Abstract
Objective
To investigate the biological role of miR-143 and miR-199a in mediating the progression of osteosarcoma (OS) by targeting cyclooxygenase (COX-2).
Introduction
COX-2 plays a crucial role in the development and progression of OS. However, the specific regulatory mechanisms of COX-2 in OS are still not well understood.
Methods
The expression levels of COX-2, miR-143 and miR-199a in OS tissues were detected using immunohistochemistry, qPCR, or western blot assays. The targeting relationship between miRNAs and COX-2 was determined. The effect of miRNA and COX-2 on OS cells was evaluated in vitro and in vivo.
Results
COX-2 expression was upregulated while miR-143 and miR-199a were downregulated in OS tissues. miR-143 and miR-199a suppressed the proliferation, migration, and invasion of OS cells. The dual-luciferase reporter gene assay showed that COX-2 was a direct target of miR-143 and miR-199a. Genetic knockdown of COX-2 significantly suppressed cell proliferation, induced apoptosis, and inhibited migration and invasion of OS cells. The expression levels of COX-2 and PGE2 were decreased after the overexpression of miR-143 and miR-199a. Additionally, COX-2 silencing inhibited the tumorigenesis of OS and the synthesis of PGE2 in vivo.
Conclusions
miR-143 and miR-199a/COX-2 axis modulates the proliferation, invasion, and migration in osteosarcoma.
Introduction
Osteosarcoma (OS) is a common and highly malignant bone tumor affecting adolescents, often leading to poor survival rates due to lung metastases. 1 Despite limited progress in clinical treatments over the past 30 years, understanding the potential mechanisms of osteosarcoma pathogenesis is crucial for improving therapeutic efficiency.
Cyclooxygenase-2 (COX-2), an enzyme regulated by various events including inflammation and carcinogenesis, has been implicated in osteosarcoma development and progression.2,3 Increasing evidence indicated that COX-2 is involved in osteosarcoma development and progression. Qu et al demonstrated that COX-2 expression was considered to be a marker of human osteosarcoma. 4 Studies have linked COX-2 expression to the prognosis and metastasis of osteosarcoma, as well as its involvement in other malignant tumors.5-8 Although previous studies demonstrated that COX-2 was an essential factor in the occurrence and progression of osteosarcoma. However, the specific regulatory mechanisms of COX-2 in osteosarcoma remain largely unclear.
MicroRNA (miRNA) is a small non-coding RNA that regulates mRNA expression post-transcriptionally and is involved in nearly every cellular process. With the continuous development of miRNA, it has become a molecular biological indicator for osteosarcoma treatment strategies and prognosis. 9 In the context of osteosarcoma, miR-143 and miR-199a-3p were found to be significantly downregulated in osteosarcoma.10,11 Studies have also indicated the impact of miRNA on COX-2 expression and its role in disease progression. For instance, miR-199a negatively regulates COX-2 expression in human chondrocytes. 12 However, the expression of COX-2 in osteosarcoma, especially the correlation with miRNA, has not been fully elucidated.
The conversion of arachidonic acid to prostaglandin H2 by COX-2 results in the production of various prostaglandins, including prostaglandin E2 (PGE2). 13 Complex potential regulatory associations remain between miRNA, COX-2, and PGE2 expression. Epigallocatechin-3-O-gallate (EGCG) inhibited PGE2 production by regulating the expression of COX-2 and miR-199a-3p in human osteoarthritis chondrocytes. 14 Inhibition of COX-2 has been linked to analgesic and anti-inflammatory effects, with PGE2 being a key mediator of inflammatory pain. 15 Thus, COX-2 inhibitors have the potential to reduce PGE2 production and alleviate inflammatory pain as a therapeutic target.
In this study, we found that miR-143 and miR-199a inhibited the proliferation, migration, and invasion and suppressed PGE2 synthesis by targeting COX-2 in osteosarcoma cells. These results may offer a novel therapeutic target for the clinical management of osteosarcoma.
Materials and Methods
Sample Collection
This study enrolled 10 osteosarcoma patients undergoing surgery in Heping Surgery Hospital of Shunde, of varying different stages, from 2015 to 2019. The inclusion criteria were as follows: osteosarcoma patients with an expected survival of over three months and no history of radiotherapy or chemotherapy before surgery. The exclusion criteria were: (i) osteosarcoma patients who received radiotherapy or chemotherapy before surgery; and (ii) patients who failed to cooperate or sign the informed consent form. Adjacent tissues were used as the negative control. The surgically removed tissue is rapidly frozen in liquid nitrogen until analyzed. The study obtained written informed consent from all participants. The present study was approved by the Research Ethics Committee of The Affiliated Shunde Hospital of Jinan University (Ethics number: 026/2019).
Cell Culture
The MG63 cell line was purchased from Shanghai Institutes for Biological Sciences Cell Resource Center. Culture medium (DMEM, Thermo Fisher Scientific, Waltham, MA, USA) with 10% fetal bovine serum (Gibco, Rockville, MD, USA). Cell culture was conducted in a 37°C, 5% CO2 incubator.
Immunohistochemistry
The immunohistochemical staining was the same as the previous study. 16 Briefly, osteosarcoma samples were fixed in 4% paraformaldehyde for 24 h, then the specimens were embedded in paraffin and cut into 5 μm thick sections. The sections were incubated with hydrogen peroxide after deparaffinization and rehydration. The segments were blocked with normal goat serum for 1 h and incubated with COX-2 antibody (Cell Signaling Technology, #12282) at 4°C overnight. The sections were incubated with the corresponding secondary antibody before DAB coloration (Abcam, #ab64238). Finally, the slides were observed under a light microscope (Olympus Corporation, Tokyo, Japan) and analyzed using Image Pro (Media Cybernetics, Silver Spring, MD).
Quantitative Real Time-PCR
The RNAs from osteosarcoma tissues or MG63 cells were collected using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Total RNA (1 μg) was reverse-transcribed using the PrimeScriptTM RT reagent Kit (TaKaRa, Tokyo, Japan). The expression levels of COX-2, miR-143, and miR-199a were detected using the DyNAmo ColorFlash SYBR-Green qPCR kit (Thermo Fisher Scientific, Inc.). GAPDH was used as internal control. The reaction conditions were: 95°C for 5 min; 95°C for 50 s; 60°C for 30 s and 72°C for 30 s, for 40 cycles. The following primers were used for qRT-PCR: COX-2 forward, 5′-CAGCCATACAGCAAATCCTTG-3′ and reverse, 5′-CAAATGTGATCTGGATGTCAAC-3′; miR-143 forward, 5′-TGTAGTTCGGAGTTAGTGTCGCGC-3′ and reverse, 5′-CCTACGATCGAA AACGACGCGAACG-3′; miR-199a forward, AAGGTGAAGGTCGGAGTCAAC and reverse, GGGGTCATTGATGGCAACAATA; GAPDH forward, 5′-CCAGG TGGTCTCCTCTGACTT-3′ and reverse, 5′-GTTGCTGTAGCCA AATTCGTTGT-3′.
Western Blot
Proteins were collected from the tumor tissue and cultured MG63 cells using RIPA buffer (Thermo Fisher Scientific, Waltham, MA, USA). The protein content was measured by the BCA Protein Assay kit (Thermo Fisher Scientific, Inc.). Total cell lysates were subjected to 10% SDS-PAGE gel and transferred to PVDF membranes (Millipore, Billerica, MA, USA). Membranes were blocked in 5% skim milk for 1 h and incubated with COX-2 (Cell Signaling Technology, #12282) and GAPDH (Cell Signaling Technology, #2118) antibodies at 4°C overnight. The membranes were washed three times with Tris buffered saline with .1% Tween-20 and were incubated with secondary antibodies for 1 h. The protein bands were detected using the Electrochemiluminescence assay and analysed by Image J (NIH, Bethesda, MD, USA).
Cell Counting Kit-8
MG63 cells (5x103 cells/well) were collected and inoculated onto a 96-well plate at 37°C overnight. Cells were transfected by Lipofectamine™ 2000 (Thermo Fisher Scientific, MA, USA) with 10 nM mimics and inhibitors of miR-143 and miR-199a for 12, 24, 48, and 72 h (mimic and inhibitors or the NC controls (mimic-NC or inhibitor-NC) were synthesized by GenePharma (Shanghai, China)). CCK-8 solution (TransGen Biotech, Beijing, China) was added into each well and incubated for 2 h at 37°C. The optical density (OD) value was detected under the absorbance of 450 nm using a microplate instrument (Corona Electronics, Tokyo, Japan).
Flow Cytometry Assay
Following transfection, cells were digested slightly and collected by trypsinization. Then cells were centrifugated and resuspended in PBS and stored at 4°C overnight, 50 μl of RNAse (1 mg/mL), 50 μl of propidium iodide (1 mg/mL) were added to the cells. Then the cells were analyzed by the FITC-Cy3 and the PerCP-Cy5 channels on a flow cytometer (Cyan ADP Analyser, Beckman Coulter). The apoptosis rate was calculated as the sum of the values of Q2-2 (late apoptotic cells) and Q2-4 (early apoptotic cells) in the 4 quadrants.
Migration and Invasion Assay
Migration and invasion assays were performed on transwells coated with or without Matrigel (BD Biosciences). Briefly, 100 μL cell suspension was added to the top chamber with serum-free medium, and 500 μL medium containing 10% fetal bovine serum was added into the basolateral chamber. Subsequently, the plate was incubated for 24 h at 37°C. The chambers were fixed with 4% paraformaldehyde for 30 min and stained with crystal violet for 20 min. The stained cell numbers were counted using a microscope (Olympus Corporation, Tokyo, Japan). The transwell chambers coated with Matrigel were used for the invasion assay.
Dual-Luciferase Reporter Gene Assay
3′UTR of WT COX-2 and Mut-COX-2 was cloned into the pMiR report vector (Promega, Madison, USA). Wildtype (WT) COX-2 and Mut-COX-2 plasmids were co-transfected with fragments of miR-143 and miR-199a using Lipofectamine 2000 (Invitrogen, Thermo Fisher Scientific, Inc.). Cell lysates were collected 48 h after transfection and luciferase activity was detected using the Dual-luciferase Reporter Assay System (Promega, Madison, USA).
Lentiviral Construction and Cell Transfection
The sh-COX-2 sequence (shRNA knockdown) was synthesized by GenePharma (Shanghai, China), and then subcloned into the lentiviral vector (Thermo Fisher Scientific, Grand Island, NY, USA). The supernatant was collected from 293T cells incubating the sh-COX-2 plasmid 48 h. MG63 cells were infected with lentiviral supernatants for 24 h and then cultured with 2.5 μg/mL puromycin (MedChemExpress, Monmouth Junction, NJ, USA) to select stable cells for 72 h. MG63 cells were transfected with plasmids using lipofectamine 2000(Invitrogen, Thermo Fisher Scientific, Inc.).
Animal Experiment
A total of 14 male BABL/c nude mice (6-8 weeks old) were purchased from southern Medical University Medical Laboratory Animal Center. Animals were divided into 2 groups, including the NC group and the sh-COX-2 group. Lentivirus-infected MG63 cells were inoculated subcutaneously into the franks of nude mice. The volume and size of the tumor were measured every 5 days. The tumors were removed from mice after 25 days treatment. All animal experiment protocol was approved by the Animal Care and Use Committee of the Affiliated Shunde Hospital of Jinan University.
Enzyme-Linked Immune Sorbent Assay
The supernatants were harvested and filtrated from cultured MG63 cells. For tumor samples from nude mice, tumors that had been excised and weighed underwent mechanical homogenization in liquid nitrogen. Following homogenization and before the powdered tissue thawed, a tumor lysis buffer was introduced. Then, the mixture was gently combined and transferred to Eppendorf tubes, then left to thaw completely for 15 minutes at 4°C. Subsequently, the lysate underwent centrifugation for 15 minutes at 20,000g and 4°C. Following removal of the lipid layer, the supernatant was separated from the solid pellet, divided into aliquots. Protein concentrations were determined using a BCA assay (Beyotime, Shanghai, China). The expression of PGE2 was quantified using a commercially PGE2 ELISA kit (Cayman Chemicals, Ann Arbor, MI, USA) according to the manufacturer’s instructions. ELISA plate was detected using a microplate instrument (Corona Electronics, Tokyo, Japan).
Statistical Analysis
Data were represented as mean ± standard deviation. One-way analysis of variance (ANOVA) was used for multiple group comparisons. All data were analysed with Graph Pad Prism 5(San Diego, CA, USA). P < .05 indicated a significant difference. Each experiment was repeated at least three times.
Results
Expression Levels of COX-2, miR-143, and miR-199a in Osteosarcoma Tissues
Previous reports have shown that the progression of osteosarcoma may be regulated by the expression of COX-2, miR-143, or miR-199a.17,18 However, the regulatory mechanism in osteosarcoma remains largely unknown, and we wondered whether the expression level of COX-2 in osteosarcoma was correlated with miR-143 and miR199a, simultaneously. To further explore the relationship between COX-2 and miR-143 and miR-199a, tissue samples from osteosarcoma patients were detected by immunohistochemistry experiments. The results demonstrated that the expression of COX-2 increased as the disease progressed (Figure 1A), suggesting the involvement of COX-2 during osteosarcoma disease development. Furthermore, the mRNA and protein expression levels of COX-2 were determined together. The qRT-PCR and western blot analysis demonstrated that the expression levels of COX-2 were significantly increased in osteosarcoma tissues (Figure 1(B)–(C)). In addition, to evaluate the role of miR-143 and miR-199a in osteosarcoma, the mRNA expression levels were detected by qRT-PCR. The results revealed that the expression levels of miR-143 and miR-199a were significantly decreased in samples of osteosarcoma, compared with the adjacent non-tumor tissues (Figure 1(D)–(E)). These results suggested that COX-2 was upregulated, while miR-143 and miR-199a were downregulated in osteosarcoma disease. Upregulation of COX-2 and downregulation of miR-143 and miR-199a in osteosarcoma. (A-C) The expression level of COX-2 was positively correlated with the severity of osteosarcoma by immunohistochemistry, qPCR, and western blot. Bar = 100 μm. The plus indicates the severity of osteosarcoma, + indicates stage I, ++ indicates stage II, +++ indicates stage III. (D-E) The expression levels of miR-143 and miR-199a were downregulated in osteosarcoma. All experiments were repeated three times. * indicates P < .05, ** indicates P < .01, *** indicates P < .001, compared to NC group.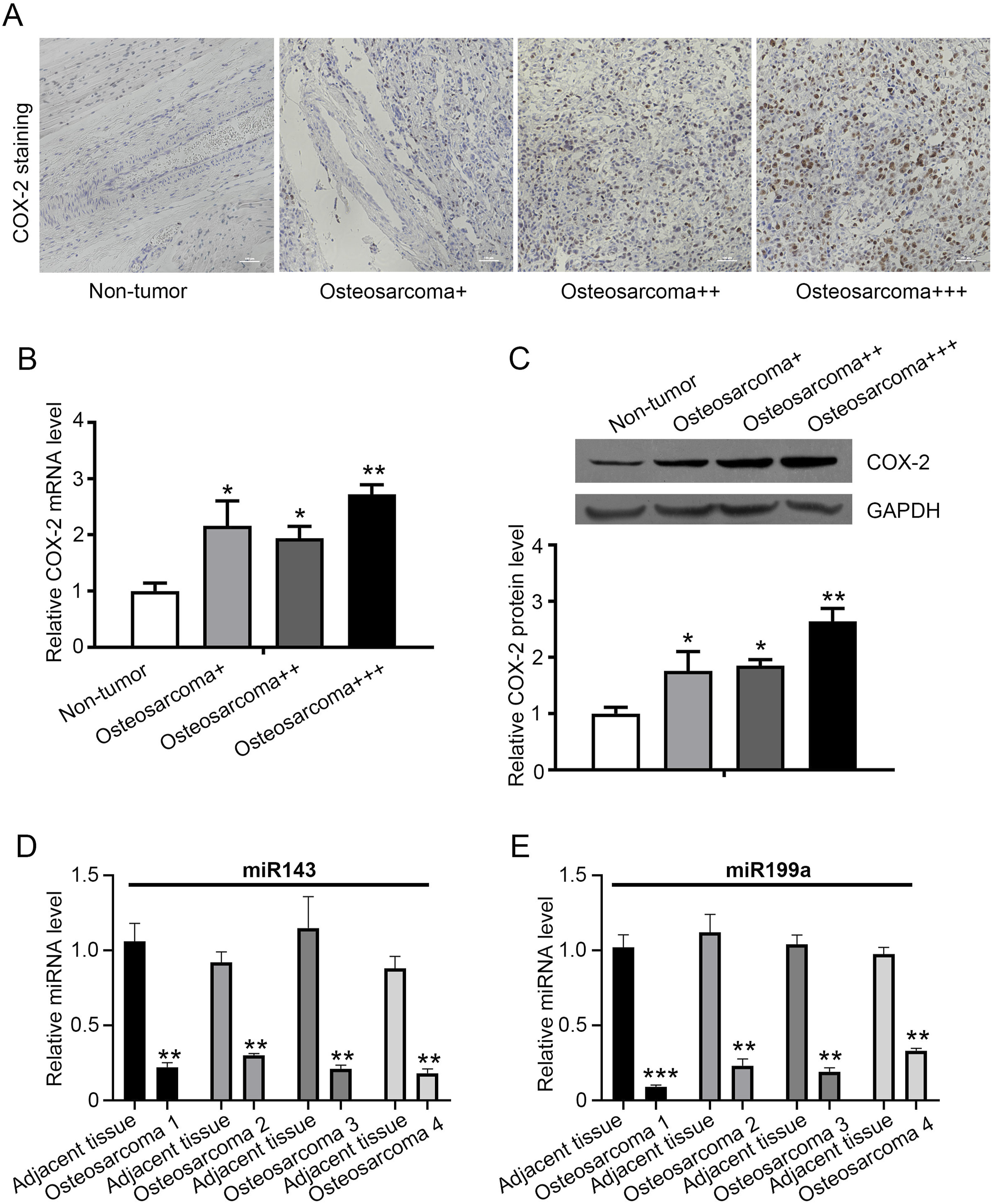
MiR-143 and miR-199a Suppress the Proliferation, Migration, and Invasion of Osteosarcoma Cells
To evaluate the biological functions of miR-143 and miR-199a, we synthesized their corresponding mimics and inhibitors. The fragments were constructed and transfected into MG63 cells and the effects on cell viability were detected (Figure 2(A)–(B)). The CCK8 assay results indicated that the proliferation ability of MG63 cells transfected with miR-143 and miR-199a mimics were significantly inhibited, suggesting that the cell viability of osteosarcoma cells was restricted. In addition, transwell assay was used to inspect their effects on the migration and invasion of osteosarcoma cells (Figure 2(C)–(F)). The results showed that the expression of miR-143 and miR-199a mimics in MG63 cells inhibited the migratory and invasive. These results suggested that miR-143 and miR-199a significantly suppressed the proliferation, migration, and invasion of osteosarcoma cells. miR-143 and miR-199a negatively regulate the proliferation, invasion, and migration in MG63 cells. (A-B) MG63 cells were transfected with miR-143 and miR-199a mimics, and then were performed to determine the cell viability by CCK-8 assay at different stages. (C-F). Migration and invasion abilities of MG63 cells were detected by expression of miR-143 and miR-199a. Scale bar, 20 μm. All experiments were repeated three times. * indicates P < .05, ** indicates P < .01, compared to mimics-NC.
COX-2 is a Direct Target of miR-143 and miR-199a
In our study, the correlation between COX-2 and miR-143 and miR-199a was investigated by dual-luciferase reporter gene assay. According to the prediction of the TargetScan database and the reported literature, COX-2 was identified as a potential target of miR-143 and miR-199a (Figure 3A). Dual-luciferase reporter plasmids (COX-2, Mut COX-2) were constructed based on the predicted target sequence and transfected into MG63 cells. The results of dual-luciferase reporter gene assay indicated that miR-143 and miR-199a mimics inhibited the luciferase activity of WT COX-2, but no significant suppression on Mut COX-2 (Figure 3(B)–(C)). These data validated that miR-143 and miR-199a were directly targeting of COX-2. COX-2 is a direct target of miR-143 and miR-199a in MG63 cells. (A) TargetScan analysis showed potential binding sites of COX-2 to miR-143 and miR-199a. (B) Dual-luciferase reporter gene assay showed that miR-143 significantly reduced the luciferase activity in MG63 cells, compared to mimics-NC. (C) Dual-luciferase reporter gene assay showed that miR-199a significantly reduced the luciferase activity in MG63 cells, compared to mimic-NC. All experiments were repeated three times. ** indicates P < .01.
Genetic Knockdown of COX-2 Affects Cell Function
Although the role of COX-2 as an oncogene during the development of osteosarcoma have been reported,4,19,20 here by shRNA approach, we tried to verify the concept. Cultured MG63 cells were infected with sh-COX-2 virus, then the cell viability was determined by CCK-8 assay, apoptosis by flow cytometry and migration and invasion by Transwell assay. As shown in Figure 4A, the cell proliferation rate was markedly suppressed when COX-2 was suppressed. Further, inhibition of COX-2 significantly induced cell apoptosis (Figure 4(B)–(C)) and abolished cell migration and invasion of osteosarcoma cells (Figure 4(D)–(E)). These data suggest that knockdown of COX-2 suppresses cell proliferation, induces apoptosis, and inhibits migration and invasion of osteosarcoma cells. Inhibition of COX-2 affects MG63 cell function. MG63 cells were infected with sh-COX-2 virus, then the cells were subjected to CCK-8 assay (A) flow cytometry (B-C) and Transwell assay (D-E) sh-NC viruses used as control. All experiments were repeated three times. * indicates P < .05, ** indicates P < .01, *** indicates P < .001. Scale bar, 20 μm.
miR-143 and miR-199a Inhibit COX-2 and PGE2 Expression
To further explore the relationship of miR-143, miR-199a, and COX-2, mimics, and inhibitors of miR-143 and miR-199a were transfected into MG63 cells, then the lysates were analyzed using western blot assay. Compared with the miRNA NC group, COX-2 was significantly decreased treated with mimics of miR-143 and miR-199a and was increased treated with inhibitors of miR-143 and miR-199a, which was consistent with the results of the dual-luciferase reporter gene assay (Figure 5(A)–(B)). The results indicated that miR-143 and miR-199a regulate COX-2 expression. According to the reported literature, COX-2 eventually converted arachidonic acid to PGE2 and PGE2 had a crucial role in pain signaling.
15
Therefore, we wondered whether miR-143 and miR-199a affect the expression of PGE2. The expression of PGE2 was detected using ELISA assay after mimics and inhibitors were transfected into MG63 cells. The results showed that the mimics of miR-143 and miR-199a resulted in a significant decrease in PGE2, while inhibitors of miR-143 and miR-199a caused an increase in PGE2(Figure 5(C)–(D)), suggesting PGE2 production was significantly reduced by inhibiting COX-2 via miR-143 and miR-199a. miR-143 and miR-199a inhibit COX-2 and PGE2 expression. (A-B) The mimics and inhibitors of miR-143 and miR-199a were transfected into MG63 cells, as detected via western blot assay. (C-D) The expression of PGE2 was detected using ELISA assay after mimics and inhibitors of miR-143 and miR-199a were transfected into MG63 cells. All experiments were repeated three times. ** indicates P < .01.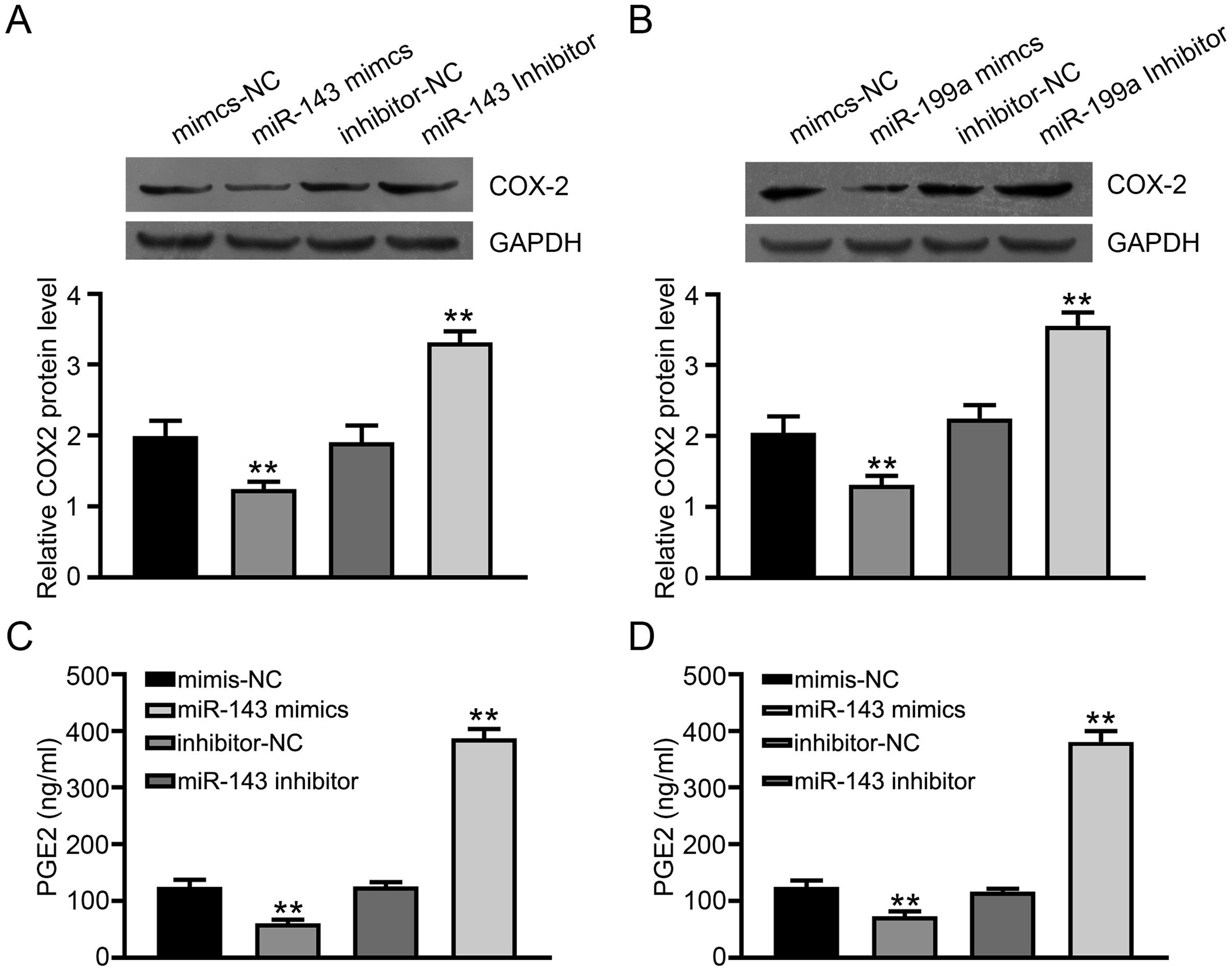
COX-2 Knockdown Inhibits Osteosarcoma Tumorigenesis and PGE2 Synthesis in Vivo
To investigate the tumorigenic effect of COX-2 in vivo, MG63 cells were subcutaneously injected with sh-COX2 into nude mice. Tumor blocks were removed from the nude mice every 5 days for a total of 25 days. The endogenous level of COX-2 was determined by immunohistochemistry with COX-2 antibody (Figure 6A) and Sh-COX-2 intuitively markedly reduced the volume of tumors, compared with the NC group (Figure 6A). Furthermore, the volume and weight of each tumor were accurately measured. The tumor volume and weight of sh-COX2 were significantly smaller than those of the NC group (Figure 6(B)–(C)). Moreover, sh-COX-2 markedly decreased the expression of PGE2 in tumor tissues (Figure 6D). These results indicated that low expression of COX-2 could inhibit the tumorigenesis of osteosarcoma and the synthesis of PGE2 in vivo. sh-COX-2 inhibit osteosarcoma tumorigenesis and PGE2 synthesis in vivo. (A) Osteosarcoma xenograft tumors were removed and photographed after 25 days. Then the samples were subjected to immunohistochemistry with COX-2 antibody. Scale bar, 50 μm. (B-C). Tumor volume and tumor weight in each animal were calculated. (D) The expression of PGE2 in tumor tissues was detected using ELISA assay. ** indicates P < .01, ***indicates P < .001.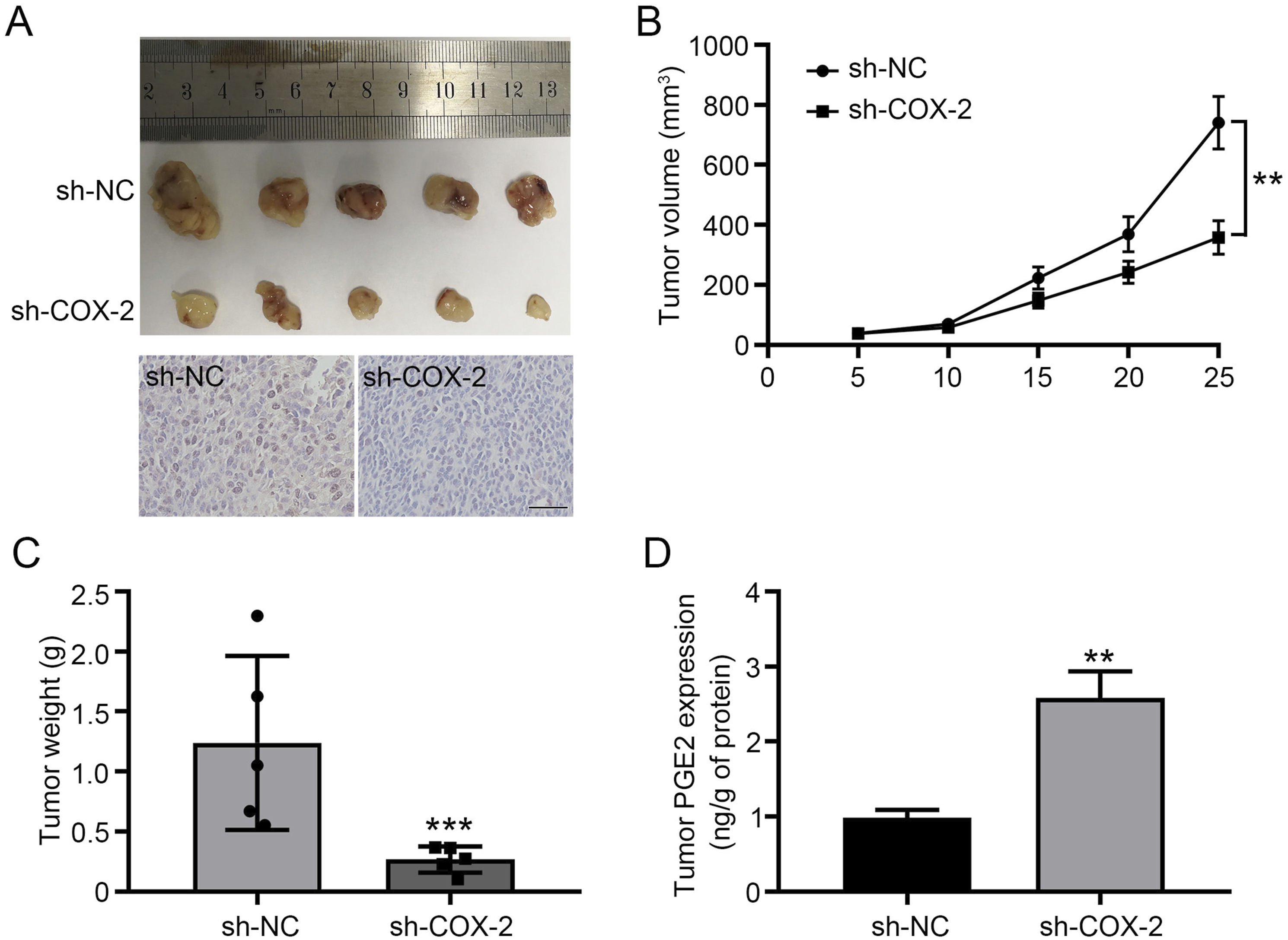
Discussion
Osteosarcoma, originating from mesenchymal cells, is the most common primary malignant bone tumor. With the disease’s invasion of bone, pain is the main clinical symptom of osteosarcoma. 21 The initial pain of osteosarcoma is intermittent, it is worth mentioning that it gradually transforms into persistent and insufferable pain. Moreover, the survival rate of osteosarcoma patients remains poor, while the side effects of treatment are serious. Therefore, the molecular mechanism of osteosarcoma needs to be urgently investigated. It contributes to identifying new biomarkers and potential targets for the diagnosis and treatment of osteosarcoma. In the present study, we revealed that miR-143 and miR-199a inhibited the proliferation, migration, and, invasion and suppressed PGE2 synthesis by targeting COX-2 in osteosarcoma cells.
COX-2, an essential molecule for the synthesis of prostaglandins, has been demonstrated to closely associate with the occurrence and progression of various types of tumors.22-24 We found that the expression of COX-2 in osteosarcoma increased with the severity of the disease. This is consistent with what was reported in the literature,5,17 indicating that COX-2 involved in the progress of osteosarcoma. Nonetheless, the exact mechanism of COX-2 in osteosarcoma has not been fully elaborated. Although, previous studies suggested that COX-2 inhibitors or knockout of the COX-2 gene could suppress the progression of osteosarcoma. 25 However, COX-2 could be induced by various substances, 26 including inflammatory cytokines, growth factors, and tumor promoters.
MiRNAs, about 19-22 nucleotides in length, are a highly conserved noncoding RNA and play a role in regulating cell proliferation, differentiation, apoptosis, and development. Previous studies have shown that miRNAs play an important role in the diagnosis, treatment, and prognosis of osteosarcoma.9,27 Therefore, it has been demonstrated that miRNA dysregulation was 1 of the main factors leading to the carcinogenesis of osteosarcoma. Li et al revealed that miR-145 inhibited osteosarcoma cell proliferation by targeting E2F transcription factor 3(E2F3). 28 Ren at al demonstrated that miR-421 promoted the development of osteosarcoma by regulating MCPIP1 expression. 29 He et al showed that miR-627 inhibited the proliferation and metastasis of osteosarcoma by targeting pleiotrophin. 30 In the present study, miR-143 and miR-199a significantly downregulated in osteosarcoma tissues. Moreover, we found that miR-143 and miR-199a reduced cell viability, proliferation, invasion, and migration in osteosarcoma cells. The expression of miR-143 and miR-199a in osteosarcoma is consistent with previous reports.31-33 Furthermore, the biological function of miR-143 and miR-199a have been demonstrated in various, including prostate cancer, ovarian cancer, colorectal cancer, and bladder cancer.34-38 These results indicated that miR-143 and miR-199a expression play a crucial role in the diagnosis of osteosarcoma and can be used as potential therapeutic targets.
Therefore, to further elaborate on the relationship between COX-2 and miR-143 and miR-199a in osteosarcoma. We revealed that miR-143 and miR-199a were directly targeted of COX-2 using dual-luciferase reporter gene assay. Moreover, the expression of COX-2 and PGE2 was regulated by miR-143 and miR-199a using Western blot assay. This was confirmed in animal studies of osteosarcoma tumorigenesis. The targeting relationship between COX-2 and miR-143 and miR-199a has been determined in various diseases, not just osteosarcoma. It was shown that miR-143 suppressed the growth and migration of gastric cancer and bladder carcinoma by targeting COX-2.39,40 He et al revealed that miR-199a plays a critical role in arsenic-induced tumor growth and angiogenesis by targeting HIF-1/COX-2. 41 Additionally, we have shown that the production of PGE2 was regulated by inhibiting COX-2 via miR-143 and miR-199a, suggesting the regulation of COX-2, miR-143, and miR-199a can be considered as a therapeutic target to relieve pain.
COX-2 is the first step in catalyzing the biosynthesis of PGE2 and plays a crucial role in the pathogenesis of inflammatory diseases, including gastric cancer, rheumatoid arthritis, and hepatocellular carcinoma.42-44 PGE2 induces the expression of COX-2 and mPGES-1 through the cAMP signaling pathway, with the involvement of EP2 receptors and PKA/CREB signaling, leading to increased transcriptional activation of COX-2 and mPGES-1 in macrophages. 45 COX-2/PGE2 signaling facilitates fracture healing by activating the Wnt/β-catenin pathway, which is evidenced by increased transcriptional and translational levels of COX-2, β-catenin, and VEGF, and improved bone healing outcomes in rats. 45 The expression of COX-2 is regulated by various extracellular signals converging on the activation of MAPKs, affecting COX-2 mRNA levels at both transcriptional and post-transcriptional levels, with involvement of transcription factors such as CREB, NFkB, and C/EBP. 46 The COX-2/mPGES-1/PGE2 cascade is critical in kidney injury, influencing fluid metabolism, blood pressure, and renal hemodynamics, and plays a role in the pathogenesis of both diabetic and nondiabetic kidney diseases as well as acute kidney injuries. 47 COX-2 signaling, particularly through the MAPK pathway, is essential in the radiation-induced bystander effect, which is an inflammatory response mechanism. 48 COX-2 is also expressed by excitatory neurons at postsynaptic sites in the cerebral cortex and is involved in synaptic signaling, suggesting a role in activity-dependent synaptic remodeling. 49 CaR signaling in medullary thick ascending limb cells mediates COX-2-derived PGE2 production through Gq- and Gi-coupled cascades, impacting ion transport and mTAL function. 50
There are several limitations to this study. Firstly, the sample size of clinical osteosarcoma cases is small due to the limited number of osteosarcoma patients in our hospital, and some patients did not consent to data publication. Secondly, the limited number of clinical samples and the lack of survival follow-up data prevented us from conducting a survival analysis between high and low COX-2 levels, which would have supplemented the clinical significance of the study. Thirdly, due to the effects of miR-143 and miR-199a on tumorigenesis in nude mice have been reported, we only observed the effect of COX-2 knockdown on tumorigenesis. The data would be more convincing if the effects of miRNA mimics were also co-operatively observed in this assay. Lastly, the molecular signaling pathways underlying the miR-143 and miR-199a/COX-2/PEG2 axis are worth investigating and will be addressed in our future work.
In the present study, we have innovative found that the expression of PGE2 in osteosarcoma was regulated by COX-2, which was a direct target of miR-143 and miR-199a. Taken together, our results indicated that miR-143 and miR-199a inhibited the proliferation, migration, and invasion and suppressed PGE2 synthesis by targeting COX-2 in osteosarcoma cells.
Conclusion
This study investigates the role of miR-143 and miR-199a in osteosarcoma progression by targeting cyclooxygenase-2 (COX-2). The findings show that COX-2 protein expression is upregulated, while miR-143 and miR-199a are downregulated in osteosarcoma tissues. COX-2 is identified as a direct target of these miRNAs, and the miR-143 and miR-199a/COX-2 axis modulates the proliferation, migration, and invasion of osteosarcoma cells. Additionally, COX-2 silencing reduces tumorigenesis and PGE2 synthesis in vivo. This study identifies a novel regulatory mechanism in which miR-143 and miR-199a inhibit osteosarcoma progression by targeting COX-2.
Footnotes
Author Contributions
Z.Z. conceived the study, contributed to data interpretation and manuscript revision. B.X. and Y.M. created the study protocol, performed the statistical analyses, and wrote the first manuscript draft. Y.L. conceived the study and critically revised the manuscript, assisted with the study design and performed the data collection. J.M. assisted with study coordination and helped draft the manuscript. All authors read and approved the final manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Foshan Self-funded Science and Technology Innovation Project, Number: 2220001003819; Key medical talents training project of Shunde District. Beijing Medical Award Foundation, Number: YXJL-2021-0307-0584.
