Abstract
Objective: This paper explores the role of DNA methylation in α-irradiation damage at the cellular level. Methods: Human normal hepatocytes L-02 were irradiated using a 241 Am α source at doses of 0, 1.0, and 2.0 Gy. The methylation levels of the six differentially methylated genes were examined by pyrophosphate sequencing, and the mRNA expression levels of the six differentially methylated genes were examined by real-time fluorescence quantitative PCR. Results: The rate of γH2AX foci positive cells was significantly higher than that of the control group after irradiation of cells in different dose groups for 1 h and 2 h respectively (P < .05). The proportion of S-phase cells was significantly increased in the 1.0 Gy and 2.0 Gy dose groups compared with the control group (P < .05). The methylation levels of CDK2AP1, PDGFRL, PCDHB16 and FAS genes were significantly increased, while the mRNA expression levels were significantly decreased (P < .05). The expression levels of CDK2Apl, PCDHB16 and FAS were significantly negatively correlated with the methylation levels (P < .05). Conclusion: The α-particle radiation can affect gene expression at the epigenetic level, which led to the speculation that altered methylation levels of CDK2AP1, PCDHB16, and FAS genes may be involved in the α radiation damage process.
Introduction
Transuranium nuclides have long half-lives, high specific activities, and a minimal load in the human body. The health hazards to humans primarily stem from the high linear energy transfer (LET) alpha particles released during their decay process when they are inhaled or enter the body through the skin. 1 Alpha particles have a short range and poor penetrating ability but interact strongly with matter, resulting in significant energy deposition over a small area and inducing a series of cytogenetic effects. 2 Current research shows that the liver is one of the important target organs for alpha particle radiation.3,4
DNA methylation is a vital form of epigenetics, which means that the DNA sequence is not altered but the level of gene expression is changed, and this change can be stably transmitted during development and cell proliferation. 1 DNA methylation is closely related to the life activity chain. Abnormal alterations in genomic DNA methylation patterns will directly contribute to the development of human diseases and even cancer.5,6
Current research has discovered that ionizing radiation can cause changes in DNA methylation, which are related to the pathological processes of cancer. CDK2AP1 is a cyclin-dependent kinase 2 (CDK2) interacting protein that is implicated in the etiology and progression of a variety of tumors through signaling pathways such as ATM-P53 and TGF-β1, which are crucial in the DNA damage repair process triggered by ionizing radiation.7,8 PDGFRL, one of the many tumor suppressor genes located on chromosome 8, functions as a tumor suppressor within the PDGF signaling pathway, antagonizing its receptor and thus affecting the binding of PDGF to its receptor and the subsequent downstream signaling. 9 Mutations in the PRKAG2 gene can result in cardiac dysfunction and are implicated in cardiovascular diseases. 10 PCDHB16, part of the protocadherin β gene cluster, has been noted in studies of DNA hypermethylation changes in late-stage liver cancer through Gene Ontology (GO) analysis, and is associated with functions related to homophilic cell adhesion, intercellular adhesion, cell adhesion, and biomembrane adhesion. 11 The FAS gene, a member of the tumor necrosis factor receptor superfamily, plays a central role in the physiological regulation of apoptosis and is linked to the pathogenesis of various malignancies and immune system disorders. 12 HOXC4, a homeobox-containing gene, is capable of enhancing the proliferation of human hematopoietic cells and may be one of the principal regulatory genes for lymphopoiesis. 13
Alterations in DNA methylation serve as a pivotal molecular signature in investigating the pathogenic mechanisms induced by ionizing radiation. This study focuses on six genes that are potentially differentially methylated: CDK2AP1, PDGFRL, PRKAG2, PCDHB16, FAS, and HOXC4. Through in vitro hepatocyte experiments, the study examines the impact of radiation on cell proliferation activity, cell cycle regulation, and the DNA methylation patterns and transcriptional expression levels of the aforementioned genes.
Materials and Methods
Cell Culture
HL-7702[L-02] is a human normal hepatocyte. Under the normal growth state, its cell growth characteristics are adherent to the wall, and its cell morphology is characterized by epithelioid, short shuttle shape. HL-7702[L-02] hepatocytes were cultured in RPMI 1640 culture medium containing 10% fetal bovine serum in an incubator with 5% CO2 at 37°C. In this study, 3-6 generations of cells could be used to perform the experiments.
Irradiation Methods
A 241 Am α source from Soochow University was used to release α particles. The whole device consists of α-particle source, source holder, slide tray, etc. The α-particle source is a 241 Am faceted source with a geometric size of 36 mm × 4 mm and an effective diameter of 20 mm, which has an activity of 5.7 × 106 Bq and a dose rate of 0.138 Gy/min. Cells in the logarithmic growth phase are taken and cultured on glass slides. After the cells have fully covered the slides, they are removed from the culture dish and placed on a slide rack. The radiation source is set up on a stand in parallel to the slides, with a 10 mm distance between them, which allows for effective irradiation of the cells on the glass slides. Schematic diagram of the 241 Am alpha particle irradiation device was showed in Supplemental Figure S1.
Cellular DNA Damage Assay
The cells were irradiated at doses of 0, 1.0, and 2.0 Gy, 14 washed three times with PBS, and fixed with 4% paraformaldehyde for 15 min. Fixation is completed, the cells were incubated with 0.5% Triton X-100 at 4°C for 15 min, followed by adding 1 mL of 0.2% BSA sealing solution at room temperature for 1∼2 h. The cells were then sequentially added with primary antibody (γH2AX mouse monoclonal antibody) and secondary antibody (rabbit/mouse IgG), and incubated at room temperature for 45 min respectively. Finally, the nuclei were stained with DAPI solution. After sealing, the cells were observed under a confocal microscope and the images were collected.
Cell Cycle Experiment
After irradiation of the cells by α-rays at doses of 0, 1.0, and 2.0 Gy, the cells were continued to be cultured for 12 h and 24 h. They were then collected in 15 mL centrifuge tubes and centrifuged at 1500 r/min for 5 min. The resulting cell precipitates were retained. Next, 300 mL of DNA staining solution and 10 μL permeabilization solution were added, vortexed for 5∼10 s, and incubated at room temperature for 30 min. Finally, the cells were detected on a flow cytometer (FACSVerse), and the results were expressed as the percentage of cells in each phase.
Pyrosequencing
Total DNA was extracted from the cells using a DNA extraction kit. The content, purity, and integrity of the extracted DNA were examined. PyroMark Assay Design 2.0 was adopted to design primers for the CpG sites in the TSS region of the six differential genes. The primers for the differentially methylated genes are shown in Supplemental Table S1.
PCR amplification was performed according to the PCR kit. It is important to note that prior to detecting the PCR amplification products, the raw PCR amplification products need to be mixed with binding beads and washed with buffer to extract the purified products. And then, the purified amplification products were detected on a pyrophosphate sequencer (QIAGEN) as described in the PyroMark Gold Q96 Reagent Pyrophosphate Sequencing Kit. The recorded experimental results were statistically analyzed.
Real-Time Fluorescence Quantitative PCR Detection
Total RNA was extracted from the cells using an RNA extraction kit. The content, purity, and integrity of the extracted RNA were examined. Reverse transcription reactions were performed. The primers involved in this test were synthesized by Sangon Biotech (Shanghai) Co., Ltd., see Supplemental Table S2.
Statistical Analysis
The experimental data are consistently expressed as the mean ± standard error. For the comparison of different experimental groups, we employ homogeneity of variance tests and one-way analysis of variance (one-way ANOVA). In cases where variances are homogeneous, the SNK test (Student-Newman-Keuls method) is utilized; when variances are heterogeneous, the Kruskal-Wallis H test is implemented. Correlation analysis was conducted using Pearson’s correlation coefficient to assess the strength and direction of the linear relationship between variables. The SPSS 20.0 software was used to analyze the data. P < .05 was considered statistically significant.
Results
Effect of α-Particle Irradiation of Cells on the Rate of γH2AX Foci Positive Cells After Irradiation
Following irradiation with different dosages (0, 1, and 2 Gy), the cells were fixed at intervals of 1 h and 2 h post-exposure, respectively. The presence of the DNA double-strand break marker, γH2AX, was then detected employing an immunofluorescence technique. the rates of γH2AX foci positive cells were (16.83 ± 1.31)%, (38.93 ± 2.25)%, and (33.83 ± 1.76)% at 1 h, respectively. The rates of γH2AX foci positive cells were (17.67 ± 2.60)%, (35.17 ± 1.99)%, and (26.77 ± 1.80)% at 2 h, respectively. The detailed results are shown Figure 1. The rate of γH2AX foci positive cells was statistically higher than that of the control group after irradiation of cells in different dose groups for 1 h and 2 h, respectively (P < .05). The rate of γH2AX foci positive cells in the 2.0 Gy dose group was lower than that in the 1.0 Gy dose group (P < .05). These results indicated that α-ray irradiation caused DNA double-strand breaks, in which the DNA double-strand breaks in the 2.0 Gy group were lower than those in the 1.0 Gy group. In addition, positive cell rate in different dose groups after 2 h of irradiation shares the same trend with that in 1 h, but it is slightly lower than the values at 1 h for the same dose group. Effect of different doses on the formation of γ-H2AX foci positive cells. (A) γH2AX foci positive cell rate after irradiation at different doses. (B) Immunofluorescence of cells irradiated with different doses for 1 h. The asterisks * and **represent P < .05 and P < .01, respectively.
Cell Cycle Assay
The results of hepatocyte cycle detection at 12 h and 24 h after irradiation with different doses were shown in Figure 2 and Supplemental Table S3. The proportion of S-phase cells in different groups after 12 h irradiation were (19.46 ± 0.86)%, (20.15 ± 0.90)%, and (21.02 ± 0.18)%, respectively (Figure 2A). The proportion of S-phase cells in the 2.0 Gy irradiation group was significantly higher than that in control group (P < .05). The proportion of S-phase cells in different groups after 24 h of irradiation increased to (22.87 ± 1.74)%, (27.16 ± 1.25)% and (31.13 ± 1.01)%, respectively (Figure 2B). Compared with the control group, the proportion of G0/G1-phase cells decreased, the proportion of G2/M-phase cells increased, the proportion of S-phase cells significantly increased (P < .05) in the 1.0 Gy and 2.0 Gy dose groups, and the proportions of cells in different phases varied in a gradient fashion in the 1.0 Gy and 2.0 Gy dose groups. The trend in dose-dependent changes was more pronounced in the results at 24 h compared to those at 12 h. The results of hepatocyte cycle detection at 12 h and 24 h after irradiation with different doses. (A) Changes in cell cycle after 12 h irradiation at different doses. (B) Changes in cell cycle after 24 h irradiation at different doses. The asterisks * and **represent P < .05 and P < .01, respectively.
DNA Methylation Level of Different Genes
Methylation level assay data of different genes between control group (0 Gy) and dose group (1.0 and 2.0 Gy) showed in Figure 3A. The gene methylation level of CDK2AP1was (90.83 ± 1.47)% in dose group, which was significantly higher than (87.33 ± 1.15)% in the control group (P < .01). Similarly, the gene methylation levels were (92.00 ± 1.00)% and (94.67 ± 1.63)% for PDGFRL, (52.67 ± 0.58)%, and (59.50 ± 3.63)% for PCDHB16, and (93.00 ± 1.00)%, and (95.50 ± 1.38)% for FAS in control and dose groups, respectively. Taken together, the methylation levels of these genes in the dose group were significantly higher than those in the control group. Unlike the above, the methylation levels of PRKAG2 were (3.33 ± 1.53)% and (2.83 ± 1.17)%, and the methylation levels of HOXC4 were (82.00 ± 3.46)% and (81.00 ± 1.90)% in control and dose groups, respectively. The methylation levels of these two genes were not significantly different from control group and dose group (P > .05). The results of methylation level of different genes. (A) Comparison of methylation levels between control group (0 Gy) and dose group (1.0 and 2.0 Gy). (B) The trend of methylation levels in different dose groups. The asterisks * and **represent P < .05 and P < .01, respectively.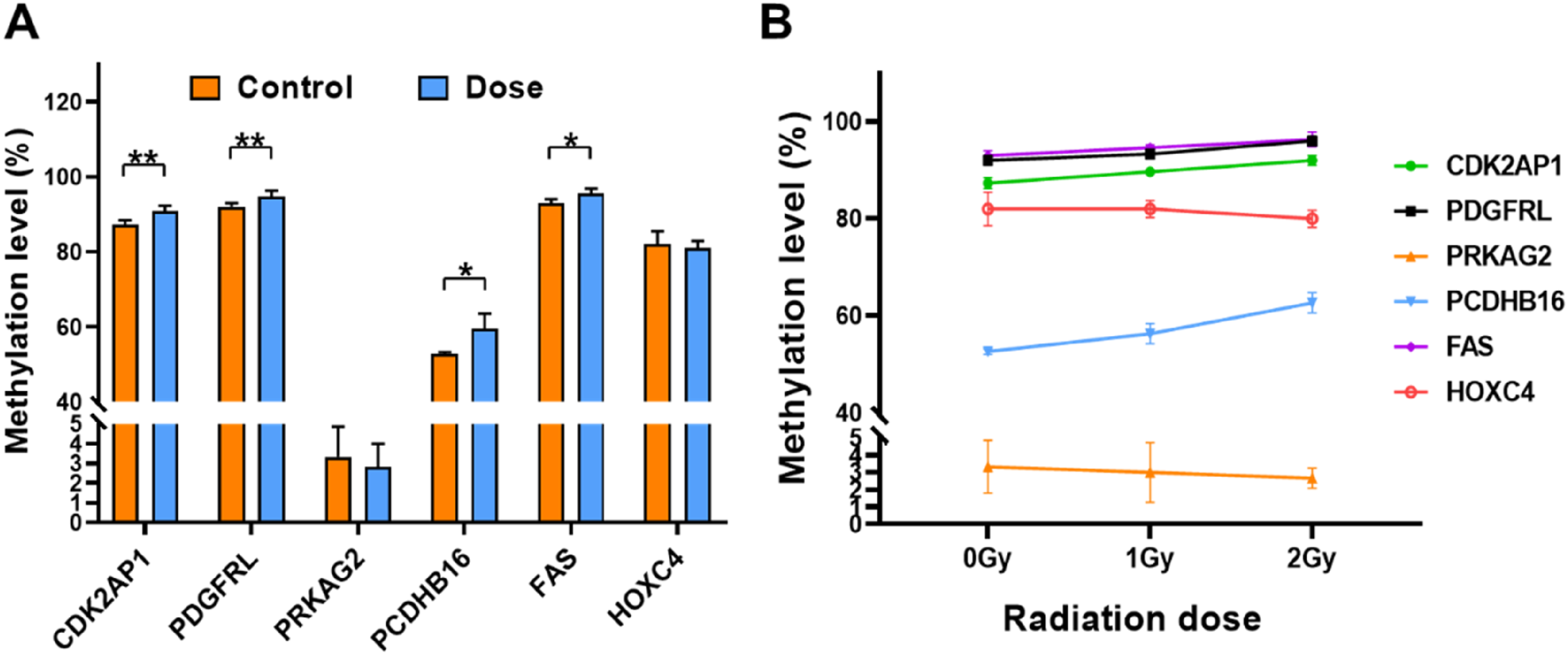
In order to demonstrate the effect of radiation dose on methylation, the dose groups were further divided into 1.0 Gy low-dose group and 2.0 Gy high-dose group. The trends in methylation levels of each gene between the three groups were shown in Figure 3B and Supplemental Table S4. The methylation levels of CDK2AP1 and PCDHB16 genes differed among the three groups of control, low-dose, and high-dose groups (P < .05), and their methylation levels tended to increase with increasing doses. Similarly, the methylation levels of PDGFRL and FAS genes tended to increase with dose elevation, but they were only significantly different between the control and high-dose group (P < .05).
Correlation Analysis of CDK2AP1 Methylation Level With DNA Damage and Cell Cycle.

The mRNA Expression Level
Relative Quantification Results of mRNA of Target Genes Among Three Groups.

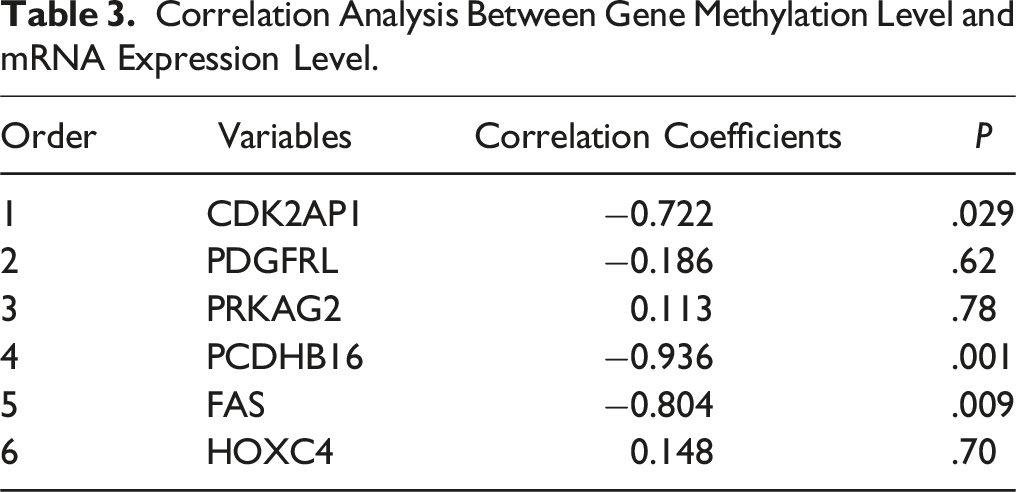
Correlation Analysis Between Gene Methylation Level and mRNA Expression Level.
Discussion
Ionizing radiation is an inducer of DNA damage and an epigenetic agent, and its genotoxicity to the organism is different from that of chemical genotoxic agents. Ionizing radiation penetrates directly into cells or tissues and deposits energy in cells in different ways, causing damage to cells and tissues of the organism. The severity of this damage and the biological consequences are related to the type of ionizing radiation, radiation dose, etc. α particles are a type of ionizing radiation and a high LET radiation source. When the body is irradiated by α particles, it is more likely to cause changes in intracellular signaling pathways, cell cycle progression, apoptosis, DNA damage repair, chromosomal aberrations, epigenetic inheritance, malignant transformation of cells, and other biological effects.15,16
α ionizing radiation can cause intracellular DNA double-strand breaks (DSBs). In this case, ATM can bind to DSBs and activate them by autophosphorylation phosphorylating histone H2AX to γH2AX. Therefore, γH2AX is one of the analytical markers of DSBs. γH2AX serves as a critical analytical marker for the assessment of DSBs, with the number of γH2AX foci correlating directly to the extent of DSBs inflicted by ionizing radiation. Ding et al found the formation of γH2AX by irradiating human peripheral blood lymphocytes with α particles. Under the same dosage conditions, high LET α-particles are more likely to induce damage to DSBs and cause cell death than low LET γ-rays. 17 In our study, the positive cell rate of γH2AX foci in the dose groups was significantly higher than that in the control group, indicating that alpha particle irradiation can induce DNA DSBs and lead to cell death. Furthermore, the observed decrease in the γH2AX foci positive cell rate at the 2-hour time point compared to the 1-hour time point suggests that cells retain a capacity for DNA repair following the occurrence of DSBs. However, the results for the 2.0 Gy exposure were lower than those for the 1.0 Gy exposure. It is speculated that the reason for this phenomenon may be that the 2 Gy irradiation conditions could lead to a greater number of cells dying directly before the occurrence of DSBs, resulting in a reduced proportion of detected γH2AX foci positive cells.
In this study, after α-particle irradiation, cells showed different levels of S-phase arrest at 12 and 24 h post-irradiation when compared to the control group. The results at the 24-hour time point were statistically significant (P < .05), indicating that α-particle irradiation induced a certain degree of controlled cell proliferation. These results are akin to the findings of Sun et al, where α-particle irradiation caused a reduction in the G1 phase duration and a prolongation of the S phase. 18 This suggests that DNA damage caused by radiation leads to compromised cell proliferation and the induction of DNA double-strand breaks. The activation of checkpoint regulatory factors in response to this damage results in S-phase arrest, preventing the damaged DNA from proceeding into mitosis.
The validation of the methylation levels of the selected six differentially methylated genes was conducted using pyrosequencing technology. The findings revealed that the methylation levels of the CDK2AP1, PDGFRL, PCDHB16, and FAS genes in the cells of the dose group were significantly higher compared to the control group (P < .05), whereas the methylation levels of the PRKAG2 and HOXC4 genes showed no significant difference when compared to the control group (P > .05). Additionally, there was a negative correlation between the methylation levels of CDK2AP1, PDGFRL, PCDHB16, and FAS and their relative mRNA expression levels, with statistically significant correlations observed for CDK2AP1, PCDHB16, and FAS (P < .05). This indicates that the expression of the CDK2AP1, PCDHB16, and FAS genes is subject to regulation by their methylation status, consistent with the common understanding that DNA methylation in the promoter region can suppress gene expression.
CDK2AP1 is a cyclin-dependent kinase 2 (CDK2). Many researches have shown that CDK2AP1 is closely related to the cyclin-dependent kinase CDK2. CDK2AP1 interacts directly with CDK2, and its overexpression inhibits the activity of CDK2-associated kinases. It has a negative regulatory role in S-phase DNA replication of the cell cycle.9,10 The results of the present study indicate that CDK2AP1 functions as an antioncogene and a promoter of cell cycle arrest. The underlying mechanism by which CDK2AP1 regulates the transition from G1 to S phase may be the CDK2AP1-mediated decrease in CDK2 activity. This can be explained by the fact that CDK2 can stimulate DNA replication by phosphorylating the DNA polymerase-α-trimer complex, and thus the CDK2AP1-mediated decrease in CDK2 activity inhibits DNA replication. 19 In addition to regulating cell cycle and DNA damage processes by modulating CDK2 activity, CDK2AP1 also serves as one of the key molecules in epigenetic regulation. In other words, CDK2AP1, a specific cell cycle regulator, has a dual role in cell cycle and epigenetic regulation. 20
The methylation level of CDK2AP1 in this experiment is significantly elevated in α-particle irradiated cells. Moreover, there exists a correlation between the methylation level of CDK2AP1 and DNA damage and cell cycle. Combined with the gene function, it was hypothesized that CDK2AP1 plays a role in DNA damage repair and cell cycle blockade caused by α irradiation through cyclin E/CDK2, TGF-β1, and other signaling pathways.
The PDGFRL gene was cloned by Fujiwara in 1995.21,22 Its protein structure has significant sequence similarity to the ligand-binding domain of platelet-derived growth factor receptor β, and it is known as the PDGFR “like” protein. Gao et al found that PDGFRL was highly expressed in liver cancer tissues and cell lines, and was closely associated with the clinical characteristics and overall survival of liver cancer patients. Through PDGFRL knockdown experiments, it was found that inhibition of the AKT/GSK-3β pathway led to enhanced phosphorylation of cell cycle protein D1 (Cyclin D1) and proteasomal degradation, which resulted in the arrest of the cell cycle in the G1 phase, and ultimately inhibited the growth of hepatocellular carcinoma tumors. 23 Accordingly, it can be concluded that PDGFRL can activate the expression of Cyclin D1 through the AKT/GSK-3β pathway and promote the proliferation of cancer cells.
Long-term low-dose ionizing radiation plays a role in promoting cell proliferation mainly by promoting the expression of Cyclin D1 gene. Overexpression of Cyclin Dl can shorten the G1-S phase transition time of the cell cycle, prompting the cells to cross the rate-limiting point and enter the S phase, which results in the uncontrolled proliferation of the cells. 24
In the present study, it was found that PDGFRL is hypermethylated in the irradiated group. This suggests that altered methylation of PDGFRL may lead to changes in its gene expression. However, there are no studies on the methylation level of this gene. Based on our findings, in conjunction with the mechanism of carcinogenesis by ionizing radiation, it is hypothesized that the PDGFRL gene suppresses its gene expression through hypermethylation, which may play a role in the early stages of alpha radiation damage.
PCDHB16 is a member of the procalcitonin β gene cluster, which is one of the three related gene clusters tandemly linked on chromosome 5. As a member of the procalcitonin genes, PCDHB16 participates in a wide range of important physiological functions and pathological processes in the body through cell adhesion. Meanwhile, PCDHB16 has also been found to play a role in autoimmune processes. Methylation alterations of the PCDHB16 gene were present in homophilic cell adhesion, intercellular cell adhesion, cellular adhesion, and bioadhesion functions in the GO functional analysis of advanced hepatocellular carcinoma. 25
FAS gene is a member of the tumor necrosis factor receptor family. It plays a central role in the physiological regulation of programmed cell death and has been implicated in the pathogenesis of various malignant tumors and immune system diseases. Abnormal methylation of the FAS gene is closely associated with the development of various tumors. Elevated methylation levels in the promoter region of the FAS gene are an important cause of low or absent expression of FAS. Meanwhile, the interaction between FAS and its ligand (FASL) induces apoptosis.
This study found that after α particle irradiation, cells undergo changes in biological effects that could potentially trigger abnormal promoter methylation of certain genes, including CDK2AP1, PDGFRL, PCDHB16, and FAS. However, the study only utilized a 241 Am source for the experiments and did not include comparisons with low-LET sources. Moreover, the experiments were validated using only hepatocytes at the cellular level, without extending to other cell types or to animal and human samples. Additionally, the DNA methylation levels of these genes and their corresponding protein translation levels have not been further confirmed. There is substantial value in pursuing further validation for genes that exhibit variations in mRNA expression, such as CDK2AP1, PCDHB16, and FAS.
Conclusion
Following alpha particle irradiation, alterations in cellular biological effects may trigger abnormal methylation of gene promoters, including those of genes related to the cell cycle (CDK2AP1), cell proliferation and development (PDGFRL), cell adhesion (PCDHB16), and tumor necrosis (FAS). mRNA expression analysis has shown a significant negative regulatory correlation between the methylation levels and the relative mRNA expression levels of CDK2AP1, PCDHB16, and FAS, which is consistent with the common understanding that DNA methylation in the promoter region can inhibit gene expression. It is speculated that changes in the methylation status of the CpG islands in the promoters of these genes may affect the regulation of mRNA expression, potentially contributing to the development of radiation-induced diseases. This suggests that the methylation alterations of the aforementioned genes could be used as future biomarkers for alpha radiation damage.
Supplemental Material
Supplemental Material - Effects of α-Particle Radiation on DNA Methylation in Human Hepatocytes
Supplemental Material for Effects of α-Particle Radiation on DNA Methylation in Human Hepatocytes by Xiangming Xue, Lixia Su, Teng Zhang, Jingming Zhan, and Xiaona Gu in Dose-Response
Footnotes
Acknowledgments
The authors are grateful to all the study participants.
Author Contributions
Conception and design of the research: Jingming Zhan and Xiaona Gu; Drafting the manuscript: Xiangming Xue and Lixia Su; Article structure design: Xiangming Xue; Statistical analysis: Teng Zhang; Samples collection and detection: Xiangming Xue and Lixia Su; Experimental management: Xiaona Gu; Acquisition of data: Xiangming Xue; All authors read and approved the final version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical Statement
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
