Abstract
Idiopathic nonobstructive azoospermia (iNOA) is defined as the absence of sperm in the ejaculate with no identifiable cause after thorough physical, hormonal, and genetic evaluations. This study aimed to investigate the differential gene expression of key epigenetic regulators—DNA methyltransferase (DNMT) 1 (DNMT1), DNMT3A, DNMT3B, and Zinc Finger CCHC-Type Containing 13 (ZCCHC13)—in patients with iNOA compared with those with obstructive azoospermia (OA). Sixty infertile men diagnosed with azoospermia were equally divided into iNOA (case group) and OA (control group). Participants with iNOA were further subdivided into iNOA⁺ (successful sperm retrieval, n = 10) and iNOA⁻ (unsuccessful retrieval, n = 20) based on microdissection testicular sperm extraction (micro-TESE) outcomes. The relative expression of DNMT1, DNMT3A, DNMT3B, and ZCCHC13 was analyzed in testicular tissue samples using quantitative real-time PCR. Expression levels of DNMT1 and ZCCHC13 were significantly lower in the iNOA group compared with OA (p = .01), while DNMT3B expression was significantly higher in iNOA, particularly in the iNOA⁻ subgroup (p < .0001). Receiver operating characteristic (ROC) analysis revealed DNMT3B had the highest diagnostic accuracy (AUC = 0.84, p < .0001), followed by ZCCHC13 (Area under the curve (AUC) = 0.69) and DNMT1 (AUC = 0.68). Our findings indicate that epigenetic dysregulation contributes to impaired spermatogenesis in iNOA, which can be molecularly distinguished from OA. Epigenetic biomarkers such as DNMT3B and ZCCHC13 may offer reliable diagnostic tools for idiopathic cases of azoospermia and serve as potential targets for future epigenetic therapies.
Introduction
Azoospermia, defined as the complete absence of sperm in the ejaculate, accounts for approximately 10%–15% of male infertility cases and is broadly categorized into obstructive (OA) and nonobstructive azoospermia (NOA) (Agarwal et al., 2021). A significant subset of NOA is idiopathic nonobstructive azoospermia (iNOA), characterized by the complete absence of sperm in the ejaculate. Crucially, in iNOA, the underlying cause for the testicles’ failure to produce sperm properly remains unknown. This diagnosis is reached after a thorough clinical, hormonal, genetic, and histological (testicular biopsy) evaluation rules out other identifiable etiologies (Zheng et al., 2024). The undefined etiology of iNOA poses diagnostic and therapeutic challenges, underscoring the importance of molecular investigations into potential genetic and epigenetic contributors (Tang et al., 2022).
Epigenetic mechanisms play pivotal roles in male infertility, as they regulate gene expression patterns essential for germ cell development and spermatogenesis (Firouzabadi et al., 2025; Marzouni et al., 2022; Firouzabadi et al., 2024). Among these, DNA methylation is a fundamental process that governs gene silencing and cellular differentiation (Kiselev et al., 2021; Talebi et al., 2016). This process is catalyzed by the DNA methyltransferase (DNMT) family, which are central to male reproductive epigenetics (Uysal et al., 2017). The DNMT family includes DNMT1, DNMT3A, DNMT3B, DNMT3C, and DNMT3L (Lyko, 2018). DNMT1 is responsible for maintaining methylation patterns during cell division, while DNMT3A, DNMT3B, and DNMT3L establish de novo methylation during spermatogenesis. DNMT3C is specifically expressed in male germ cells and plays a critical role in silencing retrotransposons, thus safeguarding genomic integrity during sperm development (Ren et al., 2018; Veland et al., 2019). The significance of DNMTs has been supported by animal model studies, in which dysregulation of these enzymes resulted in epigenetic disturbances and subfertility (Uysal et al., 2019). These findings highlight the essential role of DNMTs in maintaining appropriate DNA methylation landscapes critical for spermatogenic processes (Uysal et al., 2022). In addition to DNMTs, the ZCCHC13 gene has emerged as a key player in spermatogenesis (Li et al., 2018). Located on the X chromosome (Xq13.2), ZCCHC13 encodes a protein with six CCHC-type zinc finger domains, involved in nucleic acid binding and regulation of transcriptional and post-transcriptional processes (Aceituno-Valenzuela et al., 2020; Lee et al., 2024). ZCCHC13 is expressed in the nuclei of germ cells with normal spermatogenesis and is implicated in promoting cell proliferation and differentiation. Notably, studies have shown that ZCCHC13 expression is significantly downregulated in NOA patients due to promoter hypermethylation (Li et al., 2018). Importantly, demethylating agents such as 5-Aza-2′-deoxycitidine have been shown to restore ZCCHC13 expression and partially recover spermatogenesis, suggesting a direct link between epigenetic regulation and male infertility (Li et al., 2019).
Despite increasing recognition of iNOA and expanding insights into the molecular basis of epigenetic regulation in spermatogenesis, the underlying pathophysiology of iNOA remains largely elusive. This study aims to bridge this knowledge gap by conducting a comparative analysis of key epigenetic modulators in patients with iNOA and OA, thereby enhancing our understanding of the molecular mechanisms contributing to impaired spermatogenesis in idiopathic cases.
Material and Methods
Subjects of Study
Participant recruitment and sample collection were carried out between November 2024 and May 2025. Sixty infertile men diagnosed with azoospermia were equally divided into iNOA (case group) and OA (control group). iNOA patients were further subdivided into iNOA⁺ (successful sperm retrieval, n = 10) and iNOA⁻ (unsuccessful retrieval, n = 20) based solely on the outcomes of microdissection testicular sperm extraction (micro-TESE) reported from andrology laboratory. Inclusion criteria for OA were confirmed obstruction of the sperm transport pathway by imaging or surgical exploration and preserved spermatogenesis on testicular biopsy. Inclusion criteria for iNOA were absence of any genitourinary tract obstruction on ultrasound, MRI, or surgery, testicular histology demonstrating maturation arrest, hypospermatogenesis, or Sertoli cell-only patterns, and exclusion of identifiable etiologies after comprehensive hormonal, genetic, and imaging assessments. Common inclusion criteria for both groups were age 20 to 45 years; BMI < 30 kg/m2; absence of chronic systemic disease or active genitourinary infection; no history of epididymitis or orchitis in the preceding 6 months; no use of spermatogenesis-modulating medications in the past 6 months; no prior testicular or inguinal surgery; a normal karyotype without Y-chromosome microdeletions; and FSH, LH, and testosterone levels within normal ranges. Exclusion criteria comprised pyospermia, acute genital or systemic infection, genetic or chromosomal abnormalities, hypogonadism, and fever within the 90 days before enrollment.
Participant Demographics and Sample Collection
Demographic data were collected using a structured questionnaire, which also included information for evaluating participants’ hormonal profiles. Hormonal data were obtained from iNOA and OA patients. Testicular tissue samples were collected from all participants for gene expression analysis. Testicular tissue samples for iNOA patients were obtained exclusively via micro-TESE, while OA patients underwent TESE/PESA (percutaneous epididymal sperm aspiration) for sample collection. Following biopsy, sperm presence and characteristics were assessed in the andrology laboratory for relevant patient groups.
Gene Expression Assessments
RNA Extraction and Complementary DNA Synthesis
Total RNA was isolated from testicular tissue samples (30–50 mg) using the RNX‑Plus kit (Sinaclon Bioscience, Cat. No.: RN7713C, Iran), following the manufacturer’s protocol. RNA concentration and purity were assessed on a NanoDrop spectrophotometer (Thermo Scientific, Waltham, MA, USA) by measuring absorbance at 260 nm and calculating the 260/280 ratio (1.8–2.0). RNA integrity was verified by electrophoresis on a 1% agarose gel run at 100 V for 35 min, visualizing 28S and 18S rRNA bands under UV light. Subsequently, complementary DNA (cDNA) was synthesized from 500 ng of the isolated RNA using the cDNA Synthesis Kit (Parstous, Cat. No.: A101161, Tehran, Iran), following the kit’s reverse transcription protocol. The resulting cDNA was stored at −20°C.
Quantitative Real-Time PCR
Relative expression of target genes was quantified by quantitative real‑time PCR (qRT‑PCR) using a StepOnePlus Real‑Time PCR System thermal cycler (Applied Biosystems, Foster City, CA, USA). Primer validation was performed to select an optimal annealing temperature (58°C) (Table 1). Reactions were set up in duplicate using cDNA, primers, and Green Master Mix with high ROX (Amplicon; Cat. No.: A325402). The amplification protocol included initial denaturation at 95°C, followed by 40 cycles consisting of denaturation at 95°C, annealing at 58°C and 60°C, and extension at 72°C phases. Amplicon specificity was confirmed by melt curve analysis starting at 57°C. Data were normalized to β-Actin expression, and relative gene expression levels were calculated using the 2–ΔΔCt method.
The Sequence of Oligonucleotide Primers.

DNMT1 = DNA methyltransferase 1; DNMT3A = DNA methyltransferase 3A; DNMT3B = DNA methyltransferase 3B; ZCCHC13 = zinc finger CCHC-type containing 13; β-Actin = beta-Actin.
Statistical Analysis
Data analysis and visualization utilized GraphPad Prism version 10 (GraphPad Software, San Diego, CA, USA). Results were presented as mean ± SEM. Normality was assessed via the Shapiro–Wilk test. For comparisons between subgroups, parametric tests (independent t-test, one-way ANOVA with Tukey’s post hoc) or nonparametric tests (Mann–Whitney U test and Kruskal–Wallis with Dunn’s test) were employed based on data distribution. Correlation was evaluated using Spearman’s rank correlation. Diagnostic performance was assessed with receiver operating characteristic (ROC) curve analysis, including Area under the curve (AUC) calculation and optimal cutoff determination using the Youden index for maximizing sensitivity and specificity.
Results
As shown in Table 2, no statistically significant differences were observed between the iNOA and OA groups regarding mean age or BMI. Similarly, serum levels of sexual hormones did not differ significantly among the study groups. Within the iNOA group, sperm retrieval was unsuccessful in 20 patients (66.6%) and successful in 10 patients (33.3%).
The Baseline Demographic and Laboratory Indicators of the Study Participants.
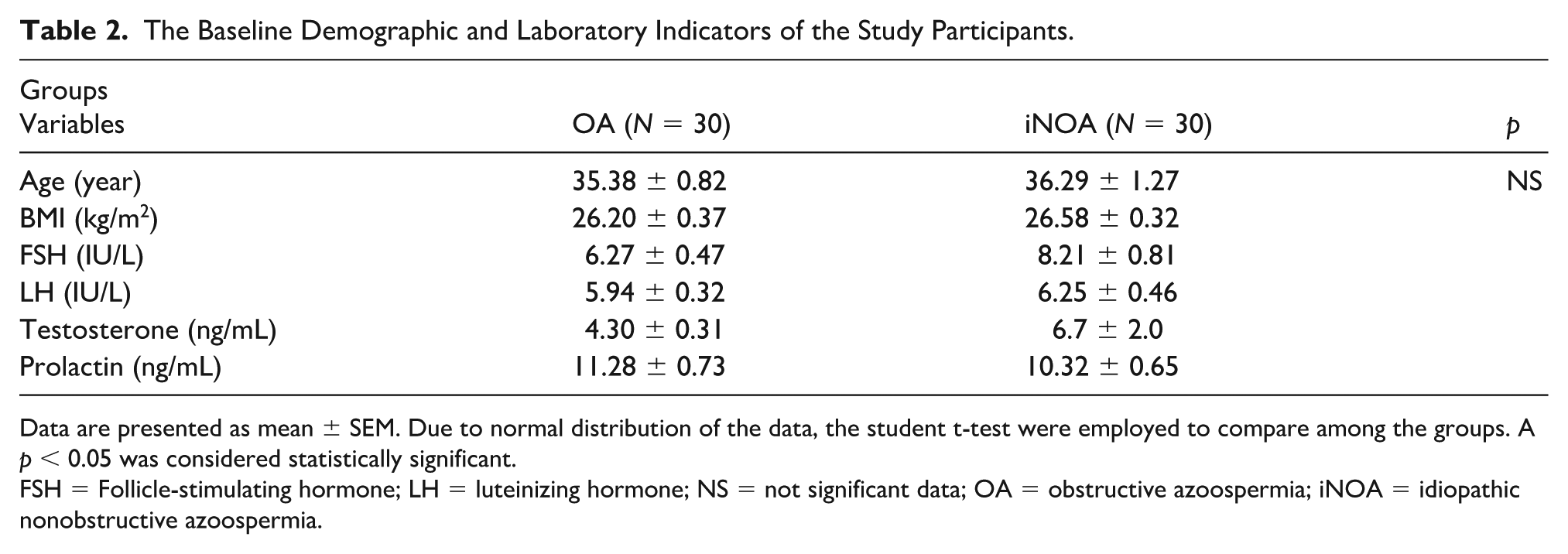
Data are presented as mean ± SEM. Due to normal distribution of the data, the student t-test were employed to compare among the groups. A p < 0.05 was considered statistically significant.
FSH = Follicle-stimulating hormone; LH = luteinizing hormone; NS = not significant data; OA = obstructive azoospermia; iNOA = idiopathic nonobstructive azoospermia.
Gene expression analysis revealed a significant reduction in DNMT1 expression in the iNOA group compared with the OA group (p
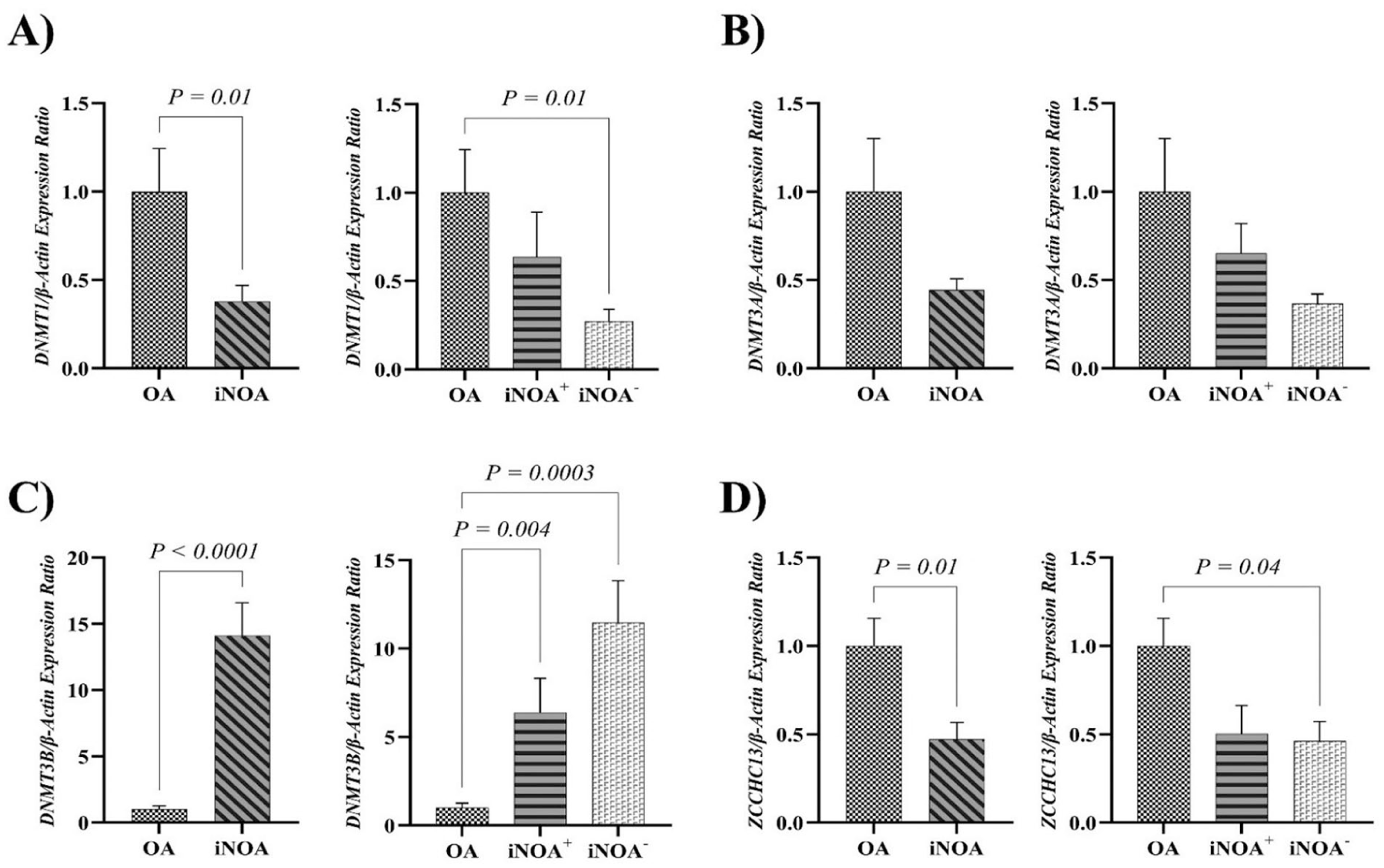
Comparison of Gene Expression Among Experimental Groups.
Subgroup analysis indicated that DNMT1 expression was significantly reduced in the iNOA⁻ subgroup compared with OA (p
ROC curve analysis demonstrated that DNMT3B had the highest diagnostic accuracy for distinguishing iNOA from OA, with an AUC of 0.84 (p

Receiver Operating Characteristic Curve of mRNA Levels of DNMT1, DNMT3A, DNMT3B, and ZCCHC13.
Correlation analysis revealed a positive association between gene expression of DNMT1 and DNMT3B (r
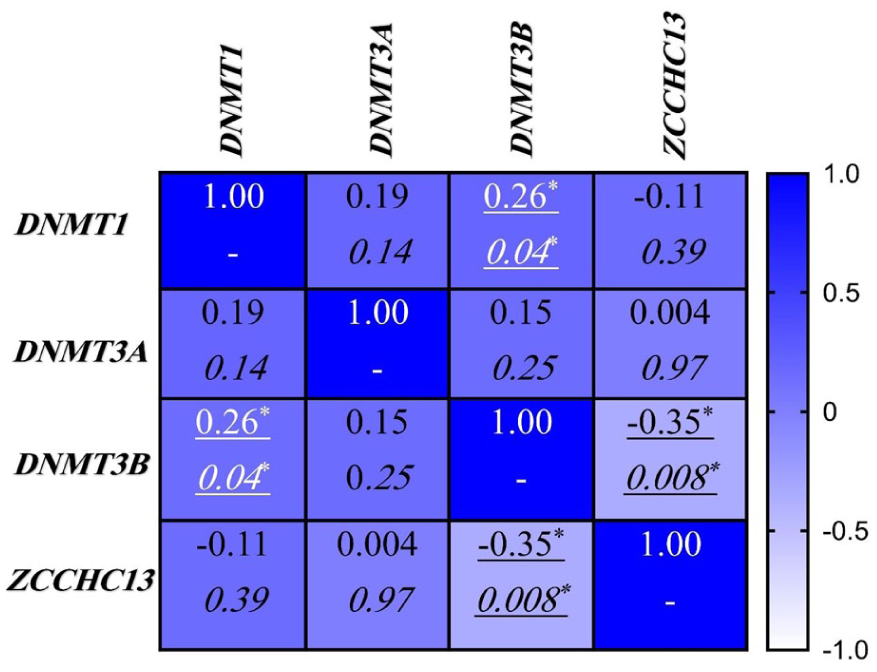
Correlation Matrix of Gene Expression in Experimental Groups.
Discussion
During DNA replication, the maintenance methyltransferase DNMT1 accurately duplicates existing methylation patterns onto the nascent DNA strand (Uysal et al., 2016). Quantitative analysis of our data revealed a significant decrease in DNMT1 mRNA levels in iNOA patients, with the most pronounced reduction observed in the iNOA⁻ subgroup compared with the OA group (Figure 1A). This downregulation aligns with previous reports showing reduced DNMT1 expression across NOA phenotypes (Uysal et al., 2019; Vahedi et al., 2022). Immunohistochemical analysis of human testicular biopsies supports these findings, demonstrating a progressive decline in DNMT1 protein levels from hypospermatogenesis through maturation arrest to Sertoli-cell-only (SCO) syndrome (Uysal et al., 2019). In addition, specific DNMT1 polymorphisms have been associated with increased risk of oligospermia (Cheng et al., 2014), and animal models have shown a postpubertal reduction in Dnmt1 expression (Washington et al., 2021). Similarly, ejaculated spermatozoa from infertile men exhibit lower DNMT1 expression, which correlates positively with normal sperm motility and morphology (Firouzabadi, 2024; Vahedi et al., 2022).
In contrast, based on our findings, DNMT3A mRNA levels showed no statistically significant difference between OA and iNOA groups (Figure 1B). This result diverges from some earlier reports of DNMT3A downregulation in specific NOA subtypes (Uysal et al., 2019), suggesting that interstudy variability or mixed histological patterns may obscure subtype-specific effects. Interestingly, our study indicated that DNMT3B expression was significantly elevated in iNOA, particularly in the iNOA⁻ subgroup compared with OA (Figure 1C). Although previous studies reported reduced DNMT3B protein levels in round spermatid arrest and SCO cases (Adiga et al., 2011; Uysal et al., 2019), it is noteworthy that baseline testicular DNMT3B expression exceeds that of other DNMT family members (Uysal et al., 2022). ROC curve analysis assigned an 84% diagnostic accuracy to DNMT3B for distinguishing iNOA from OA, highlighting its potential as a biomarker (Figure 2C).
These differential expression patterns—specifically the downregulation of DNMT1 and upregulation of DNMT3B—suggest complex regulatory mechanisms in iNOA testis. One interpretation is that increased DNMT3B expression may represent an ineffective compensatory response to underlying epigenetic disruptions, possibly including reduced DNMT1 levels (Tóth et al., 2025). Supporting this, correlation analysis showed a weak but statistically significant positive correlation between DNMT1 and DNMT3B expression (Figure 3). This suggests some degree of shared upstream regulation—such as chromatin accessibility or common transcription factors—while the divergent expression trends indicate dominant gene-specific regulatory controls within the iNOA testicular environment (Rajendran et al., 2011). Among potential regulators, the miR-29 family has been proposed to target the mRNAs of both DNMT1 and DNMT3B (Garzon et al., 2009). Previous studies indicate that pathological downregulation of miR-29 may relieve repression on DNMT3B, contributing to its elevated expression (Robaina et al., 2015; Wu et al., 2022). However, this mechanism does not fully explain the concurrent reduction in DNMT1, implying the involvement of additional regulatory factors, such as transcriptional repressors, chromatin remodelers, or distinct signaling pathways, that independently modulate each DNMT gene (Kar et al., 2012).
Expanding this epigenetic context, our study results highlighted that ZCCHC13 gene expression was found to be markedly reduced in iNOA patients, with the lowest levels detected in the iNOA⁻ subgroup (Figure 1D). Under physiological conditions, ZCCHC13 activates cellular myelocytomatosis oncogene (c-MYC), protein kinase B (AKT), and mitogen-activated protein kinase (MAPK) signaling to promote germ cell proliferation; its silencing impairs both mitotic and meiotic entries of spermatogonia, contributing to azoospermia (Li et al., 2018, 2019; Natalie & Penn Linda, 2008; Zajac-Kaye, 2001). In line with our observations, correlation analysis demonstrated a significant inverse relationship between ZCCHC13 and DNMT3B expression (Figure 3). This further supports the hypothesis that ZCCHC13 downregulation may be mechanistically linked to epigenetic disruption, potentially mediated by DNMT3B-driven promoter hypermethylation (Fanourgakis et al., 2024). Collectively, these findings indicate that ZCCHC13 silencing contributes to the molecular pathology of iNOA by disrupting germ cell proliferation and promoting spermatogenic arrest through epigenetic mechanisms (Figure 4).

A Schematic Representation of the Epigenetic Regulation Pathway Involved in Spermatogenesis.
Limitations
Although this study provides valuable insights into the role of epigenetic dysregulation in iNOA and highlights potential biomarkers such as DNMT3B and ZCCHC13, several limitations should be considered. First, the overall sample size was relatively small, and within the iNOA group, the number of patients with successful sperm retrieval was even more limited. This inevitably reduces the statistical power of the analysis, particularly given that multiple epigenetic markers were evaluated simultaneously. It should be emphasized, however, that collecting well-characterized testicular tissue from patients with iNOA poses substantial clinical challenges. Recruitment requires strict adherence to inclusion and exclusion criteria, careful matching with appropriate controls, and considerable time to ensure the integrity of the cohort. For these reasons, similar studies in the field have typically relied on comparably small sample sizes. Second, the present investigation was confined to molecular and epigenetic assessments; the absence of complementary proteomic, transcriptomic, or functional analyses limits the depth of biological interpretation. Finally, potential confounding variables cannot be fully excluded, which may affect the robustness and generalizability of the findings. Future studies with larger, multicenter cohorts, together with integrative multi-omics approaches, will be essential to validate these results and to facilitate their translation into clinically meaningful strategies for managing male infertility.
Future Directions
To build upon the findings of this study and further elucidate the molecular and epigenetic underpinnings of OA and iNOA, future research should consider the following avenues:
Comprehensive Epigenomic Profiling
Perform whole-genome bisulfite sequencing to map DNA methylation changes across the entire genome in OA versus iNOA testes. Integrate histone-modification profiling (e.g., ChIP-seq of H3K4me3 and H3K27me3) to uncover additional layers of epigenetic regulation affecting DNMT enzymes and ZCCHC13.
Single-Cell Resolution Analyses
Apply single-cell RNA-seq and single-cell ATAC-seq on testicular biopsies to distinguish gene expression and chromatin-accessibility changes in specific germ cell subpopulations. Identify cell-type–specific dysregulation of DNMT1, DNMT3A/B, and ZCCHC13 in Sertoli cells, spermatogonia, and spermatocytes.
Functional Validation In Vitro and In Vivo
Use CRISPR/Cas9-mediated knockout or overexpression of DNMT3B and ZCCHC13 in germ cell lines or organoid models to assess causal effects on cell-cycle progression and differentiation. Generate mouse models with testis-specific modulation of these genes to study impacts on fertility and potential transgenerational epigenetic inheritance.
Longitudinal and Clinical Correlation Studies
Design prospective cohorts of OA and iNOA patients sampling at multiple time points before and after treatment to track dynamic changes in gene expression and methylation relative to therapeutic outcomes. Correlate identified epigenetic markers with long-term fertility metrics and offspring health parameters.
Biomarker Development and Machine-Learning Integration
Combine expression levels of DNMT1, DNMT3B, and ZCCHC13 with circulating microRNAs or protein markers to create composite panels for noninvasive diagnosis and prognosis. Employ machine-learning algorithms on multi-omics data sets to classify OA versus iNOA and predict micro-TESE success with optimized model performance.
Therapeutic Intervention Trials
Initiate preclinical studies of epigenetic drugs—such as DNMT inhibitors or DNA methylation modulators—to evaluate their efficacy in restoring normal spermatogenesis in iNOA models. Explore targeted activation of the AKT/MAPK–c-MYC axis to rescue ZCCHC13 expression and promote germ cell proliferation.
Addressing NOA Heterogeneity and Differential Diagnosis
One crucial future direction is to address the significant heterogeneity within NOA by conducting studies that carefully stratify patients based on defined etiologies, such as genetic abnormalities, Sertoli cell-only syndrome, or specific maturation arrest patterns. This subclassification is essential to determine whether the identified epigenetic markers, including DNMTs and ZCCHC13, exhibit subtype-specific associations and to validate their utility in the differential diagnosis of these distinct conditions.
Conclusion
Our findings indicate that epigenetic dysregulation contributes to impaired spermatogenesis in iNOA, which can be molecularly distinguished from OA. Epigenetic biomarkers such as DNMT3B and ZCCHC13 may offer reliable diagnostic tools for idiopathic cases of azoospermia and serve as potential targets for future epigenetic therapies. However, further validation in larger, well-characterized patient cohorts with proteomic and functional analyses will be needed to confirm these findings and assess their diagnostic and therapeutic potential.
Footnotes
Acknowledgements
During the preparation of this work, the authors employed Gemini to assist in drafting the “Future Research Directions” section and enhancing the overall quality of the writing. Following the use of Gemini, the authors thoroughly reviewed and edited the content as needed, assuming full responsibility for the accuracy and integrity of the published material.
Ethical Considerations
The Medical Ethics Committee of Shahid Sadoughi University of Medical Sciences, Yazd, approved this study with approval number IR.SSU.SPH.REC.1403.223.
Consent to Participate
All participants signed an informed consent form before sample collection.
Author Contributions
All authors contributed to the conception and design of the study. Farzaneh Fesahat was responsible for overall supervision. Seyyed Amir Hossein Mirghanizadeh Bafghi and Farangis Demehri were responsible for sample collection. Samaneh Harimi carried out the practical work, while Amir Masoud Firouzabadi drafted the manuscript and performed the statistical analyses, with support from Parisa Heydari in the statistical work. The manuscript was revised by Farzaneh Fesahat and Amir Masoud Firouzabadi. All authors read and approved the final manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data supporting this study’s findings are available on request from the corresponding author. The data are not publicly available due to privacy or ethical restrictions.
