Abstract
The objective of the present study was to find out the association of folate genes MTR A2756G and MTRR A66G polymorphisms with the risk of male infertility. The databases of Google Scholar, PubMed, and Science Direct were searched to find relevant studies. Data were extracted from the eligible studies and were analyzed for pooled up odds ratio (OR) with 95% confidence interval (CI). Review Manager 5.4 was used for statistical analysis. Nineteen case-control studies were included in this meta-analysis which comprised 3621 cases and 3327 controls. Pooled analysis revealed that there is a significant association between MTR A2756G polymorphism with male infertility except for the dominant model. The ORs and 95% CI for each genetic model were as follows: 1.21 [1.03–1.42] for the allele model (G vs. A), 2.31 [1.38–3.96] for the additive model (GG vs. AA), 1.17 [0.98–1.38] for the dominant model (GG+AG vs. AA) and 2.10 [1.55–2.86] for the recessive model (GG vs. AG+AA). MTRR A66G has no noticeable association with male infertility. The current meta-analysis suggests that MTR A2756G polymorphism might be a potential risk factor for male infertility. In the future, the sample size should be increased to confirm the present results.
Introduction
The World Health Organization (WHO) defines infertility as a couple’s inability to conceive or carry a pregnancy following a year of routine, unprotected sexual activity (Balkan et al., 2013; Tanoomand et al., 2019; Tüttelmann et al., 2007). The prevalence of male fertility issues is between 40% and 50% (Balkan et al., 2013). Impaired semen parameters, such as aberrant sperm count, concentration, morphology, and motility, might contribute to infertility (Du Plessis et al., 2010).
Several genes regulate the intricate process of spermatogenesis, and some of these genes are implicated in the route of folate metabolism (Tanoomand et al., 2019). Folates are interconvertible co-enzymes that help in preserving the stability and integrity of the genome by helping in DNA synthesis, repair, and methylation. The methylation of lipid, protein, and nucleic acids occurs along the folate pathway, which is also critical for the metabolism of amino acids, the production of purines and pyrimidines, and the synthesis of nitrogenous bases (Balkan et al., 2013; Gava et al., 2011; Kurzawski et al., 2015; Lee et al., 2006; K. Liu et al., 2015).
Methylenetetrahydrofolate reductase (MTHFR), methionine synthase (MTR), and methionine synthase reductase (MTRR) are three essential enzymes in the folate metabolism (K. Liu et al., 2015; Tanoomand et al., 2019). MTHFR changes 5, 10-methylenetetrahydrofolate into 5-methyltetrahydrofolate. Methionine is formed by transferring a methyl group from 5-methyltetrahydrofolate, which serves as a precursor for S-adenosylmethionine. MTR and MTRR conduct the process. This methionine is converted into homocysteine by methionine synthase. In animals, MTR function is essential for maintaining adequate intracellular folate pools, and it requires vitamin B12 as a cofactor (Leclerc et al., 1996). The MTRR gene carries out the reductive methylation of the MTR gene and keeps the MTR gene alive (Farcas et al., 2009; K. Liu et al., 2015). In this manner, sufficient intracellular folate pools are preserved (Karimian & Colagar, 2016).
The MTRR gene is located on chromosome 5 at the p15.2–15 position. It has 15 exons and 14 introns (Leclerc et al., 1999). The MTR gene is located on chromosome 1 at the q43 position. It has 33 exons and 32 introns (Karimian & Colagar, 2016). Some mutations result in defective enzymes in the folate metabolism pathway, which are linked to male infertility (K. Liu et al., 2015). Single nucleotide polymorphisms (SNPs) in MTRR A66G (rs1801394) and MTR A2756G (rs1805087) have been reported (Karimian & Colagar, 2016; Kurzawski et al., 2015; Lee et al., 2006; Tanoomand et al., 2019) which can alter the amount, stability, and activity of enzymes involved in folate metabolism, which could have an impact on DNA synthesis and folate metabolism (Z. J. Ren et al., 2017). Sperm characteristics like motility, count, and morphology may be impacted by DNA damage (Singh & Jaiswal, 2013). As a result of these mutations, homocysteine accumulation may have an impact on gamete development by preventing the synthesis of nitric oxide, which is essential for sperm capacitation and the acrosome response (Karimian & Colagar, 2016). DNA and histone protein methylation changes can cause genome instability and failure of epigenetic regulation in proliferation (Godmann et al., 2009).
Numerous researchers have looked at the relationships between these SNPs and the risk of male infertility, but their findings are inconsistent. Karimian and Colagar conducted a meta-analysis in 2016 on 7 case-control studies which included 1725 cases and 1678 controls. Their results reported that MTR A2756G is associated with idiopathic male infertility in both Asian and non-Asian populations (Karimian & Colagar, 2016). The folate pathway’s associated gene variations may be a possible risk factor for male infertility. A meta-analysis was conducted regarding MTRR mutation A66G which included 14 studies. The total number of cases and controls was 2620 and 2614, respectively. The overall results revealed a lack of significant association between MTRR A66G polymorphism and idiopathic male infertility risks (S. Q. Ren et al., 2019). According to another study, infertility in males due to unexplained decreased sperm counts is not linked to MTRR A66G polymorphism. These findings imply that genetic variations in folate metabolism-related enzymes do not significantly affect sperm counts (Ravel et al., 2009). To boost the statistical power of all relevant research and to compile the available data, meta-analysis is a useful technique. The present meta-analysis based on 15 studies of MTR A2756G and MTRR A66G was performed to clarify the association of both these polymorphisms with the risk of male infertility.
Methods
Search Strategy
Meta-analysis was carried out using the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement presented in Figure 1. The databases Google Scholar, PubMed, and Science Direct were searched using the keywords “MTR,” “MTRR” and “Male Infertility.” The references of reviews and retrieved articles were also searched to find out additional pertinent publications. Studies that had been published in English only were included in the search. Since every analysis was founded on previously published research, neither patient consent nor ethical review was necessary.
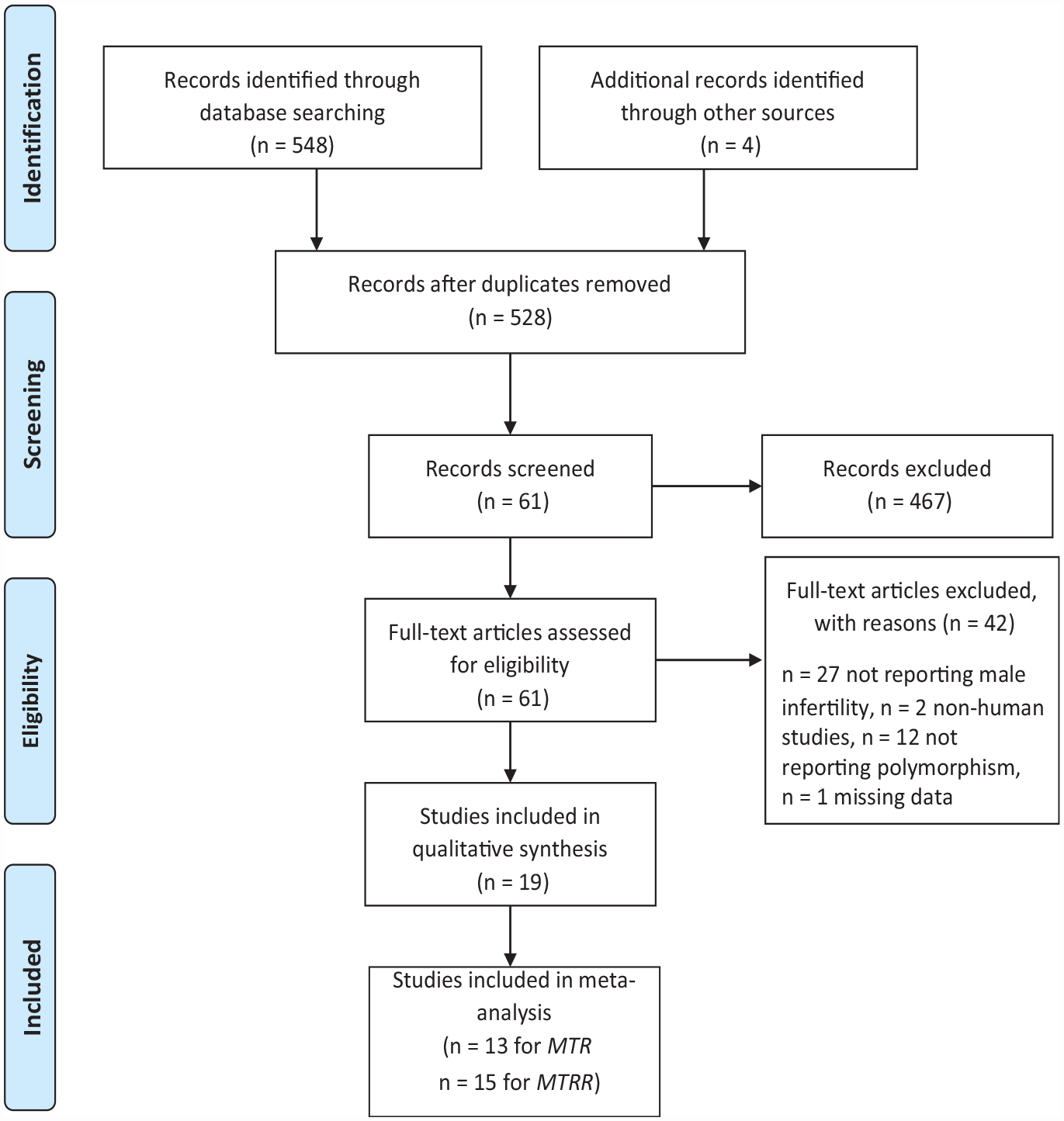
Flowchart Showing the Selected Studies
Inclusion and Exclusion Criteria
The studies included in the current meta-analysis had to satisfy the following requirements: studies with full-text articles, published in English; examining the relationship between genotype frequencies or allele frequencies for both cases and controls; case-control studies; sufficient genotype data were available for the estimation of odds ratios (ORs) and 95% confidence intervals (CIs). Non-case-control studies, duplicate publications, animal studies, reviews, repeated literature, abstracts, and comments, as well as studies with insufficient data to calculate genotype frequency, were all excluded from the consideration.
Data Extraction Strategy
From each eligible study, information was extracted according to the inclusion criteria listed above. The extracted information included the first author, publication year, country, ethnicity, age groups, inclusion and exclusion criteria, genotyping method, the sample size for the case and control groups, and genotypic and allelic frequencies. Extracted data were checked after being entered into a collection form. Disagreements were settled through discussion by two investigators (TT and AA).
Statistical Analysis
In order to analyze the relationships between the idiopathic male infertility risks and the MTRR A66G and MTR A2756G gene polymorphisms, four models were used: the allelic model, the additive model, the dominant model, and the recessive model. The ORs with 95% CIs were used to determine the strength of the association between the MTRR A66G and MTR A2756G gene polymorphism and the risks of idiopathic male infertility. To measure the degree of the heterogeneity of the study, the chi-square-based Q test and the I2 metric were applied (I2=0%-20%, no heterogeneity; I2 = 20%-50%, moderate heterogeneity; I2 > 50%, obvious heterogeneity). The pooled ORs and 95% CIs were estimated using a random-effects model. p < .10 or I2 > 50% were indicators of heterogeneity. Reviewer Manager 5.4 was used for the statistical analysis.
In-Silico Analysis of MTR and MTRR Expression
To analyze the influence of polymorphisms on MTR and MTRR expression levels, GTEx Analysis Release V8 (dbGaP Accession phs000424.v8.p2) data was used (Gibson, 2015). GEPIA (http://gepia.cancer-pku.cn/) was used to determine the differential expression of MTR and MTRR in the tumor and normal tissues.
Trial Sequential Analysis
Trial sequential analysis (TSA) was conducted by using Trial Sequential Analysis Viewer 24 software, version 0.9.5.10 Beta, Copenhagen Trial Unit, Center for Clinical Intervention Research, 25 (Gordon Lan & DeMets, 1983). A 5% percent significance was selected for type I error, and 80% power was used to calculate the sample size required and also to set the TSA boundaries.
Results
Study Characteristics
Based on literature search and inclusion criteria overall 19 case-control studies were included in this meta-analysis which comprised 3621 cases and 3327 controls. The studies were published between 2006 and 2021. The numbers of case-control studies containing the MTR A2756G mutation and MTRR A66G mutation were 13 and 15, respectively. Out of the total 19 studies, 13 studies of MTR A2756G polymorphism included 2662 cases and 2752 controls. Out of 13 studies, 8 studies were conducted on Asians and 5 were on non-Asians. For MTRR A66G polymorphism 15 case-control studies met the inclusion criteria. The number of Asian and non-Asian studies was 7 and 8, respectively. These studies included 3089 cases and 2872 controls. The main characteristics of the included studies are listed in Tables 1 and 2, respectively.
Characteristics of the Studies of MTR A2756G Included in Meta-Analysis

Note. ART = Assisted Reproductive Techology, ASA: Antisperm antibodies, BMI = Body mass index, NOA = Non-obstructive azoospermia, PCR-RFLP = Polymerase chain reaction-restriction fragment analysis, RT-PCR = Reverse transcription-polymerase chain reaction.
Characteristics of the Studies of MTRR A66G Included in the Meta-Analysis
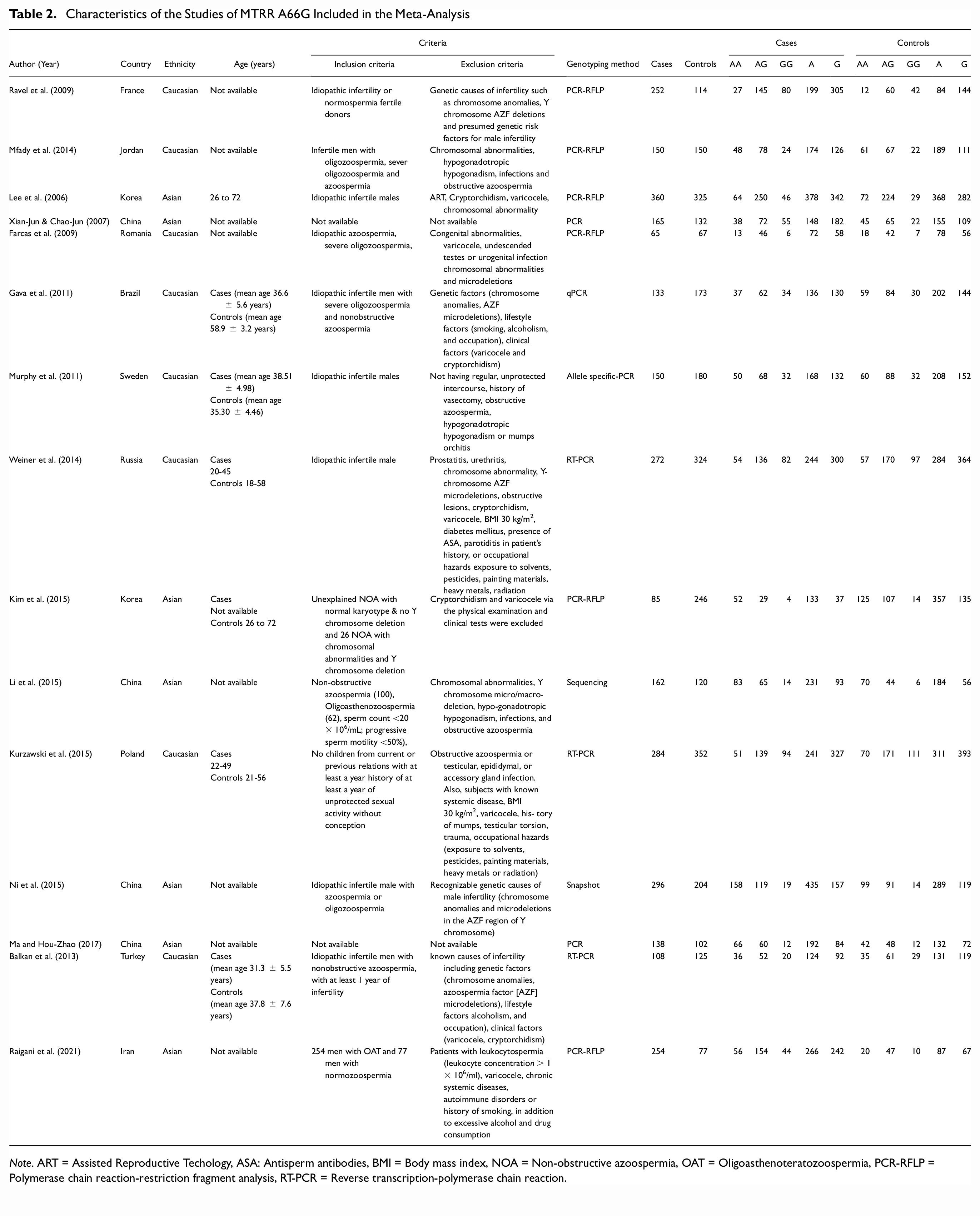
Note. ART = Assisted Reproductive Techology, ASA: Antisperm antibodies, BMI = Body mass index, NOA = Non-obstructive azoospermia, OAT = Oligoasthenoteratozoospermia, PCR-RFLP = Polymerase chain reaction-restriction fragment analysis, RT-PCR = Reverse transcription-polymerase chain reaction.
Association of MTR A2756G Polymorphism With Male Infertility
The effect of the MTR A2756G polymorphism on the likelihood of idiopathic male infertility was assessed in 11 case-control studies. The major findings of the meta-analysis of the relationships between the A2756G mutation and the risk of idiopathic male infertility are presented in Table 3. The findings of the meta-analysis are presented in Figure 2 for the allele (G vs. A), additive (GG vs. AA), dominant (GG+AG vs. AA), and recessive (GG vs. AG+AA) models. The I2 values for these models, which indicate the between-study heterogeneity, were 45%, 60%, 37%, and 0%, respectively. The results showed a strong significant association between MTR A2756G polymorphism and male infertility except for the dominant model. The ORs and 95% CIs for each model were as follows: 1.21[1.03–1.42] for the allele model (G vs. A), 2.31 [1.38–3.96] for the additive model (GG vs. AA), 1.17 [0.98–1.38] for the dominant model (GG+AG vs. AA) and 2.10 [1.55–2.86] for the recessive model (GG vs. AG+AA).
Meta-Analysis Showing Genetic Models of MTR A2756G Polymorphism With Male Infertility

Note. CI = confidence interval.

Forest Plots of the Studies of MTR A2756G Polymorphism Linked With Male Infertility. (A) Allelic Model G vs. A (B) Additive Model GG+AA (C) Dominant Model GG+AG vs. AA (D) Recessive Model (GG vs. AG+AA)
Association of MTR A2756G Polymorphism With Male Infertility in Asians
On data that had been analyzed by subgroups, based on ethnicity, the MTR A2756G polymorphism has been discovered to significantly enhance the incidence of male infertility in the Asian population, according to the additive and recessive model. The ORs and 95% CIs for each model were as follows: 1.21[0.95–1.56] for the allele model (G vs. A), 2.14[1.33–3.45] for the additive model (GG vs. AA), 1.22[0.93–1.60] for the dominant model (GG+AG vs. AA) and 1.98[1.24–3.18] for the recessive model (GG vs. AG+AA) as presented in Table 3.
Association of MTR A2756G Polymorphism With Male Infertility in Caucasians
According to the additive and recessive models of comparison, there was a significant association between MTR A2756G and the risk of male infertility. The ORs and 95% CIs were 2.96 [1.10–7.96] and 2.14 [1.32–3.47] for additive and recessive models, respectively (Table 3).
Association of MTR A2756G Polymorphism With Idiopathic Male Infertility
According to the additive and recessive models of comparison, a significant association was identified between MTR A2756G and the risk of idiopathic male infertility. The ORs and 95% CIs were 2.33 [1.59 – 3.41] and 2.42 [1.64–3.56] for additive and recessive models, respectively.
The sub-group analysis based on ethnicity presented a significantly increased risk of idiopathic male infertility in Caucasians except for the dominant model. The ORs and 95% CIs for each model were as follows: 1.41 [1.14–1.74] for the allele model (G vs. A), 2.40 [1.46–3.95] for the additive model (GG vs. AA), 1.29 [0.99–1.66] for the dominant model (GG+AG vs. AA) and 2.80 [1.71–4.59] for the recessive model (GG vs. AG+AA) as presented in Table 3. No significant association of the polymorphism was observed with idiopathic male infertility in Asians, as presented in Table 3.
Association of MTRR A66G Polymorphism With Male Infertility
Four genetic models of the MTRR A66G polymorphism had low between-study heterogeneity (I2 range: 0-21%; P-value range: 0.11-0.43). Overall, none of the comparison models revealed a significant association between the MTRR A66G polymorphism and male infertility (Figure 3). For each model, the ORs and 95% CIs were mentioned in Table 4.

Forest Plots of the Studies of MTRR A66G Polymorphism Linked With Male Infertility. (A) Allelic Model G vs. A (B) Additive Model GG+AA (C) Dominant Model GG+AG vs. AA (D) Recessive Model (GG vs. AG+AA)
Meta-Analysis Showing Genetic Models of MTRR A66G Polymorphism With Male Infertility

Note. CI = confidence interval.
Association of MTRR A66G Polymorphism With Male Infertility in Asians
On the data stratified by ethnicity, subgroup analyses were performed. Among either of the genetic models, there was no significant association found between the MTRR A66G polymorphism and the incidence of male infertility in Asians. The ORs and 95% CIs for each model were as follows: 1.09 [0.88–1.35] for the dominant model (GG+AG vs. AA), 1.37[0.89–2.13] for the allele model (G vs. A), 1.06 [0.82–1.37] for the additive model (GG vs. AA), and 1.40[0.99–1.97] for the recessive model (GG vs. AG+AA).
Association of MTRR A66G Polymorphism With Male Infertility in Caucasians
There is no significant association identified between MTRR A66G polymorphism and male infertility in Caucasians. The OR and 95% CI are presented in Table 4.
Association of MTRR A66G Polymorphism With Idiopathic Male Infertility
Overall, none of the four genetic models presented a significant association between the MTRR A66G polymorphism and the risk of idiopathic male infertility, neither in Asians nor in Caucasians. The ORs and 95% CIs for each model are mentioned in Table 4.
In-Silico Expression Analysis
By using GTExportal, a significant decrease in the expression of MTRR for rs1801394 was observed in GG as compared to AA in testis (p = 1.97 × 10−8) (Figure 4). The differential expressions of MTR and MTRR in the different tumors as well as normal tissues are also presented in Figure 5.

Graph Presenting Significant Influence of A66G (rs1801394) Polymorphism on the Expression of MTRR Gene in Testis

Gene Expression Profile (A) MTR; (B) MTRR) Across Various Tumor Samples (Red Peaks) and Paired Normal Tissues (Green Peaks). The Red Labeled Tumor Types Presents Over-Expression, Green Presents Under-Expression and Black Presents no Change in the Respective Gene Expression as Compared to Normal Tissue
Results of Trial Sequential Analysis
For the allelic model of MTR A2756G and MTRR A66G, the required sample size was 29219 and 30592, respectively, at 80% power, whereas the cumulative z-curve did not cross the trial sequential monitoring boundaries (Figures 6 and 7). However, the TSA of the allelic model of MTR A2756G crossed the conventional Z-score boundaries in favor of control. It shows that more studies, with larger sample sizes, are required for MTRR A66G to provide sufficient evidence for confirming the results of the current meta-analysis.
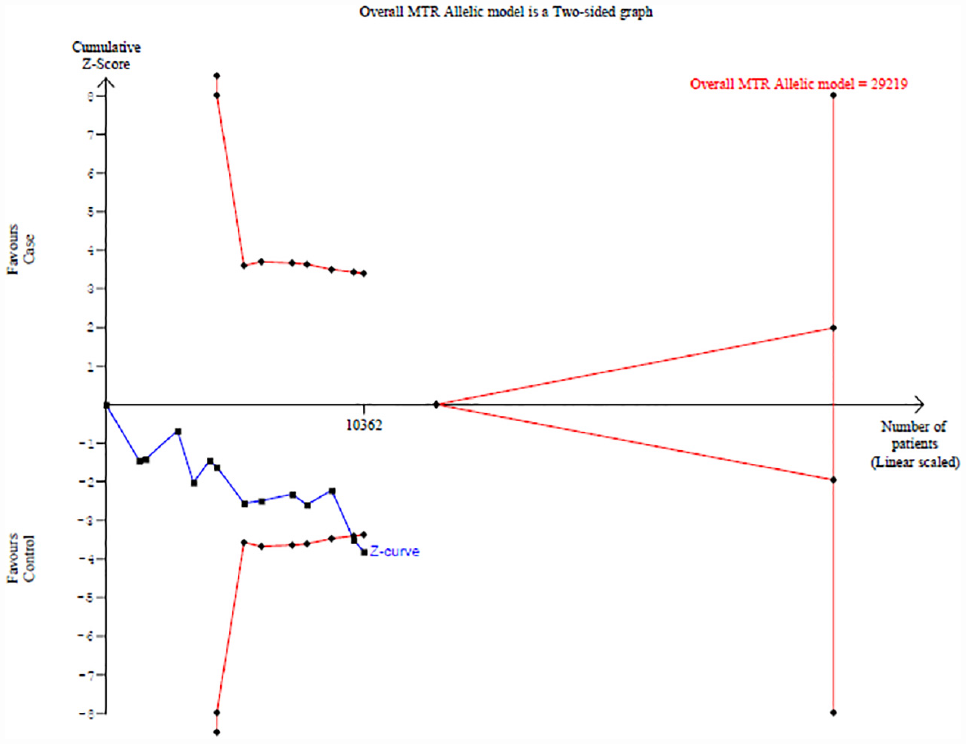
Trial Sequential Analysis (5% Type-I Error & 80% Power) for MTR A2756G Polymorphism for the Allelic Model

Trial Sequential Analysis (5% Type-I Error & 80% Power) for MTRR A66G Polymorphism for the Allelic Model
Discussion
One of the most promising areas of research in the genetics of male infertility is the complicated process of spermatogenesis, which results in haploid spermatozoa from mitotic and meiotic divisions of germ cells (K. Liu et al., 2015). Male infertility has been linked to abnormal folate metabolism. There is strong experimental support for the idea that certain folate metabolism enzymes are necessary for male infertility. The major enzymes involved in the folate metabolic pathways, essential for DNA methylation and spermatogenesis, are called folate-related enzymes. By altering the activity, stability, or concentration of the associated enzymes, the SNPs of these folate-related enzyme genes might reduce the absorption of folate or upset the equilibrium between folate derivatives (S. Q. Ren et al., 2019. The major enzymes involved in the folate metabolic pathways, MTR and MTRR, are essential for DNA methylation (K. Liu et al., 2015).
The previous literature presents a relationship between cancer and male infertility. Nagirnaja et al. (2018) identified 25 candidate genes, whereas almost half of them are associated with a predisposition to hereditary cancer. MTRR gene polymorphism was reported to be associated with malignant mesothelioma of testicular tunica vaginalis testis (Kowalik et al., 2020).
Polymorphisms in folate metabolism genes, that is, MTR and MTRR, significantly contribute to the risk of breast cancer in females in Jordan (Sadiq et al., 2019), and may also increase the risk of cervical cytological abnormalities (Silva et al., 2022). MTRR A66G was associated with the risk of developing lung cancer in the Turkish population, as compared to MTR A2756 polymorphism (Aksoy-Sagirli et al., 2017). In the case of prostate cancer, MTR A2756G was associated with increased susceptibility (Zhang et al., 2020), whereas MTRR A66G was not reported to be associated with it (Basir, 2019). MTRR A66G polymorphism plays a strong genetic role in the progression of acute myeloid leukemia in the Saudi population (Farasani, 2023).
In the current meta-analysis, the results showed a strong significant association between MTR A2756G polymorphism and male infertility. The comparison models of MTR A2756G polymorphism showed p values < .05 and have been discovered to greatly enhance the risk of idiopathic male infertility in the Asian population. However, sub-group analyses on Caucasians have revealed significant values only for the additive GG+AA and the recessive model GG vs. AG+AA. About other polymorphisms, there is no significant association found between MTRR A66G and male infertility. Sub-group analysis was also performed based on ethnicity and in both Asians and Caucasians, it has been found that there is a lack of association between this SNP and the risk of idiopathic male infertility.
Similar to these results, a meta-analysis reported that there is a strong link between the MTR A2756G polymorphism and idiopathic male infertility. A substantial connection was discovered between the polymorphism and idiopathic male infertility in non-Asians but not in Asian populations, when they conducted a subgroup analysis by ethnicity. For MTRR A66G, there was no conclusive association between the MTRR A66G polymorphism and idiopathic male infertility discovered in the overall population; however, Asians were identified to be more likely than non-Asians to have the polymorphism (Z. J. Ren et al., 2019). A meta-analysis conducted in 2016 reported that the mutation MTR A2756G may be related to a risk of male infertility. The probability of azoospermia and OAT was raised in MTR A2756G (Karimian & Colagar, 2016). Another meta-analysis performed in 2015 reported that for the pathophysiology of male infertility, MTR A2756G and MTRR A66G were viable candidates; however, further case-control studies were needed to prevent false-positive results (K. Liu et al., 2015).
According to a case-control study, it has been revealed that the SNP MTR A2756G was associated with oligozoospermia and non-obstructive azoospermia (NOA) (Karimian & Colagar, 2016). In 2011, a study on MTR A2756G reported a significant association between the GG genotype and NOA among the Brazilian population, and no connection was discovered between NOA and SO groups and controls for the MTRR A66G polymorphism (Gava et al., 2011). MTR 2756A > G polymorphism was not linked with male infertility according to the Korean (Lee et al., 2006), Russian (Weiner et al., 2014), as well as Swedish studies (Murphy et al., 2011). These findings were also supported by the case-control study conducted by Kurzawski et al., 2015 which revealed no link between Polish male infertility and the MTR 2756A > G polymorphism. However, the MTR 2756GG genotype was linked to a higher incidence of azoospermia in the Korean research by Lee et al. (2006). It was originally noted in a case-control study that the MTRR A66G mutation significantly increased the risk of male infertility in an Asian community and that was not identified to be associated with non-Asians (Lee et al., 2006). According to the current meta-analysis, similar results were reported by a case-control study in Turkey. The MTRR A66G genotype frequencies of patients and controls did not differ from one another. The MTRR A66G polymorphism allele frequencies of patients and controls reported no statistically significant difference (Balkan et al., 2013). Farcas et al. (2009) mentioned that the p-values obtained were above .05, while the OR indicates that these two polymorphisms were not the risk factors for male infertility. No significant association was identified between both these polymorphisms and male infertility (Kim et al., 2015). Regarding MTRR A66G polymorphism, the results of Mfady et al. (2014) reported a lack of its association with the risk of male infertility, which is in agreement with other reports that stated such absence of association in the Romanian (Farcas et al., 2009), French (Ravel et al., 2009) and Brazilian (Gava et al., 2011) populations. Possibly the limited number of participants who had the MTRR 66GG genotype contributed to the lack of relationship between the MTRR A66G variation and male infertility seen in the current study. The investigated polymorphisms may also interact strongly with dietary/environmental variables (such as folate status), as well as with racial and genetic factors.
Although this investigation produced generally strong statistical evidence, several shortcomings need to be addressed. Geographical variations and racial and ethnic differences in the distribution of polymorphisms in the MTRR gene might have caused discrepancies across the research. Only 19 case-control studies are included in this meta-analysis, thus the sample size was small. Further research on the potential association of MTR A2756G and MTRR A66G polymorphism with male infertility risk requires high-quality studies with bigger sample numbers. Also, the studies in this meta-analysis were based on Asians and Caucasians. A wide range of diseases, including azoospermia, oligozoospermia, and teratozoospermia, among others, are included in the idea of male infertility. Male infertility is a multifactorial illness brought on by intricate interactions between a variety of genetic and environmental variables. As a result, a more thorough subgroup analysis and examination of the cumulative impact of the many SNPs are needed. There should be more research on A2756G, A66G, and African people as there was no African study.
Conclusion
In conclusion, the current meta-analysis shows that the polymorphisms MTR A2756G and MTRR A66G may add to a gene’s vulnerability to male infertility risk. To support the conclusions presented here, large-scale, well-planned, population-based investigations are required.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical Approval
Ethical approval does not apply to the present meta-analysis.
