Abstract
The scientific literature during the last ten years has seen an exponential increase in the number of reports claiming links for genetic polymorphisms with a variety of medical diseases, particularly chronic immune and inflammatory conditions. Recently, periodontal research has contributed to this growth area. This new research has coincided with an increased understanding of the genome which, in turn, has permitted the functional interrelationships of gene products with each other and with environmental agents to be understood. As a result of this knowledge explosion, it is evident that there is a genetic basis for most diseases, including periodontitis. This realization has fostered the idea that if we can understand the genetic basis of diseases, genetic tests to assess disease risk and to develop etiology-based treatments will soon be reality. Consequently, there has been great interest in identifying allelic variants of genes that can be used to assess disease risk for periodontal diseases. Reports of genetic polymorphisms associated with periodontal disease are increasing, but the limitations of such studies are not widely appreciated. While there have been dramatic successes in the identification of mutations responsible for rare genetic conditions, few genetic polymorphisms reported for complex genetic diseases have been demonstrated to be clinically valid, and fewer have been shown to have clinical utility. Although geneticists warn clinicians on the over-enthusiastic use and interpretation of their studies, there continues to be a disparity between the geneticists and the clinicians in the emphasis placed on genes and genetic polymorphism associations. This review critically reviews genetic associations claimed for periodontal disease. It reveals that, despite major advances in the awareness of genetic risk factors for periodontal disease (with the exception of periodontitis associated with certain monogenetic conditions), we are still some way from determining the genetic basis of both aggressive and chronic periodontitis. We have, however, gained considerable insight into the hereditary pattern for aggressive periodontitis. Related to our understanding that it is autosomal-dominant with reduced penetrance comes a major clinically relevant insight into the risk assessment and screening for this disease, in that we appreciate that parents, offspring, and siblings of patients affected with aggressive periodontitis have a 50% risk of this disease also. Nevertheless, we must exercise caution and proper scientific method in the pursuit of clinically valid and useful genetic diagnostic tests for chronic and aggressive periodontitis. We must plan our research using plausible biological arguments and carefully avoid the numerous bias and misinterpretation pitfalls inherent in researching genetic associations with disease.
(I) Introduction
The periodontal diseases are initiated by microbial plaque, which accumulates in the gingival crevice region and induces an inflammatory response. This inflammation, chronic gingivitis, may progress in certain susceptible individuals to the chronic destructive inflammatory condition termed periodontitis. Whereas gingivitis is a reversible process, in periodontitis, bone and other tooth-supporting tissues are destroyed (Armitage, 1999, Armitage et al., 2000; Flemmig, 1999). While microbial and other environmental factors are believed to initiate and modulate periodontal disease progression, there now exists strong supporting evidence that genes play a role in the predisposition to and progression of periodontal diseases (Sofaer, 1990; Hart, 1994, 1996; Michalowicz, 1994; Hassell and Harris, 1995; Hodge and Michalowicz, 2001). A corollary of this realization is that if the genetic basis of periodontal disease susceptibility can be understood, such information may have diagnostic and therapeutic value.
The sequencing and annotation of the human genome ushered in the era of genomic medicine (Collins and McKusick, 2001; Guttmacher et al., 2001; Guttmacher and Collins, 2002; Khoury et al., 2003). As a consequence of the human genome project and related projects, fundamental aspects of the human genome have been characterized and are beginning to be understood. While much work remains, it is clear that a fundamental understanding of the structure and function of genes will provide a basis for the development of better disease nosology, diagnostic and susceptibility testing for the presence of disease-associated genes, and, ultimately, for the development of better treatment intervention strategies that address the etiologic basis of disease. While the application of genetic information and technology to the diagnosis and treatment of periodontitis is conceptually compelling, it is important that one maintain a realistic perspective of the clinical utility of genetic information (Friedrich, 2000; Varmus, 2002). To this end, this review evaluates current dental and medical literature reports of genes and gene polymorphisms associated with periodontal diseases. To facilitate the critical review process, an overview of the evidence for a genetic role in human periodontitis is presented, including a brief review of germane genetic principles and pertinent investigative methods.
There is now a significant body of clinical and scientific evidence that genetic factors are important determinants of periodontitis susceptibility and progression. Support for this statement comes from studies of humans and animals which indicate that genetic factors influence inflammatory and immune responses in general, and periodontitis experience specifically. Individuals may respond differently to common environmental challenges, and this differential response is influenced by the individual’s genetic profile. Specifically, different forms of genes (allelic variants) can produce variations in tissue structure (innate immunity), antibody responses (adaptive immunity), and inflammatory mediators (non-specific inflammation). Allelic variants at multiple, perhaps many, different gene loci probably influence periodontitis susceptibility. While the effects of some of these genetic variants may be large and of clinical significance, the effects of others are probably minor and not clinically significant. To understand the potential clinical relevance of genetic variability on periodontitis, we must understand how different genes can contribute to disease.
(II) Genetic Disease Paradigms
(A) Genetic variance
There are estimated to be 25,000–50,000 different genes in the human genome (Parra et al., 2003). Genes can exist in different forms or states. Geneticists refer to the different forms of a gene as allelic variants or alleles. Allelic variants of a gene differ in their nucleotide sequences. When a specific allele occurs in at least 1% of the population, it is said to be a genetic polymorphism. The genome is composed of a linear strand of nucleotides in a double-stranded helical array. The linear sequence of nucleotides in one of the strands codes for amino acids in the form of triplet codons. Most allelic variants involve a change from one of the four nucleotides (A, T, C, G) to another. These changes alter the triplet codon that codes for an amino acid. Due to the redundancy of the triplet codon, some of these codon changes still code for the same amino acid. However, some allelic variants alter the amino acid composition of the protein products of genes. When a nucleotide change is very rare, and not present in many individuals, it is often called a mutation. In contrast to mutations, genetic polymorphisms are usually considered normal variants in the population. When a nucleotide change occurs in a codon, it may or may not change the amino acid coded for by that codon. When a nucleotide change in a codon does not alter the amino acid, it is said to be “silent” and is usually of no biological consequence. Nucleotide changes that do alter the amino acid composition of a protein may be major, resulting in a dysfunctional protein, or the change may be more moderate. In the latter case, the protein function may be altered slightly. Consequently, the specific protein products of different alleles may function differently. These differences in physiological functioning of different proteins can be enhanced by certain environmental exposures (e.g., diet, smoking, microbial factors). If the affected protein functions in a biological process, e.g., inflammatory response to a specific microbial agent, certain polymorphisms may increase or decrease a person’s risk for a disease phenotype. Whether a particular gene variant contributes to a disease phenotype depends upon the magnitude of the effect in contributing to the development of disease. Thus, genetic variance is determined by allelic differences between individuals. Millions of genetic variants have been identified in the human genome (NCBI database, ftp://ftp.ncbi.nih.gov/snp/; Lander et al., 2001; Venter et al., 2001; Guttmacher and Collins, 2002). While many genetic variants do not occur in genes, a significant number do.
In summary, many single-nucleotide polymorphisms (SNPs) that occur in genes do not appear to change the protein product of genes, but a great many SNPs do have an effect on the gene product. Whether this change contributes to a disease phenotype depends on the specific consequence of the particular genetic variant and on the type of disease. Environmental exposures can also be important determinants of whether a specific allele contributes to disease risk.
Risks for many diseases, including periodontal diseases, are not borne equally by all individuals (Johnson et al., 1988; Jenkins and Kinane, 1989). A variety of microbial, environmental, behavioral, and systemic disease factors is reported to influence risk for moderate to severe periodontitis (Page and Beck, 1997). It is also increasingly evident that genetic variance is a major determinant of the differential risk for many human diseases (Friedrich, 2000; Collins and McKusick, 2001; Khoury et al., 2003). However, the contribution of an allelic variant to a disease can vary from being deterministic to having only a minor effect on the etiology. The contribution of an allelic variant to disease has major implications for the disease’s characteristics. To appreciate the effect of a genetic variant to a disease, one must understand how genes contribute to genetic diseases.
(B) Genetic basis of disease
Genetic variance and environmental exposures are the key determinants to phenotypic differences between individuals. There are extreme scenarios where either an environmental agent or a genetic factor alone can result in pathology. Environmental exposures to very high dosages of radiation have an overwhelming and pathologic effect on any individual, regardless of his/her genetic make-up. Genetic aberrations of specific chromosomes, such as trisomy chromosome 1, results in multiple system pathologies regardless of environmental conditions. However, for many diseases, human populations show differential disease susceptibilities, and the basis for this differential susceptibility may have a genetic or both a genetic and an environmental component (Fig. 1).
Most human diseases have a genetic component to their etiology. However, the extent of this genetic contribution to disease can and does vary greatly for different diseases. The manner and extent to which genetic factors contribute to disease have important implications for identifying the genetic basis of etiology and for utilizing this information for the diagnosis and treatment of disease. Geneticists have traditionally divided genetic diseases into two broad groups, “Simple” Mendelian diseases and “complex” diseases. The distinction between these broad groups is based on the pattern of transmission of the disease, which reflects the manner in which genes contribute to each disease.
(C) Simple Mendelian diseases
Diseases that follow predictable and generally simple patterns of transmission have been called “Mendelian” conditions. The name reflects the fact that these diseases occur in simple patterns in families, and in most cases a single gene locus is the major determinant of the clinical disease phenotype. These diseases follow a classic Mendelian mode of inheritance (autosomal-dominant, autosomal-recessive, or X-linked). Usually, the prevalence of these Mendelian conditions is rare (typically much less than 0.1%), with the exception of some unique populations that have been isolated from other human populations. When the genetic basis of a Mendelian condition is identified, it is often found that the condition results from the effect of a genetic mutation at a single gene locus. The disease phenotype usually occurs over a broad range of environments, and although environmental factors and other genes can modify them, in many cases they manifest in a remarkably similar way. Examples include amelogenesis imperfecta (Online Mendelian Inheritance in Man [OMIM] 104500), Crouzon syndrome (OMIM 123500), and cleidocranio dysplasia (OMIM 119600). When the gene responsible for a Mendelian disease has been identified, it is possible to develop a diagnostic test to identify individuals who carry a disease-causing mutation in the responsible gene (Table 1). Depending upon the mode of transmission, it is also possible to make fairly specific determinations of the probability of the mutant gene being passed to a child, and often it is possible to predict the course of clinical disease.
(D) Complex genetic diseases
Genetically complex diseases differ from Simple Mendelian diseases in several important ways. Genetically complex diseases are much more prevalent, and usually occur with a frequency of greater than 1% of the population. Complex diseases do not typically follow a simple pattern of familial distribution or transmission. In contrast to the single gene cause of “simple traits”, these “complex traits” are the result of the interaction of multiple different gene loci. Additionally, environmental factors are important in the disease process. Unlike simple genetic traits, which are often due to rare mutations at a single gene locus, the genetic variants that are etiologically important for most complex genetic diseases are common in the population, and occur in unaffected as well as affected individuals. Also, in contrast to mutations that can eliminate a gene product or change the protein product of a gene so significantly that it acts to disrupt other biological processes, the individual genetic variants that are important in complex diseases are much less disruptive, and usually work within the normal range of function.
This reflects the fact that these gene variants (genetic polymorphisms) are common in the population. Many disease-associated genetic polymorphisms are common in the population and can be present at allele frequencies of > 20%, with some disease-associated alleles reported in > 50% of populations studied (Eichner et al., 2002). The fact that these genetic alterations alone are insufficient to cause disease has important implications. A critically important fact is that the presence of one disease-associated allele is not sufficient to cause disease. Consequently, knowledge of the presence of one disease-associated allele in an individual does not provide enough information for a clinical diagnosis. In fact, the presence of some disease-associated alleles in a significant proportion of the unaffected general population reflects that, while these alleles may influence risk, they are not deterministic in most cases. In contrast to genetic mutations that are often diagnostic for Simple Mendelian conditions, the presence of a polymorphism in a complex trait can be difficult to interpret, and must be assessed with other information. It is important to have information about the allele frequency in the population tested, and also to be able to quantitate, in a meaningful way, the magnitude of effect a disease-associated allele has on the disease processes. For this reason, some measure of the specificity and sensitivity of a disease-associated allele to predicting disease is desirable. Currently, considerable attention is being focused on the clinical validity and clinical utility of genetic polymorphisms that have been reported to be associated with a disease.
(E) Polymorphism
vs. mutation
A major difference in the genetic basis for Simple Mendelian diseases vs. complex genetic diseases is the number of genes involved and the contribution of each gene to the overall disease phenotype. In Mendelian disease, alteration of a single gene locus can result in a mutation that has a major physiological impact and therefore may be considered to be deterministic of the disease. The fact that the genetic alteration is predictably associated with a disease phenotype indicates that there is no redundancy or compensation in the particular biological system that can overcome the effect of the underlying genetic defect. Geneticists have long termed such genetic alterations “mutations”. Even in the case of Simple Mendelian diseases, there can be a wide spectrum of clinical outcomes observed in individuals with the identical mutation (Price et al., 1998). Environmental factors as well as allelic variance at other genes may act as modifiers and contribute to variable clinical expression of a disease or trait (Chanock and Wacholder, 2002). In contrast, the genetic alterations that contribute to complex diseases are individually of much smaller effect. The types of genetic variants that contribute to complex diseases are generally called “genetic polymorphisms” because, in contrast to mutations, they are prevalent in the population. In contrast to mutations that have been causally linked with Mendelian diseases, genetic polymorphisms that are associated with complex diseases are often not directly causally linked, but rather specific alleles are reported to be found more frequently in diseased individuals than in non-affected controls. There is no one-to-one correlation of the presence of a specific genetic allele and the occurrence of disease. It is important to understand that disease alleles reported to be associated with a disease are also found in unaffected individuals, and some individuals with disease do not have the specific disease-associated allele. Thus, the presence of a disease-associated allele in an individual is not diagnostic for a disease. In contrast to monogenetic traits, genetic polymorphisms contribute a small part to the disease process which may require the contributions of multiple, perhaps 5–8, different genes to develop a disease phenotype. Environmental factors are also critical to the etiology of most common diseases. Because the disease etiology is so complex, it is extremely difficult to quantitate the specific contributions of the multiple components of complex diseases. Most chronic diseases are of adult onset, and therefore take many years to develop. Genes involved in the innate, inflammatory, and immune responses are often involved. These physiological and biological pathways have many compensatory and redundant aspects, and therefore it is extremely difficult to quantitate the effect of any one genetic variant on the disease state. Consequently, if an individual is found to have a disease-associated genetic polymorphism, its clinical significance is unclear.
(III) Methods of Genetic Analyses
Clinical and scientific data from a variety of sources suggest that genetic variants are major determinants of syndromic and non-syndromic periodontitis. The evaluation of the quality of supporting studies requires an understanding of the formal genetic analytical methods that have been used. Geneticists use a variety of techniques to demonstrate the genetic basis of disease. Some methods are general, while others facilitate the precise identification of genetic variants that cause or contribute to disease. The methods overviewed below have been important in the evaluation of genetic aspects of periodontitis.
(A) Familial aggregation
Familial aggregation of a trait or disease can suggest genetic etiology. However, families also share many aspects of common environment, including diet and nutrition, exposures to pollutants, and behaviors such as smoking (active and passive). Certain infectious agents may cluster in families. Thus, familial aggregation may result from shared genes, environmental exposures, and similar socio-economic influences. To determine the evidence for genetic factors in familial aggregation of a trait, more formal genetic studies are required. There have been many clinical reports suggesting a familial aggregation of periodontitis, but until recently the research tools to pursue these reports were lacking (Boughman et al., 1988; Hassell and Harris, 1995; Hart and Kornman, 1997).
(B) Twin studies
Through the phenomenon of twins, in particular monozygous (MZ) twins arising from one fertilized egg, nature has provided a wonderful tool for the examination of genetic influences in disease and to assess how much this is influenced by environment. MZ (monozygous) twins are genetically identical, and dizygous (DZ) twins are only as genetically similar as brothers and sisters would be, sharing, on average, ~ 50% of their genes in common (DZ twins are from two different eggs). Discordance or differences in disease experience between MZ twins must be due to environmental factors, and between DZ twins they could arise from both environmental and genetic differences. The difference in concordance between MZ and DZ twins for a particular phenotype can be used to estimate the effects of the extra shared genes in MZ twins, if the environment for twin pairs is the same. Studying disease presentation in twins is useful for differentiating the variations due to environment from those due to genetic factors and for estimating the amount of heredity in a phenotype.
(C) Segregation analysis
Genes are passed from parents to children in a predictable manner, and genes segregate in families as predicted by Mendel’s Laws (Monaghan and Corcos, 1984). Geneticists study the pattern of trait transmission in families using a method called segregation analysis. Segregation analysis evaluates the relative support for different transmission models to determine which can account for the transmission of a trait through families. By sequentially comparing models with each other, segregation analysis identifies the model that best accounts for the observed transmission of a trait in a given population. Geneticists generally apply segregation analyses to determine if trait transmission appears to fit a Mendelian or other mode of genetic transmission. When genetic models of transmission are compared, genetic characteristics—including mode of transmission (e.g., autosomal, X-linked, dominant, recessive, complex, multi-locus, or random environmental), penetrance, phenocopy rates, frequencies for disease, and non-disease alleles—are some of the characteristics included in the different models evaluated. It is important to realize that segregation analysis does not necessarily provide the true model. Since they are comparisons of two models, segregation analyses are only as good as the models tested. If important assumptions of the model tested are incorrect, this will limit the results. This limitation of segregation analysis must be realized, since it has resulted in inaccurate conclusions for the transmission of at least one form of early-onset periodontitis (Long et al., 1987). Investigators use segregation analyses to test alternative models in an attempt to develop the best characterization of transmission characteristics within a set of data. As such, this approach is most appropriately applied to datasets from many families to determine the best-fitting model. Segregation analysis does not find or aim to find a specific gene responsible for a trait.
(D) Linkage analysis
Linkage analysis is a technique used to localize the gene for a trait to a specific chromosomal location. Genetic linkage studies are based on the fact that syntenic gene loci in proximity tend to be passed together from generation to generation (i.e., segregate), as a unit. Such genes are said to be “linked” and violate Mendel’s law of independent assortment. Geneticists can apply quantitative analyses to detect this lack of independent assortment of genetic loci, and use it to map (localize) genes to specific chromosome locations. Over the past 15 years, genetic maps have been developed that show the positions of millions of polymorphic genetic loci spanning the human genome (Guttmacher and Collins, 2002; http://www.ncbi.nlm.nih.gov/genome/guide/human/). Scientists can follow a specific trait as it segregates through families of interest and determine if the trait appears to segregate with a known genetic polymorphism that has been localized to a specific chromosomal location. In this manner, scientists can test to determine if a trait appears to segregate in a manner consistent with “linkage” to a known genetic marker. Because the precise chromosomal location of the genetic marker is known, when linkage is detected, the gene responsible for the trait can be placed in the vicinity spanning the linked genetic polymorphism. Linkage can therefore prove the genetic basis of disease. Linkage is often used as a first step to determine the approximate location of a gene of interest, permitting subsequent studies to identify the mutation responsible for a disease trait. Linkage studies have been particularly effective in identifying the genetic basis of Simple Mendelian traits (OMIM, 2000). Linkage studies of complex genetic traits have not been as successful for a variety of reasons (Townsend et al., 1998; Glazier et al., 2002). A limiting factor for the traditional application of linkage to complex diseases includes the fact that complex diseases are due to the combined effects of “multiple genes of minor effect”. Fortunately, newer adaptations of the linkage approach, and the availability of Association testing approaches, offers a practical alternative (Zhao, 2000; Li et al., 2001; Marazita and Neiswanger, 2003).
(E) Association studies
Genes contributing to common, complex diseases such as periodontitis have proved to be more difficult to isolate. In the absence of specific genetic models, the etiology of complex diseases is often conceptualized as due to multiple factors—i.e., several genetic loci interacting with each other to produce an underlying susceptibility, which in turn interacts with additional environmental factors to produce an actual disease state. For complex traits such as bipolar disorder (Berrettini, 2000), obesity (Chagnon et al., 1998), and oral-facial clefting (Murray, 1995; Carinci et al., 2000), linkage analysis has produced either negative results or a plethora of weak, positive results that are not easily replicated. Theoretical research suggests several reasons for the ambiguity of the linkage results in these cases. First, if a disease gene is neither necessary nor sufficient to cause a disease, but rather is a “modifier gene” that elevates a non-zero baseline risk, conventional parametric linkage analysis may not detect the gene (Greenberg, 1993). Second, if the relative contribution of a gene to a disease phenotype is small, i.e., the disease susceptibility allele raises the risk by a factor of < 2, linkage analysis using affected sibling pairs will not be powerful enough to detect the gene, given realistic sample sizes (Risch and Merikangas, 1996). Thus, linkage analyses may not be a useful strategy for the detection of modifier genes or genes that exert small effects—precisely those genes which might be operating in chronic periodontitis and many other complex disorders. Consequently, attention has shifted away from linkage analysis to association analysis as an alternative means of locating disease susceptibility genes, especially since association studies can sometimes detect weaker effects than can linkage analysis (Hodge, 1994).
Two types of association analysis are commonly used in genetic studies: population-based and family-based approaches (Hodge, 1993). The population-based approach utilizes a standard case-control design, in which marker allele frequencies are compared between cases (affected individuals) and controls (either unaffected individuals or individuals randomly chosen from the population). When a positive association is found, several interpretations are possible: (1) the associated allele itself is the disease-predisposing allele; (2) the associated allele is in linkage disequilibrium with the actual disease-predisposing locus; (3) the association is due to population stratification; or (4) the association is a sampling, or statistical, artifact.
The first two interpretations represent the alternative hypotheses of interest in a gene-mapping context. In case 1, the marker itself is the disease-susceptibility locus. This outcome is the rationale behind candidate gene studies, in which the genes being tested have some a priori expectation of being directly involved in the disease process. Evidence of a positive association can be followed up by investigations to establish a functional role, but these are not easy. In the second case, the associated polymorphism itself does not play a functional role, but rather the polymorphism is located physically close to the gene that does contribute to susceptibility. A classic example is the human leukocyte antigen (HLA) system, in which various HLA haplotypes are associated with several diseases, including IDDM, rheumatoid arthritis, and ankylosing spondylitis (Thomson, 1988). There is currently considerable attention being directed toward the clinical use of disease-associated genetic polymorphisms for genetic testing. However, most initial reports of disease-associated genetic polymorphisms are not replicated (Hirschhorn et al., 2002), reinforcing the need for the development of acceptable criteria for determination of the clinical validity of such reports (Glazier et al., 2002). Fortunately, new approaches hold promise to identify important disease associations that may be important for our understanding of susceptibility for complex diseases. There is currently great interest in characterizing regions of the human genome in linkage disequilibrium, and this approach will undoubtedly take on much greater significance in the future.
(IV) Evidence for the Role of Genetic Variants in Periodontitis
(A) Classification of periodontitis
Currently, there are two major forms of periodontitis—chronic and aggressive periodontitis (Armitage, 1999; Flemmig, 1999). Risk for periodontitis is not shared equally by the population. It is clear that periodontitis severely affects a high-risk group representing around 10–15% of the population, in whom the disease quickly progresses from chronic gingivitis to destructive periodontitis (Johnson et al., 1988; Jenkins and Kinane, 1989). This differential risk for periodontitis is consistent with heritable elements of susceptibility, but direct evidence for a differential genetic contribution to periodontitis comes from several sources.
(B) Familial aggregation
There is literature reporting familial aggregation of periodontal diseases, but, due to different terminology, classification systems, and lack of standardized methods of clinical examination, it is difficult to compare reports directly. Although periodontal disease nosology has changed many times over the timeframe of these reports, most familial reports for periodontitis are for early-onset forms now called aggressive periodontitis (Korkhaus, 1952; Cohen and Goldman, 1960; Benjamin and Baer, 1967; Butler, 1969; Fourel, 1972, 1974; Jorgenson et al., 1975; Melnick et al., 1976; Sussman and Baer, 1978; Ohtonen et al., 1983; Saxen and Nevanlinna, 1984; Van Dyke et al., 1985; Boughman et al., 1986, 1992; Beaty et al., 1987; Long et al., 1987; Marazita et al., 1994; Stabholz et al., 1998). Reports of the familial nature of chronic forms of periodontitis are less frequent, although German studies of the familial nature of chronic forms of periodonitis from the early 20th century have been reviewed by Hassell and Harris (1995). This aggregation within families strongly suggests a genetic predisposition. It must be borne in mind that familial patterns may reflect exposure to common environmental factors within these families. Thus it is important to consider the shared environmental and behavioral risk factors in any family. These would include education, socio-economic grouping, oral hygiene, possible transmission of bacteria, diseases such as diabetes, and environmental features such as passive smoking, sanitation, etc. Some of these factors, such as lifestyle and behavior and education, may be under genetic control and may influence the standard of oral hygiene. The complex interactions between genes and the environment must also be considered in the evaluation of familial risk for the periodontal diseases.
In chronic periodontitis, the phenotype or disease characteristics do not present significantly until the third decade of life, whereas in the aggressive forms of periodontal disease, the presentation can occur in the first, second, third, and fourth decades. This variability in presentation of significant signs of disease makes diagnosis difficult, not only in declaring if a patient suffers from the disease but also in detecting patients who do not suffer from the disease, and differentiating between adult and aggressive forms of periodontitis. The problems associated with the clinical differentiation of periodontal disease are not uncommon in medical genetics, since similar problems arise in the study of other delayed-onset hereditary traits (Boughman et al., 1988; Potter, 1989).
The effects of environment—for example, plaque accumulation and smoking—have major influences on disease experience over time (Fig. 1), and these tend to confuse the diagnosis of aggressive periodontitis, which is dependent on age of presentation for its diagnosis. Huntington’s chorea is another example of a hereditary disease where the diagnosis is possible only relatively late in life (OMIM 143100). The problems of genetic model testing in aggressive periodontitis (AgP) have been highlighted by Boughman et al.(1988), who noted that AgP has a variable age of onset and is often not recognized until after puberty, that the upper age limit of expression of the disease is curtailed (artificially by a diagnostic definition that loss of attachment in patients older than 35 years is attributable to chronic periodontitis), and that difficulties exist in acquiring periodontal disease histories in edentulous family members. All of these factors create substantial problems for genetic studies of periodontal disease.
Attempts to correlate cellular, functional, and immune response variables with early-onset periodontitis phenotypes in families have been generally unproductive, except to indicate that a simple mode of transmission was not evident and that early-onset forms of periodontitis were likely to be etiologically complex and heterogeneous (Astemborski et al., 1989; Potter, 1989, 1990; Boughman et al., 1992; Stabholtz et al., 1998). Although bacterial transmission between subjects has been suggested as a feasible explanation of why aggressive periodontitis may cluster within families, the observation of bacterial transmission within families is insufficient on its own to account for familial clustering (Boughman et al., 1992). While the heterogeneity paradigm discussed by Potter (1989) is borne out in subsequent familial studies of what is now classified as aggressive periodontitis, the striking familial aggregation of the trait is consistent with a significant genetic etiology. Characterization of the genetic components of etiology requires more formal genetic analyses.
(C) Twin studies
Twin studies have been a valuable source of information about the genetic basis of simple as well as complex traits. To maximize the potential of twin studies, large, worldwide registers of data on twins and their relatives have been established (Boomsma et al., 2002). Twin studies have been used to obtain insights into the genetic epidemiology of complex traits and diseases and also to study the interaction of genotype with sex, age, and lifestyle factors. Because of their design, these registers offer unique opportunities for selected sampling for quantitative trait loci linkage and association studies. Twin studies of periodontitis have been limited in scope and generally of small numbers. However, studies of concordance for periodontitis and for clinical indices related to periodontal health and disease generally support a significant heritable component for periodontitis. Most twin studies have studied the more prevalent forms of chronic periodontitis.
A study by Corey et al.(1993) of self-reported periodontal health among 4908 twin pairs found that approximately 9% of subjects (average age = 31 years), consisting of 116 identical and 233 non-identical twin pairs, reported a history of periodontitis. The concordance rate, or level of similarity in disease experience, ranged from 0.23 to 0.38 for MZ twins, and was much lower, 0.08 to 0.16, for DZ twins. Unfortunately, factors such as sex and smoking status and other possible environmental factors were not controlled for in this analysis and may tend to introduce a bias toward finding a correlation between twins. Clearly, twin studies need to be carefully analyzed with appropriate adjustments to reduce the enormous potential for bias due to shared environmental factors.
Michalowicz et al.(1991) studied dizygous twins reared apart (DZA) and reared together (DZT), as well as monozygous twins reared together (MZT) and reared apart (MZA). The mean probing depth and attachment level scores were found to vary less for MZT than for DZT twin pairs, further supporting the role of genetics in this disease. Michalowicz et al.(1991) went on to investigate alveolar bone height in the twins from this Minnesota study and showed significant variations related to the differences in genotype. The twin groups had similar smoking histories and oral hygiene practices. It was concluded that genetics plays a role in susceptibility to periodontal disease. In a subsequent study of 117 adult twin pairs, Michalowicz and co-workers estimated genetic and environmental variances and heritability for gingivitis and adult periodontitis (Michalowicz et al., 2000). Two examiners assessed probing depth (PD), attachment loss (AL), plaque, and gingivitis (GI) on all teeth. The extent of disease in subjects was defined at four thresholds: the percentage of teeth with AL > or = 2, AL > or = 3, PD > or = 4, or PD > or = 5 mm. Genetic and environmental variances and heritability were estimated according to path models with maximum likelihood estimation techniques. MZ twins were found to be more similar than DZ twins for all clinical measures. Statistically significant genetic variance was found for both the severity and extent of disease. Adult periodontitis was estimated to have approximately 50% heritability, which was unaltered following adjustments for behavioral variables including smoking. In contrast, while MZ twins were also more similar than DZ twins for gingivitis scores, there was no evidence of heritability for gingivitis after behavioral covariates such as utilization of dental care and smoking were incorporated into the analyses. These results confirm previous studies and indicate that approximately half of the variance in disease in the population is attributed to genetic variance. The basis for the heritability of periodontitis appears to be biological and not behavioral (Michalowicz et al., 2000).
(D) Clinical diagnostic considerations
The periodontal diseases are heterogeneous and not easily slotted into firm diagnostic categories. Even within families, multiple forms of disease can co-exist (Spektor et al., 1985; Astemborski et al., 1989; Boughman et al., 1992; Hart et al., 1993; Marazita et al., 1994; Shapira et al., 1997). The occurrence of offspring with different clinical forms of aggressive periodontitis has been interpreted to indicate a close relationship among these diseases and argues in favor of a common underlying mechanism (Spektor et al., 1985). The diagnostic difficulties complicate genetic analyses of periodontitis. A complete explanation of why different clinical forms of periodontitis occur is not possible presently, but may be possible when the etiologic factors that can contribute to disease are identified and their role in the disease process clarified.
(E) Segregation analysis
While familial aggregation is consistent with a heritable component of aggressive periodontitis, and twin studies support a genetic component to chronic periodontitis, neither observation or analysis is appropriate to identify the genetic model or specific gene loci that contribute to periodontal disease. Segregation analyses can evaluate the relative support for different models to identify that which most closely represents the clinical data observed. Again, it is important to appreciate the limitations of segregation analyses, the primary being that it does not necessarily identify the true model, but rather identifies the best model of those tested. Segregation analysis is very dependent upon the assumptions of the analyses, and incorrect assumptions will likely lead to incorrect outcomes. There have been few rigorous segregation analyses of aggressive periodontitis, and many are actually studies of one or a few families and are realistically underpowered for definitive conclusions to be drawn. Genetic segregation analysis is dependent upon the accurate clinical identification of affected individuals and familial relationships as well as on genetic assumptions of the analysis. If inaccurate assumptions or data are used in the segregation analysis, the outcomes of the analysis will reflect this. Early studies of aggressive forms of periodontitis were hampered by clinical diagnostic and classification issues (particularly in older individuals) and an overrepresentation of affected females (Saxen and Nevanlinna, 1984; Hart et al., 1991). The female ascertainment bias would lead to false support for X-linked transmission. A detailed review of the topic is presented elsewhere and concludes that several reports suggesting X-linked transmission actually support autosomal-dominant transmission for aggressive forms of periodontitis in North American families when the ascertainment bias is corrected (Hart et al., 1992). These findings were consistent with linkage reports of aggressive periodontitis in an extended kindred from Maryland (Boughman et al., 1986). The most definitive segregation analysis in North American families was performed by Marazita and co-workers (1994), who studied more than 100 families, segregating aggressive forms of periodontitis, and found support for autosomal-dominant transmission. They concluded that autosomal-dominant inheritance with approximately 70% penetrance occurred for both Blacks and non-Blacks. The currently held theory on the genetics of AgP is that pre-pubertal periodontitis, localized aggressive periodontitis (LAgP), and generalized aggressive periodontitis (GagP) are probably due to a major gene locus which is transmitted in an autosomal-dominant manner with reduced penetrance. It is likely that these aggressive forms of periodontitis are genetically heterogeneous, meaning that while the mutated gene responsible for the condition is likely to be the same in any given family, there are probably several different genetic loci that, if mutated, can cause aggressive periodontitis. The expression ‘reduced penetrance’ means that some subjects with the genotype may not express the phenotype, i.e., the clinical manifestations of aggressive periodontitis, while others may express it fully, due to environmental and other genetic interactions. In the phenotypic expression of this trait, the environmental factors (such as smoking and plaque control) may play a large role in allowing the phenotype to present clinically.
Clinical evidence as well as laboratory studies of periodontitis patients suggests genetic heterogeneity. Consequently, the suggestion that aggressive periodontitis may be transmitted in different ways in different families is not a surprise. While some reviews have reported this as problematic, this is not necessarily true. There is common precedent in genetics for a heritable pathologic condition to show different modes of inheritance in different families. Often, once the genetic basis of the condition is understood, it is found that mutations in different genes can cause a clinically similar group of conditions, as occurs in the Ehlers-Danlos syndromes (EDS). These conditions involve aberrations of collagen, and several (type 4 and type 8) are known to have periodontal disease manifestations (OMIM 130050 and OMIM 130080). More than ten forms of EDS are known, and these show different inheritance patterns, including autosomal-dominant, recessive, and X-linked forms. These findings reflect that different genetic loci are capable of causing the disease in both dominant and recessive manners. The fact that some of the genes responsible are autosomal and others are X-linked accounts for the observed modes of transmission. For aggressive forms of periodontitis, the preponderance of the evidence supports autosomal-dominant transmission in North America and autosomal-recessive transmission in certain European populations (Saxen, 1980; Saxen and Nevanlinna, 1984; Hart et al., 1992; Marazita et al., 1994). These different modes of transmission may reflect genetic heterogeneity, such as is seen with EDS.
(F) Linkage studies in Ag P
To date, linkage studies have been performed on two families with localized aggressive periodontitis (LAgP). Boughman et al.(1986) identified an autosomal-dominant form of LAgP in an extended family from Southern Maryland. In this family, type III dentinogenesis imperfecta (DGI-III) and a localized form of AgP were segregating as dominant traits. Since the gene for DGI-III had been previously localized to chromosome 4, they performed a linkage analysis on this chromosome and demonstrated relatively close linkage with the suspected locus for AgP (Boughman et al., 1986). Although the support for linkage for AgP to chromosome 4 in this Brandywine kindred from Maryland was the minimum required for statistical significance (LOD score = 3.0), this was an important study, because it supported autosomal-dominant inheritance of a single major gene locus, clearly indicating a major genetic component to the disease etiology. Hart et al.(1993) evaluated support for linkage to this region of chromosome 4 in a different population of families (14 African-American and four Caucasian). Results of their linkage studies found evidence to exclude a gene of major effect from this region of chromosome 4 as being a major etiologic contributor in these families for any of the genetic models tested. They suggested that these findings supported genetic locus heterogeneity AgP. Thus, this Brandywine population appears to have a different form of periodontal disease, and a different gene is responsible for the disease in the Brandywine population than in the African-American and Caucasian families studied by Hart and co-workers. Results of linkage analyses to date have not identified a gene locus for AgP, but findings do support genetic heterogeneity, with at least one gene locus responsible for AgP located on chromosome 4.
(G) Syndromic forms of periodontitis
Significant, irrefutable clinical and laboratory data clearly indicate that genetic variants can and do predispose to disease states in humans. Severe periodontitis presents as part of the clinical manifestations of several monogenetic syndromes, and the gene mutation and biochemical defect are known for many of these conditions. A commonality of these conditions is that they are inherited as Simple Mendelian traits and are usually due to genetic alterations of a single gene locus. Examples of these monogenetic conditions are given in Table 2. The significance of these conditions is that they clearly demonstrate that a genetic mutation at a single locus can impart susceptibility to periodontitis. Additionally, these conditions illustrate that this genetic susceptibility may segregate by different transmission patterns. The fact that the altered proteins function in different structural and immune pathways indicates that genetic modulation of a variety of different genes can affect a variety of different physiological and cellular pathways, imparting susceptibility to pathological consequences in the periodontium in individuals with appropriate microbial challenges (Hart, 1996). These conditions illustrate that genetic contributions to periodontitis susceptibility are multifaceted and may potentially involve many different gene loci. However, in contrast to non-syndromic forms of periodontitis, these conditions have periodontal disease manifestations as part of a collection of syndromic manifestations. In most cases of aggressive periodontitis, individuals present with clinical manifestations of periodontitis, but do not appear to have any other clinical disease manifestations. This is not inconsistent with a genetic disease etiology. Expression of genes can vary in different tissues, and mutations of a ubiquitously expressed gene can result in a tissue specific condition. Recently, mutation of the SOS1 gene has been identified in individuals with hereditary gingival fibromatosis (Hart et al., 2002). In this condition, a significant alteration of the SOS1 gene has been identified. SOS1 is important in determining whether cells grow, divide, or differentiate. Although SOS1 is ubiquitously expressed, the only clinical manifestation of this gene defect appears in the periodontium, highlighting the tissue-specific manifestation of periodontitis. A similar tissue-specific manifestation of a gene defect may occur with most forms of non-syndromic aggressive periodontitis (Table 2).
(1) Neutrophil functional disorders
Molecular biology has highlighted the important role of several receptors on the polymorphonuclear leukocyte (PMN) surface in adhesion, and emphasized that defects in numbers of these receptors may lead to increased susceptibility to infectious disease (Anderson and Springer, 1987). Adhesion is crucial to the proper function of the PMN, since it affects phagocytosis and chemotaxis, which, if deficient, might predispose to severe periodontal destruction. Page et al.(1987) proposed that the generalized form of pre-pubertal periodontitis [this disease has generalized (GPP) and localized (LPP) forms] is an oral manifestation of the leukocyte adhesion deficiency syndromes (LAD). This is not a widely held view, and other researchers have indicated that GPP and LPP occur in otherwise healthy children (Butler, 1969; Shapira et al., 1997). Leukocyte adhesion deficiency occurs in two forms, both of which are autosomal-recessive traits. Circulating leukocytes have reduced or defective surface receptors and do not adhere to vascular endothelial cells; thus, they do not accumulate in sites of inflammation where they are needed. Reports of LAD indicate that although the blood vessels are full of neutrophils, the disease sites lack sufficient leukocytes to combat the microbial challenge, and thus infections ensue rapidly in these patients. Affected homozygotes suffer from acute recurrent infections, which are commonly fatal in infancy. Those surviving will develop severe periodontitis, which will begin as the deciduous dentition erupts (Waldrop et al., 1987).
Other disorders of neutrophil function are associated with severe forms of periodontal destruction. The Chediak-Higashi syndrome (CHS) is a rare disease transmitted as an autosomal-recessive trait. Those affected are very susceptible to bacterial infections, and this appears to be related to alterations in the functional capacity of the PMN. Man (Hamilton and Giansanti, 1974) and other animals (Lavine et al., 1976) with CHS exhibit generalized, severe gingivitis, extensive loss of alveolar bone, and premature loss of teeth (Temple et al., 1972). The PMN chemotactic and bactericidal functions are thought to be abnormal in these patients.
Clearly, these PMN functional diseases are excellent examples of how monogenic defects can cause periodontitis through a clearly attributable mechanism. The host response is made up of a vast number of processes, all of which are under genetic control and all of which are feasible candidates for variability which may result in the variability in the clinical presentation of periodontal disease seen among the population.
(2) Deficiency in neutrophil numbers (neutropenias)
A further neutrophil deficiency is that of infantile genetic agranulocytosis, a rare autosomal-recessive disease where PMN numbers are very low and which has been associated with AgP (Saglam et al., 1995). Cohen’s syndrome is another autosomal-recessive syndrome and is characterized by mental retardation, obesity, dysmorphia, and neutropenia. Individuals with Cohen’s syndrome show more frequent and extensive alveolar bone loss than do age-, sex-, and mental-ability-matched controls (Alaluusua et al., 1991). Not all neutropenias result in periodontal disease. Familial benign chronic neutropenia has variable expressivity, and although several individuals within a family may be neutropenic, not all are affected either by recurrent infections or by periodontal disease (Deasy et al., 1980). These findings might be explained by the variable genetic expression of the disorder or by the variable effects of the environment (for example, plaque levels) on these patients.
(3) Genetic defects of structural components
Papillon-Lefèvre Syndrome (PLS) is a condition in which the clinical presentation shows various degrees of periodontitis severity as well as great variation in the level of abnormal keratosis (Haneke, 1979; Hart and Shapira, 1994; Gorlin et al., 2001). Genetic linkage studies narrowed the location of the PLS gene locus to chromosome 11, and subsequent mutational analyses permitted the identification of mutations in the cathepsin C gene in patients with PLS (Hart et al., 1999; Toomes et al., 1999). Subsequent studies have identified more than 30 different cathepsin C mutations in individuals from many different ethnic groups (Hart et al., 2000). This is an excellent example of the success of genetic studies contributing to the identification of a gene defect of periodontal importance. Genetic linkage studies permitted localization of the gene defect to a specific chromosome, allowing for focused mutational analyses on genes within an area of the chromosome, and then uncovered gene variations which actually had a functional effect and high biological plausibility as a crucial etiological element of the disease. Additional work has demonstrated that PLS and Haim Munk Syndrome (HMS) (a slightly different clinical variant within the PLS group of disorders) are allelic variants of cathepsin C gene mutations, as predicted by Gorlin (2001) (Hart et al., 2000; Zhang et al., 2001, 2002).
Ehlers-Danlos syndrome refers to a collection of connective tissue disorders characterized by defective collagen synthesis. Ehlers-Danlos types IV and VIII are related to an increased susceptibility to periodontitis (Linch and Acton, 1979; Hart et al., 1997) and are inherited in an autosomal-dominant manner. Clinical characteristics of type VIII Ehlers-Danlos syndrome include fragility of the oral mucosa and blood vessels, and a severe form of generalized AgP (Apaydin, 1995). Other genetic conditions related to defects in structural components of the tissues include the very rare Weary-Kindler syndrome and hypophosphatasia. Aggressive periodontitis has been reported in Weary-Kindler syndrome, where abnormalities of the basement membrane occur (Wiebe et al., 1996). Clinical manifestations of this condition also include epidermolysis bullosa and poikiloderma congenitale. Patients with hypophosphatasia have a decreased serum alkaline phosphatase and the presence of phosphoethanolamine in the urine (Frazer, 1957). In these patients, there is severe loss of alveolar bone and premature loss of the deciduous teeth (Casson, 1969; Beumer et al., 1973), particularly anteriorly (Baab et al., 1986). There is histological evidence of enlarged pulp chambers and a disturbance in cementogenesis, the cementum being either absent or hypoplastic. The concomitant lack of or reduction in connective tissue attachment between the tooth and bone is thought to account for the early spontaneous exfoliation of the deciduous teeth. Baab et al.(1986) described a family where all three children manifested premature exfoliation of the deciduous teeth similar to that seen in pre-pubertal periodontitis (Page et al., 1983). These children were assigned a diagnosis of hypophosphatasia on the basis of alkaline phosphatase and phosphoethanolamine levels. Baab et al.(1986) noted that their data suggest an autosomal-dominant mode of transmission and suggest that hypophosphatasia might be considered in the etiology of some forms of pre-pubertal periodontitis and as a possible explanation for certain site-specific periodontal destruction.
(H) Animal models
While most attention has been devoted to human studies, there is an important body of literature on animal studies of genetic modulation of susceptibility to inflammation in general, and of periodontitis specifically. Studies of animal models, particularly the murine model, demonstrate that genetic factors modulate immune responses to microbial infections (Baer and Lieberman, 1960; Malo and Skamene, 1994; Malo et al., 1994; Nikonenko et al., 2000; Olsson et al., 2000; Niederman et al., 2001; Houpt et al., 2002).
(V) Polymorphism Studies in Periodontitis
(A) Introduction
Most common diseases have a complex genetic etiology. In contrast to the relatively simple monogenetic diseases discussed and illustrated in Tables 1 and 2, complex genetic diseases are not due to a single gene defect. In complex diseases, genetic variants at multiple gene loci contribute to overall disease liability. As such, a cause-and-effect relationship between a particular genetic allele and a disease is not possible. In these cases, a genetic allele is found to be statistically associated with disease more than is found in unaffected individuals. This mathematical association is not necessarily biological or physiological. Studies reporting association vary in design and rigor from reports of an association in only a few individuals in a family (which are not statistically validated to be associated in a general population) to large population-based studies. Association studies ideally evaluate large numbers in population-based studies and thus have power to detect a significant association. Issues of allele frequency in the population studied, case-control design, and population stratification are very important but unfortunately are often omitted from dental studies. This is not acceptable, and studies performed without sufficient rigor need to be interpreted in this light (Glazier et al., 2002; Hirschhorn et al., 2002; Lohmueller et al., 2003). Overstatement of results has become commonplace, such that rigorous, scientifically principled approaches are needed to guard against unfounded and erroneous conclusions.
(B) Gene polymorphisms of host response elements and periodontitis
Our current understanding is that periodontal disease (gingivitis and periodontitis) is initiated by the microbes within the plaque which accumulates in the gingival crevice region. Gingivitis will progress in many individuals to periodontitis, but this progression is governed by the subject’s host response. The host response is determined to some extent by previous experience (acquired immunity) but is predominantly influenced by the person’s genetic make-up. Individuals respond to different antigens in ways predicted by their genes. A good example of this is in the case of the atopy diseases (viz. eczema, hay fever, asthma). Sufferers of hay fever have specific IgE antibody responses to antigens, such as those in pollen, which initiate a mass release of inflammatory mediators from mast cells in the respiratory system. This excessive inflammation is seen as hay fever. Hay fever, asthma, and eczema are genetically related conditions that are grouped within families and show how responses of the immune system can be affected by genetics.
Differences in host response between subjects are not solely confined to differences in immune response, as is the case in the above example, but may also be manifested through differences in the inflammatory response (e.g., complement C1 deficiency that produces angio-edema) or in basic innate immune aspects (an example is the dysfunction of sweat glands which predisposes to infection in cystic fibrosis patients). Thus, a range of genetic deficiencies or genetic variations in the host response can increase the likelihood of periodontitis if microbial plaque is allowed to accumulate in the gingival crevice region.
The genetic basis of many aspects of the periodontal host response has been discussed in reference to genetic disorders predisposing to periodontal disease. The aim of this section is to summarize the potential influence of innate, inflammatory, and immunological genetic variations and to consider where the most promising candidates lie from the viewpoint of a genetic diagnostic approach to periodontitis.
Several features of the host’s innate immune system which may contribute to genetic susceptibility to AgP have already been outlined and include epithelial, connective tissue, and fibroblast defects. Functional defects or deficient numbers of PMN, as discussed previously, have profound effects on the host’s susceptibility to periodontitis. Other aspects of the host inflammatory response—namely, cytokines—have attracted much attention as potentially crucial variants influencing the host response in periodontitis.
(1) Immunological polymorphisms and periodontal disease
Variations in IgG2 levels influence the immune response to periodontal pathogens (Tew et al., 1996), and IgG2 antibodies are considered to be most effective in dealing with microbial carbohydrate moieties. A segregation analysis of IgG2 levels in AgP families has suggested an autosomal-co-dominant mode of inheritance (Marazita et al., 1996). Class II MHC molecules are part of the process of recognition of bacterial antigens and could therefore feasibly influence susceptibility to AgP (Shapira et al., 1994). The MHC or HLA genes determine our response to particular antigens and may thus influence our response to periodontal pathogens and, thus, the host response to periodontitis. Molecular biological techniques are now available to investigate, in detail, genetic polymorphisms, such as those demonstrated by the HLA gene cluster. A Japanese study of AgP patients has found a significant association for these patients with an atypical BamHI restriction site in the HLA.DQB gene (Takashiba et al., 1994). A study performed on Caucasian AgP patients found no association between this restriction site and AgP subjects (Hodge and Kinane, 1999). A further study carried out by the same Japanese group has not been able to corroborate fully this previously reported association (Ohyama et al., 1996). Another group has investigated HLA.DR polymorphisms in patients with GAgP and found a significant association between several DRB1 alleles and the disease (Takashiba et al., 1999). These alleles have previously been associated with rheumatoid arthritis.
Choi et al.(1996) found that allotypes of heavy and light immunoglobulin chains had different frequencies in localized and generalized AgP. The authors consider that immunoglobulin allotypes are linked with IgG2 responses to bacteria, but the link among the allotypes, the immunoglobulin levels, and disease susceptibility is not clear and requires additional evidence. In addition, Gunsolley et al.(1997) added the smoking environment risk factor to this hypothesis and further indicated that Black and Caucasian subjects may have different responses. Ishihara et al.(2001) has further complicated the picture by suggesting that the pro-inflammatory cytokine IL-1 may regulate IgG2 responses.
(2) IL-1 gene polymorphisms in periodontal disease
The IL-1 gene polymorphisms associated with periodontitis provide a useful example for arguing the strengths and limitations of gene polymorphism in disease association studies in the periodontal diseases.
In 1997, Kornman et al. found an association between polymorphisms in the genes encoding for IL-1a (-889) and IL-1β (+3953) and an increased severity of periodontitis. This initial study has since spawned numerous publications and has been the most influential in creating interest in gene polymorphisms and periodontal disease. The specific genotype of the polymorphic IL-1 gene cluster (periodontitis susceptibility trait, PST) was associated with severity of periodontitis in only non-smokers, and distinguished individuals with severe periodontitis from those with mild disease (odds ratio 18.9 for ages 40–60 years, but wide confidence intervals of 1.04 to 343.05). Functionally, the specific periodontitis-associated IL-1 genotype consists of a variant in the IL-1B gene that is associated with high levels of IL-1 production (Pociot et al., 1992, 1993). Kornman et al.(1997) found that 86.0% of the severe periodontitis patients were accounted for by either smoking status or by the IL-1 genotype. Similar results were reported by McDevitt et al.(2000) and from smaller studies by McGuire and Nunn (1999), Laine et al.(2001), and Gore et al.(1998) (who showed that the two polymorphisms within the composite genotype may be in linkage disequilibrium). Other contradictory reports, such as Meisel et al.(2002), stated that the composite genotype showed a strong interaction with smoking [one of the established risk factors for periodontitis (Kinane and Chestnutt, 2000)], whereas non-smokers, even genotype-positive, were not at any increased risk. A similarly contradictory study (Papapanou et al., 2001) of 132 periodontitis patients who were age- and sex-matched with controls did not show any association with the composite genotype and periodontitis. The prognostic utility of the IL.1 genotype on chronic periodontitis progression following non-surgical therapy was studied by Ehmke et al.(1999). Of the 33 patients studied, 16 had the susceptible composite genotype reported by Kornman et al.(1997). Following two years of periodontal maintenance care, no differences in tooth or attachment loss were detected between those with and those without the genotype. Equivocal studies, such as those of Cullinan et al.(2001), demonstrated an interaction of the IL-1-positive genotype with age, smoking, and P. gingivalis, which suggests that the IL-1 genotype is a contributory but non-essential risk factor for periodontal disease progression in this population. Cattabriga et al.(2001) reported no significant differences in tooth loss in patients with the IL-1 genotype in a non-smoking, well-maintained periodontal population after 10 years. De Sanctis and Zucchelli (2000) demonstrated that genotype expression did not affect GTR treatment response at one year, but had a great impact on long-term stability (year 4). In a three-year period, patients with positive IL-1 genotype lost about 50% of the CAL gained in the first year and were about 10 times more likely to experience > 2 mm CAL loss when compared with oral-hygiene-matched genotype-negative patients.
The polymorphisms in the interleukin-1 (IL-1) gene cluster linked with periodontitis (Kornman et al., 1997) are found in approximately 30% of the European population, but the prevalences are dramatically lower in Chinese subjects (2.3%), and thus the usefulness of the composite genotype of allele 2 of both IL-1A +4845 and IL-1B +3954 for determining susceptibility in Chinese patients is dubious (Armitage et al., 2000).
(3) The interleukin-1 polymorphism in aggressive periodontitis
Hodge et al.(2001) examined IL-1A and IL-1B genetic polymorphisms in unrelated European white Caucasian patients with generalized early-onset periodontitis (GEOP) and found no significant differences between patients and controls for any of the composite genotypes described by Kornman et al.(1997). No significant differences were found between patients and controls regardless of whether smoking was included as a covariate. It was concluded that there was a lack of association between the IL-1 polymorphisms and aggressive periodontitis, which questions the utility of these candidate genes as markers of susceptibility. This was a relatively homogeneous Scottish population, and the results, although negative, merely reflect the lack of utility of this composite genotype test in this population.
Other studies on the composite genotype reported by Kornman et al.(1997) and aggressive periodontitis have had similarly mixed results. For example, the studies by Diehl et al.(1998, 1999) actually found that allele 1 rather than 2 of the IL.1B+3953 exhibited polymorphism. Furthermore, Parkhill et al.(2000) investigated the frequency of polymorphisms in the genes encoding interleukin-1β (IL-1β) in Caucasians with aggressive periodontitis compared with controls. The frequency of IL-1β genotypes homozygous for allele 1 of the IL-1β+3953 SNP was found to be significantly increased in AgP patients (p = 0.025). Upon stratification for smoking status, a significant difference was found in the IL-1β genotype distribution in AgP smokers compared with control smokers (F-exact test, p = 0.02), but not between AgP non-smokers and control non-smokers. The IL-1β 1/1 genotype occurred at a higher frequency in AgP smokers (odds ratio = 4.9) compared with control smokers. These results of Parkhill et al.(2000), in contrast to those of Hodge et al.(2000), found that an IL-1β genotype in combination with smoking is associated with aggressive periodontitis.
Similar negative findings for this composite genotype and both chronic and aggressive periodontitis populations from different racial and ethnic backgrounds have been demonstrated, and thus the diagnostic utility of the composite genotype may be restricted to specific populations, i.e., the results do not appear to be applicable globally and across ethnic populations, and certainly not for aggressive periodontitis.
(4) Biological plausibility for the composite IL-1 polymorphisms
Many investigators have suggested a role for IL-1 in the initiation and progression of periodontitis and have quoted in vitro and in vivo studies showing that IL-1 activates the degradation of the extracellular matrix and bone of the periodontal tissues, and elevated tissue or gingival fluid levels of IL-1β have been associated with periodontitis. Kornman et al.(1997) quotes abstracts of in vitro studies from Pociot et al.(1992) and others (e.g., Larsen et al., 2001) which claim that the IL-1 polymorphism associated with severe periodontitis in their studies is also known to correlate with a two- to four-fold increase in IL-1β production. A problem with this line of reasoning is that IL-1 is a pro-inflammatory cytokine intimately involved in all inflammatory reactions as well as in immune and reparative or healing responses, and any perturbation of its level which would not be homeostatically controlled could have widespread consequences not limited to periodontal disease. In addition, IL-1 is one of many pro-inflammatory cytokines (IL-6, TNFα, etc.) which have overlapping activities, and thus some redundancy exists in the cytokine system. Furthermore, IL-1 has many controlling mechanisms which include inhibition of transcription, release controls, receptor antagonists, etc., and thus is highly regulated such that any polymorphism coding for increased production of this molecule could readily be controlled by the elaborate positive and negative feedback loops associated with its regulation.
The claimed association of severe periodontitis with smoking and the IL-1 genotype (in that the composite IL-1 genotype did not influence susceptibility in smokers) poses further problems: Do smoking and the overproduction of IL-1 work along the same pathogenic pathway? If so, the action of both factors is not additive but renders the other redundant, or the overall effect of smoking is so overriding that the composite genotype has little or no effect. These explanations, while feasible, require much more mechanistic knowledge for both risk factors, but especially for the genotype, given that the association with periodontitis is not as established as the literature on smoking (Kinane and Chestnutt, 2000).
Socransky et al.(2000) investigated the association between the composite genotype and carriage of periodontal species. They found that the mean counts of specific species were higher in general in IL-1 genotype-positive compared with -negative subjects. The species detected at higher levels were those frequently associated with measures of periodontal inflammation. A further study aimed at studying the composite genotype and inflammation was performed by Lang et al.(2000). Genotype-negative subjects had significantly lower percentages of bleeding on probing (BOP) (p = 0.0097), and it was concluded that the increased BOP prevalence and incidence observed in IL-1 genotype-positive subjects indicate that some individuals have a genetically determined hyper-inflammatory response that is expressed in the clinical responses of the periodontal tissues.
(5) Functional studies on the composite IL-1 polymorphisms
Shirodaria et al.(2000) took the research focus farther by attempting an assessment of the functional effect of the composite genotype in terms of the quantity of IL-1α protein in the gingival crevicular fluid of severe chronic periodontitis patients. These researchers found that allele 2 at position -889 of the IL-1A gene [one of the alleles that Kornman et al.(1997) linked with susceptibility to periodontitis] was associated with a four-fold increase in IL-1α, as determined by enzyme-linked immunoassay. This technique does not demonstrate activity but merely protein presence or absence and would not differentiate protein bound to receptors from that bound to inhibitors. Furthermore, it is feasible that inhibitors of pro-inflammatory cytokines may be concomitantly produced to dampen this effect. The authors noted reduced levels of IL-1α protein in heavy smokers regardless of genotype, but this may be related to the reduced gingival crevicular fluid noted in smokers (Kinane and Chestnutt, 2000). This is a useful study, given that it addresses the in vivo effects of the polymorphism on IL-1 protein quantities, but the variation in local gingival crevicular fluid production among patients, sites, and smokers and across gender add considerable variance to such a study; these factors must be considered when the data are interpreted.
Engebretson et al.(1999) also found elevated levels of IL-1β in the gingival crevicular fluid in shallow sites in patients who were positive for the composite genotype reported by Kornman et al.(1997). Smoking was not considered in the study, and no statistically significant differences were noted for deeper pockets. Interestingly, in the 22 chronic periodontitis patients examined, only seven were positive for the susceptible genotype.
Mark et al.(2000) studied peripheral blood monocytes from composite genotype-positive and -negative patients to examine whether the IL-1β polymorphism correlated with increased IL-1β expression by monocytes in response to periodontal bacterial stimulus. Contrary to previous reports, these workers found no significant differences in IL-1β production in response to any stimulant tested. They went on to report marked inter-individual variation in production of IL-1β within both the genotype-positive and -negative patients. Clearly, either the genotype is not important in monocyte production of IL-1 or other genetic loci may determine the monocyte IL-1 responses.
(6) Summary of the findings on the IL-1 composite genotype in periodontitis
It appears that this IL-1 composite genotype has equivocal ability in detecting susceptibility to periodontitis and may be limited in its utility to only specific populations at best. It would appear, from the mixed reports on this composite genotype, that:
it is unlikely to be relevant in aggressive periodontitis; it is, at best, in linkage disequilibrium with the gene contributing susceptibility to chronic periodontitis; it confers risk independent of that attributable to smoking; the polymorphism is at best one of several involved in the genetic risk to chronic periodontitis, which is likely to be a disease in which multiple genes may confer risk; the polymorphism is a useful marker in only defined populations, is relatively absent in some (Armitage et al., 2000), and is too prevalent (Walker et al., 2000) in others to be a genetic marker with utility; demonstration of the functional significance of this gene polymorphism has yet to be confirmed; and clinical utilization of these composite polymorphisms for risk assessment and prognostic determination is currently premature.
(7) Tumor necrosis factor
α
(TNF
α
)
The TNFα cytokine is crucial to both the immune and inflammatory responses. For example, TNF up-regulates host defenses and has other effects on tissue physiology, including bone resorption (Mundy, 1993). Over-expression of TNFα in the periodontium may be harmful to the host. Normally, TNFα and other pro-inflammatory agents are regulated by IL-10, suggesting that some deficiency in this regulation mechanism may be linked with disease. Genotypic variations in cytokine response have been shown in vitro for TNF, and specific alleles implicated in diseases such as systemic lupus erythematosus (SLE) and rheumatoid arthritis (RA). One microsatellite at the TNF locus, TNFα, was recently analyzed in 77 GAgP patients (Kinane et al., 1999). Due to the highly polymorphic nature of the microsatellite loci, a statistical comparison with ethnically matched healthy controls (TNFα n = 91) was conducted by means of a Monte Carlo simulation for each marker. No significant differences were observed between cases and controls for carriage of these TNFα microsatellite polymorphisms. Galbraith et al.(1998) determined TNF genotypes in chronic periodontitis patients and healthy controls and found no differences in the three bi-allelic polymorphisms of TNFα (-238, -308, +252). More recently, Craandijk et al.(2002) also found no significant associations between a different series of four TNFα gene polymorphisms and periodontitis patients.
(8) Interleukin-10 genes
A study of the distribution of genes related to interleukin-10 (IL-10) found no association between the genes for this cytokine and aggressive periodontitis compared with healthy controls (Kinane et al., 1999). As for TNFα genotypic variations, IL-10 polymorphisms have been implicated in diseases such as SLE and RA. Two microsatellites at the IL-10 locus, IL10.R and IL10.G, were typed for 77 GAgP patients (Kinane et al., 1999). A statistical comparison with ethnically matched healthy controls (IL10.R n = 94, IL10.G n = 102) revealed no significant differences for any of the IL-10 markers, although there were possible indications of trends similar to those observed in SLE for the IL10.G marker.
There is little to suggest that variations at the IL10.R locus are relevant to the genetic transmission of GAgP, whereas in RA a significant trend toward IL10.R2 has been shown (Eskdale et al., 1998). It seems that the genotype represented by this allele plays little if any role in GAgP. IL10.G showed the greatest variation between GAgP and control populations, which seems mainly due to a 12% decrease in IL10.G9 occurrence vs. that of other alleles in the GAgP population. While this is insignificant and has subsequently been corroborated (Hennig et al., 2000), it shows a trend which is similar to SLE (not RA) (Eskdale et al., 1997). GAgP may have underlying genetic influences similar to those of other chronic inflammatory disorders, but the data are not sufficiently consistent to warrant classifying GAgP with any other disease genotype. A further lack of association between periodontitis patients and IL-10 polymorphisms in the gene-promoter regions was found by Yamazaki et al.(2001) in a Japanese population which had haplotype frequencies quite different from those reported in studies of Chinese and Caucasians. Again, caution in interpretation is needed here, because not finding an association cannot be claimed as evidence of no association for IL-10 and periodontal diseases. At the moment, however, this appears unlikely.
It is possible that GAgP has no single clear mode of inheritance, because GAgP may represent a heterogeneous group of diseases (Baelum et al., 1996) and may be multigenic rather than having a single major genetic locus, or may have different major loci with the same clinical appearance. Different immunological defects coded by different genes could render individuals susceptible to GAgP through different mechanistic pathways but manifesting the same clinical phenotype. Complex and multifarious interactions among host response, the environment, and bacteria make elucidation of genetic factors difficult. This may be particularly pertinent to the periodontal arena, given the sensitive nature of the gingival environment to imbalance in bacterial and immuno-inflammatory stimuli. It should be borne in mind that microsatellites are only markers; they are highly unlikely in themselves to contribute to disease. Screening microsatellite markers provides a useful tool for focusing on areas of the genome which may then be subjected to more detailed molecular genetic investigations.
(9) Fc-gamma receptor
The Fc-gamma receptor (FcγR) is the receptor present on phagocytes which binds immunoglobulin G (IgG) and is thus crucial in the opsono-phagocytosis of bacteria. Polymorphisms that influence the binding affinity between the Fcγ receptors and IgG of different subclasses are considered important in susceptibility to periodontal disease. Two studies of adult periodontitis patients have investigated associations between FcγR polymorphisms and susceptibility to chronic periodontitis (Kobayashi et al., 1997; Van Schie et al., 1998). No associations were found for FcγRIIa and FcγRIIIb genotypes between maintenance patients and healthy controls (Kobayashi et al., 1997). The authors also considered the influence of various covariates together with the FcγRIIIb allele 2, which included serum IgG subclass, baseline and follow-up clinical indices, and smoking. None of these covariates was found to be significant. It was concluded that the presence of the FcγRIIIb allele 2 may be a risk factor for recurrent periodontitis.
Kobayashi et al.(2000) found functional polymorphisms of IgG Fc receptors (FcγR) in Japanese patients with AgP. They found that FcγRIIIb (NA2 allele) was linked with AgP and possibly also a FcγRIIIa allele. Sugita et al.(2001) reported that FcγRIIIb has an NA1-NA2 polymorphism. NA1 is a more efficient opsono-phagocytic agent than NA2. NA1 is found to a greater extent in Japanese patients who are resistant to chronic periodontitis (OR 1.87, 95% CI 1.07–3.28). Kobayashi et al.(2001) reported that 2 FcγRIII alleles may be associated with severity of chronic periodontitis in a Japanese population.
Meisel et al.(2001) analyzed the association between FcγRIIIa (high-affinity receptor) and FcγRIIIb (low-affinity receptor) and chronic periodontitis (CP). They found that FcγRIIIa was associated with CP (OR 5.3, 95% CI 1.4–26.2) but that this is in linkage disequilibrium with FcγRIIIb, which is also associated with CP. There is a major difficulty here in explaining the effects of these polymorphisms on the disease, since one is associated with a high-affinity binding of the immunoglobulin molecule and one with a low-binding strength: Both polymorphisms are found together (possible linkage disequilibrium), and both are considered to confer risk for chronic periodontitis.
Van Schie et al.(1998) studied a US Caucasian population with moderate to severe periodontitis and age-matched healthy controls for the presence of specific FcγRIIa and FcγRIIIb genotypes. A composite genotype of two FcγR alleles was found more frequently in patients than in controls, and the frequency of another composite genotype was reduced in patients. When the genotype pairs were examined individually, they showed no association with disease. These results are odd, in that one pair of FcγR alleles is part of both composite genotypes, and one of the genotypes, FcγRIIa-H/H131, is considered to have immune functional protective advantages (Sanders et al., 1995). It is feasible that the combined genotypes are in linkage disequilibrium (found close together on the chromosome) with more crucial alleles, and that the FcγRIIb combined genotypes are thus part of a gene pattern or haplotype which may predispose subjects to periodontitis.
The studies from Japan, Europe, and the US tend to associate both FcγRIIIa and FcγRIIIb alleles with both aggressive and chronic periodontitis. These alleles may be in linkage disequilibrium with the real gene causing disease. It is unclear, however, if the functional defect is through reduced or even increased affinity binding conferred by a specific Fcγ receptor.
Colombo et al.(1998) have investigated serum levels of IgG2, Gm(23) allotype, and FcγRIIa and FcγRIIIb allotypes in 32 refractory, 54 successfully treated, and 27 periodontally healthy subjects. No significant differences in serum IgG2 levels, Gm(23) allotypes, or FcγR genotypes were found among the three groups, and these variables could not be related to the clinical status of the subjects.
(VI) General Considerations
(A) Clinical diagnoses
Many confounding factors have been recognized in population association studies of periodontal disease. It is only recently that we have, within the discipline of Periodontology, more carefully defined and classified periodontal disease. Prior to the AAP Workshop in 1999, there was much variation in the literature with respect to disease definition (Armitage, 1999). Thus, there are numerous differences in what we define as a case and indeed what constitutes a control. These variable definitions make comparisons across studies virtually impossible. Other complications arise related to the variable presentation over time of periodontal diseases, notably the aggressive forms of periodontitis which generally appear prior to the age of 35 years of age but are not confined to this age. In many cases, the aggressive forms will appear very early in the teenage years or quite late in the early 30s. Chronic periodontitis is similar in that its presentation and its categorization are quite complex and very much dependent on environmental exposures of the patient over his/her lifetime. Exposures such as smoking, microbial plaque deposition (oral hygiene), and other factors such as systemic disease modifiers (e.g., diabetes) or general factors (e.g., socio-economic status) have a large influence on the expression of the phenotype.
An additional criticism arises in the suggested utility of these population association studies, in that there are often methodological problems within the reported studies. These include the fact that multiple polymorphisms may be tested, but only few are reported, and for the few that are reported, there is often a lack of understanding of the importance of multiple range testing, i.e., the reduced statistical inference that can be made in such exploratory studies. On many occasions, the results are reduced such that only the positive ones are reported, with obvious statistical implications. The bias against publishing ‘negative’ results is one that has tremendous impact on our literature, and investigators need to recognize the importance of such results.
Many studies fail to report the sensitivity and specificity of their tests, nor do they report the many associated environmental aspects which need to be considered. A further criticism is that many studies have numbers of cases and controls insufficient to make robust pronouncements on the association between particular polymorphisms and disease. A feature which should also be included is a precise account of the racial and ethnic background of both cases and controls, because biases within the case and control populations, which could occur in multiracial areas of different socio-economic classes, would render the inference from such study questionable.
(B) Environmental variables
There are numerous tacit assumptions made regarding the populations researched in periodontal disease population association studies. Are the controls truly not susceptible to the disease? Could it be that some of the control subjects are performing high standards of oral hygiene (e.g., educated and motivated subjects), and thus the environmental factors required to interact with the genotype to express the phenotype are missing? In addition, these subjects may have excellent access to health care and health education and may not smoke, thus avoiding other known risk factors which may be needed to reveal the genotype. In this situation, misplacing the subject(s) in the control group will dilute the chances of seeing any disease association. Similarly, it is accepted that periodontal destruction tends to be incremental, and thus the disease experience and the clinical features increase with age. Thus, the ages of the cases and controls have to be considered and matched. Gender and, more importantly in genetic studies, race have to be considered and documented carefully—that is, the genetic heterogeneity of the populations. Thus, the importance of environmental factors in revealing the genotype cannot be overstated in the categorization of cases and controls, and prior knowledge of racial or genetic heterogeneity is needed. A further difficulty is knowing a priori which environmental factors will have an influence, and thus simultaneous epidemiological studies are needed. Twin studies can be particularly useful here in sorting out genetic and environmental factors (discussed in detail later).
(C) Smoking and susceptibility genes
The interplay between smoking and genes in the risk of periodontitis has been discussed in most studies on genetic risk, and clearly these environmental and genetic risk factors have to be considered. The attributable risk of periodontitis from smoking may be influenced by the polymorphism of the N-acetyltransferase (NAT2) gene, which may change the subject’s metabolism to produce more damaging smoking-derived xenobiotics. Meisel et al.(2000) hypothesized that a NAT2 genotype could be a risk factor for periodontal disease. Severe chronic periodontitis patients were predominantly slow acetylators (OR = 2.38–5.02). Thus, the slow acetylator phenotype may be associated with a higher risk of periodontitis.
(D) Biologic plausibility
Researchers must also consider the biological plausibility of the effect of the gene polymorphism in question having a role in the periodontal disease process. There are, however, a great many unknown aspects which provide considerable leeway for investigators in this area. The pathogenesis of periodontal disease is to some extent known. It is a chronic inflammatory condition which progresses to the destruction of bone and supporting structures and, ultimately, the loss of teeth, but the reason that some patients are more susceptible than others is not understood. Any of the innate, inflammatory, or immune processes may be involved in the etiology of periodontal disease. In addition, it is feasible that structural aspects and aspects which may influence the healing process may be similarly involved. This leaves open an extremely wide range of factors which could be considered fortuitous from a research justification viewpoint, but frustrating in the search for the etiology of the periodontal diseases.
(E) Penetrance
The allele or polymorphism in question may not always be active and may need environmental or other gene activity to be brought into play. This is particularly the case in multigene conditions but may also occur in single gene disorders and helps to explain the phenomenon of ‘penetrance’. The aggressive forms of periodontal disease are considered to be inherited as autosomal-dominant conditions with reduced penetrance (Hodge et al., 2000).
(F) Logic of association studies
A hypothetical example of the logic in play in the study of gene polymorphism associations with diseases such as periodontal disease might be the following: In periodontitis (condition A), bone is lost (abnormality B), and since bone is removed by osteoclasts, which are influenced by a specific gene (gene C), we then study the association of a known polymorphism (polymorphism D) of gene C with periodontal disease. Obvious problems in this logic are apparent even at this hypothetical stage. In particular, that the osteoclast has a pivotal role in the disease process is unknown and would be purely speculation. Nevertheless, the link between A and B generally results in the search for a gene C and a polymorphism D which can be investigated. Linking the gene C with periodontal disease is a task for molecular geneticists, who would use linkage analysis techniques and positional cloning together with robust statistical approaches including a precise genetic model and knowledge of the population frequency, penetrance of the disease, mutational searches, and laboratory pathogenesis investigations.
Gambaro et al.(2000) have stressed the crucial importance of demonstrating that allelic variants or polymorphisms are actually functional prior to engaging in population polymorphism association studies. It is unusual that investigators will have chosen to investigate a particular polymorphism with the understanding that it is functionally active. They are more likely to choose polymorphisms of candidate genes (of which there is an extremely wide range in periodontal disease, due to our limited understanding of the etiology) on an arbitrary or convenience basis. A highly illustrative example exists in the medical literature on how the I/D polymorphism associated with serum angiotensin-converting-enzyme (ACE) concentrations was researched. Gambaro et al.(2000) recount that, in 1989, a polymorphism had been related to serum concentrations of ACE, an enzyme possibly involved in atherosclerosis. In 1992, this polymorphism was clearly associated with an atherosclerosis-dependent clinical event. For the next five years, hundreds of researchers investigated thousands of patients to look at new associations with various diseases, based on a simplistic view of the pathophysiology of the way in which this polymorphism (I/D) may influence the disease process. Many studies were spawned to reconfirm other researchers’ findings, and it was as late as 1997 that basic findings appeared to support the pathogenic and pathophysiologcal relationship between the polymorphism and the disease. The disease in question turned out to be renal disease, which, although not cardiovascular disease, does influence hypertension and, thus, cardiovascular disease indirectly. Thus, it was only in 1997 that the rationale fortuitously appeared to support studies which had previously been published. The I/D polymorphism story had a happy ending in that the link between hypertension and renal disease was related to the gene polymorphism, but the original population association studies were never based on a clear demonstration that ACE was linked with atherosclerosis and that the polymorphism (I/D) was relevant to the ACE gene in a functional way. This polymorphism was and is extensively studied and is an example of a useful polymorphism emerging from a weak research foundation. Most other polymorphisms with similarly weak foundations have not been so fortunate (Gambaro et al., 2000), and thus much research effort has been wasted.
(G) Reporting bias
A further concern related to polymorphism population studies which are not based on firm assumptions is that once they are in the literature they tend to take on a veracity which may not be justified. The reason for this is largely due to conscious or subconscious reporting bias. Simply put, authors tend not to believe data in, or submit for publication, papers which do not agree with already-published findings unless they are established researchers and have a track record which commands support and respect of editors and reviewers, or that they have other hypotheses which they are pursuing (note publication bias described previously). In addition, to reliably disprove a hypothesis may require a much larger study than the original study, which was likely significant at the 5% or even the 1% level (approximately four times the n number of the original study may be required).
(H) Interpretation of results
An interpretive problem with the logic sequence that disease A is related to abnormality B, which is influenced by gene C, which has a polymorphism D, is the possible oversimplification by clinicians to gene C → problem B → disease A, or, worse, polymorphism D → problem B → disease A, or, infinitely worse, that polymorphism D → disease A. A commonly overseen but consistent mistake is to interpret studies which do not show an association between polymorphism D and disease A as meaning that gene C is not involved in the pathogenesis or is not a risk factor for disease A, when in fact only one of possibly thousands of candidate polymorphisms may have been ruled out (Fig. 2).
Linkage disequilibrium could also confuse the clinical researcher into considering that because polymorphism D is found to be associated with disease A, then gene C, which contains polymorphism D, is involved in the etiology of the disease. It may be that polymorphism D is in linkage disequilibrium with another allele (polymorphism E) which is crucial to the proper function of gene C, or is in fact part of another unrelated gene X. Another issue which arises is that if the association is found in one population, it has to be tested in other populations if population stratification biases are to be avoided. If polymorphism D is in linkage disequilibrium with polymorphism E, but polymorphism D is the actual susceptibility allele, it should exist in every population. Conversely, if the susceptibility locus is polymorphism E, the association between D and E may occur only in the original homogeneous population, due to a phenomenon termed the ‘founder effect’—that is, the ancestors have the linkage disequilibrium which is passed on to offspring, but the same linkage is not seen in completely distinct populations. Two points are worth considering here. First, if polymorphism D is in linkage disequilibrium with E, polymorphism D is still a useful marker for the homogeneous population. Second, testing polymorphism D in other populations will truly test whether this polymorphism has a crucial influence on gene C and also the disease in question. Of course, the biological plausibility of the polymorphism or marker, if checked initially, would have been crucial evidence in clarifying the association (Table 3).
(I) Proving the association
Recommendations emanating from population polymorphism association studies are invariably overstated in the periodontal and medical literature and are not based on a proper interpretation (conservative or ambitious) of the findings. If researchers are lucky and find an association between a polymorphism and periodontal disease, their immediate priority should be to determine if the polymorphism actually does anything, i.e., to address whether the polymorphism is non-neutral. A non-neutral finding might be that the polymorphism results in the production of a protein with enhanced activity, as has been claimed for the IL-1 polymorphisms (Pociot et al., 1993) which Kornman and colleagues (1997) have associated with periodontal disease. A further approach might be to consider to what other genes the polymorphism is close on the chromosome and whether these may plausibly have a role in the etiology of periodontal disease.
Other important considerations should be to check for population biases, case/control selection, possible confounders (and correction for these), and whether their probability values are strong enough to discount false-positive associations which may creep in if allowance has not been made for multiple comparisons. The typical next stage is to roll out the investigation to consider the association in larger and diverse populations which would be independently assessed and the results reported in the literature (whether they confirm or deny the positive association first claimed). Associations of polymorphisms with periodontal disease studies are at best hypothesis-generating exercises, and clinicians should be clear about the extreme limitations of these approaches when trying to develop robust associations which can be used as clinically relevant risk evaluators for patients. In the vast majority of cases, associations will not relate to the function of the gene, but proving a functional relationship will require considerable multidisciplinary effort in population, pathogenesis, and molecular genetics research.
(J) Susceptibility profiles
An interesting approach which is emerging in other fields of medicine is the development of the susceptibility profile concept in diseases such as Alzheimer’s. The risk of Alzheimer’s has been considered to be substantially influenced by a total of ten genetic polymorphisms of inflammation-related molecules, including pro-inflammatory cytokines (IL-1α, IL-1β, and IL-6, and TNFα) and the protease inhibitors (α-2-macroglobulin, α-1-antitrypsin). McGeer and McGeer (2001) consider that several of these relatively common high-risk polymorphisms may be inherited by an individual giving them a susceptibility profile which reflects the combined influence of the high-risk polymorphisms. A similar high-risk profile may yet emerge for periodontal disease, but clearly much testing across different populations will be needed. If such information was available, therapeutic intervention strategies could be envisaged, aimed at preventing the development of periodontal disease.
(K) Genetic screening for periodontitis risk
The current practical clinical utility of genetic knowledge in periodontics is limited. However, performing clinical periodontal assessments of siblings of AgP probands is one of the most useful actions we can perform to ensure the early diagnosis of this disease. By careful clinical diagnostic procedures, we may detect susceptible patients early and instigate therapy which may prevent the more significant disease aspects from occurring. In the pursuit of better genetic diagnostic tests for chronic and aggressive periodontitis, we must plan our research using plausible biological arguments and carefully avoid bias and misinterpretation of genetic associations with disease.
(VII) Conclusions
Reports on the genetic polymorphisms associated with periodontal disease are increasing, but the limitations of such studies have not yet been fully appreciated. Although geneticists warn clinicians on the over-enthusiastic use and interpretation of their studies (Kidd, 1993), there is a large disparity in the significance placed on genes and genetic polymorphism associations between geneticists and clinicians. One can conclude that, despite major advances in the awareness of genetic risk factors for periodontal disease, we are still some way from determining the genetic basis of both aggressive and chronic periodontitis. A strong plea in the discovery of better genetic diagnostic tests for chronic and aggressive periodontitis is that we plan our research using plausible biological arguments and carefully avoid bias and misinterpretation of genetic associations with disease.
Many studies fail to quantitate, in a meaningful way, the magnitude of contribution of a particular disease-associated allele to disease risk. This failure to quantitate the sensitivity and specificity of tests precludes determination of the clinical validity and ultimately the clinical utility of the association. Additionally, associated environmental aspects which need to be considered are not adequately addressed. Typically, studies have numbers of cases and controls insufficient to make robust pronouncements on the association between particular polymorphisms and disease. Rarely is a precise account of the racial and ethnic backgrounds of both cases and controls included. Biases within the case and control populations, which can occur in multiracial areas of different socio-economic classes, make the inference from such studies questionable.
An interesting approach which is emerging in other fields of medicine is the development of the susceptibility profile concept for specific diseases. For example, the risk of Alzheimer’s has been considered to be substantially influenced by a total of ten genetic polymorphisms of inflammation-related molecules, including pro-inflammatory cytokines and protease inhibitors. It is feasible that a similar situation may pertain in periodontitis, where several relatively common high-risk polymorphisms could be inherited by an individual, giving them a cumulative high-susceptibility profile. For such a high-risk profile to be proved in periodontal disease, much testing across different populations would be needed. Such useful genetic information would be invaluable in therapeutic intervention strategies aimed at preventing the development of periodontal disease.

The classic relationship among phenotype, environment, and genotype. For the periodontal disease phenotype, environmental risk factors include: smoking status, plaque control, socio-economic status, diabetes, etc. G x E is the interaction between environment and genotype. * includes gene-gene interactions.
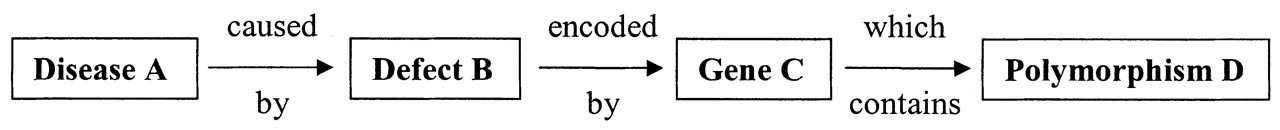
Steps in determining a link between a polymorphism and a disease. (1) Does Polymorphism D influence Gene C functionally? (2) Does Gene C uniquely result in Defect B? (3) Does Defect B have mechanistic plausibility for Disease A?
