. 4 MiRNAs appear to be significant in the pathophysiology of DM being one of the primary epigenetic modifications. 5
The class of endogenous RNAs known as microRNAs (MiRNAs) is made up of about 22 nucleotide-long molecules that play a variety of roles in cellular functions, such as cell differentiation, proliferation, and apoptosis. 6 MiRNAs are important regulators of immune responses and immune cell growth, according to available data. 7 In the course of miRNA processing, the primary transcript of miRNA (pri-miRNA) is first transformed by Drosha (RNase III) into a double-stranded precursor miRNA (pre-miRNA), which is then transported to the cytoplasm by exportin-5 (XPO5) and RAN.8,9 Once the pre-miRNA is formed, the GEMIN3 and GEMIN4-containing RNA-induced silencing complex (RISC) will choose one strand as the mature miRNA and guide it to its target mRNA location and its partner protein TRBP (transactivator RNA binding protein) are in the cytoplasm, and the Dicer will cleave pre-miRNA into a 21-bp miRNA.10,11 The target mRNA is degraded or will have its translation repressed when the mature miRNA binds to the target mRNA’s 3′ UTR, close to SET8 and KRT81. 12 Single nucleotide polymorphisms (SNPs) in the miRNA genes, miRNA binding sites, and miRNA processing machinery are known as miRNA-related SNPs (miR-SNPs), and they have the ability to influence the gene regulation involved in immune system and inflammatory processes.13–15
Innate immunity, adaptive immunity, and non-immune variables interact in a complicated way throughout the development of DM. 16 Unknown environmental stimuli cause the innate immune system to respond by releasing chemokines, activating particular Toll-like receptors (TLRs) and engaging in complex cytokine interactions, all of which result in the activation of the type 1 interferon pathway, the maturation of dendritic cells (DC), and their recruitment.17,18 An immune-activating positive feedback mechanism is created when type 1 interferons activate B cells, T cells, and DC, which in turn stimulates the production of type 1 interferons through B cell-derived autoantibodies.3,17 Additionally, oxidative stress has a role in the pathogenesis of DM as a non-immune component. 19 Reactive oxygen species (ROS), through decreased force production and muscular catabolic and autophagic pathways are activated, appears to play a crucial role in skeletal muscle weakening, one of the symptoms of DM. 19
SNPs in the miRNAs have been suggested to affect DM susceptibility, 20 despite the fact that few studies have explicitly looked at the association between DM risk and SNPs in the genes that encode the miRNAs' binding sites and processing machinery. Six miR-SNPs for genes in the miRNA processing machinery were genotyped, including XPO5 (rs11077), RAN (rs14035), Dicer (rs3742330), TNRC6B (rs9623117), GEMIN3 (rs197412) and GEMIN4 (rs2740348), and two miR-SNPs for microRNA binding site, including SET8 (rs16917496) and KRT81 (rs3660), DM risk was examined using a case-control approach to assess the effect of these miR-SNPs. All these genes are involved in miRNA formation and functional execution. Many studies have reported the significance of selected SNPs in the course of multiple diseases,21,22 and some of them may also cause changes in the function of the corresponding genes.23,24 Then we assessed cytokine expression and ROS levels in DM to determine the relationship between risk-related miR-SNPs and cytokines.
Materials and methods
Collection of Samples and DNA Extraction
76 people with DM participated in this case-control research between November 2021 and March 2022 at the Department of Rheumatology and Immunology. All DM patients were diagnosed based on the Bohan-Peter’s classification criteria.25,26 Age, sex, muscle pain, muscle weakness, heliotrope rash, Gottron’s sign, Shawl’s sign and V-sign, interstitial lung disease (ILD), dysphagia, and laboratory test data such as myositis-specific autoantibodies (MSAs), electromyography (EMG), muscle pathology, CK, LDH, ESR, and CRP, were all noted in DM patients. In addition, 88 age-matched healthy controls were chosen from the Physical Examination Center who did not have cancer, an immunological problem, or any other chronic conditions. Total DNA was extracted from blood samples by using a genomic DNA extraction kit (Tiangen, Beijing, China). Every procedure was overseen by the Ethics Committee and authorized in line with the Declaration of Helsinki (2021-R490). The subjects' written, fully informed consent was acquired. Subjects signed the informed consent for this study in advance.
MiR-SNP genotyping
Primers and probes used for genotyping of miR-SNPs.

MiR-SNPs:MicroRNA-related single nucleotide polymorphisms
Measurement of cytokines
The Human TH1/TH2 Panel (8-Plex) with Filter Plate V02 (Biolegend, San Diego, CA) was used to assess the levels of interleukin-5 (IL-5), interleukin-13 (IL-13), interferon-γ (IFN-γ), interleukin-2 (IL-2), interleukin-6 (IL-6), interleukin-10 (IL-10), tumor necrosis factor-α (TNF-α), and interleukin-4 (IL-4). Then, 25 μL of streptavidin-phycoerythrin (SA-PE) was added to each medium, and the mixtures were shaken for 30 min at room temperature, in the dark, at 800 r/min. The PE fluorescence signal of the analyte-specific bead region was quantified using a flow cytometer, the MACSQuant Analyzer 10 (Miltenyi Biotec, Bergisch Gladbach, Germany), and the concentration of the specific analyte was determined using a standard curve generated by BioLegend’s LEGENDplexTM Data Analysis Software (Biolegend, San Diego, CA).
ROS measurement
The BBOXiProbe ® Plasma Active Oxygen Detection Kit was used to measure the ROS levels (BestBio Technology, Shanghai, China). Briefly, 10 μL of O12 probe and 100 μL of serum were incubated for 30 min at 37°C. The presence of ROS was measured using a fluorescent microplate reader (BIOTEK, Winooski, VT, USA) with excitation and emission wavelengths of 488 nm and 520 nm, respectively.
Statistical analysis
The mean and standard deviation of the data were shown. When the data met the requirement for normality, Student's t test was applied to continuous variables; if not, the Wilcoxon rank sum test was used. Dichotomous variables were analyzed using the chi-square test, including the clinical characteristics and the miR-SNPs between DM patients and healthy controls. All statistical evaluations were performed using the SPSS 25.0 software (SPSS, Inc., Chicago, IL, USA). The sample size was assessed by PASS 15.0. p values <0.05 were regarded as statistically significant for all statistical tests.
Results
Clinical characteristics of DM patients and controls.

DM: dermatomyositis; ILD: interstitial lung disease; MSAs: myositis-specific autoantibodies; anti-Tif1-γ: anti-transcription intermediary factor 1 γ; anti-NXP2: anti-nuclear matrix protein 2; anti-MDA5: anti-melanoma differentiation antigen 5; anti-SAE: anti-small ubiquitin-like Modifier activating enzyme; EMG: Electromyography; χ2: Chi-square.
a32 patients recorded MSAs.
Distribution frequency of miR-SNPs between DM patients and healthy controls.

MiR-SNPs:MicroRNA-related single nucleotide polymorphisms; DM: dermatomyositis.
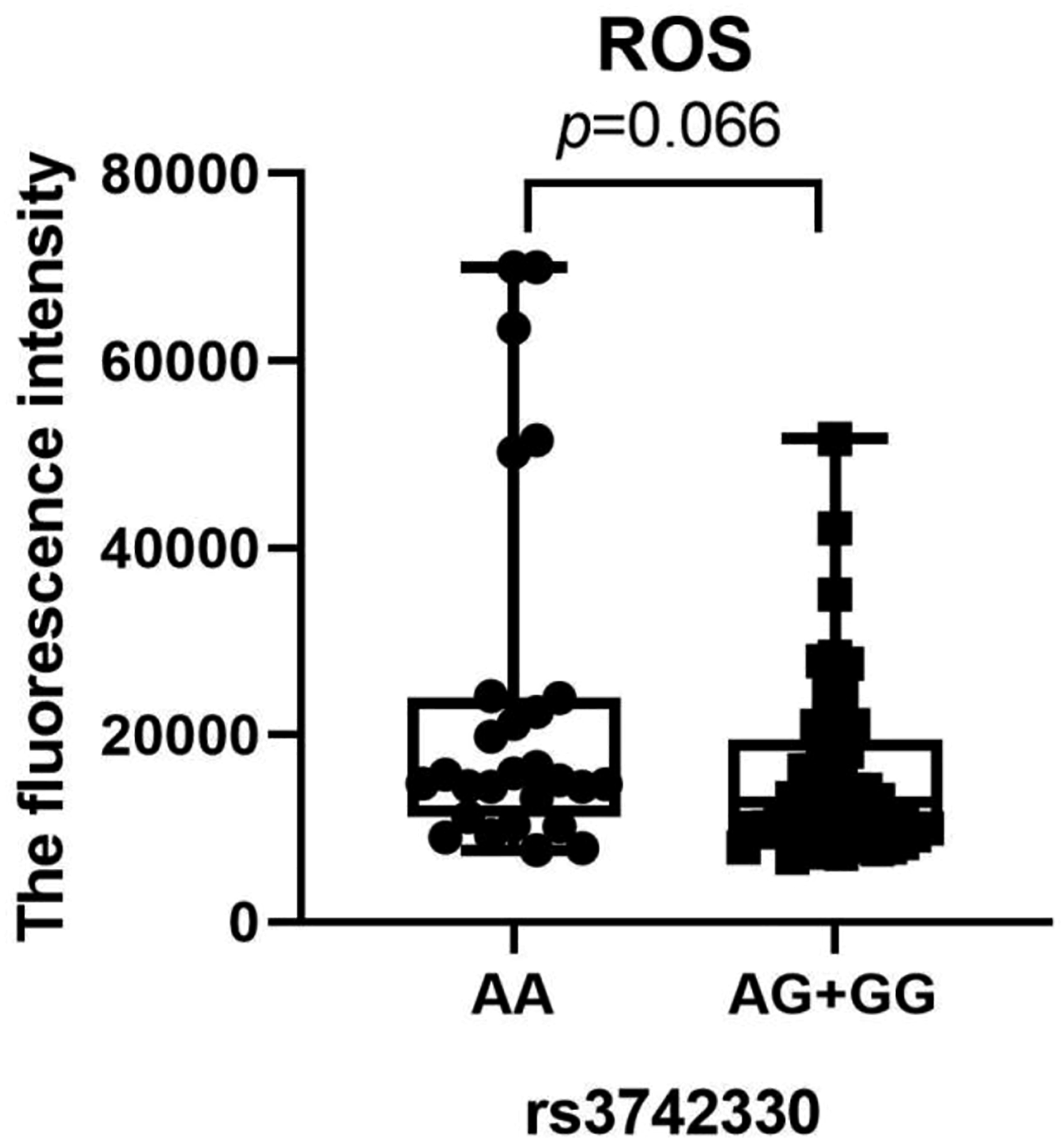
To examine the possible correlation between cytokine levels, including IL-5, IL-13, IFN-γ, IL-2, IL-6, IL-10, TNF-α, and IL-4 and genotypes of rs3742330, the Wilcoxon rank sum test was used (Figure 1). The AA genotype carrier had greater levels of IL-4 (p = 0.034). According to this research, miR-SNP may act as a moderator of DM development by influencing cytokine expression and serving as a predictor of DM risk. Following genotyping analysis, no significant correlation between ROS levels and rs3742330 genotypes was found (p = 0.066, Figure 2). Boxplot of IL-5, IL-13, IFN-γ, IL-2, IL-6, IL-10, TNF-α and IL-4 levels in DM genotypes of rs3742330. Wilcoxon rank sum test was used to determine significance. IL-5: Interleukin-5; IL-13: Interleukin-13; IFN-γ: interferon-γ; IL-2: Interleukin-2; IL-6: Interleukin-6; IL-10: Interleukin-10; TNF-α: Tumor Necrosis Factor-α; IL-4: Interleukin-4; DM: dermatomyositis. Boxplot of ROS levels in DM genotypes of rs3742330. Wilcoxon rank sum test was used to determine significance. ROS: reactive oxygen species; DM: dermatomyositis.
Discussion
In the present study, MiR-SNPs, including XPO5 (rs11077), RAN (rs14035), Dicer (rs3742330), TNRC6B (rs9623117), GEMIN3 (rs197412), GEMIN4 (rs2740348), SET8 (rs16917496), and KRT81 (rs3660) were examined for their ability to predict DM risk. The findings indicated that the rs3742330 SNP of the Dicer gene is a potential risk biomarker for DM.
Dicer is an enzyme belonging to the RNase III family, and it plays a crucial part in the converting the pre-miRNA into mature cytoplasmic miRNA. 27 According to several researches, the Dicer gene is essential to the formation and function of immune cells. 28 Dicer-deficient regulatory T cells (Tregs) could also lose their ability to inhibit the immune system and cause autoimmune responses in sick mice, according to animal studies. 29 MiRNA levels and activity are directly influenced by Dicer. 30 The expression levels of miRNAs are altered as a result of miR-SNPs’ influence on the choice of functional strands during pre-miRNA processing, including Dicer enzyme processing, which results in varied phenotypes and the emergence of diseases.31,32 rs3742330 has been found to be related to an increased risk of colorectal cancer in earlier studies. 14 It is unknown by what mechanism SNP influences DM occurrence. Due to its location in the 3′-untranslated region of the enzyme, this SNP may have an effect on mRNA stability and Dicer expression.
We discovered a connection between higher IL-4 levels and the Dicer AA genotype. 33 Dicer deletion caused the upregulation of IL4 responsive fusion protein Tm7sf4.34,35 IL-4 controls macrophages to inhibit immune responses, repair damaged tissue, and take part in parasite defense. IL-4 enhances muscle self-repair and suppresses IFN-induced inflammation, appearing to be resistant to DM. 36 Our results suggested that the AA genotype of Dicer induces higher IL-4 levels by lowering immune responses.
This study has some limitations. First, the study was conducted in a single center lacking a validation group, and data from multiple centers should be evaluated in further studies. Second, the sample size was small. The larger sample size was valuable for multiple comparisons to maintain the rigor of the article. Therefore more cases and repeat test of gene should be conduct in the future. Third, due to the large time span of our study, the restriction of economic level or medical development, and the update of electronic medical record system, only some patients completed the MSAs. We will collect detailed information of MSAs to perform stratified analysis. However, to our knowledge, the present study is the first on the relationship between miR-SNPs and the risk of DM, and the further discovery of a potential role of cytokines in it is encouraging for subsequent scientists to explore the pathogenesis of DM.
Conclusion
To our knowledge, this is the first study to look into the associations between miR-SNPs with the risk of developing DM. The SNP of Dicer (rs3742330) holds significant promise as a biomarker for DM’s early diagnosis and prophylaxis. Although researches on miR-SNPs is still new to our field of study, our findings will most likely prove to be useful, since the SNP of Dicer (rs3742330) maybe an attractive predictor of DM, moreover the cytokine of IL-4 may act as the factor that distinguishes SNP of Dicer (rs3742330) into AA and AG + GG.
Footnotes
Author contributions
Yufei Zhao designed the original experiment. Chenxing Peng and Jingjing Zhang collected the tissue specimens. Shasha Zhang and Xiaoyun Zhang conducted the experiments. Yufei Zhao interpreted the data and drafted the manuscript. All authors contributed to the article and approved the submitted version.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
