Abstract
In the contemporary research world, the intestinal microbiome is now envisioned as a new body organ. Recently, the gut microbiome represents a new drug target in the gut, since various orthologues of intestinal drug transporters are also found present in the microbiome that lines the small intestine of the host. Owing to this, absorbance of sulpiride by the gut microbiome in an in vivo albino rats model was assessed after the oral administration with a single dose of 20mg/kg b.w. The rats were subsequently sacrificed at 2, 3, 4, 5 and 6 hours post oral administration to collect the gut microbial mass pellet. The drug absorbance by the gut microbiome was determined by pursuing the microbial lysate through RP-HPLC-UV. Total absorbance of sulpiride by the whole gut microbiome and drug absorbance per milligram of microbial pellet were found significantly higher at 4 hours post-administration as compared to all other groups. These results affirm the hypothesis that the structural homology between membrane transporters of the gut microbiome and intestinal epithelium of the host might play an important role in drug absorbance by gut microbes in an in vivo condition.
Introduction
Sulpiride is a dopamine antagonist prescribed to treat schizophrenia and associated depressive disorders 1 (Figure 1). After oral administration, sulpiride shows a dose-dependent erratic absorption and low with a bioavailability of as low as 30% of the oral dose. 2 Absorption of orally administered drugs occurs in the gut mucosa as they pass through the gut lumen before reaching blood circulation. Absorption characteristics of an orally administered drug show complex dynamics that depend upon physicochemical and physiological factors of the drug and gut mucosa 3 and probably an interaction with the gut microbiome. 4

Sulpiride (-).
Intestinal microbiome harbors a diversified microbial genome which is 150 times greater than human genes and regarded collectively as a microbial organ that performs many functions which the human host is unable to process. 5 -7 Currently, in literature archive, membrane transporters have been recognized to be important determinants in regulating drug pharmacokinetics and pharmacodynamics 8 and play key roles in the influx and efflux of various nutrients and drugs in gut epithelial cells. 9 Human peptide transporter (hPepT 1) present on the apical side of the enterocyte membrane mediates sulpiride transport in the systemic circulation. 10 Such transporters could also be present in the gut microbiome analogous to different eukaryotic cell transporters. 11 hPepT 1 share 23% sequence similarity to YdgR (proton-coupled oligopeptide transporter) present in the inner membrane of E. coli. 12
We hypothesize that transporters in the membrane of the gut epithelial cells and microbial cells embedded in the intestinal mucus layer 13 may compete for the uptake of a orally administered drug. 4 Results presented by Prabhala et al. (2017) 13 proposed that commonly used drugs, like sulpiride, enters the bacterial cell by the bacterial POTs (proton-coupled oligopeptide transporters) in case of E. coli. Therefore, in the current study, we postulated that the sulpiride may also serve as a substrate for gut microbiome since several gut microbes contain POT gene in their genome.
Materials and Methods
Animals, Housing and Diet
Healthy Wistar albino male rats (n = 36) aged 8 to 10 weeks and weigh around 160 ± 20 grams were selected for the current study. The rats were raised in the Animal Research Lab, Department of Physiology, Government College University, Faisalabad. The selected animals were transferred to experimental cages in an isolated room with 25 ± 2 °C temperature, 12-hour light/12-hour dark period and with 40-60% humidity maintained for 24 hours at the animal workstation.
Chow maintenance diet composed of 76% starch, 10% protein and 10% oils was served, which was consumed on an average of 46g/kg of body weight daily. Autoclaved water was provided, and water bottles were changed every day during acclimatization and experimentation. The rats were fasted 8 hours before the oral administration of drugs to avoid the influence of food. Rats were simply decapitated with a sharp knife at the end of each experiment. All the planned experiments and drug administration protocols were followed by the written consent of the Ethical Review Board, Government College University, Faisalabad with Reference No. GCUF/ERB/131.
All 36 rats were divided into 6 groups namely control group (E1) and sulpiride treated groups E2, E3, E4, E5, and E6 with 6 rats in each group based upon transit time of drug in small intestine. A single oral dose of sulpiride at 20 mg/kg of BW was administered to E2-E6. All the doses were given orally by a 16-18 gauge feeding tube about 0.79-1.18 cm in length. Following oral drug administration, rats were allowed to feed ad libitum till decapitated. Treated groups were decapitated with a sharp knife at intervals of 2, 3, 4, 5 and 6 hours after drug administration. The microbial mass from the digesta was isolated to obtain microbial lysate. Untreated animals in the control group (E1) were killed at the very beginning of the experiment to collect the whole small intestinal length to harvest microbial mass for analysis.
Isolation of the Microbial Mass and Microbial Lysate
Methods described by Mukhtar et al. (2019) 4 for the isolation of gut microbiota from the small intestine of rats were adopted. The supernatant was isolated and dried with the application of nitrogen gas. Resulting dried mass was dissolved in 800 µL of required mobile phase and filtered (pore size, 0.45 µm, Milli Pore®, USA). The microbial lysates obtained from treated and control groups were preserved at −20°C till subsequent HPLC analysis.
HPLC System and Conditions
Method for HPLC analysis was adopted from described previously 14 with some amendments. For separation of sulpiride, the mobile phase consisted of a mixture of methanol: ACN: water: B-R (Britton Robinson buffer) universal buffer of pH 9 (20:20:40:20 v/v/v/v).
Liquid chromatography consisted of the HPLC system (Perkin Elmer®, USA) attached with Flexer Binary LC pump, UV/VIS LC Detector (Shelton CT®, 06484 USA) and reverse phase C18 column (5 µm, 250 × 4.6 mm) accompanying oven set at 30°C. Chromera software version. 4. 1. 2. 6410 was used to analyze data. Fresh mobile phase was constituted daily for the analysis of drug and filtered (cellulose nitrate filter paper pore size 0.45 µm, 47 mm, Sartorious®, Germany) through assembly under vacuum to remove the residues in the solutions. Protocol was adopted throughout the study period. Filtered mobile phases and solutions were degassed by ultrasonic sonicator for 30 min. For sulpiride, volume of 10 µL was inserted by syringe injection keeping controlled flow rate of 0.6 ml/min to find the drug concentration. The drug concentration was estimated from calibration curve (Figure 2) obtained against sulpiride standards (0.5, 1, 5, 10, 15, 20, 30 and 50 µg/ml) at given HPLC conditions with referenced wavelength at 225 nm showing retention time of 3.87 ± 0.2 min. Validation methods included Linearity r 2 (0.991), Precision (0.61, 1.01, 0.88 for running 10 samples each of 5, 10 and 15 µg/mL sulpiride). Accuracy and recovery of sulpiride (10 µg/ml) from spiked samples was 91.6% within the range of 92-100% (Table 1). Precision of the method was evaluated by both intra-day and inter-day precision. Intra-day precision (1.23,0.58, 1.55) determined by injecting volume of 10 µL of 3 level of standards sulpiride 5, 10 and 15 µg/mL respectively to calculate the peak area values. Same procedure was repeated after 5 days with the same standard of sulpiride to determine interday-precision 0.61, 1.01, 0.88 respectively.
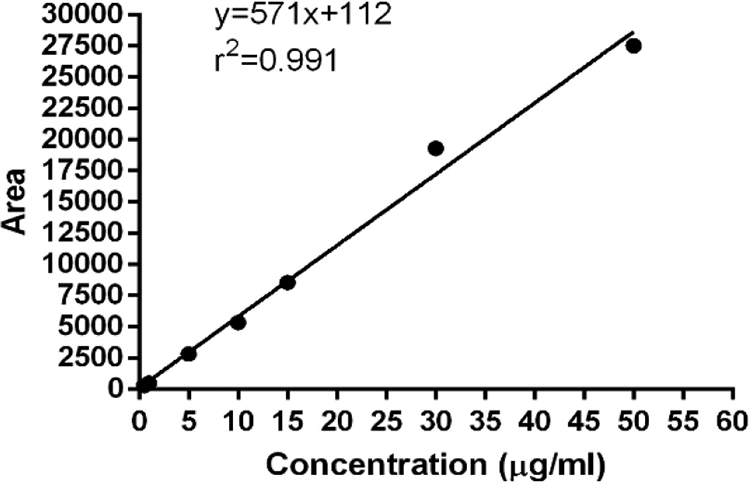
Sulpiride calibration curve (0.5-50 µg/mL) studied.
Linearity Data of Sulpiride.

See Figure 3 for Representative chromatogram of sulpiride standard solutions.

Representative chromatogram of sulpiride standard solutions (15 µg/mL).
See Figure 4 for Chromatogram of sulpiride-treated sample.

Chromatogram of sulpiride-treated sample (n = 6) at 4 hours post-sulpiride oral treatment (20 mg/kg of BW) sampling time detected sulpiride (127.2 ± 7.53 µg) in whole microbial mass.
See Figure 5 for Chromatogram of blank microbial lysate samples.

Chromatogram of blank microbial lysate samples spiked with 10 µg of sulpiride.
Statistical Analysis
Data obtained was statistically analyzed by applying ANOVA (one way analysis of variance) by GraphPad Prism 6 San Diego, CA 92108, USA. Duncan Multiple Range test was applied as Post hoc test to check the significance among different groups by setting (P ≤ 0.05) through Costat 6.4 software.
Results
Parameters such total body weight, small intestine length, small intestine weight, wet content weight and total microbial mass (Table 2) were found similar in control and treated groups. Sulpiride was not detected in the gut microbiome of the groups E1 (control), E2, E3, E5 and E6 at 0, 2, 3, 5 and 6 hours post-drug administration sampling, respectively. Sulpiride absorbance was observed in group E4 at 4 hours post-drug administration sampling time. Total sulpiride absorbance was significantly (P ≤ 0.05) higher in group E4 (4 hours) post-drug administration sampling as compared to control (E1) and all other sulpiride-treated groups; E2, E3, E5 and E6 at 0,2, 3, 5 and 6 hours post-drug administration sampling time, respectively (Figure 6). Maximum sulpiride absorbance per mg of microbial mass (Figure 7) was observed in group E4 as compared to all other groups and was significantly (P ≤ 0.05) higher. Sulpiride percent dose recovery was significantly (P ≤ 0.05) higher in group E4 at 4 hours post-drug administration sampling time as compared to the remaining groups (Figure 8). The maximum transit time of sulpiride at which maximum concentration (127.2 ± 7.53 µg) of the drug was absorbed by the microbiome was 4 hours post-sulpiride administration after which no drug was detected in any of the sample. Principle component analysis (PCA) of observed parameters (Figure 9) signifies the contribution of component F1 and F2 maximally. Component F1 contributed (40.61) followed by F2 (22.65) with total contribution of 63.26%. Total sulpiride absorption show a significant positive correlation with drug absorption per mg of microbial mass (0.996***) and given dose of sulpiride (0.964***).
Means of Body Weight (gm ± SE), Small Intestine Weight (gm ± SE), Small Intestine Length (cm ± SE), Wet Content (gm ± SE), and Microbial Mass (mg ± SE) of Control (E1) and Sulpiride Treated Groups Based Upon Post-Sulpiride Oral Treatment (20 mg/kg of Body Weight) at Different Sampling Times; E2 = 2 Hours, E3 = 3 Hour, E4 = 4 Hours, E5 = 5 Hours and E6 = 6 Hours Were Found Non-Significant (P ≥ 0.05).

N* = 6.

Total sulpiride absorbance (µg ± SE, N = 6) by the whole small intestine microbiome measured in various groups: E1 = Control group without treatment and drug treated groups summarized by post-sulpiride oral treatment (20 mg/kg of body weight) at different sampling times; E2 = 2 hours, E3 = 3 hour, E4 = 4 hours, E5 = 5 hours and E6 = 6 hours. Alphabets on mean bars show significant difference between groups (P ≤ 0.05).

Sulpiride absorbance per mg of microbial mass (µg ± SE, N = 6) measured in various groups: E1 = Control group without treatment and drug treated groups based upon post-sulpiride oral treatment (20 mg/kg of body weight) at sampling times; E2 = 2 hours, E3 = 3 hour, E4 = 4 hours, E5 = 5 hours and E6 = 6 hours. Alphabets on mean bars show significant difference between groups (P ≤ 0.05).

Percentage dose recovery (% ± SE, N = 6) from administered dose of sulpiride measured in various groups: E1 = Control group without treatment and drug treated groups based on post-sulpiride oral treatment (20 mg/kg of body weight) at different sampling times; E2 = 2 hours, E3 = 3 hour, E4 = 4 hours, E5 = 5 hours and E6 = 6 hours. Alphabets on mean bars show significant difference between groups (P ≤ 0.05).

PCA of various parameters observed for a given dose (20 mg/kg of body weight) of sulpiride measured in different groups based upon different intestinal transit times.
Discussion
In the current study, we aimed to assess the potential of gut microbes to competitively absorb the orally administered drug, sulpiride, in laboratory rat models. Sulpiride is transported by PepT1, OCTN1, and OCTN2 present in the apical membrane of Caco-2 cells. 10 The sulpiride contains an N-terminus in its structure that equals the size of a typical dipeptide (Figure 1). In the current study, sulpiride seems to be a poor substrate of POT in the gut microbiome since the percent dose recovery was estimated as 3.91% of the administered dose. Generally, as a potential substrate of POT, peptides with L-amino acids are preferred over D-amino acids and are transported in their trans conformation as L-isomers over the D-isomers. 15,16 The L-isomer of sulpiride was used in the current study. The current study demonstrates that orally administered sulpiride is absorbed by the gut microbiome. Previously, paracetamol has also been detected as a good substrate of the gut microbiome with percent dose recovery (13.16 ± 0.55%) 4 hours post drug administration in a rat model. 4 Absorption of sulpiride in intestine might have occurred passively, actively or both and we believe similar process of absorption in the microbial cells as well. In vitro trials with E. coli, confirms that POTs can transport orally administered drugs such as sulpiride, ampicillin and bestatin across the bacterial cell wall. It was subsequently suggested that similar POTs and porins might also be present in other gut microbes, which can indeed present a similar type of interaction of the gut microbiome with the orally-administered drugs. 13,17
Human hPepT1 and POT (YdgR) transporters in bacteria are both specific for di- and tripeptides and have preferences for positive side chains on the N-terminal position of the peptide. In addition, D-amino acids are not as preferred as the L-amino acids. 12 Moreover, a homology exists between transporting mechanisms in microbial cell membranes and the enterocytes of the gut of the host, which further raises the possibility of drug uptake as a competitive interplay between these 2 transport systems which may lead to decreased absorption of some drugs in the systemic circulation. Sulpiride is also a substrate of hPepT 1 which is present in the apical membrane of enterocytes of the small intestine. 13 hPepT1 is structurally homologous with membrane transporter YdgR, which is present in microbial cells like E. coli. 12 YdgR in E. coli also is involved in the uptake of ampicillin, sulpiride, bestatin and chloramphenicol. 13,18 POTs have been identified in E. coli and Gram-negative bacteria (Salmonella) and Bacillus. subtilis and L. lactis. 19 Sulpiride mimics the size of a dipeptide and was observed to be a substrate of YdgR found in the membrane of Gram-positive microbes that predominantly inhabited the small intestine of rats. 20 Previously, it was found that lactic acid-producing bacteria contain genes for proton-coupled oligopeptide transporter (POT) in their genome. 21
In the current study, maximum dose recoveries from the rat gut microbiome were 3.91 ± 0.25% (sulpiride-treated group). Since the bioavailability of sulpiride has been reported to be only 30% 22 of administered drug, the percent dose recovery of sulpiride from the intestinal microbiome can be a possible answer for the missing amount of drugs from systemic circulation for effective therapeutic response. Similar findings were also reported in our previous study showing uptake of 13.64% of administered dose 4 while bioavailability of paracetamol was 70 to 90%. 23,24 We propose that other orally administered drugs should also be tested for their absorption in the gut microbiome and addressed in drug monographs by reviewing the pharmacokinetic profile in context to microbial interaction during transit through the small intestine. Unabsorbed or free amounts of drugs in the gut should be quantified by adopting some strategies to find out the missing links in the absorbed and unabsorbed amounts of drugs in the gut to further address pharmacokinetic during drug development.
Conclusion
In the current in vivo microbial drug absorption assay, the human targeted drug sulpiride was postulated to serve as a substrate of the gut microbiome due to structural analogy between transporting proteins present in the enterocytes and the gut microbiome. This phenomenon limits the bioavailability of sulpiride hence affects its therapeutic response. Therefore, the researchers need to explore the new chemical entities to inhibit microbial absorbance and improved bioavailability of orally administered drugs.
Footnotes
Acknowledgments
Research work is a complementary part of PhD thesis of Mr. Imran Mukhtar supervised by Haseeb Anwar.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research was completed after utilizing the available resources and financial support in the Department of Physiology, Government College University Faisalabad, Pakistan.
