Abstract
Objectives
To investigate the potential mechanisms of Nagab-9 in alleviating acute lung injury (ALI) by integrating network pharmacology analysis with in vivo and in vitro validation experiments.
Methods
Active compounds of Nagab-9 were identified using TCMSP and ETCM databases. ALI-related targets were collected from relevant disease databases, and an intersection of these targets was used to construct a protein-protein interaction (PPI) network to identify core targets. Functional analysis through Gene Ontology (GO) and KEGG pathway enrichment was performed. The key targets of Nagab-9 intervention in ALI were further validated in LPS-induced ALI mouse models and in mouse alveolar epithelial cell injury models.
Results
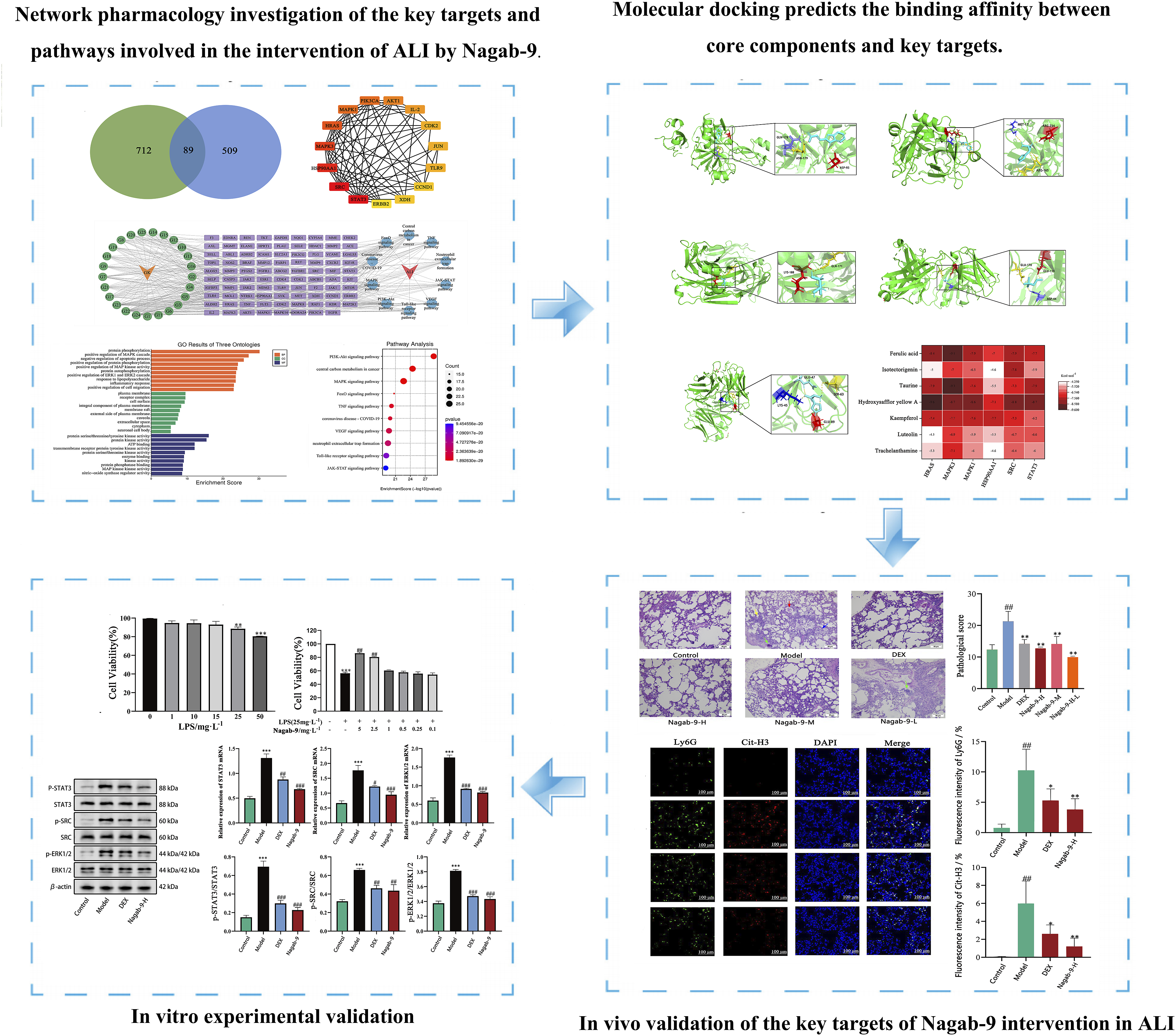
A total of 25 active components were identified from Nagab-9. PPI network analysis highlighted core targets, and GO and KEGG pathway analyses identified significant pathways involved. Six core components were selected based on topological parameters of the “compound-target-pathway-disease” network. In vivo, Nagab-9 was shown to alleviate ALI-induced lung damage, inhibit inflammatory infiltration, and modulate inflammatory factors by downregulating Ly6G, Cit-H3, and phosphorylated proteins SRC, ERK1/2, and STAT3 in lung tissue. In vitro experiments demonstrated that Nagab-9 effectively inhibits LPS-induced inflammatory responses, protecting lung tissue and suppressing neutrophil infiltration and NET formation, likely through the SRC/ERK1/2/STAT3 pathway.
Conclusion
Nagab-9 exerts a protective effect against ALI by modulating inflammatory responses and reducing neutrophil infiltration and NET formation, primarily via the SRC/ERK1/2/STAT3 signaling pathway. This study supports Nagab-9 as a promising therapeutic agent for ALI intervention.
Introduction
Acute Lung Injury (ALI), characterized by an acute inflammatory response, represents a respiratory disorder with a substantial mortality rate, exerting significant impacts on public health. The prevailing approach to managing patients with ALI involves interventions such as protective mechanical ventilation, administration of anti-inflammatory agents, and corticosteroid medications. The hallmark pathological features of ALI include the release of inflammatory mediators in the lungs, the exudation and infiltration of inflammatory cells, pulmonary edema, and disruption of alveolar architecture. Consequently, targeting the aberrant inflammatory response is conducive to the recuperation of individuals afflicted with ALI.1,2
Nagab-9 is one of the traditional Mongolian medicinal therapies, utilized for treating “lung heat year” diseases, a condition recognized within European medicine. Nagab-9 comprises nine ingredients, including musk, myrrh, borneol, processed aconite, typha, realgar, bezoar, safflower, and single clove garlic. It is primarily used for gastrointestinal and chest pain symptoms caused by epidemics, influenza, diphtheria, and anthrax. Preliminary studies have shown that Nagab-9, by preserving lung function and mitigating excessive inflammatory responses and tissue damage, can effectively improve the symptoms and prognosis of patients with ALI. 3 Experimental studies indicate that Nagab-9 treats ALI by inhibiting inflammation and regulating immunity. For instance, Nagab-9 can reduce the expression of Cluster of Differentiation 14 (CD14), Lipopolysaccharide Binding Protein (LBP), Nuclear Factor-kappa B (NF-kB), and Toll-like Receptor 4 (TLR4) in lung tissues. However, its mechanism of action in treating ALI remains unclear. 4
Traditional Chinese Medicine (TCM) and other herbal medicines are renowned for their multi-component, multi-target, and multi-effect characteristics. Network pharmacology constructs the complex relationships among each component, target, disease, and molecular pathway. 5 Molecular docking represents a computational strategy for predicting the binding affinity and mode between proteins and ligands, predicated on the “lock and key” model of interactions between proteins and small molecule ligands. It involves the computation and prediction of ligand conformations and orientations at the protein’s active site, thereby assessing their binding intensity. This methodology is pivotal in the prediction of drug targets within biological entities, underscoring its critical role in pharmacological research. Consequently, integrating network pharmacology with molecular docking techniques to explore the potential active components and molecular mechanisms of Traditional Chinese Medicine and other herbal medicines represents a beneficial strategy. 6
This study employs network pharmacology and molecular docking tools to predict the mechanism of action through which Nagab-9 intervenes in Acute Lung Injury. It further validates the pharmacological effects and mechanisms using a lipopolysaccharide (LPS)-induced acute lung injury model in mice, aiming to provide experimental evidence for the subsequent development and application research of this medication.
Methods
Candidate Compounds of Nagab-9
With the aid of the Traditional Chinese Medicine Systems Platform (TCMSP) (https://tcmspw.com/), this study identified the chemical constituents of Nagab-9, excluding mineral-based compounds such as realgar. Utilizing the TCMSP database platform, this study evaluated the pharmacokinetic properties (Absorption, Distribution, Metabolism, Excretion, ADME) of the principal compound components. 7 Chemical constituents that simultaneously met the criteria of an Oral Bioavailability (OB) ≥ 40% and Drug-Likeness (DL) ≥ 0.18 were selected as candidate active ingredients. Here, OB is directly related to bioavailability, while DL refers to the similarity between the molecule under examination and known drug molecules, essentially indicating the molecule’s likelihood of being a viable drug. 8 Molecular structures of each active compound were confirmed through literature mining and platforms such as PubChem (https://pubchem.ncbi.nlm.nih.gov/).
Candidate Targets Related to Nagab-9 and ALI
The compound-related targets of Nagab-9 were collected from TCMSP with each candidate compound. The Human Gene Database (GeneCards) and UniProt were used for standardizing the names of target proteins to Homo sapiens. The targets failed to meet the condition were not selected for further analysis. Meanwhile, the disease-related targets were collected from three databases as follows: GenCards, Online Mendelian Inheritance in Man (OMIM) database, and DisGeNET. Two keywords “acute lung injury” and “acute respiratory distress syndrome” were used for searching, and only “Homo sapiens” proteins linked to the disease were selected. Finally, the candidate targets were obtained from the overlaps between the compound related targets and the disease-related targets.
Functional and Pathway Enrichment Analysis
The intersecting targets identified above were imported into the DAVID database for GO and KEGG pathway enrichment analysis, specifying “Homo sapiens” as the species. A P-value < .05 was used as the selection criterion, and the ChiPlot online platform was employed for visualization.
“Compound-Target-Pathway-Disease” Network Construction
Data from KEGG pathway analysis, identifying the top 10 pathways related to ALI and their targets, as well as compounds interacting with these targets, were imported into Cytoscape software to construct a “Compound-Target-Pathway-Disease” network. Based on degree centrality, the top six compounds were selected as the core components of Nagab-9 for ALI treatment.
PPI Analyses and Kernel Target Genes
By intersecting the targets of the compounds with disease-related targets, the potential targets of Nagab-9 for intervening in ALI are identified. Intersecting targets were uploaded to the STRING database to create a Protein-Protein Interaction (PPI) network. Following the removal of isolated nodes, network topology was analyzed using Cytoscape 3.9.1, and topological parameters of network nodes were assessed with the Network Analyzer plugin. The top six targets, selected based on degree centrality, were identified as key targets for Nagab-9 in the treatment of ALI.
Molecular Docking
The selected core components were subjected to molecular docking validation with the top six key target proteins. Initial steps involved downloading 3D structures of active components in SDF format from the PubChem database and 3D structures of key target proteins in PDB format from the RCSB database. Molecular optimization was conducted using SYBYL-X 2.0 software with settings as follows: Tripos force field application, Gasteiger–Hückel charge assignment, a maximum iteration limit set to 10 000, and an energy gradient confined to 0.005 kcal mol−1 Å−1, with all other parameters defaulted.9,10 Core target proteins were rectified using mgltools_win 32_1.5.6 software, followed by receptor pre-treatment such as the removal of water molecules and metal ions, subsequently saved as PDBQT files. The affinity of docking between key target proteins and core components was assessed using AutoDockTools 1.5.7, quantified by binding energy (kcal mol−1), with final molecular docking data visualization rendered by Discovery Studio (Figure 1). The Flow Chart of Network Pharmacology and Molecular Docking.
Animal Grouping, Administration, and Modeling
Forty-eight SPF-grade male healthy ICR mice were adaptively fed for 7 days and then stratified by body mass and randomly divided into control, model, low, medium, and high-dose Nagab-9 groups, and a dexamethasone(DEX) group, with seven mice per group. The dosages of Nagab-9 for mice were calculated as 5, 10, and 20 times the adult (60 kg) clinical daily dosage of 62 g, equivalent to 63, 126, and 252 mg/kg, respectively. Mice were administered orally once daily for 10 consecutive days, and the DEX group received 3.5 mg/kg on days 1, 3, 5, 7, 9, and 10. Both the control and model groups received an equal volume of saline daily, with a gavage volume of 0.1 mL/10 g. Prior to modeling, mice were fasted but not water-restricted for 12 h. On the day of modeling, 30 min after administration, mice in all groups were anesthetized with an intraperitoneal injection of sodium pentobarbital (30 mg/kg), fixed in a supine position, and after disinfection of the ventral neck area, a small incision was made to expose the trachea, into which a sterile injection needle was inserted at an angle for dripping. The control group received an equivalent volume of sterile saline dripped into the trachea, while the remaining groups received 100 μL of LPS (5 mg/kg) to replicate the mouse ALI model. After tracheal instillation, mice were held upright to facilitate the distribution of the LPS solution into the lungs, followed by suturing of the incision and application of iodine for disinfection. Three hours post-modeling, mice in all groups were administered a dose of their respective treatments, and six hours post-modeling, mice were euthanized to collect bronchoalveolar lavage fluid (BALF).
11
The entire lungs were lavaged three times with 0.5 mL of pre-cooled phosphate-buffered saline, the collected BALF was centrifuged at 4°C, 3000 rpm for 15 min, the supernatant was collected, and stored at −80°C for later use. This study was conducted at the Central Laboratory of the Hohhot Hospital of Traditional Chinese Medicine and Mongolian Medicine in Inner Mongolia Autonomous Region, China, from January 2023 to October 2023. The nature of the study is basic research (Figure 2). Schematic Illustration of the Experimental Protocols of the Study.
Pulmonary Function Testing in Mice
At 2 h after the second administration (28 h after modeling), a non-invasive pulmonary function testing system was employed to measure tidal volume (TV), minute ventilation (MV), peak inspiratory flow (PIF), peak expiratory flow (PEF), and mid-expiratory flow (EF50) in each group of mice to evaluate lung function.
Histological Morphological Observation of Lung Tissues
Lower lobes of right lung tissue were fixed in 10% neutral buffered formalin, dehydrated using an automated dehydration instrument, embedded in paraffin, and sectioned. Sections were stained with hematoxylin and eosin (HE) and structural alterations in lung tissue were examined under an optical microscope at 400× magnification. The percentage of inflammatory cell infiltration in lung tissues was quantitatively assessed using ImageJ software.
Ratio of Lung Tissue Wet Weight to Dry Weight (W/D)
The left lung was rinsed with saline to remove any debris, then blotted dry to measure its wet weight. Subsequently, the lung was dried overnight in an oven set to 65°C to determine its dry weight. The wet-to-dry weight ratio (W/D) was calculated based on these measurements.
Determination of Relevant Parameters in BALF
The supernatant of bronchoalveolar lavage fluid (BALF) was collected and processed according to the instructions of the respective reagent kits. The total protein content was quantified using the bicinchoninic acid (BCA) method. Nitric oxide (NO) levels were determined by the Griess assay. Myeloperoxidase (MPO) levels were assessed using an enzyme-linked immunosorbent assay (ELISA). Levels of TNF-α, IFN-γ, IL-1β, IL-6, VEGF, and CXCL1 were measured using Luminex xMAP technology.
Expression of Ly6G and Cit-H3 in Lung Tissues
Lung tissue sections were dewaxed, rehydrated, subjected to antigen retrieval, and blocked with 10% donkey serum. They were then incubated overnight at 4°C with primary antibodies against Ly6G (1:500) and Cit-H3 (1:100). Following TBST buffer washes, sections were incubated with secondary antibodies, and nuclei were stained with DAPI (1:500). The slides were coverslipped, and the activation of neutrophils and the formation of neutrophil extracellular traps (NETs) in lung tissue were observed using a TSC SP8 STED super-resolution confocal microscope. The fluorescence intensity of Ly6G and Cit-H3 in lung tissue was quantitatively analyzed using ImageJ software.
Protein Expression Detection of Cit-H3, SRC, p-SRC, ERK1/2, p-ERK1/2, STAT3, and p-STAT3 in Lung Tissues
Mouse lung tissues were homogenized in RIPA lysis buffer, followed by centrifugation to collect the supernatant. Protein concentration was measured using a BCA assay kit, and samples were normalized to the same concentration. Proteins were separated by 10% SDS-PAGE and transferred to a PVDF membrane, which was blocked with 5% non-fat milk. Primary antibodies against Cit-H3 (1:1000), STAT3 (1:2000), p-STAT3 (1:2000), SRC (1:2000), p-SRC (1:1000), ERK1/2 (1:2000), p-ERK1/2 (1:2000), β-actin (1:2000), and vinculin (1:3000) were incubated overnight at 4°C. After washing the membrane five times, each for 6 minutes, it was incubated with HRP-conjugated goat anti-rabbit IgG antibody (1:5000) for 90 min. Following a final wash, the membrane was developed using ECL substrate and exposed in an imaging system, with band data analyzed using Image Lab software version 6.0.1.
Culturing and Grouping of Mouse Lung Epithelial Cell Line (MLE-12)
In a humidified incubator at 37°C and 5% CO2, cells were cultured in DMEM supplemented with 10% FBS and 1% penicillin-streptomycin. After 2-3 days of culture, the cells were passaged. Cells in good growth condition and in the logarithmic phase were selected for further experiments. MLE-12 cells were divided into four groups: a blank group, a model group, a Nagab-9 group, and a DEX group. The model, Nagab-9, and DEX groups were treated with LPS (5 mg of LPS was dissolved in 1 mL of DMSO to prepare a 5 g·L−1 solution, stored at −20°C).
Cell Viability Assessment
MLE-12 cells in the logarithmic growth phase were seeded in a 96-well plate at a density of 5 × 103 cells/well. After cell adhesion, they were treated with various concentrations of LPS along with the addition of drugs, and then incubated at 37°C and 5% CO2 for 24 h. The culture medium was removed, and 10 μL of CCK-8 solution was added to each well, followed by further incubation for 30 min. Finally, cell viability was determined by measuring the absorbance (A) at 450 nm. Cell survival rate = (A measured − A blank) / (A control − A blank) × 100%.
Analysis of Relative Expression of SRC, p-SRC, ERK1/2, p-ERK1/2, STAT3, and p-STAT3 mRNA by qRT-PCR
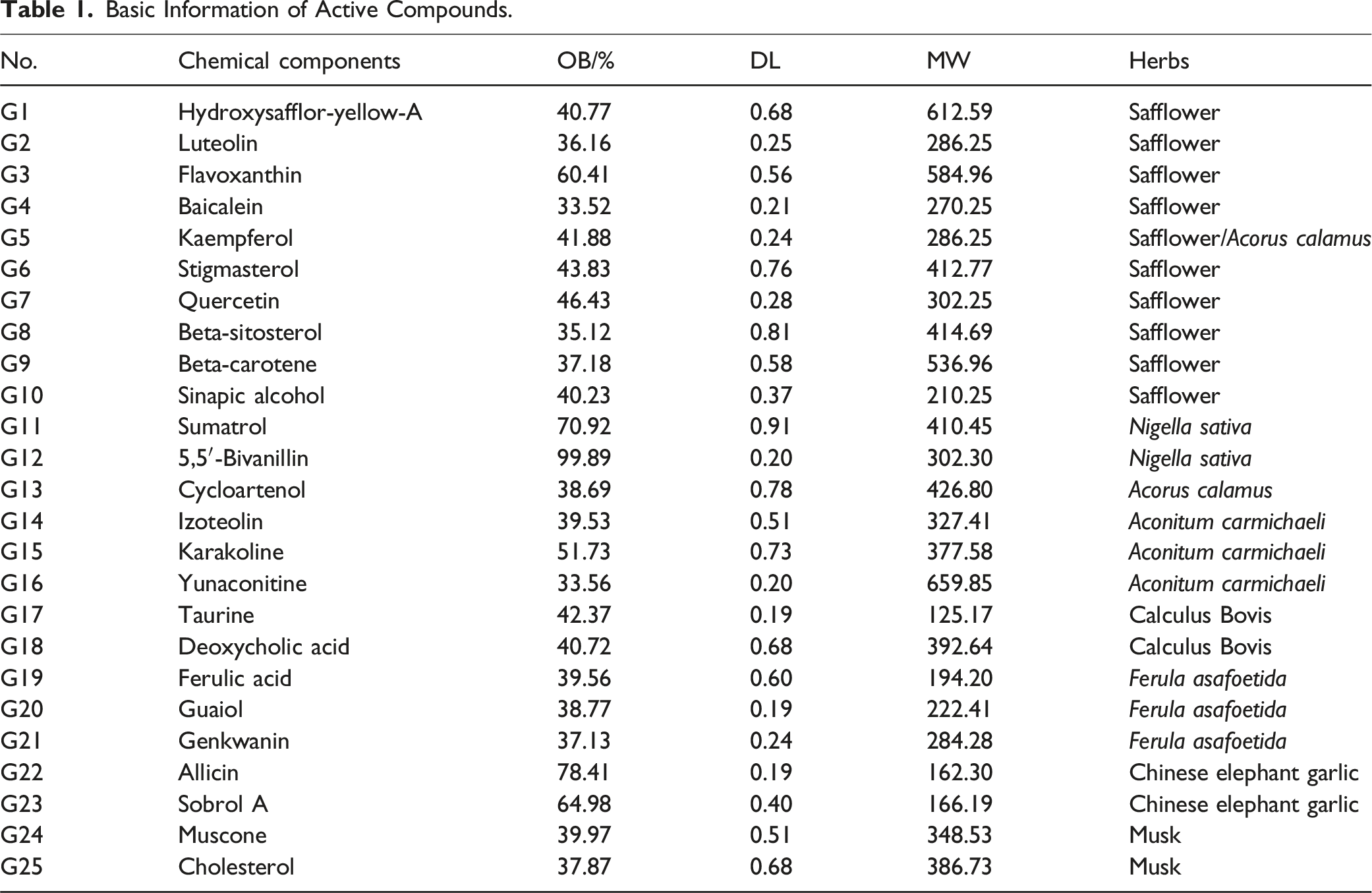
Basic Information of Active Compounds.
Western Blot Analysis of Protein Expression in MLE-12 Cells
Total proteins were extracted from MLE-12 cells using RIPA buffer, and their concentration was determined using a BCA protein assay kit. Following normalization to the lowest protein concentration, samples underwent SDS-PAGE and were subsequently transferred to a PVDF membrane. The process included blocking, washing, incubation with the primary antibody, washing again, incubation with the secondary antibody, and chemiluminescent detection. The relative expression levels of target proteins were quantified by analyzing the grayscale values of the target and reference proteins using ImageJ software.
Statistics
For bioinformatics analyses, the data were analyzed using the hypergeometric distribution test and Fisher’s exact test. Te Benjamini-Hochberg method was used to correct the false discovery rate (FDR). Otherwise, the data were expressed as means ± standard deviation (SD) and analyzed using the statistical package for the social sciences (SPSS) 21.0 software (IBM, NY, USA). Te means of multiple groups were compared with one-way analysis of variance (ANOVA), which is preliminary performed by checking homogeneity of variance, and then followed by the least-signifcant diference (LSD) method. A P < .05 was considered statistically significant.
Results
Active Ingredients and Target Screening
Based on the active compounds listed in the TCMSP and ETCM databases, this study confirms the molecular structure of each active compound using the TCMSP and PubChem databases. Additionally, by evaluating oral bioavailability, drug-likeness, and Lipinski’s rule, a total of 25 effective compounds within Nagab-9 were identified. The basic information of the active compounds is presented in Table 1. Target prediction for the active compounds in Nagab-9 was conducted using the SWISS and Superpred websites, resulting in a total of 801 unique targets after deduplication. By merging and removing duplicates across the DisGeNET and GeneCards databases, and selecting targets with a relevance score above the median, a total of 598 disease targets were ultimately identified. By intersecting the 801 targets of active compounds in Nagab-9 with the 598 ALI-related targets, 89 potential targets for Nagab-9 intervention in ALI were identified,shown in Figure 3A. (A) Venn Diagram of “Component-Disease” targets; (B) PPI Network of the Top 10 targets; (C) Network Analysis of “Active Component-Target-Pathway-Disease”; (D) GO Enrichment Analysis (BP: Biological Process; CC: Cellular Component; MF: Molecular Function); (E) KEGG Pathway Analysis.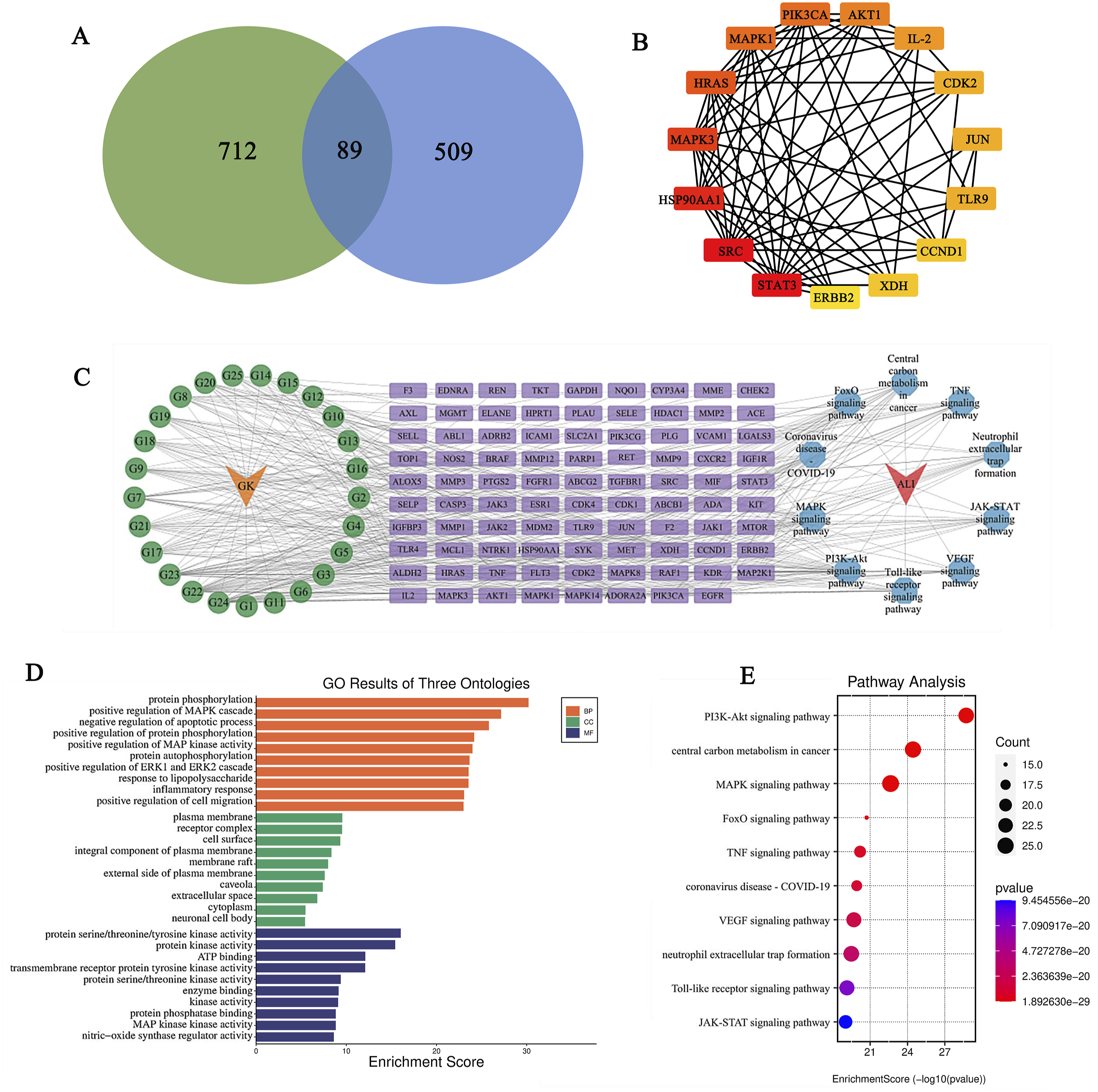
Construction of PPI Network
The intersection of 89 common targets was imported into the STRING database, with isolated targets hidden, to construct a PPI network of common targets comprising 73 nodes and 329 edges at the highest confidence level (0.900). Here, nodes represent common targets; node size and color intensity indicate the degree of connectivity, with larger and darker nodes denoting higher degrees and greater target relevance. Lines between nodes signify interactions between two common targets. The common target PPI network of Nagab-9 active components and ALI disease was visualized using Cytoscape 3.9.1 software, and topological parameters of network nodes were analyzed with the Network Analyzer plugin. If degree values were identical, targets were ranked by betweenness centrality, selecting the top 6 targets (STAT3, SRC, HSP90AA1, MAPK3, HRAS, MAPK1) as key targets for Nagab-9 intervention in ALI. Results are shown in Figure 3B.
GO Enrichment Analysis and KEGG Pathway Analysis
Utilizing the DAVID 6.8 database, a GO functional enrichment analysis was conducted on 89 common targets (P < .05), yielding 668 entries, including 514 biological processes (BP), 67 cellular components (CC), and 87 molecular functions (MF). The top 10 entries for each were visually represented based on P-value. BPs primarily involved protein phosphorylation, positive regulation of MAPK cascade, positive regulation of ERK1 and ERK2 cascade, inflammatory response, and response to lipopolysaccharide. CCs mainly concerned plasma membrane, receptor complex, components of the plasma membrane, extracellular space, and cytoplasm. MFs predominantly related to protein serine/threonine/tyrosine kinase activity, protein kinase activity, ATP binding, enzyme binding, and nitric oxide synthase regulator activity, reflecting the multifaceted effects of Nagab-9 intervention in ALI. KEGG pathway enrichment analysis identified 131 related signaling pathways (P < .05), from which the top 10 were ranked and visualized in a bubble chart according to P-value. These pathways involve viruses, inflammation, cancer, etc., with the PI3K-Akt signaling pathway, MAPK signaling pathway, NET formation, VEGF signaling pathway, and JAK-STAT signaling pathway being among the top ranked, suggesting a potential association of Nagab-9 intervention in ALI with these pathways. Results are shown in Figure 3D and E.
Construction of “Compound-Target-Pathway-Disease”” Network
The top 10 KEGG pathways identified as related to ALI, along with their targets and interacting compounds, were imported into Cytoscape to construct a “compound-target-pathway-disease” network. Using the Network Analyzer plugin, a topological analysis of network nodes was performed to visually demonstrate the relationships between the components. Based on the degree of centrality, active components such as hydroxysafflor yellow A, ferulic acid, taurine, sinapic alcohol, luteolin, and genkwanin were ranked highly, suggesting their potential as active components in ALI intervention. In terms of targets, MAPK3, MAPK1, STAT3, SRC, and AKT1 were among the top, indicating their significance as disease treatment targets. Pathways such as the MAPK signaling pathway, NET formation, JAK-STAT signaling pathway, and VEGF signaling pathway were identified as crucial in Nagab-9 treatment of ALI. This reveals the multi-component, multi-target, and multi-pathway characteristics of Nagab-9’s intervention in acute lung injury.
Molecular Docking
Based on the degree values, six core components were selected: hydroxysafflor yellow A, ferulic acid, taurine, sinapic alcohol, luteolin, and genkwanin. These core components were docked with six key targets: STAT3 (PDB ID: 5AX3), SRC (PDB ID: 4U5J), HSP90AA1 (PDB ID: 1BYQ), MAPK3 (PDB ID: 4QTB), HARS (PDB ID: 6G54), and MAPK1 (PDB ID: 8AOJ), to predict their binding affinities. Binding energies lower than −4.25 kcal mol−1 indicate good binding activity between the drug components and targets; energies less than −5.0 kcal mol−1 suggest strong binding activity. The smaller the binding energy, the better the docking results. The binding energies of Nagab-9’s six core components with the six key targets were all below −4.25 kcal mol−1, with MAPK3, MAPK1, SRC, and STAT3 showing binding energies below −5.0 kcal mol−1 with the six core components, indicating a strong association between the six core components and the four key targets (MAPK3, SRC, MAPK1, STAT3). Interactions between core components and key targets include van der Waals forces, π-alkyl, π-π stacked, π-cation, and hydrogen bonds.
The Effects of Nagab-9 on Survival and Pulmonary Function in ALI Mice
Compared to the model group, the 7-day survival rate of mice in both the Nagab-9 and DEX groups significantly increased (P < .01). Regarding body mass, the control group showed a steady increase in body mass, while the model group exhibited a significant decrease post-modeling. The rate of body mass decrease was notably slower in the Nagab-9 and DEX groups compared to the model group, as shown in Figure 5A–F. Further assessment of mouse lung function through a non-invasive lung function measurement system revealed that, compared to the control group, the TV, MV, PIF, PEF, and EF50 in the model group were significantly reduced (P < .01); compared to the model group, these parameters significantly improved in the Nagab-9 and DEX groups (P < .05).
The Effects of Nagab-9 on Lung Tissue W/D Ratio and Histopathology in ALI Mice
The lung tissue wet-to-dry weight ratio (W/D) reflects changes in capillary permeability and the degree of pulmonary edema. Compared to the control group, the model group exhibited a significant increase in lung tissue W/D; compared to the model group, both the Nagab-9 and DEX groups showed a decrease in lung tissue W/D, with the Nagab-9 group showing a more pronounced decrease, as shown in Figure 5G. Histological sections of lung tissues showed that, in the control group, the lung structure was intact with no significant inflammatory infiltration or fibroconnective tissue proliferation; the structure of the bronchi at all levels was clear and intact, with normal morphology of bronchial epithelial cells and alveolar epithelial cells. In the model group, bronchial epithelial cells were shedding (see green arrow), the alveolar septa were slightly thickened, and inflammatory cells were visibly present in the alveolar space and interstitium, primarily infiltrated by neutrophils and lymphocytes. Compared to the model group, the pathological structure of lung tissues in the Nagab-9 and DEX groups was significantly improved. Pathological scoring showed that, compared to the control group, the pathological score of the model group significantly increased (P < .001); compared to the model group, the pathological scores of the Nagab-9 and DEX groups were significantly reduced (P < .01), as shown in Figure 4A. (A) The Effect of Nagab-9 on Histopathological Changes in the Lungs of Mice With Acute Lung Injury (H&E Staining, ×400; x ± s, n = 5), With Green Arrows Indicating Bronchial Epithelial Cell Detachment; Red Arrows Indicating Alveolar Wall Thickening; Blue Arrows Indicating Neutrophils; Fluorescent Yellow Arrows Indicating lymphocytes; (B) The Effect of Nagab-9 on the Expression of Ly6G and Cit-H3 in Lung Tissue of Mice With Acute Lung Injury (x±s, n = 3), With Green Fluorescence Labeling Ly6G; Red Fluorescence Labeling Cit-H3; Blue Fluorescence Labeling Nuclei. Compared With the Control Group *P < .05, **P < .01, ***P < .001; Compared With the Model Group #P < .05, ##P < .01, ###P < .001.
The Effects of Nagab-9 on Total Protein, NO, MPO, and Related Inflammatory Factor Levels in BALF of ALI Mice
Compared to the control group, the model group mice exhibited significantly increased levels of total protein, NO, and MPO in the BALF, along with significantly elevated levels of cytokines IL-6, IFN-γ, TNF-α, IL-1β, and VEGF. The total protein content reflects the degree of protein leakage caused by damage to the alveolar capillaries. MPO is a specific marker reflecting inflammation, neutrophil infiltration, and tissue damage. CXCL1 is a chemokine for neutrophils, and the recruitment of neutrophils to damaged tissue requires the involvement of CXCL1. VEGF is an indicator for assessing damage to alveolar capillary endothelial cells and the extent of pulmonary edema. NO, IL-6, IFN-γ, TNF-α, and IL-1β are all inflammatory mediators, indicating the successful establishment of the mouse ALI model. Compared to the model group, all dosage groups of Nagab-9 were able to reduce the levels of total protein, NO, MPO, CXCL1, IFN-γ, and VEGF in the BALF. The high-dose Nagab-9 group significantly decreased the levels of IL-6, TNF-α, and IL-1β in the BALF, indicating that this formulation can exert a beneficial preventive and therapeutic effect on mice with ALI models by alleviating pulmonary inflammatory responses. Results are shown in Figure 5(H–P). (A) Changes in Body Weight Among Different Groups of Mice; B-F The Effects on Survival and Lung Function in Different Groups of Mice (x±s, n = 7); (G) The Impact of Nagab-9 on the Wet/Dry Weight Ratio of Lung Tissue in Mice With Acute Lung Injury (x±s, n = 7); (H-P) The Effects of Nagab-9 on the Total Protein, NO, Related Inflammatory Factors, MPO, and CXCL1 Contents in BALF of Mice With Acute Lung Injury (x±s, n = 5∼7). Compared With the Control Group *P < .05, **P < .01, ***P < .001; Compared With the Model Group #P < .05, ##P < .01, ###P < .001.
The Effects of Nagab-9 on Neutrophil Infiltration and NET Formation in Lung Tissues of ALI Mice
Based on the results predicted by network pharmacology and the outcomes of previous pharmacological experiments, the high-dose group of Nagab-9 was selected as the representative group to investigate its effects on NETs. Ly6G is a surface marker of neutrophils, reflecting the degree of neutrophil infiltration. NETs are web-like structures formed by neutrophils upon stimulation (e.g., LPS, viruses) with DNA as the scaffold. The primary components are double-stranded DNA and histones, followed by granular enzymes and peptides, including NE, MPO, etc., which are closely related to the defensive function of NETs. NETs are responsible for capturing and eliminating pathogenic microorganisms that damage the body, but their excessive activation triggers more NET formation, leading to lung damage in the body. Cit-H3 serves as an important marker of NET formation. Compared to the control group, mouse lung tissues in the model group showed increased secretion of Ly6G and Cit-H3, along with elevated protein expression levels of Cit-H3. After treatment with DEX and Nagab-9, secretion of Ly6G and Cit-H3 was reduced, and the protein expression level of Cit-H3 was downregulated, indicating that both treatments could lessen the degree of neutrophil infiltration and NET formation, as shown in Figures 6B and 7A. (A-H) Molecular Docking of the Eight Core Components With the Lowest Binding Energies to Key targets; (I) Heatmap of the Lowest Binding Energies for the Docking of Nagab-9 Core Components With Key Targets. (A) The Effect of Nagab-9 on Cit-H3/β-Actin Protein Expression in Lung Tissue of Mice With ALI (x±s, n = 4); (B) The Effect of Nagab-9 on the Ratio of p-SRC/SRC, p-ERK1/2/ERK1/2, and p-STAT3/STAT3 Protein Expression in Lung Tissue of Mice With ALI (x±s, n = 4); (C) The Effect of Different Concentrations of LPS and Nagab-9 on the Viability of MLE-12 Cells (x±s, n = 4); (D) The Effect of Nagab-9 on SRC, ERK1/2, and STAT3 mRNA Expression in MLE-12 cells; (E) The Effect of Nagab-9 on the Ratio of p-SRC/SRC, p-ERK1/2/ERK1/2, and p-STAT3/STAT3 Protein Expression in MLE-12 Cells (x±s, n = 4). Compared With the Control Group *P < .05, **P < .01, ***P < .001; Compared With the Model Group #P < .05, ##P < .01, ###P < .001.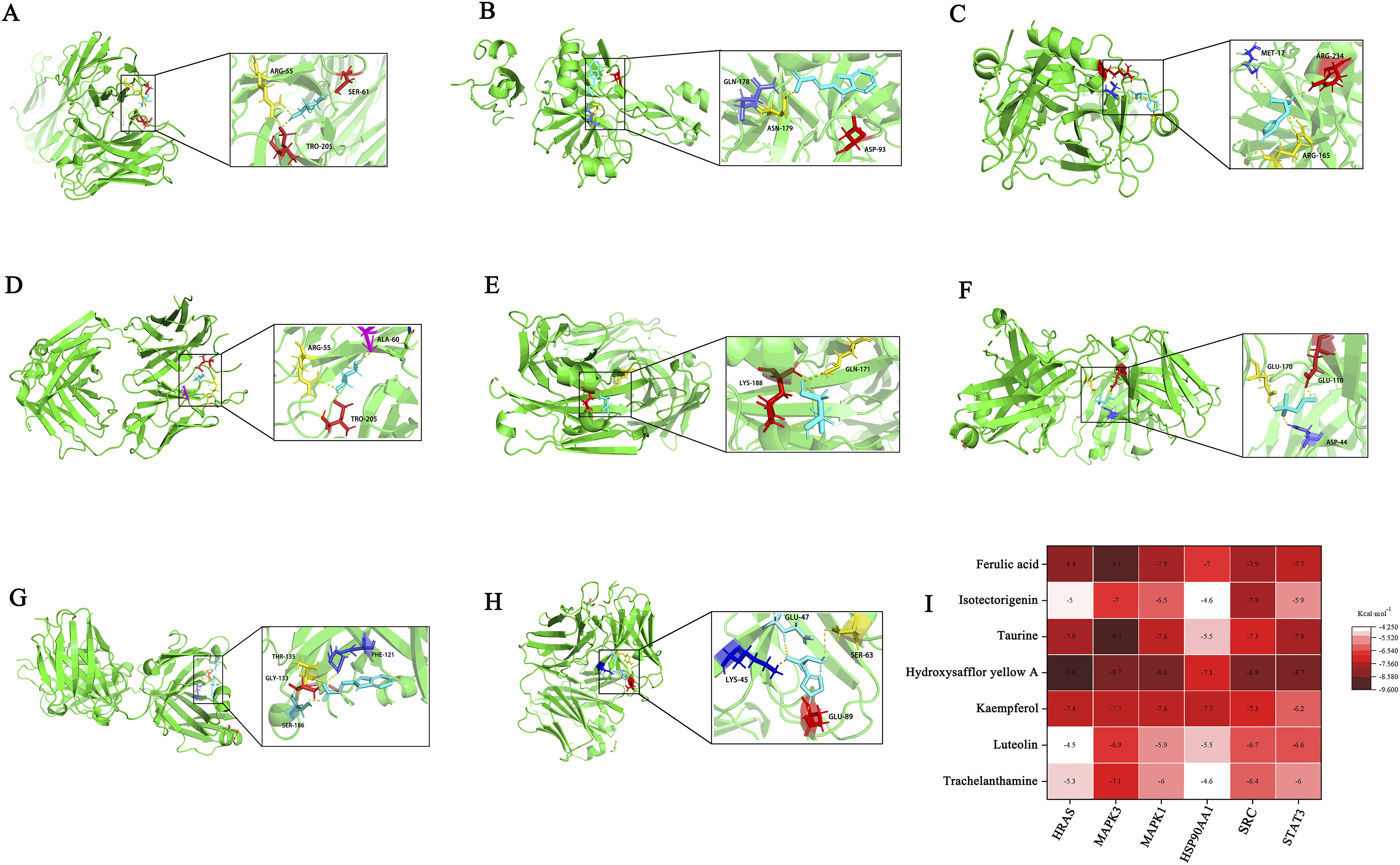

The Effects of Nagab-9 on SRC, ERK1/2, and STAT3 Protein Expression in Lung Tissues and MLE-12 Cells of ALI Mice
The ratio of phosphorylated protein to total protein reflects the impact of the high-dose group of Nagab-9 on the expression of SRC, ERK1/2, and STAT3 proteins, as shown in Figure 7B. Compared to the control group, the expression levels of p-SRC, p-ERK1/2, and p-STAT3 proteins in the lung tissues of the model group mice were significantly upregulated. In mice pre-treated with DEX and Nagab-9, the expression levels of p-SRC, p-ERK1/2, and p-STAT3 proteins in lung tissues were significantly downregulated compared to the model group. Alveolar epithelial cells were stimulated with different concentrations of LPS, and the results showed that LPS at concentrations of 25 and 50 mg L−1 significantly inhibited cell activity (P < .01). Therefore, 25 mg·L-1 LPS was selected for subsequent experiments. Alveolar epithelial cells stimulated with LPS were treated with different concentrations of Nagab-9. As the concentration of Nagab-9 increased, the LPS-induced decrease in cell viability gradually reversed. When the concentration of Nagab-9 reached 2.5 mg L−1, a significant increase in cell viability was observed (P < .01), as shown in Figure 7C. Therefore, the concentration of 2.5 mg L−1 was selected for subsequent in vitro intervention experiments with Nagab-9. Analysis of the expression of SRC, ERK1/2, and STAT3 proteins revealed that, compared to control group, the LPS-stimulated group showed significantly upregulated expression levels of p-SRC, p-ERK1/2, and p-STAT3 proteins (P < .001); after the administration of Nagab-9, protein activation was inhibited (P < .001). These results indicate that the protective effect of Nagab-9 on alveolar epithelial cells is achieved through or partly through the regulation of SRC, ERK1/2, and STAT3 protein expressions.
Discussion
ALI presents a significant challenge in critical care medicine, characterized by acute respiratory distress and refractory hypoxemia due to various causes. The primary feature of ALI is an excessive inflammatory response leading to diffuse alveolar-capillary membrane damage, resulting in pulmonary edema, atelectasis, increased capillary permeability, and impaired gas exchange in the lungs. 12 In this study, an ALI model was established in mice by tracheal instillation of LPS to simulate the occurrence of clinical ALI. 13 Given that corticosteroids are frequently used in the clinical treatment of ALI, we chose Dexamethasone as a positive control drug, providing a relevant benchmark for evaluating the therapeutic effect of Nagab-9 on ALI. 14
The LPS-induced ALI animal model shares several key features with clinical ALI patients, including pulmonary edema, respiratory dysfunction, extensive infiltration of inflammatory cells in the lungs, and the excessive release of early inflammatory mediators. In preliminary studies, animals in the LPS group exhibited rapid breathing 4 days after intraperitoneal injection of LPS; blood gas analysis indicated metabolic acidosis combined with respiratory acidosis and hypoxemia; HE staining pathology results revealed an acute inflammatory response in lung tissue. 15 These manifestations are consistent with lung damage and clinical presentations in ALI patients, indicating the success of our ALI animal model.
The traditional efficacy of the Mongolian medicine “Nagab-9” is well-documented, yet modern research is still in its exploratory phase, particularly in clinical studies, mechanisms of action, and identification of drug targets. Literature searches have shown that Liang Cun and colleagues determined the content of hydroxysafflor yellow A in “Nagab-9” using HPLC, finding a good linear relationship within the range of 54-2160 ng. 16 In recent COVID-19 research, Zhang Jizhong and others investigated ethnopharmacological measures against COVID-19, noting Mongolian medicine methods to prevent COVID-19, including wearing “Nagab-9” as a preventive measure. 17 The second and third editions of the “Mongolian Medicine Prevention and Treatment Plan for Novel Coronavirus Pneumonia” issued by the Health Commission of Inner Mongolia Autonomous Region mention wearing Nagab-9 as a preventative method against novel coronavirus pneumonia. 18 Preliminary studies involving smoke-induced acute bronchitis in rats found that Nagab-9 could downregulate lung tissue levels of PI3K and p-Akt proteins, reduce the secretion levels of inflammatory cytokines IL-1β, IL-6, and TNF-α, effectively alleviating lung inflammation damage. Thus, it can be considered that Nagab-9 has a good anti-inflammatory effect, which forms an important pharmacological basis for its use in respiratory inflammatory diseases. However, whether Nagab-9 intervenes in acute lung injury and its mechanism of action remain unclear.
Network pharmacology is considered a bridge connecting the active components of traditional Chinese medicine (TCM), relevant targets, and associated diseases. It features a systemic and holistic approach, aiding in predicting the action targets of TCM for treating specific diseases. Furthermore, through molecular docking methods, it is possible to infer the binding sites and affinity of active components with target proteins. Oral administration is a common method for administering TCM and its compounds, with the absorbed components being considered the active ingredient group that enters the body and directly relates to the pharmacological effects. 19 This study utilizes network pharmacology to identify targets based on the effective compounds of Nagab-9, intersecting these with ALI-related targets to construct a PPI network. The results identify STAT3, SRC, HSP90AA1, MAPK3, HRAS, and MAPK1 as six targets with higher degree values.
GO function and KEGG pathway analyses indicate that most targets are enriched in pathways related to inflammatory response, ERK1 and ERK2 cascade, response to lipopolysaccharide, as well as MAPK, NET formation, VEGF, JAK-STAT, among others. In conjunction with the “compound-target-pathway-disease” network, it is shown that hydroxysafflor yellow A, ferulic acid, taurine, sinapic alcohol, luteolin, and genkwanin are ranked high in terms of degree value, suggesting these six components may be the core constituents of Nagab-9 in intervening ALI. Further prediction of the binding possibilities between six core components and key targets through molecular docking techniques revealed a strong association between MAPK3, SRC, MAPK1, STAT3, and the six components. Additionally, according to existing literature, safflor yellow (SY) is the main effective part of safflower, and hydroxysaffloryellow A (HSYA) is the most abundant active monomer in safflor yellow, accounting for about 70% of its content. 16 The structure of HSYA was first determined by Japanese scholar Meselhy et al in 1993, with a relative molecular mass of 612. Wang’s research demonstrates that HSYA can alleviate symptoms of pulmonary congestion and edema, suppress the reduction of arterial oxygen partial pressure, decrease the incidence of acidosis, protect alveolar type II epithelial cells, downregulate the expression levels of TNF-α and ICAM-1 mRNA in lung tissue, and reduce the plasma levels of IL-1β and IL-6. 20 They suggest that HSYA may alleviate ALI by inhibiting the expression of pro-inflammatory cytokines and endothelial cell adhesion molecules, thereby reducing the accumulation, adhesion, activation of inflammatory cells in pulmonary vessels, and the release of inflammatory mediators during the development of ALI.21,22 Ferulic acid (FA), a polyphenolic compound extracted from natural plants, possesses anti-inflammatory, anti-edematous, and antioxidant pharmacological properties. 23 Studies have reported that FA can alleviate LPS-induced oxidative stress responses by reducing the activity of superoxide dismutase and glutathione peroxidase in pulmonary epithelial cells. 24 Additionally, FA has been proven to mitigate acute respiratory distress syndrome in mice induced by LPS through its anti-inflammatory and antioxidant activities. Most research on FA in treating pulmonary injuries and other respiratory diseases has focused on inflammatory infiltration and free radical scavenging, with less attention to the levels of pulmonary epithelial ion transport. 25 Consequently, it is preliminarily considered feasible that Nagab-9, as predicted by network pharmacology, intervenes in ALI by regulating four key targets: MAPK3, SRC, MAPK1, and STAT3, to inhibit the inflammatory response.
In animal experiments, an ALI model in mice induced by intratracheal instillation of LPS is selected, which has been applied in numerous fundamental research studies on acute lung injury. This model is mature, reliable, and highly reproducible. LPS, as a component of the outer membrane of Gram-negative bacteria, is pathogenic. Upon entering the organism, it binds to cell surface receptors, stimulating the recruitment and infiltration of inflammatory cells such as macrophages and neutrophils into lung tissue. This increases the synthesis and release of inflammatory mediators (such as IFNs, TNF-α, ILs, etc.), inducing an inflammatory cascade that leads to lung tissue damage and functional impairment. MAPK3 and MAPK1, as extracellular signal-regulated kinases (ERK1/2), are vital components of the MAPK family. 26 ERK1/2 regulates processes such as cell proliferation, differentiation, and apoptosis, and can also intervene in inflammatory diseases. After LPS enters the body, it can mediate through Toll-like receptor 4 (TLR4) and CD14, leading to the activation and increased phosphorylation of ERK1/2, which results in elevated levels of inflammatory mediators (NO, IL-1β, IL-6, TNF-α, VEGF, MPO, CXCL1, etc.) and induces an inflammatory cascade. STAT3 is a major part of the JAK/STAT signaling pathway, playing a broad regulatory role in the body, including cell survival, proliferation, and inflammation. Research shows that in LPS-induced inflammation models, STAT3 is rapidly activated, its phosphorylation level increases, and this promotes the release of TNF-α, CXCL1, and exacerbates inflammatory cell infiltration. STAT3 is downstream of ERK1/2, and the ERK1/2/STAT3 signaling pathway can be activated in LPS-induced inflammation models, participating in the development of the inflammatory response. Increased phosphorylation levels of ERK1/2 and STAT3 lead to abnormal levels of pro-inflammatory cytokines such as TNF-α, IFN-γ, IL-1β, and IL-6, resulting in inflammatory lesions. 27 SRC, a tyrosine kinase, can regulate cell metabolism, proliferation, and inflammatory responses. In lung tissue of LPS-induced ALI models, increased phosphorylation levels of SRC further upregulate the expression of phosphorylated proteins such as ERK1/2 and STAT3, promoting the increase of inflammatory mediators like MPO, IL-6, and TNF-α, as well as exacerbating the extensive infiltration of neutrophils, excessive release of NETs, leading to lung tissue damage. 28 Inhibiting the phosphorylation levels of SRC and ERK1/2 can effectively reduce neutrophil infiltration in ALI and regulate Cit-H3 expression levels, suppress the formation of NETs, thereby alleviating disease progression. SRC is upstream of the ERK1/2/STAT3 pathway. Upon cellular entry, LPS activates the SRC/ERK1/2/STAT3 signaling pathway, promoting the rapid recruitment of innate immune cells (such as macrophages and neutrophils) to inflamed damaged tissues, synthesizing and releasing excessive inflammatory cytokines (TNF-α, IL-1β, IFN-γ, etc.), and exacerbating the body’s inflammatory response. 29 In this study, pretreatment with Nagab-9 significantly reduced the levels of inflammatory mediators NO, IL-1β, IL-6, TNF-α, IFN-γ, MPO, CXCL1 in the BALF of ALI mice, as well as decreased the expression levels of Ly6G, Cit-H3 in lung tissue, inhibited excessive NET formation, alleviated lung tissue pathological damage, and inflammatory cell infiltration; it also significantly reduced VEGF levels in BALF, decreased pulmonary vascular endothelial cell permeability, and alleviated pulmonary edema. Additionally, Nagab-9 significantly reduced the expression levels of phosphorylated proteins SRC, ERK1/2, and STAT3 in lung tissue of ALI model mice, preliminarily validating the key target sites for Nagab-9 intervention in acute lung injury as predicted by network pharmacology and molecular docking, namely MAPK3, SRC, MAPK1, STAT3, thus indicating the direction for further mechanism research.
Although this study explored the potential intervention mechanisms of Nagab-9 in ALI and confirmed its anti-inflammatory effects through in vivo and in vitro experiments, there are still some limitations. First, Nagab-9 is a compound preparation containing multiple active ingredients. Although key components were identified through network pharmacology, the specific effects of individual components were not thoroughly validated, limiting a comprehensive understanding of its pharmacologically active basis. Secondly, this study utilized an LPS-induced ALI model, which simulates the inflammatory response but may not fully capture the complex clinical pathophysiological state, potentially limiting the representation of Nagab-9’s effects under various ALI-inducing conditions. Finally, as this study’s experiments were primarily conducted in a mouse model, certain interspecies differences may exist. Future research should include larger-scale animal studies and clinical trials to further verify the efficacy and safety of Nagab-9.
Conclusion
In summary, Nagab-9 can effectively intervene in LPS-induced ALI, alleviate inflammatory responses, reduce the release of inflammatory factors, inhibit neutrophil infiltration and NET formation, and improve lung tissue pathology. The mechanism of action may be related to inhibiting the activation of the SRC/ERK1/2/STAT3 signaling pathway.
Footnotes
Informed Consent
The manuscript is approved by all authors for publication.
Author Contributions
S.L. designed and performed the experiments and analysed the data; Z.C. provided technical assistance; Y.J. provided intellectual input; L.B. Supervised the overall study and advised on study design and data interpretation; J.Y. wrote the manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research is funded by the following projects: National Natural Science Foundation of China (No. 81760748 and 81550047); The Major Science Foundation of Affiliated Hospital of Inner Mongolia Medical University(NYFY ZD 010); Natural Science Foundation of Inner Mongolia Autonomous Region(No. 2019MS08110); Inner Mongolia Autonomous Region “Prairie excellence” Project. Western Light Young Scholars Program of the Chinese Academy of Sciences; Inner Mongolia Medical University Youth Program (YKD2023QN037) for this research.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
All analyses were generated with existing packages, and no original code was created. The packages associated with this analysis can be found below. Data analysis was performed using a variety of biostatistical platforms Perseus 1.6.15.0 (https://www.coxdocs.org/), R (https://www.r-project.org/) and Python package (![]() ).
).
