Abstract
Millions of adults have diabetes across the globe and the overall cost for managing diabetic patients has reached up to approximately 250 million. A major constraint in existing ontology-based systems for diagnosing and treating diabetes is the presence of semantic inconsistencies and lack of a comprehensive clinical approach primarily due to consideration of a limited number of classes in the model. In this research, we are focused on building an ontology-based model for diabetic patients by collecting detailed diabetic knowledge of subjects for further diagnosis and treatment. The concept of semantic resources to electronic health record standards is an essential factor for semantic interoperability in remote health monitoring. This study applies semantic web ontology language for developing ontology-based model for diabetic patients to aid doctors in reaching an efficient diagnostic decision about the status of diabetes by applying Semantic Web Rule Language. A total of 766 medical records from clinical environment were selected in this study, and 269 of them were known for developing diabetes. The experimental results suggest that the proposed solution is more accurate in managing diabetes compared to other medical applications. The performance analysis of the ontology-based model for diabetic patients regarding the accuracy of disease prediction, diagnosing diabetes, and recommending medicine is 95%, 98%, and 85%, respectively.
Keywords
Introduction
Recent developments in medical devices and communication technologies have revolutionized healthcare systems.1–5 In remote healthcare, sensor nodes measure patient’s vital signs and aggregate these data into medical files which are uploaded to the cloud for storage and access by healthcare workers as shown in Figure 1. These healthcare systems are applied for several applications including diabetes management, hypertension, and rehabilitation, among others. However, the incidence and prevalence of diabetes mellitus worldwide have been observed due to increasing urbanization, obesity, aging populations, and decrease in the levels of physical activities. 6 A report by World Health Organization (WHO) 7 predicts that, by 2030, approximately 600 million people will be suffering from diabetes, and diabetes will be the seventh deadliest disease. Diabetes mellitus often leads to complex coronary heart diseases, cerebrovascular diseases, blindness, and other serious complications. Since diabetes is a critical and complex disease, its diagnosis and treatment require proper clinical guidelines for physicians to make adequate diagnostic and therapeutic recommendations for patients. This approach presents a lot of problems for physicians who need to memorize a variety of cases and diagnostic items. Thus, any negligence can cause incorrect diagnosis of diabetes, and it is difficult to cope with a large number of diabetic patients. Hence, by exploring the emerging computational intelligence methods, the development of a clinical decision support system (CDSS) has brought new directions for the diagnosis and treatment of diabetes and other diseases.8–10

Ontology models for patients in remote healthcare systems.
Electronic health records (EHR) ensure that individuals can access their medical records and laboratory test outcomes to track, monitor, and manage their prolonged diseases. 11 The increasing use of EHR at global level causes subjects’ medical information to be transferred to several clinics and hospitals. 12 This condition requires semantic interoperability of scientific data for providing accurate and reliable transmission of data across medical networks. The lack of such interoperability usually results in wastage of enormous resources across the globe on a yearly basis. 13 Rea et al. 14 developed a semantic health platform based on electronic records; they utilized ontology and standard terminology to attain the anticipated semantic interoperability. Recently, several researchers have a tendency toward developing ontology-driven models for health monitoring applications. Ontology is an organized system designed to categorize and help explain the relationships between various entities in the same area of knowledge and research. 15 There are several applications of ontology-based systems including neural systems, 16 cultural variations in interpersonal communication, 17 information retrieval for mobile learning resources, 18 diabetes, 19 epilepsy, 20 and others.
Many researchers are focusing on developing ontology-based frameworks and techniques for diabetic patients. 21 Various studies have developed different analytical frameworks which could specify the main symptoms of the illness and examine diabetes by analyzing the medical information of patients.22–25 Lee et al. 22 proposed an ontology-based approach which provides a particular recommendation in diet menu to the patients. It is a well-known fact that diet menu is essential for the treatment of diabetic patients. In addition, the menu plays an important role in offering valuable suggestions to the patients. It can efficiently assist in handling medical information and acquired knowledge is applied for recommending a meal plan for diabetic patients. But the main drawbacks of this method are the inability to provide adequate exercise and treatment plans to aid patients’ recovery as soon as possible.
S Mekruksavanich 23 developed a medical expert system to diagnose diabetes and used Web Ontology Language (OWL) 26 with nine different subclasses for analyzing the performance of the system on 65 patients. The method is applicable to multiple biosignals for diagnosing diabetes having an accuracy of 90.7%. Nevertheless, early and proper analysis is the prerequisite for managing diabetes mellitus. However, the system’s diagnosis process requires several input variables to be processed and they do not have an integrated system with the EHR system; therefore, it is often difficult to utilize such systems for health management in real time. Schriml and Mitraka 24 designed a disease ontology (DO) based on cross-domain clinical knowledge incorporation using the general type of illness concept. They claimed that DO is a universal ontology method which gathers the notion linked to several diseases such as diabetes mellitus. A major limitation of DO is that it provides an integrated ontology model of various illnesses, but, for the actual healthcare case, a model needs to be established for one specific disease. It is clear that different diseases have their own associated conditions and measurement parameters. Hence, an integrated model will influence its quality, capability, and efficiency. Thus, an independent ontology framework is significant for diagnosing diabetes. Moreover, in the literature a few scholars have concentrated on diabetes ontology within a specific domain, but they are merely inclined toward one or two aspects of diabetes mellitus. El-Sappagh et al. 25 proposed an encoding method for medical information by means of SNOMED CT (SCT) in which the case-based textual characteristics are transformed to SCT IDs for supporting the generation of semantic case reuse mechanism. However, their method does not include the concept of developing a complete ontology which can be useful for diabetes knowledge base.
The main limitation in existing research is the lack of a complete ontology model for diabetic patients. Furthermore, previous studies have not adopted a systematic approach to developing ontology. Also, their standards for top-level ontology are not globally recognized. Due to these drawbacks, such ontology does not provide a viable solution for diagnosing diabetes accurately and efficiently, as they do not incorporate the symptoms, drug information, and laboratory tests that may affect the doctors’ judgment. To address these problems, the contributions of this research are as follows: first, an ontology-based model for diabetic patients (OMDP) that is aimed at providing standard, consistent, comprehensive, and accurate means of representing diabetes is developed in this study; second, we elaborated how OMDP could be efficiently integrated with EHR to enable semantic interoperability for the mapping of patient’s medical information; third, we have implemented the proposed OMDP by applying OWL and Semantic Web Rule Language (SWRL) based on different classes related with diabetes to obtain a knowledge-based representation of diabetic patients; fourth, we analyzed and compared the performance of the proposed OMDP with previous ontology regarding reusability, extensibility, and completeness. Consequently, this study has developed an ontology-based prototypical model for diabetic patients, namely, OMDP, to resolve the ambiguity and semantic inconsistency associated with the existing conventional methods and provide support for clinical diagnosis by integrating diverse classes of diabetes knowledge for active diagnosis and treatment.
This research article is organized as follows: section “The proposed ontology model for diabetic patients” presents the OMDP structure and construction mechanism as well as ontology evaluation. Section “System implementation and experimental results” presents the implementation and results of the OMDP. The conclusion and future work are presented in section “Conclusion and future work.”
The proposed ontology model for diabetic patients
Structure of the proposed model
In this study, an ontology model, namely, OMDP is developed, which aims to provide risk prediction, screening, and treatment of diabetes mellitus. Figure 2 presents the architecture of the established model. This model includes a user interface, an EHR database, an OMDP SWRL rule base, an OMDP knowledge base, and an inference engine. Primarily, the user interface may be used to gather medical data without taking much of the physician’s time. Since the EHR database houses a huge amount of structured and unstructured medical data, there is always a need to adopt an appropriate preprocessing technique.

An ontology-based model for diabetes patients (OMDP) for medical applications.
Therefore, the missing data were handled by replacing them with their corresponding mean attribute value. Verbal attributes can be translated into standardized terminology using the leading natural language process. We used the test method proposed by Grubbs27,28 to analyze outliers. The fuzzy values were also handled in the preprocessing stage by categorization using different numerical ranges. 29 Then the preprocessed EHR database for accumulating the patient’s data that are obtained from the medical record and the user’s information from user inference is transformed into a semantic file. Our proposed model can impulsively gather an individual’s information from the database in the user semantic profile data. The OMDP rules and medical guidelines about diabetes for ontology development are given by the domain experts and the related literature as well as a clinical guideline for facilitating knowledge extraction. Moreover, medical information resulting from OMDP and basic rules is transferred to the interference engine to assist in providing appropriate treatment on the basis of the acquired knowledge. Meanwhile, OMDP is integrated with the semantic file which stores information about patients. And the final diagnosis result and new information about the patient is stored in the EHR database. The proposed OMDP aims to provide a warning to those high-risk patients who have a high chance of having diabetes. Besides, this system assists diabetes diagnosis and offers treatment guidelines to patients based on their diabetic level.
OMDP construction mechanism
This section includes the mechanism for constructing OMDP for healthcare applications in order to provide necessary guidance to diabetic patients. In this study, we have merged top-level ontology features with the proposed OMDP to support semantic interoperability for domain-specific ontology such as diabetes. A top-level ontology offers notions that do not depend on the domain including object, relation, and property. 30 The basic terms in the diabetes ontology are placed under the terms of the top-level ontology. In this study, we use basic formal ontology (BFO) and ontology for general medical sciences (OGMS) as the top-layer ontology. Based on top-level ontology concepts for constructing domain-specific ontology, it will overcome the chances of errors and provide a better understanding of how medical information is integrated into the proposed OMDP. Besides, providing a general top-level structure to the domain ontology experts via BFO and OGMS would help facilitate interoperability between our proposed models and the existing systems.
The OMDP construction has four stages including the OMDP preparation stage, OMDP class initialization stage, merging top-level ontology, and OMDP evaluation as shown in Figure 3. The OMDP preparation stage has three main parts such as ontology domain confirmation, detailed analysis of patients, and reuse of standard ontology. The OMDP class initialization stage is based on the hierarchy of OMDP and OMDP properties. The top-level ontology stage has two main sequential levels such as BFO and OGMS. In the final level, we have evaluated the proposed OMDP.
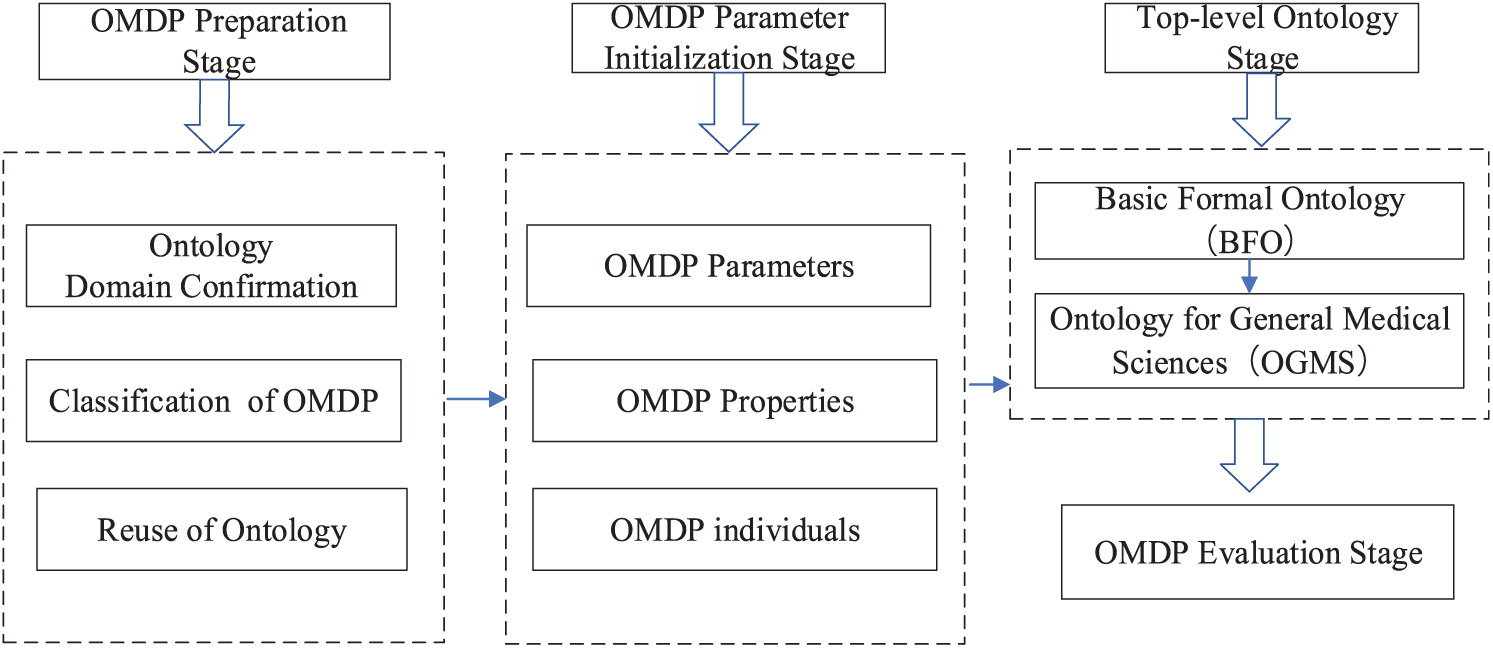
OMDP construction mechanism based on top-level ontology.
OMDP preparation
Ontology domain confirmation
One of the fundamental steps in developing an ontology for any application is to confirm its domain and scope. Generally, the domain knowledge is extensive. Hence, it is significant to validate the specific domain at an initial stage before building an ontology. The underlying reason for ontology domain confirmation is that the performance of different domains varies in distinct applications. Also, within the same application, the prominence of the ontology concepts fluctuates with a different goal. In this study, our domain for ontology modeling is associated with diabetic patients to solve the ambiguity and semantic inconsistency issues that are observed in traditional knowledge-based ontology methods.
Ontology classification
Considering a specific case of a patient, the patient’s health status with respect to diabetes is determined using the proposed OMDP model to obtain a set of health-related information to which a structured reasoning mechanism is applied. For example, patient’s clinical information including symptoms, complications, medical history, laboratory examination, physical examination, and genes is considered by the OMDP model for diagnosis of diabetic patients. Afterwards, our proposed OMDP model provides a possible treatment plan for the patient which could be in the form of drugs, diet as well as exercise for the patient. A use case diagram is used to illustrate the concept as shown in Figure 4.

Use case analysis of diabetes based on OMDP.
Reuse of ontology
In order to develop an ontology, it is a good option to adopt some features from the existing ontology rather than developing the model from scratch. The current healthcare research already comprises many interesting ontology models such as SCT which is widely applied in several medical applications. After reusing SCT, an inclusive model can be developed to facilitate computer processing, to provide convenience for clinical care, and to monitor data acquisition and coding. The subsequent step is to produce a mid-level ontology to analyze symptoms and signs of diabetes. We build disease classes based Human Disease Ontology; Drugs classes based RxNorm Ontology, symptom classes based on Symptom Ontology, Gene classes based on gene ontology and add physical ontology and food ontology. 31 After that, the mid-level ontology is connected with a top-level ontology to overcome the chances of errors by providing a better understanding of integration of medical information and OMDP as shown in Figure 5.

Description of mid-level and top-level ontologies for reuse ontology.
OMDP classes and properties confirmation
OMDP classes
In this section, we have defined classes utilized for OMDP as shown in Table 1. The first step is to know the medical condition of the patient through an electronic medical record (EMR), for example, medical history, allergy, and family history. In order to diagnose a patient, the foremost step is to understand the patients’ symptom from the lab test or physical examination. Patients have some symptoms when they are diabetic, for instance, polydipsia, polyphagia, and polyuria, among others. The most common type of diabetes is type 2 diabetes, whereas the other types are type 1 diabetes and gestational diabetes mellitus.
Description of OMDP classes.
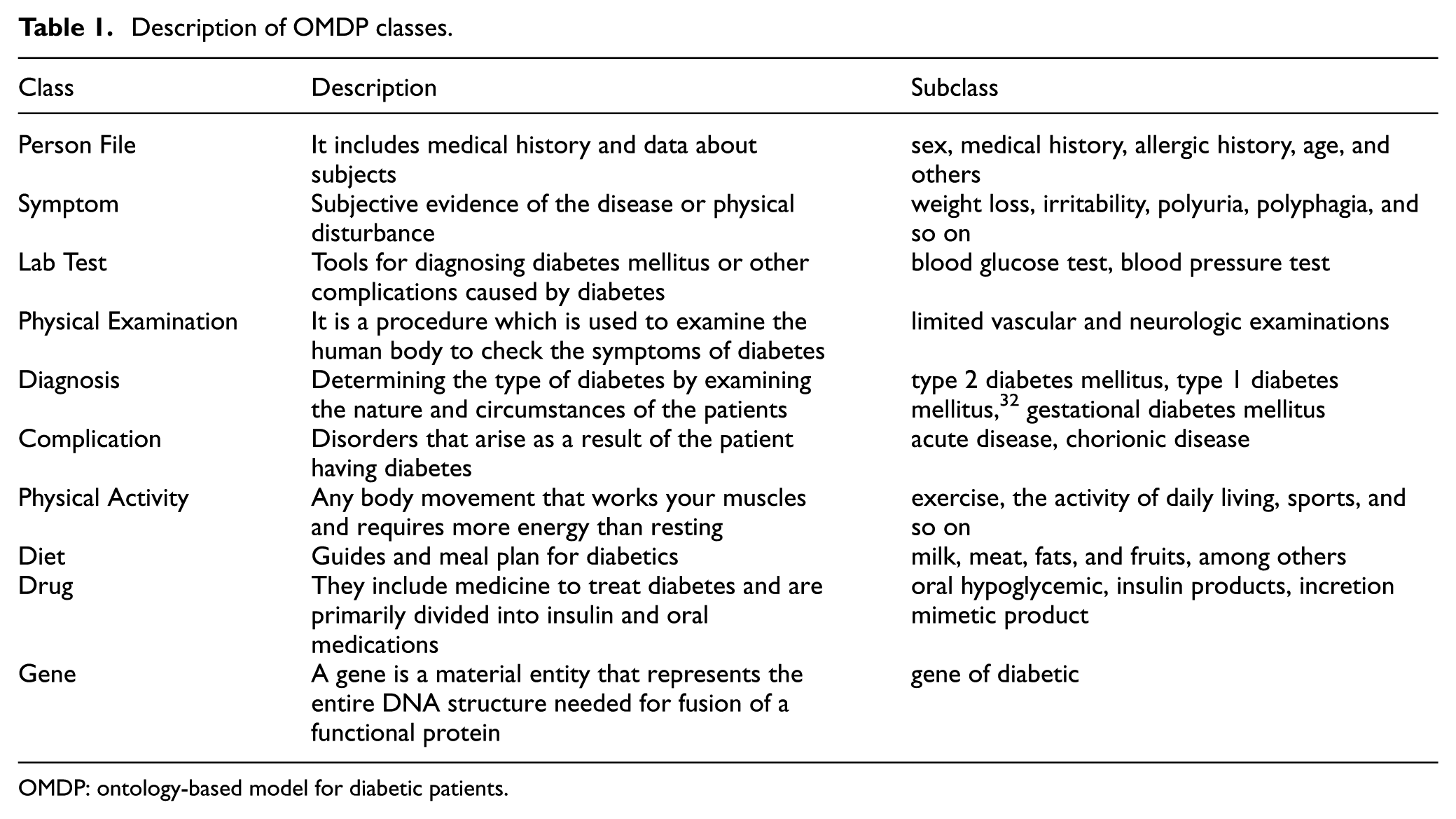
OMDP: ontology-based model for diabetic patients.
If
OMDP properties
Ontology properties include both object and data attributes. Object properties are used to define the objects, so it may be beneficial to provide the association between classes. For example, the history of diabetic patients can be used to specify the object property, has_history, and the ontology of the object properties is used to provide further explanation to individuals about diabetes. The list of OMDP objects is shown in Table 2.
OMDP object properties.

OMDP: ontology-based model for diabetic patients.
Data attributes are utilized to calculate the values, for example, the patient’s fasting plasma glucose (FPG) is 7.9, and we may apply the data attribute (has_quantitiave) to describe has_quantitiave (FPG, 7.9), as expressed in Table 3.
OMDP data properties.

OMDP: ontology-based model for diabetic patients.
OMDP instances
The data are usually elaborated in the form of instances for all software applications. In this section, we discuss all cases for OMDP. To suitably recognize the semantics of the data applied in OMDP, it is mandatory to understand the sharing and exchange of information between different entities. This study uses D2RQ as a mapping tool for connecting the medical data with OMDP. Different kinds of features were extracted to establish mapping operations among class names, attribute names, class instances, and so on. Also, classes were selected for mapping preparation in both source ontology and object ontology, respectively. Finally, we calculated the similarity score between the selected classes and features, measured the similarity between entities, and then optimized the results until satisfactory outputs were obtained. Finally, we use an EMR database for storing data of medical history, lab test, and other information about patients. This database is associated with OMDP as an instance.
Merging top-level ontology
A top-level ontology is an ontological structure which is used for upkeep of broad semantic interoperability by offering a general term between huge amounts of domain-specific ontology. In our study, two top-level ontologies, namely, BFO and OGMS, are merged in the proposed OMDP. Actually, it is a good practice to reuse terms from top-level ontology in another domain ontology. It has been found in the literature that several researchers apply this practice.33–35 For example, the term “entity” is taken from BFO and the term “symptom” from OGMS, which are reused to develop our OMDP model.
BFO
BFO includes three subclasses: entity, continuant, and occurrence, which serve as the top structure in the hierarchy of the developed OMDP. BFO supports the incorporation of medical information acquired using systematic research. It is intentionally miniaturized to facilitate representation in a consistent style from top-level ontology to domain ontology for numerous applications. Another reason is that it can fulfill the modularity condition that permits the division of knowledge so that a top-level ontology should suitably fit into a domain-specific ontology. 36 Furthermore, it differentiates the following types of independent entrants, boundaries of substances, fiat parts of substances, aggregates of substances, and cavities.37,38
OGMS
The main responsibility of OGMS is to symbolize the entities utilized in a clinical domain. It may consist of diseases and their symptoms and causes, as well as diagnostic procedures which are beneficial for recognizing and interpreting the disease for healthcare. It was developed to avoid conflations that often occur in medical discourse between entities. 39 A fundamental axiom of OGMS is that each disease is based on some physical reasoning. For instance, if blood glucose is higher for a few patients, it is due to the disordered nature of the physical structure in the organism, and there occurs a disposition for the organism to behave abnormally. This disposition is realized in pathological procedures comprising practices that may be identified as symptoms of the disorder and assistance can be provided by conducting lab tests or imaging processes.
Ontology evaluation stage
It is an important stage to develop OMDP that characterizes the validity and quality of ontology. 40 Ontology evaluation validates the consistency check of ontology and guarantees that the developed OMDP meets some defined criteria. It widely adopts the ontology of semantic web and semantically facilitated approaches.
In this study, the proposed ontology framework evaluation was performed in two main parts: one is to pay attention to check the competency and consistency of the explanation ontology and the second is to have an integrality comparison between existing diabetes ontologies and on the proposed OMDP. We used Pellet reasoner developed for an OWL DL to assess the consistency and classification of the ontologies. Moreover, we consulted medical experts in the domain of diabetes diagnosis and they certified that the content coverage of the proposed OMDP satisfies the competency requirements for the treatment of diabetic patients and realized that the proposed model would be potential in practical applications.
System implementation and experimental results
In this section, discussion about the implementation of OMDP is included. In this research, we used Protege editor and Pellet for developing the OMDP, and OWL 2 language is utilized to express OMDP. Furthermore, the performance of the proposed OMDP in terms of accuracy, reusability, and reliability in managing diabetic patients is also elaborated.
Language and tools for OMDP
Constructing ontology needs not only the support of experts in the knowledge domain and the suitable edifice method, but also the appropriate ontology expression language. The diabetes ontology is developed using OWL2. It is the latest version of OWL. The example in Figure 6 shows the relationship that exists between different classes.

Coding example in OWL.
Here, the blood glucose test is a subclass of the laboratory tests, while fasting and random blood glucose and glucose tolerance test are used as the examples of blood glucose tests. In this research, Protege is used to construct the OMDP; its working principle is shown in Figure 7.

Construction of the proposed OMDP using Protege software.
The knowledge base in a multifaceted domain needs to be arranged into an ontological hierarchy, in which each level delivers a specific description of the entity and a different set of realistic inferences.41,42 A hierarchy of OMDP is significant to offer the direct or indirect connection between objects of ontology (Figure 8). It facilitates organizing information in a human–machine acceptable method for reusing and maintaining the data. 43 In this research, the hierarchy of OMDP is classified by adding the basic information of patients and symptoms such as physical activity, exercise, and drugs to process the hierarchical structure of the standard ontology SCT and alignment technique.

A detailed hierarchy example of OMDP.
The implementation of SWRL rules
SWRL comprises a high-level abstract syntax for Horn-like rules in both the OWL DL and OWL Lite sub-languages of OWL. SWRL rule provided in our model mainly covers everything from diagnosis to treatment and can complete the entire process of screening, diagnosing, and treating diabetic patients. According to the symptoms and laboratory indexes of diabetes screening and the determination of parting, according to the specific condition of patients for an individualized treatment plan, including the exercise therapy, diet therapy, and drug therapy. Some pre-diabetes or no medication, the right diet, and exercise therapy can make your blood sugar return to normal. Since genes are not readily available in patients’ daily lives, genes are not taken into account in treatment recommendations. Examples of SWRL rules are given in Table 4.
Illustration of SWRL rules for OMDP.

SWRL: Semantic Web Rule Language; OMDP: ontology-based model for diabetic patients.
In this study, Pellet is used as the reasoner in OMDP having widespread support for constructing ontologies. First, we transformed the data stored in the relational database of patients by applying D2RQ into the Resource Description Framework (RDF) triple (subject–predicate–object) format. Second, the reasoning result is produced based on the SWRL rule that is applied on the preprocessed medical information of patients. Part of the reasoning result in Protege reasoning is shown in Figure 9.

Reasoning results based on OMDP.
Reusability measurement
Reusability has always been a defining characteristic of ontology. An ontology framework can be adapted and be used in the existing models for producing a new ontology, rather than developing a new ontology from scratch.44,45 In this study, we reused some standard ontology such as RxNorm ontology and SCT. Table 5 demonstrates the reusability evaluation of the developed OMDP with existing models. The results reveal that OMDP has a better capability to be used in existing systems than the compared approaches.
Reusability comparison between different ontology models.

AACEMG: American association of clinical endocrinologists medical guidelines for clinical practice for the management of diabetes mellitus; IRS-T2D: individualize recommendation system for Type2 diabetes medication; SWRL: Semantic Web Rule Language; OMDP: ontology-based model for diabetic patients.
Integrality analysis
Ontology-based diabetic assisted diagnostic systems exhibit far more performance than the traditional expert systems in terms of reusability, extensibility, and semantic interoperability. Although there are many auxiliary systems based on an ontology of diabetes, the contents are not complete. Several researchers are inclined toward a physical examination of glucose sugar to diagnosis of diabetes.46,49,50 However, various other critical factors are not completely overlooked which may be useful in diagnosing and treating diabetes such as the significance of subject information (diet or physical activity).47,51,52 Table 6 presents the integrality comparison of different ontology models for diabetic patients.
Integrality comparison of different ontology models.

OMDP: ontology-based model for diabetic patients.
Performance evaluation
In this section, we evaluate the performance of the proposed approach for different ontology models as demonstrated in Figure 10. Accuracy is a very critical factor to check the performance of ontology. The accuracy of the proposed model was assessed using the following performance metrics: true positives (TPs), false positives (FPs), false negatives (FNs), and true negatives (TNs) for classifying the extracted data. TP is the number of correctly classified individuals with diabetes, FN is the number of individuals without diabetes that are misclassified as having diabetes, TN denotes the number of individuals without diabetes that are correctly classified, and FP represents the number of patients without diabetes that are misclassified as having diabetes. The formula is shown in equation (1). The performance analysis for OMDP in terms of accuracy for disease prediction, diagnosing diabetes, and recommending medicine is 95%, 98%, and 85%, respectively
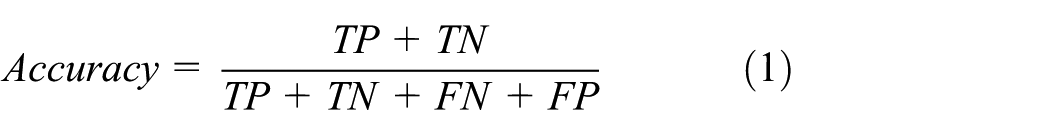
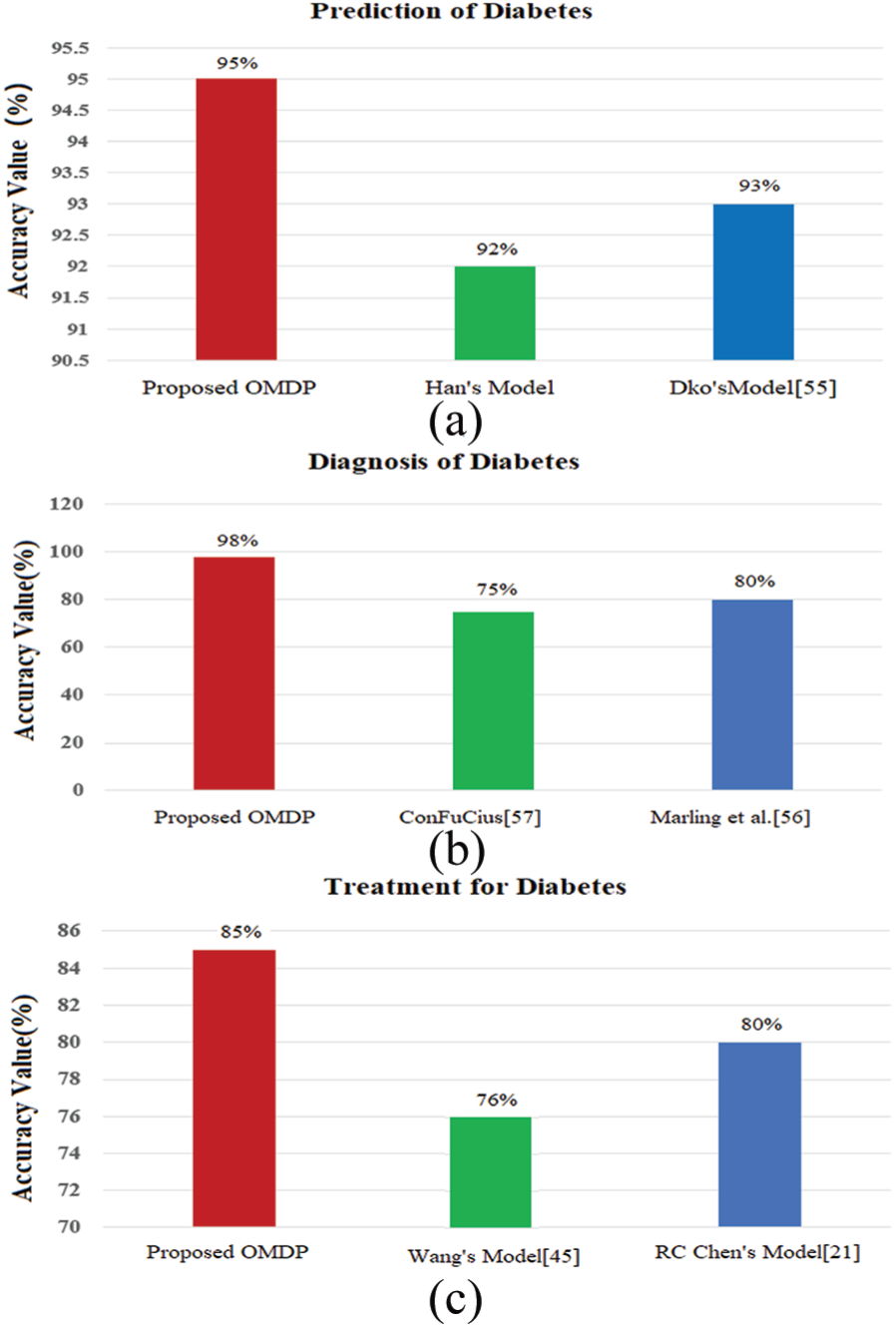
Performance evaluation in terms of accuracy for different ontology models: (a) prediction of diabetes; (b) diagnosis of diabetes; and (c) treatment for diabetes.
In this study, a total of 766 medical records were selected from the First Affiliated Hospital, Sun Yat-sen University. The records include medical history, sex, symptom, family history, and other information, and 269 of them were known to develop diabetes. The accuracy of the prediction of diabetes using the proposed OMDP is 95% for all the risk factors, that of the model developed by Han et al. is near 92%, and that of the model of diabetes knowledge–based ontologies (DKOs) 48 is 93%, as presented in Figure 10(a). The main reason due to which the proposed OMDP provides better results than the existing models is that it inputs many classes such as smoking, alcohol, physical activity, and sexual and cardiovascular diseases accounting for predicting diabetes. Once it is predicted that subjects have chances of diabetes, the next step is accurate diagnosis of diabetes.
The diagnosis of diabetes on the basis of different ontology models is shown in Figure 10(b). For the diagnosis, we consider factors and values specified in the clinical recommendation, mainly according to the value of fasting blood glucose. The experimental results show that our proposed OMDP has an accuracy of 98%. The proposed model outperforms in terms of diagnostic accuracy than the compared approaches including ConfuCius model (75%) and Marling et al.’s model (80%).
The developed OMDP model offers a higher diagnosis accuracy for patients because it is closely integrated with the detailed analysis of the patients, for example, medical history, family history, laboratory test, and physical examination. In the medical field, it is important for doctors to have more precise data and history of patients so the possibility to diagnose the diseases is increased. Existing researchers have overlooked the fundamental requirements of including detailed classes in their developed models, so their approaches have less accuracy for diagnosing diabetes.
The proposed OMDP not only provides better performance for predicting and diagnosing diabetes for individuals, but also has better accuracy for recommending useful treatment to patients than the compared models as demonstrated in Figure 10(c). To evaluate system efficiency for the recommendation, we consulted the hospital staff taking care of the diabetic patients to assess the correctness of the system to determine the retrieval efficiency. From Figure 10(c), it can be revealed that the treatment recommendation accuracy for the proposed model is 85%, while Wang and Chen’s models have the treatment recommendation accuracies of 76% and 80%, respectively. The main reason for having a better recommendation accuracy for our developed OMDP is that we combined the patient’s present situation, that is, medical history, and laboratory test in the proposed model. It will assist concerned physicians in recommending suitable exercises and an appropriate diet plan. Also, the proposed OMDP recommends patients on what medications they should take and which drugs are contraindications to them.
Along with accurate comparisons of the proposed OMDP with the related work systems, it is also necessary to compare the performance of our developed model in diagnosing diabetes regarding specificity and sensitivity with the existing approaches.
Specificity is the true-negative rate which measures the proportion of actual negatives that are correctly identified. In medical tests, specificity is the extent to which actual negatives are classified. The calculation method of specificity is shown in equation (2). From the experimental results, it can be revealed that the diagnosis specificity for the proposed model is 99.2%, while the models of A Rahimi et al. 53 and Yilmaz et al. 54 have the diagnosis specificity values of 99.18% and 95.06%, respectively (Figure 11)

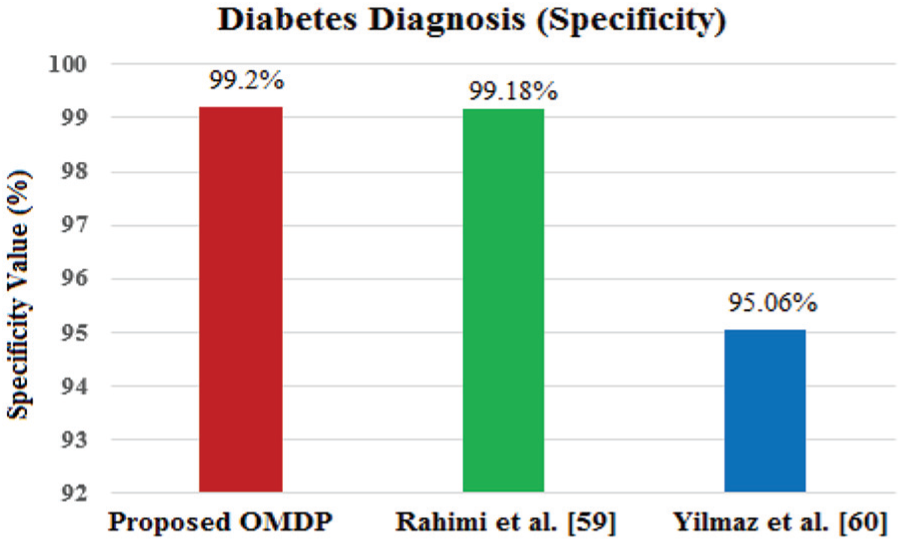
Performance evaluation in terms of specificity for different diabetes diagnosis models.
Sensitivity is the true-positive rate that measures the proportion of actual positives that are correctly identified, and it represents the extent to which actual positives are not overlooked. The calculation method of sensitivity is shown in equation (3). The results demonstrate that our proposed OMDP has a sensitivity of 98.2%. However, sensitivity values for the diabetes diagnosis models presented in A Rahimi et al. 53 and Yilmaz et al. 54 are 97.5% and 97.31%, respectively (Figure 12). Therefore, it can be stated that the proposed OMDP exhibits better performance from the aspect of sensitivity, specificity, and accuracy than the existing approaches


Performance evaluation in terms of sensitivity for different diabetes diagnosis models.
Conclusion and future work
In this study, we have developed an OMDP; it combines different kinds of diabetes knowledge for diagnosis and treatment of diabetes. The proposed OMDP solves the ambiguity and semantic inconsistency issues by performing a detailed analysis of patients in the system. To validate the performance of OMDP, it is implemented using OWL and SWRL rules in our study to provide an efficient knowledge-based model to aid doctors in reaching an efficient diagnostic decision for diabetic patients. From the experimental results, OMDP not only offers better performance for predicting and diagnosing diabetes in individuals, but also has better accuracy for recommending useful treatment to patients. Therefore, it can be concluded that the proposed OMDP has greater scientific significance to be applied in the real-time clinical environment. The main limitation of the proposed OMDP model is that it takes relatively little time to reach a decision and this is due to the fact that it adopts the inference engine with several rules for its reasoning. Therefore, the model may not be effective for patients with severe status that require urgent diagnosis/therapy.
The aim of our future work is to develop a more effective clinical auxiliary diagnosis system, which provides remote healthcare to diabetic patients. We also plan to apply the proposed ontology model on the Internet of Medical Things (IoMT)-based applications, since IoMT measures patient’s vital signs and aggregates the collected data into medical files, which are further uploaded to the cloud for storage and access by healthcare workers. Therefore, an IoMT-based ontology model will be more beneficial for real-time healthcare applications to improve accuracy and efficiency for not only hospitalized patients but also remote available subjects.
Footnotes
Handling Editor: Shinsuke Hara
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (81601575).
