Abstract
Objective:
Hepatocellular carcinoma (HCC) is the most common cancer of the digestive system. Recently, transmembrane proteins (TMEMs) have been extensively studied in different types of malignant tumors. However, the influence of TMEM206 in hepatocellular carcinoma is unclear.
Methods:
The UALCAN database was used to investigate the expression of TMEM206 mRNA in liver cancer tissues and used the Human Protein Atlas (THPA) to study the expression of TMEM206 in HCC tissues. The expression of TMEM206 was measured in normal liver HL-7702 cells and HepG2, SMMC-7721, and Bel-7402 liver cancer cells. Next, a lentivirus was used to knockdown TMEM206 in HepG2 cells. Furthermore, after verifying knockdown, we studied the effect of TMEM206 downregulation on the malignant behavior of HepG2 and on the PI3K/AKT pathway.
Results:
TMEM206 was highly expressed in liver cancer cells (p < 0.001). Downregulation of TMEM206 significantly inhibited the proliferation, migration and invasion of HepG2, significantly promoted the apoptosis of HepG2, and inhibited the expression of P-PI3K and P-AKT.
Conclusion:
TMEM206 can affect the malignant behavior of HCC. And the PI3K/AKT pathway was affected. This study provides new ideas for the treatment of HCC.
Introduction
Hepatocellular carcinoma (HCC) is a common malignant tumor of the digestive system. 1 Previous studies have shown that the progression of HCC may be a multistep process involving multi-factorial etiology and multi-gene alteration. 2 Therefore, a better understanding of the underlying molecular mechanisms of HCC metastasis will be of great significance for the prognosis and targeted therapy of HCC.
Transmembrane protein 206 (TMEM206), also known as proton-activated chloride channel (PAC), is a transmembrane protein. 3 Transmembrane proteins (TMEM) are mainly involved in the composition of the cell membrane and play an significant role in regulating cell invasion and migration. 4 TMEM206 is over expressed in colorectal cancer tissues and can promote the migration and invasion of colorectal cancer cells by activating AKT and FAK. 5 However, there are no studies investigating whether TMEM206 plays a role in the various malignant behaviors of liver cancer cells. Therefore, we used a lentivirus to knock down TMEM206 in HCC cells and studied the effect of TMEM206 downregulation on the malignant behavior of HCC and the related mechanisms.
Materials and methods
Bioinformatics analysis
The UALCAN (http://ual can.path.uab.edu/index.html) database is an interactive network resource. Not only can a primary tumor be compared with normal tissue samples but different tumor subgroups can also be compared according to the pathological cancer stage, tumor grade, and patient ethnicity. 6 The GEPIA is a newly developed interactive web server for analyzing the RNA sequencing expression data, using a standard processing pipeline. 7
In the present study, UALCAN and GEPIA was used to evaluate the TMEM206 gene expression in HCC, including HCC tissue differential analysis and patient survival analysis. The Human Protein Atlas (THPA) online tool (https://www.proteinatlas.org/) shows images of antibody staining in samples of normal and cancer tissue. 8 We downloaded TMEM206 immunohistochemical staining data of normal liver cells and liver cancer tissues from THPA.
Lentiviral vector constructs and cell culture
Lentiviral vectors knocking down human TMEM206 (NM_001198862) and containing a green fluorescent protein tag were constructed by GeneChem (Shanghai, China). The infection procedure was described in the Lentiviral Handbook of GeneChem. Subsequently, cells stably expressing the target gene were obtained by 2 μg/mL puromycin selection (Cayman Chemical, Ann Arbor, MI) for 7 days. Cells were cultured in DMEM/high-glucose medium (Gibco) or RPMI 1640 medium (Gibco) containing 10% foetal bovine serum (FBS) (Gibco) at 37°C in a 5% CO2 humidified incubator.
Real-time PCR assay
Total RNA was extracted from each group using TRIzol reagent, and the concentration and purity were determined using a NanoDrop™ One ultramicro UV-visible spectrophotometer (Thermo, USA). Then, qRT-PCR was performed after total RNA was reverse transcribed into cDNA using the Prime Script™ RT Reagent Kit. qRT-PCR was performed on the Bio-Rad real-time PCR system (Bio-Rad, USA) by SYBR Premix Ex Taq™. The relative expression of TMEM206, MMP2, MMP9, E-CAD and N-CAD was normalized to the mRNA level of GAPDH and calculated using the formula 2−ΔΔCT. Primer design and synthesis were completed by Servicebio (Wuhan, Hubei, China).
Western blotting assay
The cells to be tested were seeded into a 6-well plate, cultured, and this was repeated three times for each group of cells. After 48 h, the confluency of the cells in each well reached 80%. Then, we collected the cells and extracted the total cell protein. Proteins were separated by SDS-PAGE and transferred to a PVDF membrane. The proteins in the PVDF membrane were incubated with the target protein-specific primary antibody and then incubated with the fluorescent secondary antibody. The expression of the target proteins was scanned and imaged on the Odyssey (LICOR) dual-color infrared imaging system.
Cell counting Kit 8 assay and clone formation assay
CCK-8 assay: Cells were seeded in a 96-well plate to ensure that the cell density was 2000 cells/100 μl, and this was repeated three times for each group of cells. The culture plate was placed in a 37°C and 5% CO2 incubator and incubated for 4 h. After the cells adhered to the wall, the old medium was aspirated, and 10 μL of 10% CCK8 medium was added to each well. The cells were incubated for 2 h in the dark in an incubator, and the 0 h absorbance (OD) was measured at 450 nm with a microplate reader. The same procedure was used to measure the OD values at 24 h, 48 h, 72 h, and 96 h and compare cell proliferation.
Clone formation assay: Cells were seeded in a 6-well plate to ensure that the cell density was 5000 cells/well, and this was repeated three times for each group of cells. The media was changed every 2 days, and the cells were cultured for 7 days. The old medium was aspirated; the cells were washed 3 times with 2 mL of PBS, fixed with 2 mL of 2% paraformaldehyde for 30 min, then washed 3 times with 2 mL of PBS. Cells were then stained with 2 mL of 0.5% crystal violet solution for 30 min, and then the crystal violet was aspirated. The solution was dried, and the cells were photographed to compare the number of cell colonies formed.
Transwell migration and invasion assay
Transwell invasion assay: Matrigel (50 μl, BD Matrigel: serum-free medium = 1:8) was added to the upper chamber and spread evenly, and the chamber was placed in a 37°C incubator for 4–5 h to solidify. Six hundred microlitres of medium containing 10% serum was added to the lower chamber, and the upper chamber was slowly placed into the lower chamber to ensure that there were no bubbles. Then, 200 μL of cells resuspended in serum-free medium (cell density = 105 cells/mL) was added to the upper chamber, and this was repeated three times for each group of cells. After 24 h, the upper and lower chamber medium was discarded, the upper chamber was washed with PBS, the cells were fixed with a tissue fixative solution for 15 min, and the cells were stained with 0.5% crystal violet solution for 30 min. The cells were washed with PBS until the purple color faded, and the cells in the chamber were wiped with a cotton swab, leaving the invaded cells outside the membrane, which were observed and photographed under a microscope. Transwell migration assays: The Matrigel coating step was eliminated, and the rest of the steps mentioned were the same as those in the Transwell invasion assays.
TUNEL assay
The slides were placed into a six-well plate, the cell suspension to be tested was added, and the cells were incubated in a 37°C and 5% CO2 incubator until the cell confluency reached 60%. This was repeated three times for each group of cells. The old medium was aspirated, and 2% paraformaldehyde was added to fix the cells for 30 min. Cells were washed twice with PBS, the slide was taken out and spun dry, a circle was drawn with a histochemical pen at the position where the cells were evenly distributed in the middle of the cover glass, 50-100 μLof rupture working solution was added, and the cells were incubated at room temperature for 20 min and then washed twice with PBS. After spin-drying, buffer was added dropwise into the circle to cover the tissue and incubated at room temperature for 10 min. The appropriate amount of TDT enzyme, dUTP, and buffer in the TUNEL kit was mixed at a ratio of 1:5:50, added to the circle to cover the cells, and incubated at 37°C for 2 h. The cells were washed twice with PBS, DAPI staining solution was added dropwise to the circle, and the cells were incubated for 10 min at room temperature in the dark. The slide was washed twice with PBS, dried slightly, and mounted on the slide with an anti-fluorescence quenching mounting tablet facing down with the cell side. Cell apoptosis was observed under a fluorescence microscope.
Immunofluorescence
The slides were placed into a six-well plate, the cell suspension was added, and the cells were incubated at 37°C and 5% CO2 until the cell confluency was 60%. This was repeated three times for each group of cells. The old media was aspirated, cells were washed twice with PBS, 2% paraformaldehyde was added, and cells were fixed for 30 min then washed twice with PBS. After the slide was dried slightly, a circle was drawn with a histochemical pen at the position where cells were evenly distributed in the middle of the cover glass. A total of 50–100 μL of rupture working solution was added, incubated at room temperature for 20 min, and then washed twice with PBS. After spin-drying, 3% hydrogen peroxide solution was added dropwise to the circle, incubated for 20 min at room temperature in the dark, and washed twice with PBS. After the slides were slightly dry, the specific primary antibody was added to the circle to cover the cells, and the cells were placed flat in a wet box and incubated overnight at 4°C. Next, the cells were washed twice with PBS, and after spin-drying, a fluorescent secondary antibody was added to the circle to cover the cells, and the cells were incubated at room temperature for 50 min. Cells were washed twice with PBS, 50–100 μl of DAPI dye solution was added dropwise to each well, and the cells were incubated at room temperature for 5 min. After washing twice with PBS, an appropriate amount of anti-fluorescence quencher was dropped onto the cells, the coverslip was mounted, and the expression of pathway proteins was observed under a fluorescence microscope.
Statistical analysis
GraphPad Prism 8 (version 8.4.3), SPSS (version 26.0) and ImageJ (version 1.53) software were used to analyze the data. All experiments were performed in independent triplicates, and the results are expressed as the mean±standard deviation (x ± s). Quantitative variables were compared using Student’s t-test.
Results
Expression and survival analysis of human TMEM206 in HCC
The expression analysis of TMEM206 in HCC was downloaded from the UALCAN database. The results showed that TMEM206 was significantly overexpressed in HCC tissues (p < 0.001, Figure 1(a)), and the overexpression of TMEM206 was significantly related to the higher tumor stage of HCC (p < 0.01, Figure 1(b)). At the same time, the high expression of TMEM206 was significantly related to the poor prognosis of HCC patients(p < 0.01, Figure 1(c)). TMEM206 expression in normal liver tissues/cells and HCC tissues/cells. (a). The expression of TMEM206 is significantly increased in HCC tissues compared with normal liver tissues; (b). TMEM206 expression shows significant difference in tumor staging; (c). TMEM206 overexpression is related to the poor prognosis of liver cancer patients; D-E. TMEM206 is overexpressed in HCC tissues (e) than normal tissues (d); (f-g). TMEM206 is overexpressed in liver cancer cells compared with normal liver cells. *p < 0.01, **p < 0.001, ***p < 0.0001, ns is not statistically significant compared with the normal/HL-7702 group)
To further confirm the difference in TMEM206 protein expression levels, the THPA online tool was used to determine the protein expression level of TMEM206 in normal liver tissue and liver cancer tissue. The results showed that the TMEM206 protein is mainly distributed in the cell membrane and is clearly highly expressed in liver cancer tissues (Figures 1(d) and (e)).
A normal hepatocyte cell line HL-7702 and three HCC cell lines, HepG2, SMMC-7721, and Bel-7402 were selected for analysis the expression of TMEM206 in HCC cells. The mRNA expression of TMEM206 in different cell lines was measure by qRT-PCR. Western blotting was used to determine the protein expression of TMEM206 in different cell lines. The results showed that TMEM206 was significantly highly expressed in HCC cells, and TMEM206 expression was the highest in HepG2 among the three HCC cell lines (Figures 1(f) and (g)).
TMEM206 knockdown cell lines were successfully created
To confirm that TMEM206 was knocked down in HepG2 after lentivirus transfection, the transfection efficiency was observed with an inverted fluorescent microscope, the mRNA and protein expression of TMEM206 in HepG2 was measured by qRT-PCR and Western blot. The results showed that the transfection efficiency was higher than 95% (Figure 2(a)). Compared with the NC group, the mRNA and protein expression of TMEM206 in the LV group was significantly reduced (p < 0.001, Figures 2(b) and (c)). The mRNA of TMEM206 was knocked down to 14.98% and the protein of TMEM206 was knocked down to 43.20%. Cell lines of TMEM206 knockdown were successfully created. (a. Transfection efficiency is higher than 95%; b-c. The expression of TMEM206 in the LV group was significantly suppressed. ***p<0.0001, compared with the NC group).
Knockdown of TMEM206 inhibits the malignant behavior of HepG2
The CCK-8 and plate cloning methods were used to measure the proliferation ability of HepG2. The results showed that compared with the NC group, the proliferation ability of the LV group was significantly inhibited (Figures 3(a) and (b)). Knockdown of TMEM206 inhibited the malignant behaviour of HepG2. (a-b. Knockdown of TMEM206 inhibited the proliferation of HepG2; c: Knockdown of TMEM206 inhibited the migration and invasion of HepG2; D-E. Knockdown of TMEM206 significantly inhibited the expression of MMP2 and N-CAD in HepG2 and significantly increased the expression of E-cad in HepG2; d-e. f. Knockdown of TMEM206 increased apoptosis of HepG2. *p < 0.01, **p < 0.001, ***p < 0.0001, ns is not statistically significant compared with the NC group.)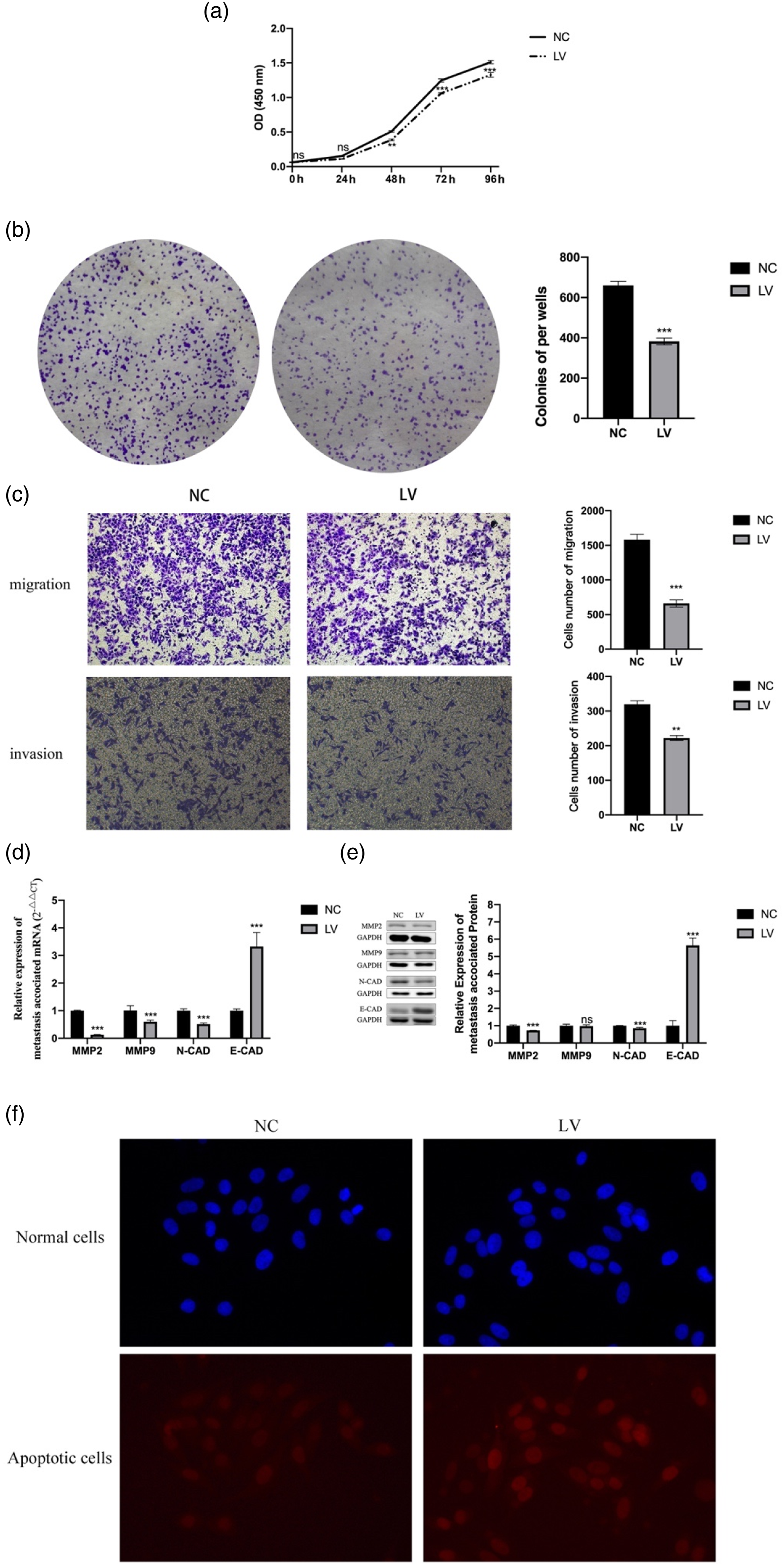
Furthermore, the transwell assays were used to measure the migration and invasion ability of HepG2. The results showed that compared with the NC group, the migration and invasion ability of the LV group were significantly inhibited (Figure 3(c)). At the same time, we also measured the expression of proteins related to tumor migration and invasion. Compared with the NC group, the expression of MMP2 and N-CAD protein in the LV group was significantly reduced, and the expression of E-CAD gene and protein in the LV group was significantly increased (Figures 3(d) and (e)).
The TUNEL assay was used to measure the apoptotic ability of HepG2. The results showed that compared with the NC group cells, the apoptotic ability of the LV group cells was significantly enhanced (Figure 3(f)).
TMEM206 can affect P-PI3K and P-AKT
The immunofluorescence and WB were used to measure the expression of PI3K and AKT proteins of HepG2. The immunofluorescence results showed that the expression of P-PI3K and P-AKT in the LV group was significantly reduced (Figure 4(a)). Further WB results showed that the TMEM206 gene can inhibit the expression of P-PI3K and P-AKT. Knockdown of TMEM206 affected the PI3K/AKT pathway in HepG2. (a. Knockdown of TMEM206 inhibited the expression of P-AKT in HepG2; b. Knockdown of TMEM206 inhibited the expression of P-PI3K and P-AKT in HepG2. ***p < 0.0001, compared with NC group).
Discussion
The high morbidity and mortality of HCC is a serious threat to human life and health; there were 841080 new cases of HCC worldwide and 781631 new deaths in 2018. 1 The incidence of HCC is increasing yearly, and the majority of cases (90%) occur in the context of chronic liver diseases, such as latent cirrhosis and chronic hepatitis B or C, accounting for 75% of all liver cancer incidences. Most of the HCC cases are at an advanced stage once diagnosed, and patients’ survival rate is very low.9–11 Considering the complexity of HCC, the treatment method used mainly depends on the stage of the disease, the location and size of the tumor, whether there is metastasis, and the potential liver function. At present, the preferred treatment for HCC is surgical resection and orthotopic liver transplantation. For patients who do not meet the treatment criteria, such as unresectable tumors, treatment options include systemic radiotherapy and chemotherapy and molecular targeted therapy. Due to the low specificity of these methods for tumors cells, the risk of serious drug-related adverse effects and acquired drug resistance is increased, and the survival rate of patients has not significantly improved.12,13 Therefore, to reduce treatment resistance and improve the prognosis of HCC patients it is of great importance to test or develop new and efficient therapeutic drugs, discover new molecular tumor targets for operable tumors, and increase the specificity for tumor cells to inhibit the malignant progression of HCC.
The protein of transmembrane (TMEM) family was involved in the composition of the cell membrane. Recent studies have found that the expression of TMEM is abnormal in a variety of malignant tumors.5,14–16 Some studies believe that some protein of TMEM family can be used as prognostic markers for different types of cancer, and some of that can be used as a potential target for cancer treatment. 16 TMEM16 A, as a calcium-activated Cl− channel, is highly expressed in a variety of cancers, such as head and neck squamous cell carcinoma, glioma, gastrointestinal cancer, lung cancer, oral cancer, oesophageal cancer and breast cancer.5,14,15 TMEM17 is expressed at low levels in lung cancer, and TMEM25 is expressed at low levels in breast cancer and is associated with poor survival of patients.17,18
In the tissue level, through the UALCAN database analysis, the mRNA expression of TMEM206 in HCC was found to be significantly higher than that in normal liver tissue. The result of the expression of TMEM206 protein was also consistent from the THPA. In the cell level, the expression of the TMEM206 gene and protein was measured in a normal liver cell line, HL-7702, and three liver cancer cell lines, HepG2, SMMC-7721, and Bel-7402. The expression of TMEM206 in liver cancer cells was significantly higher than that in normal liver cells. The result was consistent with the database analysis. It can be speculated that TMEM206 may play an important role in the occurrence and development of HCC.
The expression of TMEM206 in HepG2 was knocked down by lentiviral vector with a fluorescent tag. After puromycin selection, fluorescence microscopy was used to visualize the transfection efficiency, and it was confirmed that a high transfection efficiency was obtained in the stably transfected NC and LV groups. Then, the expression of the TMEM206 gene and protein in the two groups of cells was measured, and it was confirmed that TMEM206 in the cells from the LV group was significantly knocked down. Therefore, TMEM206 knockdown cell line was generated successfully.
The development and growth of tumor metastasis require that neoplastic cells either shift genetically or epigenetically between proliferative and invasive phenotypes or simply express both phenotypes simultaneously. 19 In this study, the CCK-8 assay and plate cloning experiments were performed to measure cell proliferation in the NC and LV groups. We found that the proliferation of HepG2 was significantly reduced after knocking down TMEM206. Therefore, we speculated that TMEM206 can affect the proliferation of HCC cells.
Matrix metallopeptidases (MMPs) are an important class of proteolytic enzymes that can degrade the components of the extracellular matrix and basement membrane. 20 Studies have found that MMP2 and MMP9 can degrade collagen IV in the basement membrane of cells and promote cell migration and invasion. 21 E-cadherin (E-cad) is a protein necessary for the formation of cell-to-cell connections. A decrease in E-cad may lead to tumor migration and invasion. 22 In other words, an increase in E-cad may inhibit cell migration and invasion. An increase in N-cad may promote the migration and invasion of tumor cells. E-cad and N-cad play a key role in the normal formation of mammalian embryos and in the maintenance of tissue homeostasis by dynamically regulating cell adhesion. 23 In this study, we measured the migration and invasion of the NC and LV groups by using Transwell assays. The migration and invasion capabilities of HepG2 were significantly reduced after knocking down TMEM206. By measuring the expression of MMP2, MMP9, E-cad and N-cad in HepG2, we found that following TMEM206 knockdown, the expression of MMP2 and N-cad in the cells was significantly reduced, and the expression of E-cad was significantly increased. This further shows that knocking down TMEM206 can significantly inhibit the migration and invasion of HepG2.
Defects in the mechanism of apoptosis play an important role in tumor pathogenesis. 24 Selective killing of cancer cells has long been pursued in the development of effective cancer treatment. 25 In this study, we found that knockdown of TMEM206 can promote apoptosis as measured by TUNEL assays, suggesting that TMEM206 may be related to apoptosis.
PI3K is a heterodimeric lipid kinase composed of regulatory and catalytic subunits encoded by different genes. 26 AKT, a downstream PI3K effector, regulates many biological processes, including proliferation, apoptosis and growth. The PI3K/AKT pathway is dysregulated in many cancers.27–32 PI3K/AKT phosphorylation and activation are necessary for the progression and metastasis of HCC. Interestingly, our study found that knockdown of TMEM206 can significantly reduce the phosphorylation of PI3K and AKT. We speculated that TMEM206 can affect the PI3K/AKT pathway.
There were still some limitations in this research. There was not calculation for estimation of sample size selected for this study. The effect of overexpression of TMEM206 was not studied in this research, although we think it is more important to study knockdown of TMEM206 for the treatment of HCC, since TMEM206 is overexpressed in HCC.
Conclusion
The present research found that TMEM206 is highly expressed in HCC, and that knockdown of TMEM206 can significantly inhibit the proliferation, migration and invasion of HCC cells. At the same time, knockdown of TMEM206 inhibited the phosphorylation of PI3K and AKT. These findings provide new ideas for the development of treatments for HCC. Whether it is possible to specifically treat HCC by knocking down TMEM206 is worthy of future studies.
Footnotes
Acknowledgements
The authors wish to thank the reviewers for their careful, unbiased and constructive suggestions, which led to this revised manuscript.
Authors’ contributions
SL and ZH conceived the study, analyzed and interpreted the data, and wrote the manuscript. FD and TJJ analyzed the artificial dataset and assessed the results. All authors revised and approved the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Availability of data and materials
The datasets presented in this study can be found in online repositories.
Ethics approval and consent to participate
Competing interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
