Abstract
Motile bacteria often use sophisticated chemotaxis signaling systems to direct their movements. In general, bacterial chemotactic signal transduction pathways have three basic elements: (1) signal reception by bacterial chemoreceptors located on the membrane; (2) signal transduction to relay the signals from membrane receptors to the motor; and (3) signal adaptation to desensitize the initial signal input. The chemotaxis proteins involved in these signal transduction pathways have been identified and extensively studied, especially in the enterobacteria Escherichia coli and Salmonella enterica serovar typhimurium. Chemotaxis-guided bacterial movements enable bacteria to adapt better to their natural habitats via moving toward favorable conditions and away from hostile surroundings. A variety of oral microbes exhibits motility and chemotaxis, behaviors that may play important roles in bacterial survival and pathogenesis in the oral cavity.
(1) Preface
The first observation of bacterial movement was made by Antony van Leeuwenkoek in 1683. While examining dental plaque from the mouth of an old man, Leeuwenhoek found “an unbelievably great company of living animalcules, a-swimming more nimbly than any I had ever seen up to this time. The biggest sort...bent their body into curves in going forwards.... Moreover, the other animalcules were in such enormous numbers, that all the water...seemed to be alive....” (Fig. 1). Some of the bacteria described by van Leeuwenhoek are now recognized as the highly motile oral spirochetes whose corkscrew-like motility results in a very remarkable swimming behavior that will be addressed in more detail later.
(2) Bacterial Movement
Since van Leeuwenhoek’s first report on bacterial motility, the majority of bacterial species were found to be motile during at least a part of their life cycle (Fenchel, 2002). Bacterial motility can be categorized into flagellum-dependent motility and flagellum-independent motility. The flagellum is a bacterial motility apparatus that, in most motile species, can be observed on the cell surface as long filamentous cellular appendices (Macnab, 1996). The species variety within the bacterial kingdom is reflected by an impressive diversity in flagellation patterns that commonly serves as means of classification: Single or multiple flagella can be found at one cell pole (e.g., Helicobacter pylori, Pseudomonas sp., Vibrio sp., or Chromatium okenii), at both cell poles (e.g., spirochetes or Spirillum sp.), in the middle of a cell body (e.g., Rhodobacter spheroides or Selenomonas sp.), or all over the cell body (Escherichia coli, Salmonella sp., or Bacillus subtilis).
While, for the most part, bacterial motility is associated with the presence of at least one flagellum, some bacteria do translocate without the aid of flagella. The best-studied flagellum-independent types of motility include the gliding motility of myxobacteria, cyanobacteria, mycobacteria and the Cytophaga-Flavobacterium group, the twitching motility of Pseudomonas, Neisseria, and Synechocystis (McBride, 2001), and the swimming motility of Synechococcus (Ehlers et al., 1996).
Flagellum-dependent motility, which generates bacterial movement via rotation of the flagellar filaments in aquatic environments, has been extensively studied, revealing remarkable amounts of detailed structural and functional information (Macnab, 1996, 1999; Aldridge and Hughes, 2002). Much less is known about flagellum-independent motility, which is mainly involved in surface translocation. Recent studies suggest that several independent mechanisms are involved (McBride, 2001; Bardy et al., 2003). Substantial evidence has been generated supporting a model for twitching motility of Pseudomonas aeruginosa and Neisseria gonorrhoeae as well as for cell-group gliding motility of Myxococcus xanthus. In these species, movement is mediated through the extension, adherence, and retraction of type IV pili (Merz et al., 2000; Sun et al., 2000; Skerker and Berg, 2001; Li et al., 2003). It appears that the single-cell gliding motility of cyanobacteria and myxobacteria is associated with secretion of carbohydrate containing slimes (Hoiczyk and Baumeister, 1995, 1998; Wolgemuth et al., 2002). Surface translocation of the bacteria in the Cytophaga-Flavobacterium group may involve adherence of moving outer membrane components to the surface (Lapidus and Berg, 1982). However, various other motility models—including wave generation in the outer membrane and muscle-like expansion/contraction mechanisms—have been proposed (Duxbury et al., 1980; Burchard, 1984). The gliding motility of mycobacteria requires unique membrane proteins that modulate surface attachment/detachment by altering the hydrophobicity of the cell envelope (Recht et al., 2000), whereas the swimming motility of Synechococcus appears to rely on structures analogous to the cilia of eukaryotic organisms (Samuel et al., 2001).
(3) Bacterial Chemotaxis
More than two centuries after the detection of motile micro-organisms, Theodor Engelmann and Wilhelm Pfeffer found that bacterial movement was not random and arbitrary (Engelmann, 1883; Pfeffer, 1884). Instead, bacterial cells exhibited directed movement toward certain stimuli and away from others, a behavior they termed ‘chemotaxis’. The discovery of chemotaxis gave a completely new meaning to the ability of bacteria to move within their environment. With the recognition of chemotaxis-guided bacterial motility, it became apparent that bacteria can respond efficiently to environmental changes. Almost every motile bacterial species studied so far has been found to possess chemotactic abilities which enable them to adapt better to their natural habitats. For example, the tumor-inducing Agrobacterium tumefaciens enhances colonization by moving chemotactically toward injured plant roots guided by specific phenolics released by the wound sites (Hawes and Smith, 1989; Shaw, 1991), Rhizobium sp. are attracted by substances exuded by plant roots during nodulation (Ames et al., 1980; Parke et al., 1985; Armitage et al., 1988; Pandya et al., 1999), Halobacterium sp. use phototaxis that is triggered by light of the wavelengths required for photosynthesis (Schimz and Hildebrand, 1979; Sundberg et al., 1990; Krohs, 1994; Cercignani et al., 2000), and motile E. coli and Salmonella cells move toward optimal nutrients such as sugars, amino acids, and oxygen (Stock and Surette, 1996). In addition to responses to environmental signals, motility and chemotaxis have been found to be involved in virulence of many pathogenic bacteria (Freter, 1981; Ottemann and Miller, 1997; Lux et al., 2000; Josenhans and Suerbaum, 2002).
(4) Understanding the Molecular Mechanism of Chemotaxis
Since its discovery, chemotactic behavior has stimulated the curiosity of numerous investigators. However, it wasn’t until the groundbreaking work by Julius Adler—using genetic, biochemical, and behavioral approaches to analyze the chemotactic behavior of E. coli—that a detailed understanding of this phenomenon became possible (Adler, 1965, 1969, 1973, 1975; Armstrong et al., 1967; Armstrong and Adler, 1969a,b). The intensive research efforts by numerous laboratories in the ensuing 40 years have yielded a comprehensive understanding of the molecular mechanism of bacterial chemotaxis. In general, the bacterial chemotactic signal transduction pathway has three basic elements (Fig. 2): (1) signal reception by bacterial chemoreceptors located on the membrane; (2) signal transduction to relay the signals from membrane receptors to the motor; and (3) signal adaptation to desensitize the initial signal input.
(4.1) Signal reception by membrane receptors
Bacterial chemoreceptors are membrane proteins that can be modified by methylation at specific glutamate and glutamine residues, and are therefore also called methyl-accepting chemotaxis proteins (MCPs) (Kort et al., 1975). MCPs are very sensitive detection devices that can perceive the extracellular concentrations of specific ligands at low concentrations in the μM to nM range (Clarke and Koshland, 1979; Biemann and Koshland, 1994; Lin et al., 1994). Most chemoreceptors bind only a small number of specific ligands but do not transport them. The occupancy with ligand is constantly monitored by the cell and transformed into appropriate motor responses. Lack of a specific MCP does not affect the cell’s general ability to perform chemotaxis but only abolishes the response to the chemo-effectors perceived by the missing MCP (Adler, 1969).
Recently, it has become evident that bacterial chemoreceptors are not isolated entities but rather function as ‘signaling lattices’ that facilitate ‘teamworking’ of various receptor types for stronger signal amplification (Ames et al., 2002; Gestwicki and Kiessling, 2002). The individual receptor types form very stable homodimers that are organized in higher-order structures in groups of three (trimers of dimers) (Kim et al., 1999). These ‘trimers of dimers’ tend to cluster in large patches at the cell poles in a variety of bacteria (Alley et al., 1992; Maddock and Shapiro, 1993; Harrison et al., 1999; Gestwicki et al., 2000). This higher-order clustering is thought to have an important biological role in signal amplification, since the clustered chemoreceptors can sense very small changes in stimulus concentration (1-10%) that would otherwise not be detected (Mesibov et al., 1973; Jasuja et al., 1999; Kim et al., 2001). The clustering and polar localization of the chemoreceptor that facilitates sensitive responses to small environmental changes is highly conserved among bacteria. Phylogenetically very distant species like E. coli, B. subtilis, or Spirocheta aurantia exhibit very similar patterns (Lamanna et al., 2002).
A ‘classic’ chemoreceptor is a membrane-spanning protein that can be divided into three major parts: an ‘extra-cytoplasmic’ ligand-binding domain and a transmembrane segment that connects to a cytoplasmic portion consisting of the signaling and methyl-accepting domains. The MCPs of E. coli and S. enterica serovar typhimurium have been intensively studied by genetic, biochemical, and structural analysis techniques for several decades, and detailed structural and functional information has been revealed. Structural analysis uncovered a mainly helical composition of the MCPs. X-ray crystallography showed that the periplasmic ligand-binding or sensory domain of the enteric chemoreceptors consists of a four-helix bundle (Fig. 3) (Milburn et al., 1991; Yeh et al., 1993). Two of these helices extend into the cytoplasmic membrane as the transmembrane portion. Ligand-binding surfaces have been identified via genetic mapping of the residues involved (Kossmann et al., 1988; Wolff and Parkinson, 1988; Mowbray and Koshland, 1990; Gardina et al., 1992; Scott et al., 1993) and co-crystalization of the ligand with the ligand-binding domain (Milburn et al., 1991). The structural changes that are induced upon ligand-binding to the sensory domain of the receptor and transmitted via the transmembrane segment to the cytoplasmic domain have been analyzed in great detail by solid-state nuclear magnetic resonance (NMR) and spin-labeling electron paramagnetic resonance (EPR) techniques (Ottemann et al., 1998).
Recently, deviations from the ‘standard’ ligand-binding domain have been discovered in a variety of bacterial species: Caulobacter crescentus, R. spheroides, the archeon Halobacterium salinarium, B. subtilis, and possibly others that contain ‘soluble’ receptors missing the transmembrane part (Harrison et al., 1999; Larsen et al., 1999; Hou et al., 2000, 2001; Potocka et al., 2002; Wadhams et al., 2002). These receptors are membrane-associated or found in the cytoplasm rather than integrated into the membrane. Interestingly, similar to myoglobin, the soluble chemoreceptors of H. salinarium and B. subtilis carry heme-groups in their sensory domains that allow for oxygen-binding and are involved in aerotactic signaling.
The membrane-proximal part of the cytoplasmic domain, the so-called ‘linker region’, which consists of an extension of one of the transmembrane helices, contains a motif that appears to be highly conserved among signaling proteins in bacteria as well as in archaea (Aravind and Ponting, 1999; Galperin et al., 2001). This domain is a member of a newly emerging family of signaling domains commonly found in various signaling pathways, the HAMP domain (Aravind and Ponting, 1999). A possible function of this domain may involve dimer formation. The remaining part of the cytoplasmic domain is composed of two helices carrying conserved glutamine and glutamate residues that are required for the ability to adapt to a given chemo-effector concentration, a process that will be explained in section (4.3). The membrane-distal portion of the cytoplasmic chemoreceptor domain can bind the main chemotaxis proteins CheA/CheW that relay the signal to the motor via the phosphotransfer reactions detailed below.
(4.2) Signal relay from membrane receptors to the motor
The environmental signal perceived by the chemoreceptors triggers an excitation response that is relayed to the motor via a cognate pair of bacterial sensory proteins, CheA (a histidine kinase) and CheY (a response regulator). Both proteins are representative members of the two-component-system super-family which executes most signal transduction events in bacteria (Stock et al., 2000; West and Stock, 2001). The histidine kinase CheA forms a tight ternary complex with the cytoplasmic signaling domain of MCP in conjunction with the coupling factor CheW (Gegner et al., 1992; Boukhvalova et al., 2002). CheA has been studied extensively, and many functional aspects are now well-understood (Stock and Surette, 1996; Stock et al., 2000). In contrast, the exact function of CheW still remains largely unknown, despite detailed genetic and structural analyses (Conley et al., 1989; Gegner and Dahlquist, 1991; Gegner et al., 1992; Griswold and Dahlquist, 2002; Griswold et al., 2002).
CheA autophosphorylates at a highly conserved histidine residue with ATP as a substrate, after which the phosphoryl group is transferred from CheA to a conserved aspartate residue of its cognate response regulator CheY (Hess et al., 1988; Borkovich et al., 1989). In its phopshorylated state, CheY can bind to the motor and reverse the direction of flagella rotation or modulate the speed of flagella rotation, depending on the type of flagellated motility used by the individual species (Welch et al., 1993). Another cognate response regulator for CheA is the methylesterase/amidase CheB that is required for adaptation (Hess et al., 1988), which will be discussed later.
As mentioned earlier, conformational changes induced by ligand-binding to the sensory domain of MCP are transmitted to the signaling domain and result in subsequent structural changes that control CheA autophosphorylation activity. In many species, the presence of a repellent (negative chemo-effectors) greatly stimulates CheA autophosphorylation, whereas attractant (positive chemo-effectors) binding to the chemoreceptor results in an almost complete shutdown of autokinase activity (Borkovich et al., 1989; Simon et al., 1989; Ninfa et al., 1991).
Similar to the chemoreceptors, CheA of the model organisms E. coli and S. enterica serovar typhimurium has been extensively characterized for more than two decades by genetic, biochemical, and structural approaches. CheA is a unique histidine kinase (Dutta et al., 1999) and can be divided into five distinct functional domains (Fig. 4): the N-terminal phosphotransfer domain (HPt), the YB or S domain for response regulator recognition, a separate dimerization domain (unlike other histidine kinases), a catalytic kinase domain (CA), and the C-terminal receptor/CheW binding domain (R). Like all histidine kinases, CheA functions as a dimer, with the kinase domain of one subunit phosphorylating the conserved histidine of the other and vice versa (Schuster et al., 1993; Swanson et al., 1993; Wolfe and Stewart, 1993).
The next interacting partner in the signal transduction chain of the excitation response is CheY, one of the cognate response regulators for CheA. CheY is an unusual response regulator, since it consists only of the phosphorylatable response regulator domain but is lacking the DNA-binding domain or any other catalytic subunit that is characteristic of the majority of response regulators (Stock et al., 2000). CheY directly interacts with the switch protein, FliM, of the flagellar motor and induces alterations in swimming behavior (Roman et al., 1992; Sockett et al., 1992). Upon CheY binding to the YB domain of CheA, the phosphoryl group is transferred from the conserved histidine to the conserved aspartate of CheY (Hess et al., 1988; Wylie et al., 1988). Phosphorylation of CheY results in a conformational change that decreases its affinity for CheA (Schuster et al., 1993; Swanson et al., 1993) but increases its affinity for the motor protein FliM (Welch et al., 1993). The release of phosphorylated CheY from CheA and subsequent binding to the motor switch is an important part in the signal transduction process.
Ultimately, the environmental signals perceived by the chemoreceptors are relayed to a motor to enable the organism to perform displacements corresponding to the type of stimulus perceived. The flagellar motor of the enteric bacteria E. coli and S. enterica serovar typhimurium has been studied in great detail for several decades (Fig. 5) (Macnab, 1996, 1999). As mentioned earlier, flagella are thin helical cellular appendices that can be localized as a single structure or as a bundle at one or both cell poles, evenly distributed over the cell surface, or even reside in the periplasmic space (Eisenbach, 2000). Typically, the long helical flagellum is connected to the membrane-integral basal body that extends to a bell-shaped structure into the cytoplasm via a curved hook (Macnab, 1996, 1999). This basal body, in combination with a ring of transmembrane stators formed by the MotA/B complexes (Khan et al., 1988), is thought to be the rotary device that powers flagella rotation via H+ or Na+ gradients (Larsen et al., 1974; Manson et al., 1977; Shioi et al., 1978; Goulbourne and Greenberg, 1980; Imae et al., 1986; Kawagishi et al., 1995). Many flagellar motors are reversible rotary devices whose direction of rotation is controlled by the intracellular ratio of CheY to CheY~P. Some motors, however, are unidirectional, and CheY~P binding modulates the speed rather than the direction of rotation (Gotz and Schmitt, 1987; Packer and Armitage, 1993; Platzer et al., 1997). In all these different systems, the ratio of CheY to CheY~P is modulated by the MCP/CheW/CheA/CheY signaling complex in accordance with the input signal (attractants or repellents). Depending on the input signal, MCPs will relay positive or negative chemotactic signals, thus decreasing or increasing CheY~P levels.
The flagella motor is encoded by more than 50 genes that are expressed following a strictly controlled hierarchy to ensure proper assembly of this complex structure (Macnab, 1996; Kalir et al., 2001). In E. coli and S. enterica serovar typhimurium, a master operon on top of this hierarchy is absolutely necessary for expression of the remainder of the flagellum-related genes that are organized into 15 and 17 operons, respectively (Komeda et al., 1980; Kutsukake et al., 1988).
(4.3) Sensory adaptation and desensitization
Sensory adaptation (or desensitization) describes the ability of an organism to re-set and desensitize its signal transduction system to the original stimulus. This important feature allows the organism to respond to higher concentrations of the same chemo-effectors or other types of chemo-effectors. Adaptation is a time-delayed secondary mechanism that sets in after the excitation response is accomplished. The ‘excited’ chemoreceptors are desensitized via reversible methylation events of their cytoplasmic signaling domain (Springer et al., 1979). Binding of an attractant results in increased methylation levels, whereas demethylation occurs when the ligand is a repellent. These actions are performed by two enzymes: CheR, a methyltransferase (Springer and Koshland, 1977; Simms et al., 1987; Djordjevic and Stock, 1997); and CheB, which exhibits deamidase as well as methylesterase activity (Stock and Koshland, 1978; Djordjevic et al., 1998) (Fig. 2). Substrates for these enzymes are conserved glutamate and glutamine residues within the signaling domain of MCP that regulates CheA activity (Stock and Surette, 1996). The glutamine residues require deamidation (to glutamate) by CheB to become a substrate for methylation by CheR in a S-adenosyl-methionine-dependent reaction (Rollins and Dahlquist, 1981; Sherris and Parkinson, 1981). Upon methylation of these sites, the signaling domain of MCP activates CheA autokinase activity similarly to the structural changes induced by repellent binding to the ligand-binding domain of the chemoreceptor (Ninfa et al., 1991; Borkovich et al., 1992). Demethylation by the CheR-antagonist, CheB (Simms et al., 1987), has the opposite effect.
These chemo-effector-specific shifts in chemoreceptor methylation levels are achieved by the regulatory and catalytic properties of CheR and CheB: The methyltransferase CheR methylates the chemoreceptor cytoplasmic domain with a constant slow turnover rate of about 6 min-1 and does not appear to be regulated (Simms et al., 1987). The methylesterase activity of the catalytic domain of CheB, in contrast, is strictly controlled by its regulatory domain (Hess et al., 1988; Lupas and Stock, 1989; Stewart et al., 1990). As mentioned earlier, similar to CheY, CheB constitutes a cognate response regulator for CheA. The methylesterase activity of CheB is regulated according to the autophosphorylation levels of its histidine kinase (Hess et al., 1988; Lupas and Stock, 1989; Stewart et al., 1990) and exhibits a strong intrinsic autophosphatase activity (Lukat et al., 1991; Stewart, 1993). CheB activity increases over 10-fold upon phosphorylation with a basal level of activity in the unphosphorylated protein below the CheR methyltransferase rate (Hess et al., 1988; Lupas and Stock, 1989; Stewart et al., 1990). Both CheB and CheR were found to be intimately associated with the ternary signaling complex (MCP, CheA, and CheW) (Wu et al., 1996; Djordjevic and Stock, 1998b), and their respective structures have been solved (Djordjevic and Stock, 1997, 1998a,b; Djordjevic et al., 1998). In an unstimulated cell, both enzymes maintain an equilibrium of methylation/demethylation.
In brief, adaptation to a chemo-effector (e.g., an attractant) involves the following scenario: During the excitation response (signal reception and signal transduction), the binding of attractant molecules to the MCPs abolishes the autokinase activity of CheA completely, triggering a smooth swimming response of the cell as a result of decreasing CheY~P levels. Simultaneously, phosphorylation levels of CheB, the other cognate response regulator of CheA, decrease as well, resulting in low CheB methylesterase activity. In the meantime, CheR, whose activity appears not to be regulated, keeps adding methyl groups to the cytoplasmic portion of the MCP. This combination of reduced CheB activity and constant CheR-mediated methylation leads to a shift of the methylation equilibrium in the chemoreceptors of an unstimulated cell toward the methylated state. Chemoreceptor methylation overrides the structural changes in the signaling domain that originally caused the inhibition of CheA activity. As a result, CheA autophosphorylation increases to pre-stimulation levels. The phosphorylation levels of CheY and CheB rise as well, and the cell eventually balances its behavior to an unstimulated random walk, even in the presence of an attractant. Adaptation to repellents involves opposite changes in activity and chemoreceptor methylation levels.
In some bacterial species, including the enteric bacteria E. coli and S. enterica serovar typhimurium, adaptation is enhanced by a CheY-specific phosphatase, CheZ (Hess et al., 1988). However, most bacterial species do not contain this protein and have evolved other strategies, such as using additional CheY-homologues as a phosphate sink (see section 4.4) (Armitage and Schmitt, 1997) or requiring only the action of CheB and CheR for adequate adaptation.
In summary, during chemotaxis of E. coli and S. enterica serovar typhimurium (the enterobacterial paradigm), environmental gradients are perceived by (typically transmembrane) chemoreceptors that form a complex with the protein kinase CheA via the coupling protein CheW. CheA autophosphorylates itself at a histidine residue with ATP as a substrate and is able to transfer this phosphor group to a highly conserved aspartate in CheY. CheY is a small protein that can freely diffuse in the cell, binds to a motor switch upon phosphorylation, and modulates the bacterial swimming behavior according to the stimulus detected. The phosphorylation level of CheY ultimately determines the motor response. Attractant binding suppresses CheA autophosporylation, resulting in a smooth swimming behavior, whereas repellents greatly stimulate autophosphorylation of this protein, and the subsequent rise in intracellular CheY~P levels causes the cells to tumble. Adaptation to such stimuli involves methylation and demethylation events that are carried out by the methylase CheR and the methylesterase CheB, another cognate response regulator for CheA.
(4.4) Variations of the enterobacterial paradigm
The chemotaxis in E. coli and S. enterica serovar typhimurium has been intensely investigated for several decades, providing very detailed understanding of many of the underlying mechanisms. Since these two species have been the focus of chemotaxis research for so long, they eventually became the ‘paradigm’ for bacterial chemotaxis. Initially, other bacterial species—such as B. subtilis or R. spheroides—that were coming into play but would not ‘follow the rules’ were regarded somewhat as oddballs in the swimming world. With more and more different bacterial species studied outside of the enteric world, and especially with genome information becoming available at an almost exponential rate over the past couple of years, it has become apparent that even though the basic mechanisms outlined earlier are highly conserved, the variations on the theme are manifold and could be the rule rather than the exception.
Unique chemotaxis proteins are found in the spirochetes that contain some members of the oral community in the form of oral treponemes. The chemotaxis operons of spirochetes contain a novel gene cheX in addition to cheA and cheY, as well as an interesting gene fusion between cheW and cheR (Greene and Stamm, 1999). CheX is required for chemotaxis, since cheX gene inactivation mutants fail to respond to chemo-effectors (Lux et al., unpublished results). A possible role in controlling cellular reversal has been implied. Homologues of cheX can be found in other bacterial genomes, but they are not localized in chemotaxis operons. Since functional studies of this protein in bacteria other than the spirochetes Treponema denticola and Borrelia burgdorferi are still lacking, its general function in other species remains unclear.
So far, the Gram-positive soil bacterium B. subtilis seems to contain the most novel or unusual chemotaxis proteins besides the chemoreceptors and the proteins CheA, CheW, and CheY, that form the backbone of the general chemotactic signal transduction pathway. Interestingly, in this bacterium, the default rotation of its motor causes tumbling rather than the smooth swimming behavior observed in many other bacterial species (Garrity and Ordal, 1995). Chemotaxis genes that are required for proper function of the chemotaxis pathway in B. subtilis but do not have counterparts in the E. coli genome are comprised of cheC, cheD, and cheV. All three proteins (CheC, CheD, and CheV) appear to be involved in particular aspects of adaptation to specific attractant stimuli (Rosario and Ordal, 1996; Zimmer et al., 2000; Karatan et al., 2001; Kirby et al., 2001; Kristich and Ordal, 2002) and are needed in addition to CheB and CheR, which are known to mediate adaptation (Kirsch et al., 1993a,b). CheC has been found to interact with McpB and CheA of the ternary signaling complex, CheD, and the motor protein FliY. CheC action appears to be involved in the regulation of chemoreceptor methylation in combination with CheD by an unknown mechanism. This protein possibly shuttles between the input (ternary complex of MCP, CheA, and CheW) and output (motor) domains of the excitation pathway to relay adaptation events. Despite extensive genetic analysis that revealed an array of distinct phenotypes, no specific functional mechanism has been found so far for this protein; therefore, the exact role of CheC in adaptation remains speculative. In addition to its role in receptor methylation, CheD is thought to facilitate CheA autophosphorylation, even though a direct interaction between these two proteins was not observed (Kristich and Ordal, 2002). CheV exhibits strong homology to CheW at its N-terminus and contains a response-regulator domain at the C-terminal portion (Rosario et al., 1994). This protein was recently found to be required for adaptation to specific amino acids and appears to be activated upon phosphorylation by CheA, thus constituting a third cognate response regulator for this autokinase (Karatan et al., 2001). Mutant strains defective in CheV phosphorylation or lacking the protein still showed normal excitation responses but were missing adaptation to certain stimuli.
Another well-studied example of ‘variation of the enterobacterial paradigm’ is the soil bacterium, Sinorhizobium meliloti. Unlike B. subtilis or the spirochetes, this member of the α-subclass of proteobacteria does not contain any novel chemotaxis proteins, but does contain the same linear excitation pathway known for E. coli and S. enterica serovar typhimurium. S. meliloti lacks the phosphatase CheZ but features a second CheY homologue (Greck et al., 1995; Sourjik and Schmitt, 1996). This CheY homologue is not involved in the excitation pathway but serves as a phosphate sink, thus substituting for CheZ function (Sourjik and Schmitt, 1998). Interestingly, many motile members of the same subclass of proteobacteria—including R. spheroides, C. crescentus, and A. tumefaciens—appear to feature a similar ‘phosphate sink’ mechanism, since they encode homologues of this cheY pair while lacking cheZ (Armitage and Schmitt, 1997). In addition to CheY homologues that may serve as a phosphate sink rather than transducing the excitation response, R. spheroides contains multiple complete chemotaxis pathways, including 13 chemoreceptors, four CheW, four CheA, six CheY, two CheB, and three cheR homologues, most of which are essential for normal chemotaxis responses (Martin et al., 2001). Recent evidence suggests that these pathways are not independent of each other, but instead, are interconnected in a very complex signaling network whose detailed interactions remain to be elucidated (Porter and Armitage, 2002).
(5) The Role of Motility and Chemotaxis in Microbial Pathogenesis
Tested with in vivo pathogenesis model systems, many motile pathogens were found to require their flagella and, in many cases, normal motility for efficient infection (Ottemann and Miller, 1997; Josenhans and Suerbaum, 2002). Interestingly, a variety of these organisms also exhibit a striking co-regulation of motility/chemotaxis genes and virulence factors, underscoring the in vivo significance of (directed) movement during various stages of the infectious process (Akerley et al., 1992, 1995; Givaudan and Lanois, 2000; Hay et al., 1997; Lee et al., 2001; Sperandio et al., 2001; Krukonis and DiRita, 2003; Liaw et al., 2003; Xu et al., 2003). More than two decades ago, chemotaxis as the navigation system for motility had already been contemplated to be an important feature for the virulence of many motile pathogens (Freter, 1981; Freter et al., 1981a,b; Freter and O’Brien, 1981a,b). Until very recently, however, little attention has generally been paid to this interesting idea compared with that devoted to the emerging studies on the involvement of motility in pathogenesis.
For important pathogens such as H. pylori (the primary causative agent of chronic gastric diseases) or Campylobacter jejuni (one of the major factors in food-borne diseases), the significance of chemotaxis for pathogenesis in addition to motility has been established (Takata et al., 1992; Yao et al., 1997; Foynes et al., 2000). Mutant strains lacking the central proteins of the chemotaxis pathway, CheA or CheY, failed to colonize the gastric/intestinal mucus layer of their host successfully in vivo and establish infection. Recently, two chemoreceptors were found to promote H. pylori-induced stomach infection (Nakazawa, 2002). Both H. pylori (Foynes et al., 2000) and C. jejuni (Hugdahl et al., 1988) are known to perform chemotaxis toward components present in mucin, which could play a role in identifying suitable colonization sites (Nakazawa, 2002). Similarly, chemotactic motility toward mucin was shown to be indispensable for the fish pathogen Vibrio anguillarum to initiate infection in its host (O’Toole et al., 1996, 1999). Other mucus chemotaxis-associated infections with V. cholerae and Brachyspira (Serpulina) hyodysenteriae have also been reported (Freter et al., 1981b; Freter and O’Brien, 1981b; Kennedy and Yancey, 1996). Recent studies in V. cholerae revealed that even though mutants defective in chemotaxis were more successful in colonization than the wild-type parent, the colonization pattern was erratic and spread throughout most of the gastro-intestinal tract, whereas the wild-type settles in specific parts of the small intestines (Lee et al., 2001). The same group also found that chemotaxis genes are the main regulators of cholera toxin production in vivo (Merrell et al., 2002); mutants lacking chemotaxis genes do not produce the toxins that trigger the disease. Interestingly, V. cholerae, which is shed from the human host, was found to suppress chemotaxis gene expression. Based on the above findings and earlier data by Freter and O’Brien (1981b), these authors then proposed a model whereby chemotaxis might be important in the early stages of infection to guide the bacterium to a suitable niche within its host and induce optimal virulence gene expression. Later, chemotaxis gene expression is down-regulated to produce motile but non-chemotactic cells that can easily be shed from the host and re-enter the infectious cycle.
The most controversial observations on the significance of motility and chemotaxis for virulence have been reported for the enterobacterium S. enterica serotype typhimurium. ‘Salmonellosis’ is one of the most widespread gastro-intestinal infections and the second most common food-borne illness. Some authors claimed that the presence of flagella was not necessary for virulence (Lockman and Curtiss, 1990), or that flagella served only as an anchor for attachment but not as means of locomotion during the pathogenic events (Carsiotis et al., 1984). Most investigators, though, acknowledged the importance of flagellar motility for in vivo infection and recognized that motility did enhance the invasion process in in vitro animal model systems. Chemotaxis, however, was either not addressed in these studies (Betts and Finlay, 1992; Lee et al., 1996; Methner and Barrow, 1997) or was found to be unnecessary (Carsiotis et al., 1984). Some chemotaxis mutants that result in constantly smooth-swimming phenotypes were more invasive than wild-type mutants, whereas constantly tumbling derivatives exhibited a pronounced decrease in tissue invasiveness, suggesting that the physical orientation of the flagella is important for the invasion process (Jones et al., 1992). Most recently, one group provided evidence that both motile and chemotaxis mutants failed to colonize successfully in an animal model (Lovell and Barrow, 1999). Additionally, co-regulation of motility and virulence genes has been described (Lucas et al., 2000).
P. aeruginosa is a bacterium that exhibits flagellated as well as type IV pilus-dependent motility. It is an important opportunistic pathogen that causes chronic pulmonary infections in about 70% of patients with cystic fibrosis and in immuno-compromised hosts. Colonization of P. aeruginosa on the mucus surface requires flagella and type IV pili. Chemotaxis of P. aeruginosa toward mucin has been found (Nelson et al., 1990) and was shown to be involved in the initial stages of infection (Feldman et al., 1998).
In addition to the above-mentioned gastro-intestinal pathogen B. hyodysenteriae that causes infections via intimate attachment to mucosal surfaces, many spirochetes are highly virulent or at least opportunistically pathogenic (Lux et al., 2000). Direct in vivo evidence demonstrating the involvement of chemotaxis in the pathogenic process of these bacteria is still lacking, but motility has been shown to be a crucial virulence factor in vivo and in vitro (Sadziene et al., 1991; Lux et al., 2001). Most of these pathogenic spirochetes were also found to exhibit strong chemotaxis responses toward substances that they would encounter during infection of their respective hosts: B. burgdorferi, the causative agent for Lyme disease, has a life cycle involving persistence in the gut of ticks and entry into the bloodstream of their prospective mammalian hosts via the salivary glands of the tick. A study showed that the bacterium perceives salivary gland extract and serum as attractants (Shih et al., 2002). Other examples are that Leptospira interrogans performs chemotaxis toward hemoglobin, and that the oral spirochete T. denticola performs chemotaxis to serum and a variety of sugars and amino acids (Mayo et al., 1990; Yuri et al., 1993; Kataoka et al., 1997; Li et al., 1999) and requires motility and chemotaxis for effective tissue penetration in vitro (Lux et al., 2001).
Bacteria that use chemotaxis and motility in their pathogenic processes also include some important plant pathogens such as A. tumefaciens or P. fluorescens (Hawes and Smith, 1989; Shaw, 1991; Singh and Arora, 2001; de Weert et al., 2002). Symbiotic events between Rhizobium sp. or Azospirillum brasilense and their respective hosts rely equally on motility and chemotaxis (Schmidt, 1979; Gulash et al., 1984; Parke et al., 1985; Caetano-Anolles et al., 1988; Bakanchikova et al., 1989; Vande Broek et al., 1998; Pandya et al., 1999; ).
(6) Motility and Chemotaxis in Oral Bacteria and Their Possible Roles in Pathogenesis
The most striking type of motility that can be observed in bacterial plaque samples is without a doubt the corkscrew-like movement exhibited by the oral spirochetes. More than 50 different species of these interesting micro-organisms have been identified in the oral cavity, and all were found to be members of the genus Treponema (Paster et al., 2002), although most of them cannot be cultivated in vitro for further analyses. The flagellated motility of spirochetes is unique, since the flagella do not extend as cellular appendices from their very long and thin cell bodies but rotate within the periplasmic space. Similar to the sheathed flagella of H. pylori (Geis et al., 1989), this interesting feature enables rapid motility to occur in very viscous gel-like environments (Charon et al., 1992b; Li et al., 2000) where rotation of unprotected external flagella is greatly impaired (Schneider and Doetsch, 1974). For example, T. denticola, the model oral spirochete, increases its swimming velocities from less than 5 μm/sec to up to 19 μm/sec in response to increased viscosity and temperature (Ruby and Charon, 1998). Earlier measurement by other investigators demonstrated a significant increase of translocational speed under higher viscosity conditions, but the velocity reported was in the range of 19 μm/min (Klitorinos et al., 1993). The subpolar flagella at both cell ends rotate around the cell bodies to generate translocation. This movement of the flagella around the helical cytoplasmic cylinder creates the thrust for the bacterium to swim through its environment (Berg, 1976; Charon et al., 1992b). The motility pattern of spirochetes involves translocation in one direction that can be reversed, as well as an occasional bending of the cell in the middle of its body, the so-called ‘flexing’ (Fosnaugh and Greenberg, 1988). Models explaining how flagellar rotation at opposing cell ends can result in unidirectional movement have been proposed for various spirochete species, including the human pathogens L. interrogans and B. burgdorferi, as well as the free-living bacterium S. aurantia (Berg, 1976; Fosnaugh and Greenberg, 1988; Goldstein et al., 1994). As in most other free-swimming bacteria, the flagella motor is driven by proton motive force (Goulbourne and Greenberg, 1980), and the direction of flagellar rotation appears to be bi-directional, as found in the enteric bacteria (Berg and Anderson, 1973; Charon et al., 1992a). The flagellar components of numerous spirochetes, including the oral spirochete T. denticola, are encoded by various gene clusters with strong homology to motility genes of other motile organisms (Stamm and Bergen, 1999, 2001; Charon and Goldstein, 2002).
Treponemes have been associated early on with the etiology of periodontal disease, since they were always found in intimate association with inflamed gingival epithelium and appeared to be the only bacteria capable of penetration into the tissue (Listgarten et al., 1975; Soames and Davies, 1975). This predominance of oral spirochetes (in combination with fusiformen bacteria) within diseased sites is especially pronounced in patients with acute necrotizing ulcerative gingivitis (ANUG) (Rosebury and Sonnerwirth, 1958; Listgarten, 1965; Listgarten and Lewis, 1967; Loesche et al., 1982). Interestingly, Loesche (1976) hypothesized that nutrients present in the serum transudate of the gingival crevice may attract the motile spirochetes and place them at the advancing margin of the plaque in the subgingival sulcus. Later, the oral treponemes T. denticola, T. medium, and T. vincentii were actually found to perform chemotaxis toward different sera, including rabbit, goat, horse, bovine, and human (Umemoto et al., 2001). Interestingly, T. denticola, which has been implicated as one of the causative agents for periodontal disease, exhibited the strongest response to all types of serum, whereas T. medium and T. vincentii were significantly attracted only by rabbit serum. The active component in serum was identified as albumin. Another study showed that T. denticola is also attracted by a variety of sugars and amino acids (Mayo et al., 1990). The chemotaxis response to serum and albumin by treponemes could indicate the role of chemotaxis in the pathogenesis of periodontal disease, since these components are exuded from the gingival crevice and could attract treponemes to gingival pockets for colonization and initiation of periodontal disease.
Versatile genetic tools have recently become available for T. denticola, allowing for a more detailed examination of motility and chemotaxis and their potential role in pathogenesis (Girons et al., 2000; Hardham and Rosey, 2000; Tilly et al., 2000). In addition to the above-mentioned motility genes, an operon encoding the general chemotaxis genes cheA, cheW, the novel chemotaxis gene cheX, and cheY, as well as two genes that exhibit strong homology to known chemoreceptors, has been found in T. denticola (Kataoka et al., 1997; Li et al., 1999; Greene and Stamm, 1999). Further genome search of the unfinished T. denticola genome database (www.tigr.org) revealed the presence of about 16 additional putative chemoreceptors and homologues for cheB and cheR (Lux and Shi, unpublished results). Thus, T. denticola has the complete set of chemotaxis genes required for signal reception, transduction, and adaptation. Gene inactivation of these chemotaxis genes results in loss of chemotactic responses and altered motility patterns (Lux et al., 2002; Lux and Shi, unpublished results), indicating that these are indeed the chemotaxis genes. Most interestingly, these chemotaxis mutants are deficient in penetration of in vitro tissue layers formed by human gingival keratinocytes (Lux et al., 2001). In vitro tissue penetration has been established as a model system for assessment of the ability of pathogenic spirochetes to migrate through endothelial or epithelial barriers (Thomas et al., 1988, 1989; Comstock and Thomas, 1989, 1991; Riviere et al., 1989, 1991; Szczepanski et al., 1990; Thomas and Higbie, 1990; Sadziene et al., 1991; Haake and Lovett, 1994; Peters et al., 1999; Lux et al., 2001). Strains unable to penetrate these in vitro tissues were generally found to be non-virulent as well.
In addition to T. denticola, many other motile oral bacteria (Table)—including species of Campylobacter (Wolinella), Selenomonas, and Capnocytophaga—were frequently isolated from the deep pockets of patients with aggressive forms of periodontal disease (Socransky, 1977; Slots, 1979; Tanner et al., 1979). The chemotactic properties of several oral Campylobacter (Wolinella) species have been compared with those of non-oral members of the same genus (Paster and Gibbons, 1986). The oral Campylobacter strains were found to be strongly attracted by formate at concentrations similar to those measured in dental plaque (Gilmour et al., 1976), whereas intestinal species did not perceive formate as a stimulus. Unidentified motile species isolated from dental plaque were also found to perceive formate, lactate, and n-valeric acid as positive chemo-effectors (Paster and Gibbons, 1986). The authors proposed that the observed chemotaxis response toward these microbial metabolites would allow motile oral species to identify suitable colonization sites in periodontal pockets by localizing already-established microbial communities.
(7) Concluding Remarks
Motility/chemotaxis is a very important feature that allows bacteria to identify optimal growth conditions, avoid harmful situations, or target specific tissues for interaction or invasion of a host. Even though a vast amount of information leading to detailed understanding of motility and chemotaxis at a molecular level has been collected for many bacterial species, still very little is known about these features in oral bacteria. Pioneering studies indicate that motility and chemotaxis may assist oral bacteria to survive the complex oral flora environment better, e.g., via directing the bacteria to favorable colonization sites for nutrition or to damaged tissue areas for invasion. With dental and periodontal disease still affecting the majority of the population worldwide, a more in-depth understanding of the role of motility and chemotaxis in pathogenesis of oral bacteria may further our understanding of pathogenic events in the oral cavity.
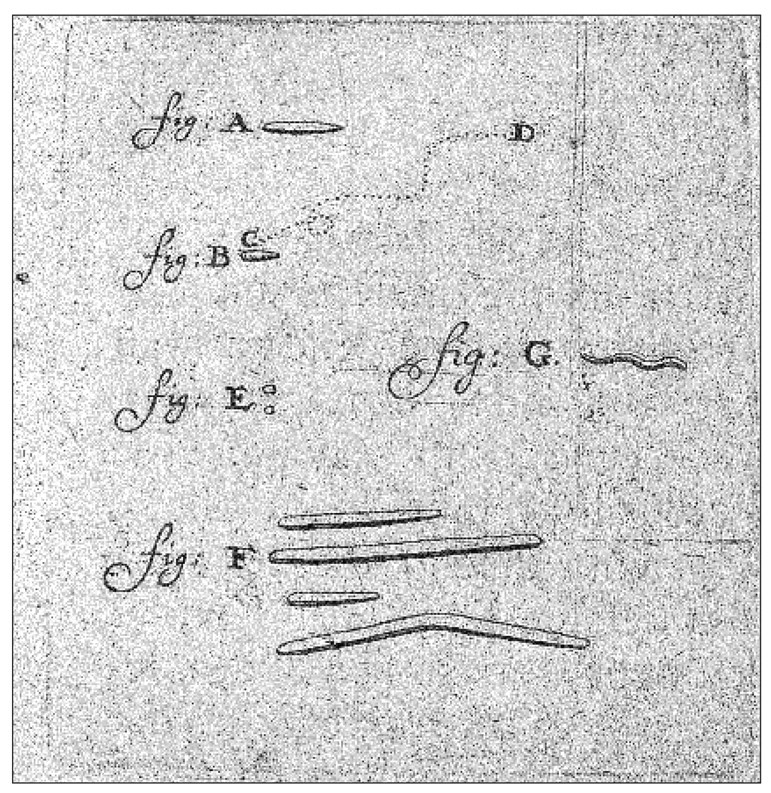

Original drawing from Antony van Leeuwenhoek’s notes. Shown are some types of bacteria that he observed in a dental plaque sample. Among them appear to be cocci (E), fusiforme bacteria (F), and spirochetes (G)
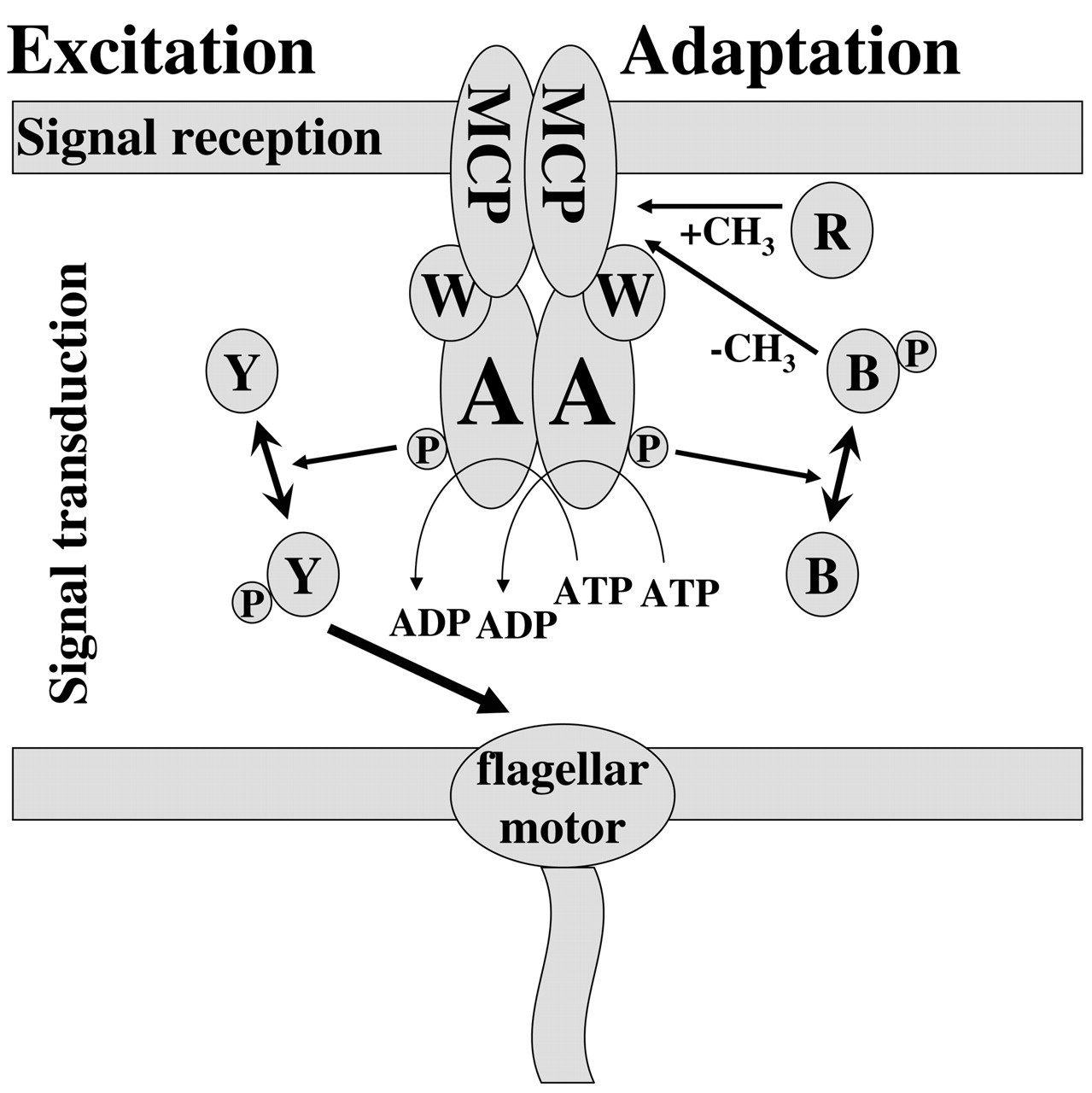
Schematic illustration of basic chemotactic signal transduction pathway. The proteins involved in the basic chemotactic signal transduction pathway are shown. The excitation response involves signal reception by the chemoreceptors (MCP) and further signal transduction via CheW, CheA, and CheY to the flagella motor. Adaptation is achieved via methylation/demethylation of the MCPs by CheR and CheB. See text for details.
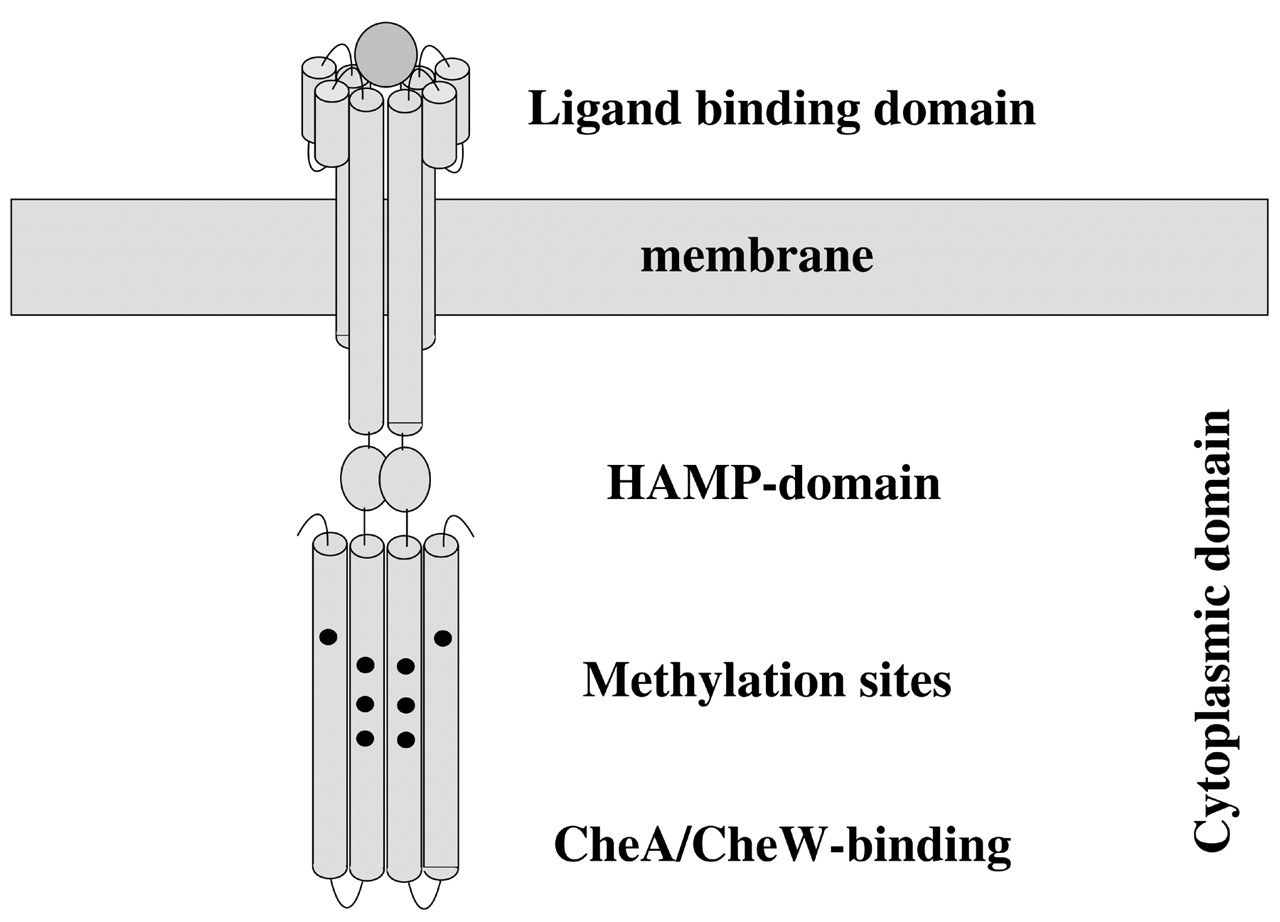
Schematic illustration of chemoreceptor structure. The ‘classic’ chemoreceptor consists of an extracytoplasmic (periplasmic) ligand-binding domain and a transmembrane portion that connects to the cytoplasmic domain via a HAMP domain. The cytoplasmic domain contains the CheA/CheW binding domain and methylation sites that are substrate for CheR and CheB. See text for more details.
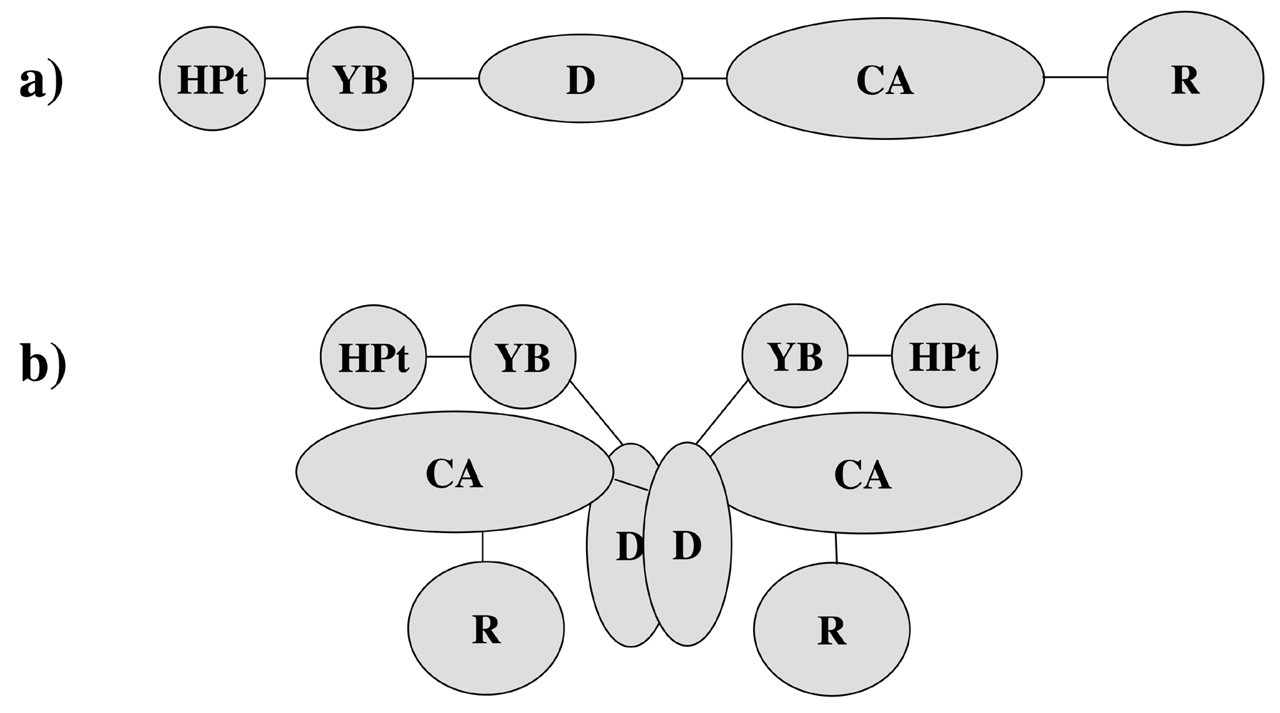
Schematic illustration of CheA domain organization. CheA is organized into five distict domains:
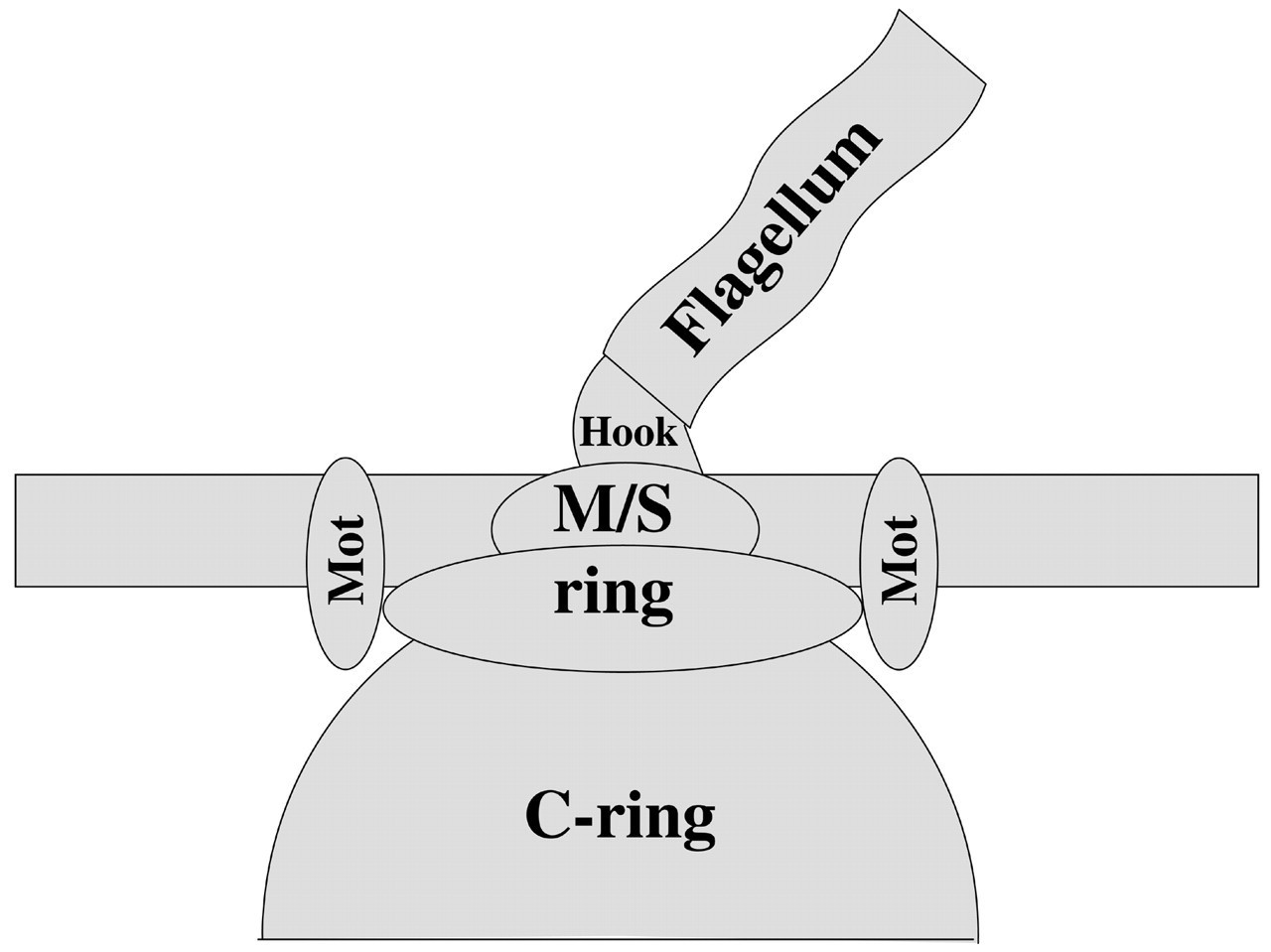
Schematic illustration of flagella motor. The basic elements of the flagella rotary motor are shown. The basal body is formed by the membrane integral M/S-ring and its cytoplasmic extension, the C-ring, and connects via a curved hook to the flagellum. These parts are thought to comprise the rotary part of the flagella motor. The membrane-spanning Mot complexes build the stator that allows for torque generation. For more detailed information on architecture and function, see Macnab (1996, 1999).
Footnotes
Acknowledgements
We gratefully acknowledge the critical comments and valuable advice of Drs. Fengxia Qi, Melissa Sondej, and Richard Ellen. The review was prepared with funding support from the National Institutes of Health, Washington Dental Service, C3 Scientific Corporation, and BioSTAR of the University of California.
