Abstract
The oral cavity is a complex ecosystem in which several hundred microbial species normally cohabit harmoniously. However, under certain special conditions, the growth of some micro-organisms with a pathogenic potential is promoted, leading to infections such as dental caries, periodontal disease, and stomatitis. The physiology and pathogenic properties of micro-organisms are influenced by modifications in environmental conditions that lead to the synthesis of specific proteins known as the heat-shock proteins (HSPs). HSPs are families of highly conserved proteins whose main role is to allow micro-organisms to survive under stress conditions. HSPs act as molecular chaperones in the assembly and folding of proteins, and as proteases when damaged or toxic proteins have to be degraded. Several pathological functions have been associated with these proteins. Many HSPs of oral micro-organisms, particularly periodontopathogens, have been identified, and some of their properties—including location, cytotoxicity, and amino acid sequence homology with other HSPs—have been reported. Since these proteins are immunodominant antigens in many human pathogens, studies have recently focused on the potential contributions of HSPs to oral diseases. The cytotoxicity of some bacterial HSPs may contribute to tissue destruction, whereas the presence of common epitopes in host proteins and microbial HSPs may lead to autoimmune responses. Here, we review the current knowledge regarding HSPs produced by oral micro-organisms and discuss their possible contributions to the pathogenesis of oral infections.
(1) Oral Microbial Ecosystem
The oral cavity is one of the most complex microbial ecosystems of the human body and is colonized by several hundred microbial species, predominantly bacteria and, to a lesser extent, yeasts, protozoa, and viruses. The oral microbial community is structured in such a way that each specific population contributes to its overall stability. However, under certain special conditions, the growth of some micro-organisms with a pathogenic potential is favored over other members of the ecosystem, leading to infections such as dental caries, periodontal disease, and stomatitis.
The oral cavity is subjected to a wide range of environmental changes that may affect the biology of resident micro-organisms. Large temperature variations occur when hot and cold foods are consumed. Although the variation is smaller, inflammation at periodontal sites also causes an increase in temperature (Baab et al., 1990; Kung et al., 1990; Fedi and Killoy, 1992). Bacterial products (organic acids and proteases) can influence the pH of the oral cavity. More particularly, the metabolic activity of asaccharolytic bacteria (such as Porphyromonas gingivalis and Fusobacterium nucleatum) that colonize periodontal pockets is associated with a rise in the pH of gingival crevicular fluid to 8.5, compared with 6.9 in the healthy sulcus (Marsh and Martin, 1992). The redox potential, which is largely influenced by oxygen levels, is a critical parameter for the growth of anaerobic bacteria. Certain oral bacterial species are able either to decrease or to increase the redox potential of culture media (Leke et al., 1999). Kenney and Ash (1969) reported redox values of +14 to -157 mV in affected periodontal sites, as opposed to +74 mV in the healthy gingival sulcus. Last, the availability of nutrients (carbohydrates, amino acids, iron) can also significantly influence the survival and growth of oral micro-organisms.
The physiology and other biological properties, including pathogenicity, of micro-organisms are influenced by the environmental parameters mentioned above. Indeed, signal transduction systems that respond to specific environmental cues are often used by pathogens to ensure that genes required for survival, multiplication, and virulence are appropriately expressed during infection (Dirita and Mekalanos, 1989; Straley and Perry, 1995; Guiney, 1997; Konkel and Tilly, 2000). For instance, Spitznagel et al.(1991, 1995) reported that leukotoxin expression in Actinobacillus actinomycetemcomitans is regulated by several environmental signals. Studies on P. gingivalis also revealed the role of environmental parameters in the expression of putative virulence factors, including fimbriae and proteases (McKee et al., 1986; Chu et al., 1991; Smalley et al., 1991; Amano et al., 1994, 2001; Marsh et al., 1994; Percival et al., 1999). Individuals may thus harbor latent pathogens in their subgingival plaque for years until changes in the local environment stimulate their growth, activate the expression of virulence factors, and lead to disease. In addition, changes in in vivo environmental conditions may affect the survival of oral bacteria and induce the accumulation of damaged or denatured proteins, resulting in the expression of bacterial stress proteins.
(2) Microbial Stress Response
When exposed to a wide range of environmental stresses (temperature, pH, redox potential, etc.), prokaryotic and eukaryotic cells respond by inducing or accelerating the synthesis of specific proteins known as stress proteins, including heat-shock proteins (HSPs), which have a high degree of homology. Genes coding for HSPs are among the most conserved in nature (Lindquist and Craig, 1988; Ray, 1999). HSPs are commonly grouped into families based on molecular weight: small HSPs, GroES-homologue proteins or HSP10 (~ 10 kDa), DnaJ-homologue proteins or HSP40 (~ 40 kDa), GroEL-homologue proteins or HSP60 (~ 60 kDa), DnaK-homologue proteins or HSP70 (~ 70 kDa), HptG-homologue proteins or HSP90 (~ 90 kDa), and Clp ATP-dependent proteases (HSP100) (Lindquist and Craig, 1988; Watson, 1990; Lathigra et al., 1991; Shinnick, 1991; Ellis, 1996; Ray, 1999). HSPs that belong to the same family share strong amino acid sequence identity.
Many HSPs are expressed constitutively under normal conditions but are quickly up-regulated under stressful conditions. Induction, which is an emergency response of the micro-organisms, is rapid. Slight changes in environmental conditions—including temperature, pH, redox potential, and nutrient availability—induce the synthesis of HSPs (Lindquist and Craig, 1988; Watson, 1990; Lathigra et al., 1991). For instance, a shift in temperature from 30°C to 37°C results in a two- to 20-fold increase in the synthesis of at least 17 proteins in Escherichia coli (Neihardt et al., 1984). E. coli HSP60 (GroEL) and HSP70 (DnaK) make up 1.6% and 1.4%, respectively, of the total protein content under normal conditions, but increase to approximately 15% of total protein content when a stress is applied (Shinnick, 1991). HSPs act as molecular chaperones in the assembly and folding of proteins, and as proteases when damaged or toxic proteins have to be degraded (Lindquist and Craig, 1988; Lathigra et al., 1991; Ellis, 1996; Ray, 1999). HSPs thus protect cells from the damaging effects associated with stressful conditions. In many micro-organisms, sub-lethal exposure to a stress can confer resistance to a lethal exposure to the same stress (adaptive response) or to other stresses (cross-protection response) (Zeuthen and Howard, 1989; Ellis, 1996; Flahaut et al., 1997). This phenomenon involves an increase in the synthesis of HSPs, which are an integral part of the protective response. Recently, we showed that A. actinomycetemcomitans cells pre-stressed at 43°C were transiently protected against a lethal temperature stress (Goulhen et al., 2003b). This response involved an overexpression of GroEL and DnaK proteins. A sublethal pH stress also transiently protected A. actinomycetemcomitans cells from a lethal pH of 4.5.
(3) HSPs of Oral Micro-organisms
As mentioned above, oral micro-organisms are subjected to a wide range of stresses that may affect their growth and virulence and induce a stress response. The stress response to heat shocks has been studied in several oral micro-organisms. Tables 1 and 2 list the proteins of oral micro-organisms either up- or down-regulated following a heat stress. From that, it appears that the syntheses of GroES, GroEL, DnaK, and HtpG are all up-regulated following a heat stress. Most studies on HSPs produced by oral micro-organisms have involved GroEL, DnaK, and HtpG. A simple adenosine 5′-triphosphate (ATP)-agarose affinity column chromatography procedure has been used to purify A. actinomycetemcomitans, Bacteroides forsythus, P. gingivalis, and Campylobacter rectus GroELs from cellular extracts (Hinode et al., 1995, 1996, 1998b). B. forsythus, P. gingivalis, Streptococcus mutans, Candida albicans, and Prevotella spp. HSP70s (natural or recombinant) have also been purified (Hinode et al., 1995, 1996, 1998a; Jayaraman and Burne, 1995; La Valle et al., 1995; Kadri et al., 1998). In this review, we will focus on the GroES, GroEL, DnaK, and HtpG families, since studies reporting the stress responses of oral micro-organisms related to other HSP families are poorly documented.
(3.1) Gro ES and Gro EL proteins
A. actinomycetemcomitans cells subjected to a heat stress (42 or 50°C) in the presence of radiolabeled amino acids have been shown by autofluorography to up-regulate the synthesis of a major 64-kDa protein belonging to the GroEL family (Koga et al., 1993). This was confirmed by two-dimensional SDS-PAGE (Goulhen et al., 2003a) and Western immunoblotting (Vayssier et al., 1994). Immunolocalization by transmission electron microscopy of A. actinomycetemcomitans GroEL shows that this HSP is located on the cell surface, in surface-associated material, and on outer membrane vesicles produced by the bacterium (Fig. 1) (Kirby et al., 1995; Goulhen et al., 1998; Paju et al., 2000). Native A. actinomycetemcomitans GroEL consists of two stacked heptameric rings (Fig. 2) with 64-kDa subunits (Goulhen et al., 1998). To date, all known GroELs interact with a smaller co-chaperone protein with 10-kDa subunits (GroES). In A. actinomycetemcomitans, the genes coding for GroES and GroEL are arranged in an operon (http://www.genome.ou.edu/act.html). A. actinomycetemcomitans GroEL shares 85% identity with E. coli GroEL (Goulhen, 2001). In addition, the N-terminal amino acid sequence of A. actinomycetemcomitans GroEL shares a high degree of homology with E. coli GroEL (Nakano et al., 1995; Hinode et al., 1998a). A. actinomycetemcomitans GroEL cross-reacts with monoclonal antibodies raised against homologues from Mycobacterium bovis, Mycobacterium leprae, E. coli, and Synechococcus spp. (Løkensgard et al., 1994; Vayssier et al., 1994). Polyclonal antibodies raised against A. actinomycetemcomitans GroEL react strongly with E. coli GroEL and, to a lesser extent, with P. gingivalis and B. forsythus GroELs (Hinode et al., 1998a).
The heat-shock response of P. gingivalis was first examined by Lu and McBride (1994) using one- and two-dimensional SDS-PAGE after labeling the cells with 14C-amino acids. Four proteins were up-regulated (62, 74, 80, and 92 kDa), and one (50 kDa) was down-regulated following a heat stress (Lu and McBride, 1994). Western immunoblotting analysis was used to confirm that P. gingivalis can increase production of GroEL and DnaK when subjected to a temperature shift from 35°C to 43°C (Vayssier et al., 1994). The amino acid sequence of P. gingivalis GroEL shares 59% identity with E. coli GroEL and 58.7% with M. tuberculosis GroEL (Goulhen, 2001). The N-terminal amino acid sequences of the A. actinomycetemcomitans, P. gingivalis, and B. forsythus GroEL proteins share a high degree of homology (Maeda et al., 1994; Nakano et al., 1995; Hinode et al., 1998a). Similarly, polyclonal antibodies raised against P. gingivalis GroEL cross-react with B. forsythus GroEL and, to a lesser extent, with E. coli GroEL and human HSP60 (Hinode et al., 1998a). P. gingivalis GroES shares 42% identity with E. coli GroES (Goulhen, 2001).
B. forsythus GroEL was first studied by Hinode et al.(1998a), who showed that it shares a high degree of similarity (83%) with the 68-kDa P. gingivalis GroEL protein. B. forsythus GroEL shares 50% to 81% identity with the predicted amino acid sequences of the GroELs of several bacterial species (Hinode et al., 1998b). Indeed, the N-terminal amino acid sequences of A. actinomycetemcomitans, P. gingivalis, and B. forsythus GroELs share a high degree of homology (Maeda et al., 1994; Nakano et al., 1995; Hinode et al., 1998a).
The S. mutans stress response was studied by Burne’s group (Jayaraman and Burne, 1995; Jayaraman et al., 1997; Lemos et al., 2001). GroEL is a part of the general stress response of S. mutans, and the genes coding for GroES and GroEL are arranged in an operon that is expressed from a stress-inducible sigma(A)-type promoter located immediately upstream from a CIRCE element. GroEL protein and mRNA levels rise dramatically in cells exposed to a variety of stresses, including acid shock. Induction of stress proteins in S. mutans, including GroEL, following a temperature or acid stress has also been reported by Hrimech et al.(2000).
A few studies have reported that C. rectus and F. nucleatum express GroEL following a stress. Hinode et al.(1998b) studied C. rectus GroEL and showed that it shares 83% homology with Helicobacter pylori GroEL, 54% with A. actinomycetemcomitans GroEL, 54% with E. coli GroEL, and 51% with human HSP60 over the first 35 N-terminal amino acids. The P. gingivalis and A. actinomycetemcomitans GroES proteins share, respectively, 42% and 85% identity with E. coli GroES (Goulhen, 2001). C. rectus GroEL cross-reacts with a monoclonal antibody directed against the 383- to 419-amino-acid residues of human HSP60 (Hinode et al., 1998b). Polyclonal antibodies directed against A. actinomycetemcomitans, B. forsythus, and P. gingivalis GroELs, but not polyclonal antibodies directed against E. coli GroEL, react with C. rectus GroEL (Hinode et al., 1998b). A classic heat-shock response in F. nucleatum was demonstrated by SDS-PAGE and autofluorography (Mayrand et al., 2001; Skar and Bakken, 2001). Among the proteins overexpressed, the 62-kDa protein reacted with antibodies raised against Synechococcus spp. and P. gingivalis GroELs, whereas the 70-kDa protein reacted with antibodies directed against E. coli DnaK.
(3.2) Dna J and Dna K proteins
Yoshida et al.(1997) characterized two open reading frames (ORF) in A. actinomycetemcomitans that code for DnaK and DnaJ. The first ORF codes for a segment of 633 amino acids that has an apparent molecular weight of 68.4 kDa and that shares a high degree of homology with E. coli DnaK. The second ORF codes for a segment of 376 amino acids that has an apparent molecular weight of 41.3 kDa and that shares a high degree of homology with E. coli DnaJ. These results have been confirmed by Minami et al.(1998). The complete amino acid sequence of A. actinomycetemcomitans DnaK shares 84%, 55%, and 48% identity with E. coli, M. tuberculosis, and human homologues, respectively (Yoshida et al., 1997), while the complete amino acid sequence of A. actinomycetemcomitans DnaJ shares 70%, 36%, and 32% identity with the E. coli, M. tuberculosis, and human homologues, respectively (Yoshida et al., 1997). Comparisons of the amino acid sequence alignment of A. actinomycetemcomitans DnaK with the alignment of DnaKs from other bacteria confirm the unique features previously found for other members of the HSP70 family. First, a 27-amino-acid segment is present in the N-terminus domain of Gram-negative bacteria and absent in Gram-positive bacteria. Second, there is a gap of four amino acids in the middle of the DnaK protein sequence, as well as a gap of three amino acids at the C-terminus domain in bacterial DnaK but not in human HSP70. Third, there is a four- to six-amino-acid insertion in the middle of the coding sequence that is unique to prokaryotic HSPs (Minami et al., 1998; section 4 of this article).
N-terminal amino acid sequence analysis of the 75-kDa protein (DnaK) of P. gingivalis showed that it shares significant homology with homologues in both prokaryotic (E. coli [80%] and Chlamydia pneumoniae [90%]) and eukaryotic cell (Drosophila melanogaster [75%]), suggesting that the structure of the N-terminus domain is well-conserved (Hinode et al., 1995). Unlike A. actinomycetemcomitans DnaK, which reacts strongly with monoclonal antibodies against E. coli DnaK (Vayssier et al., 1994), P. gingivalis and B. forsythus DnaKs react poorly (Hinode et al., 1998a). Similarly, antibodies to P. gingivalis and B. forsythus DnaKs react weakly with E. coli DnaK (Hinode et al., 1998a). A. actinomycetemcomitans, P. gingivalis, and B. forsythus DnaK PAbs do not react with recombinant human HSP70 (Hinode et al., 1998a). The first 16 amino acids in the N-terminus of B. forsythus DnaK share a high degree of homology with the sequences of the Haemophilus influenzae, E. coli, and Borrelia burgdorferi homologues. This region also shares a great deal of similarity with P. gingivalis DnaK (87.5%) (Hinode et al., 1998a). F. nucleatum also overexpresses DnaK following a heat stress (Mayrand et al., 2001; Skar and Bakken, 2001).
Stress proteins, including DnaK and GroEL, are also induced in S. mutans by acid and temperature stresses (Jayaraman and Burne, 1995; Hrimech et al., 2000; Lemos et al., 2001). Interestingly, exposure to xylitol but not to other polyols induced a decrease in both DnaK and GroEL levels in xylitol-sensitive strains of S. mutans (Hrimech et al., 2000). It was speculated that the observed reduction (rather than the expected increase) in these HSPs under xylitol exposure contributes to the reduction of the number of mutans streptococci in patients consuming xylitol for caries prevention. A detailed analysis of the transcription of the S. mutans dnaK gene and its regulation under heat and acid shocks has been reported by Burne’s group (Jayaraman et al., 1997; Lemos et al., 2001).
(3.3) HSP90 proteins
The HSP90s of even the most distantly related eukaryotes (human and yeast) shareover 40% identity with E. coli HtpG (HSP90) (Lindquist and Craig, 1988). Lopatin et al.(1999a) found a 68-kDa HSP90 protein in P. gingivalis that cross-reacts with antibodies to human HSP90. They also reported the presence of antibodies in the sera of periodontitis patients that cross-react with P. gingivalis HSP90 (Lopatin et al., 1999b). P. gingivalis HSP90 is mostly membrane-associated, and significant levels of this protein have been detected in extracellular vesicles (Lopatin et al., 1999a, 2000). It has been suggested that membrane-associated HSP90 may act as cell receptors or serve to stabilize or transport bacterial virulence factors. A. actinomycetemcomitans HtpG shares 77% amino acid identity and 83% similarity with E. coli HtpG over the entire length of the ORF (Winston et al., 1996). F. nucleatum also overexpresses HSP90 after a heat stress, as demonstrated by Mayrand et al.(2001) and by Skar and Bakken (2001).
C. albicans HSP90 is present as a 47-kDa protein (heat-stable) both inside and outside the cell (Chaffin et al., 1998). The predicted mass of the protein from the hsp90 gene is 81 kDa (Chaffin et al., 1998). It is interesting to note that homozygous null-HSP90 mutants cannot be constructed, because HSP90 seems to be essential for cell viability (Chaffin et al., 1998). The sera of candidiasis patients recognize both conserved and unconserved epitopes of the 47-kDa protein, which is part of HSP90 (Chaffin et al.,1998).
To conclude this section, we highlight several important features. Various methods have been used to study the stress responses of oral micro-organisms (see Tables 1 and 2). The expression, identification, and characterization of the genes and biological functions of HSPs from oral bacteria have been extensively studied. On the other hand, the location and biological activities of chaperones require further study. Last, the inventory presented here is worthwhile, since the reports are dispersed across multiple subdisciplines.
(4) Phylogenetic Aspects of Oral Microbial HSPs
HSPs are good candidates for evolutionary studies, since they are highly conserved, constitutively expressed, maintained via natural selection for stress resistance, and ubiquitously distributed in prokaryotic and eukaryotic cells (Macario, 1995; Feder, 1999; Feder and Hofmann, 1999). HSPs and regulatory hsp genes such as hrcA (Ahmad et al., 1999) may be of interest in the investigation of phylogenetic relationships among bacteria. hsp genes can also be used to speciate different micro-organisms and thus represent an alternative to DNA-DNA hybridization and 16S RNA analyses (Kwok et al., 1999). The phylogenetic study of HSP families is also of interest, since these ancient genes are now much more diversified, are located in different cellular compartments, and exhibit various functions (Feder and Hofmann, 1999). Viale et al.(1994) proposed that the HSP60 family be used as an evolutionary chronometer, while Goh et al.(1996) suggested that degenerated primers could be used in phylogenetic investigations. This is probably also applicable to HSP90s and HSP70s because of their large size, their high degree of conservation, and their presence in all eubacteria and eukaryotic organelles. Germot and Philippe (1999) also suggested that HSP70s could be reliable phylogenetic markers. Previous phylogenetic analyses revealed that eukaryotic HSPs could be distinguished from prokaryotic HSPs (Boorstein et al., 1994; Rensing and Maier, 1994; Gupta, 1995, 1998; Gribaldo et al., 1999). Phylogenetic analyses of HSP10s have been performed mostly in plants (Waters, 1995).
Although the phylogenetic relationships of HSP60s from various micro-organisms have been previously examined, nothing is known about HSP60s from oral micro-organisms. A global alignment of HSP10, HSP60, HSP70, and HSP90 protein sequences reported in the literature (listed in Table 3) was carried out, and a neighbor-joining distance matrix tree was constructed to evaluate the positions of oral micro-organisms in a global phylogenetic tree. The PHYLIP package software version 3.57 (University of Washington) (Felsenstein, 1993) was used to construct the four phylogenetic trees, with branch lengths proportional to phylogenetic distances. Phylogenetic analyses of HSP10s, HSP60s, HSP70s, and HSP90s from several Gram-positive and Gram-negative bacteria, including several oral species, are presented in Fig. 3. The trees were rooted using the Gram-positive bacteria group, since an analysis of unrooted trees showed a clear branch clustering all Gram-positive bacteria (data not shown). The phylogenetic HSP60 analysis clustered Gram-positive bacteria, chloroplasts and cyanobacteria, and proteobacteria, bacteroides, chlamydiae, and spirochetes in separate groups (Viale et al., 1994). For HSP10, Bacillus subtilis lies in the Gram-negative cluster for HSP10s (Fig. 3, panel A). Bootstrap values for this family were less confident than for the other HSPs.
An analysis of the alignments revealed that HSP families share several features. HSP60s have repeated GGM or GGGM sequences in their C-terminal regions, except for micro-organisms with more than one HSP60 (M. leprae) (Gupta, 1995). Gram-negative bacteria have a conserved, 20- to 25-amino-acid insert in their HSP60s (beginning at position 80 in E. coli) that is not present in the Gram-positive cluster. Bacterial HSP60s and HSP70s share approximately 60% identity with the E. coli sequences, while HSP10s and HSP90s share approximately 40% identity. Gram-positive bacteria lack a single amino acid in the residue 150 region of E. coli HSP60 (Gupta, 1995). All Gram-positive bacterial HSP60s also have a gap in their C-terminal regions.
The affiliation shown for the HSP families in Fig. 3 provides confident bootstrap values. The number of mutations per site indicates that HSP10s and HSP90s are more prone to mutations than HSP60s and HSP70s, suggesting that the HSP60 and HSP70 families were more conserved during evolution. The phylogenetic analyses of HSP10 and HSP60 presented in Fig. 3 are in agreement with the results reported by Gupta (1995) and with a microbial phylogenetic 16S RNA-based analysis (Ribosomal data project II, http://www.cme.msu.edu) clustering Gram-positive bacteria, spirochetes, bacteroides, β-proteobacteria, ε-proteobacteria, and γ-proteobacteria. These clustering patterns, which are similar to 16S RNA trees, were also observed in other studies of mycobacterial HSP60s (Swanson et al., 1996). HSP70 and HSP90 phylogenies also cluster the micro-organisms in a pattern similar to that of 16S RNA trees. P. gingivalis HSP90 is very distant from the HSP90s of other Gram-negative bacteria (Fig. 3, Panel D). P. gingivalis HSP10, HSP60, and HSP 60 are, on the other hand, closely related to the respective HSPs of other Gram-negative bacteria (Fig. 3, Panels A, B, and C, respectively).
The HSPs of oral bacteria and other pathogens possess specific biological activities that might be related to their phylogenetic position. Waters (1995) suggested that the small HSPs in plants diversified not only in function but also in location within the cell. It is interesting to note that Legionella pneumophila, Haemophilus ducreyi, Vibrio cholerae, H. pylori, and A. actinomycetemcomitans, all of which possess HSP60s located outside the cell, are grouped in the same cluster (Fig. 3, Panel B). This was not observed with the 16S RNA clustering analysis. Phylogenetic analyses based on HSP protein sequences may thus be more a reflection of a functional approach. Nevertheless, further studies will have to be performed to evaluate the relationships between the specific phylogenenetic positions and biological activities of HSPs.
(5) Potential Roles of HSPs in Oral Infections
(5.1) Expression of HSPs in diseased periodontal tissues
HSPs may be expressed during periodontal diseases. Ando et al.(1995) reported that inflamed human gingival tissues from periodontitis patients reacted with monoclonal anti-bovine brain HSP70 and monoclonal anti-human HSP60 antibodies. No reactions were found with samples from healthy control patients. Blood vessel endothelial cells, gingival basal cells, and infiltrated inflammatory cells from periodontitis patients also react with monoclonal anti-bovine brain HSP70 and monoclonal anti-human HSP60 antibodies (Ando et al., 1995). Tabeta et al.(2000) reported that gingival tissue extracts from healthy or periodontitis patients contain antibodies to the GroEL protein of P. gingivalis. They also demonstrated that diseased periodontal tissues reacted more strongly to P. gingivalis GroEL and human HSP60 compared with healthy controls. Shelburne et al.(2002) evaluated the gene expression in vivo of P. gingivalis using quantitative reverse-transcription polymerase chain-reaction analysis. They found correlations between probing depth and increased transcription of hsp genes. Yamazaki et al.(2002) demonstrated that periodontitis patients have a higher proliferative response of peripheral blood mononuclear cells to human HSP60 but not to P. gingivalis GroEL compared with healthy subjects. They suggested that the HSP60-reactive T-cells accumulate in periodontitis lesions. Analysis of these observations suggests that bacterial HSPs are produced in periodontal sites and that they may form immunocomplexes with specific antibodies.
(5.2) HSPs as immunodominant antigens
Microbial HSPs appear to be important biological molecules, since they have been implicated in thermotolerance, pathogenicity, immunodominance, and autoimmunity phenomena (Lindquist and Craig, 1988; DeNagel and Pierce, 1993; Young et al., 1993; Kaufmann and Schoel, 1994; Schoel and Kaufmann, 1996; Flahaut et al., 1997; Multhoff et al., 1998; Zügel and Kaufmann, 1999). HSPs are highly immunoreactive proteins (Polla, 1988; Dubois, 1989). The host immune system targets HSPs because (i) they have been highly conserved during evolution (as discussed in section 4), (ii) they are abundant cellular proteins (see section 2), (iii) they are likely presented to the immune system frequently, and (iv) they may act as virulence factors (Shinnick, 1991). The immune system might avoid acting against HSPs because they are highly conserved, but since the biosphere expresses HSPs, the immune system could be misled. Moreover, since HSPs are very abundant during stress conditions and are not exclusively intracellular proteins, as demonstrated with A. actinomycetemcomitans (Goulhen et al., 1998), they may be presented to macrophages as foreign antigens by lymphocytes (Lindquist and Craig, 1988). Some stress proteins are known to be immunodominant antigens of many human pathogens, including etiological agents of malaria (HSP70 and HSP90), schistosomiasis (small HSPs, HSP70, and HSP90), trypanosomiasis (HSP70 and HSP90), filariasis (HSP70), syphilis (HSP60), tuberculosis (small HSPs, HSP10, HSP70, and HSP90), leprosy (small HSPs, HSP10, HSP70, and HSP90), Legionnaire’s disease (HSP60), Lyme disease (HSP60 and HSP70), and Q fever (HSP60) (Shinnick, 1991; Schoel and Kaufmann, 1996). Moreover, HSPs might act as danger-signaling molecules by carrying peptides to elicit antigenic specificity. Thus, some HSPs could activate cells of the innate immune system (Wallin et al., 2002) (this will be discussed in section 5.3).
The first immunodominant HSP antigen identified in patients affected by candidiasis was a 47-kDa protein, which is part of the HSP90 of C. albicans (Chaffin et al., 1998). Antibodies to this protein have been found in the sera of candidiasis and AIDS patients (Chaffin et al., 1998). In addition, sera from patients with oral or esophageal candidiasis react with heat-inducible and ATP-binding proteins of C. albicans, which are probably HSPs (Swoboda et al., 1993; Chaffin et al., 1998). Patients with candidiasis have also been shown to have higher titers of IgA directed against mycobacterial GroEL than do healthy subjects (Ivanyi and Wong, 1992; Chaffin et al., 1998).
Several streptococcal species (S. sanguis, S. pyogenes, S. faecalis, and S. salivarius) that share highly conserved HSPs (Mizushima, 1988; Watson, 1990; Yoshikawa et al., 1996) have been implicated in the etiology of Behçet’s disease, a multisystem disorder presenting recurrent oral and genital ulcerations (Lehner et al., 1991; Lehner, 1997). The sera from patients with Behçet’s disease recognize mycobacterial HSPs (65 to 70 kDa) (Lehner et al., 1991) and contain high levels of anti-S. pyogenes and anti-retinoblastoma cell line GroEL antibodies, compared with healthy controls (Tanaka et al., 1999). Lymphocytes from patients with Behçet’s disease can be stimulated by four peptides derived from mycobacterial GroEL that have been identified by T-cell mapping (Lehner, 1997) and that share strong homology with human HSP60 (Lehner, 1997). Last, patients with oral or vulvovaginal candidiasis lesions have stronger responses to mycobacterial HSP65 compared with healthy subjects and patients without lesions (Ivanyi and Ivanyi, 1990; Ivanyi and Wong, 1992).
Several groups (Koga et al., 1993; Meghji et al., 1993; Tabeta et al., 2001) showed that A. actinomycetemcomitans GroEL is an immunodominant antigen in humans. Sera from ten healthy individuals did not react with A. actinomycetemcomitans GroEL, whereas sera from nine of 29 patients with periodontitis reacted strongly (Koga et al., 1993). Ando et al.(1995) also demonstrated that serum from a periodontitis patient contained antibodies that reacted with A. actinomycetemcomitans, F. nucleatum, and P. nigrescens HSP60 and/or HSP70, suggesting that these proteins may be involved in the pathogenesis of periodontitis. Inbred strains of mice immunized with sonic extracts of A. actinomycetemcomitans showed a primary immune response against a 65-kDa protein, which is likely GroEL (Nitta and Ishikawa, 1993). This protein also strongly reacted with sera from early-onset periodontitis patients (Nitta and Ishikawa, 1993). Patients with localized juvenile periodontitis had higher antibody titers to A. actinomycetemcomitans surface-associated material, which contains GroEL, than did patients with severe generalized periodontitis, while those with severe generalized periodontitis had higher antibody titers to surface-associated material from P. gingivalis (Meghji et al., 1993). Similarly, Maeda et al.(1994) showed that sera from three of nine healthy subjects and eight of ten periodontitis patients reacted with recombinant P. gingivalis HSP60. The positive reactions obtained with healthy subjects might be due to cross-reactivity with antibodies directed against HSP60s of other bacteria (oral and non-oral) that may have infected the patients in the past (Tabeta et al., 2001). Maeda et al.(2000) reported that sera from periodontitis patients all reacted with truncated P. gingivalis HSP60, and epitopes were recognized along the entire protein. Tabeta et al.(2000) showed that periodontitis patients had a higher antibody titer against P. gingivalis GroEL than did healthy subjects.
Lopatin et al.(1999b) demonstrated that patients with healthy periodontal tissues had higher antibody titers to several HSPs (human HSP90, bacterial GroEL and DnaK) than did periodontitis patients, suggesting protective role associated with these antibodies. They also found a significant association between increased colonization of periodontal pockets by P. gingivalis, T. denticola, and B. forsythus, but not A. actinomycetemcomitans, and anti-HSP90 antibody concentrations (Lopatin et al., 1999b). Schett et al.(1997) reported higher mycobacterial, E. coli, and human HSP60 salivary antibody concentrations (IgA predominance) in gingivitis patients compared with healthy and periodontitis patients. Petit et al.(1999) suggested that the higher responsiveness to HSP60 and HSP70 observed in gingivitis subjects may prevent the conversion from gingivitis to periodontitis.
Sims et al.(2002) showed that insulin-dependent mellitus (IDDM) patients with periodontitis had higher antibody titers to commercial HSP90 and HSP70 compared with control patients, whether diabetic or not. They did not observe any differences in HSP60 titers. Our group also evaluated the immunological responses of periodontitis, non-insulin-dependent diabetic (NIDD), periodontitis-NIDD, Sjögren’s, and periodontitis-Sjögren’s patients to HSPs purified from a range of periodontopathogens. Briefly, the combination of diabetes and periodontitis was associated with a lower immunological response to P. gingivalis HSP60 compared with the healthy group. Also, Sjögren’s patients had weaker immunological responses to P. gingivalis GroEL, while diabetics patients had stronger responses to P. gingivalis GroEL, but only when they also had periodontitis (Goulhen, 2001). To conclude, patients with systemic diseases had immunological responses to HSPs.
(5.3) Microbial HSPs as virulence factors
The GroELs and, to a lesser extent, the DnaKs of several pathogenic bacteria may become major antigens, since they are strongly expressed under stressful conditions. These HSPs are antigenic, and the recognition of specific epitopes on such highly conserved antigens may contribute to protective immunity or may have pathological autoimmune consequences (Dubois, 1989; Res et al., 1991; Shinnick, 1991; Yamaguchi et al., 1992; Tabeta et al., 2000). Because of the similarities between microbial and mammalian HSPs, a humoral response against microbial HSPs may be destructive for the host due to antigen mimicry leading to an autoimmune response. Three models have been proposed to link microbial infections to subsequent autoimmune reactions involving HSPs (Rose, 1998). These models are based on (i) molecular mimicry between microbial HSPs and HSPs or constitutive proteins from the host, (ii) inflammation-induced exposure of cryptic cell epitopes that could be a target for immune reactions, and (iii) antigen persistence in infected sites leading to chronic immunological reactions. For a long time, HSP autoimmune responses were thought to be the result of cross-reactivity between bacterial and host HSPs. Recently, Srivastava et al.(1998) discovered that HSPs could act as carriers of antigenic peptides derived from tumors or virus-infected cells. These HSP-peptide complexes “shuttle” antigenic peptides to the MHC class I presentation pathway of antigen-presenting cells. Wallin et al.(2002) reported that HSPs also activate the innate immune system.
Molecular mimicry has been well-documented. Molecular mimicry between bacterial and human HSPs may allow micro-organisms to evade the host defenses (Dubois, 1989). The immune response to HSPs might be directed against autologous molecules and involve T-lymphocytes if common epitopes are presented by MHC class II molecules (Aguas et al., 1990). Yamazaki et al.(2002) showed that HSP60-reactive T-cells accumulate in the gingival tissues of periodontitis patients and that this response is inhibited by anti-MHC class II antibodies. Lo et al.(2000) demonstrated that MHC class Ib molecules are involved in autoimmune infections with Gram-negative bacteria and identified an immunodominant epitope (GMQFDRGYL) derived from the HSP60 family that is present in Salmonella typhimurium, Salmonella typhi, E. coli, Y. enterolitica, Klebsiella pneumoniae, and H. pylori. A database analysis indicated that this peptide sequence pattern is also present in the HSP60s of oral micro-organisms such as P. gingivalis, A. actinomycetemcomitans, B. forsythus, C. albicans, and S. mutans, as well as in human HSP60 (Goulhen, 2001). Moreover, interactions of non-oral HSPs with the immune system might play a role in activating responses against new infections caused by oral bacteria. This is an interesting possibility and should be investigated. Molecular mimicry between bacterial GroELs and human HSP60 or constitutive human proteins may lead to autoimmune diseases (De Graef-Meeder et al., 1991; Hotokezaka et al., 1994; Handley et al., 1996; Ueki et al., 2002). Tanaka et al.(1999) reported that HSP60s from S. pyogenes, a retinoblasma cell line, a bovine retinal extract, and Y. enterolitica have common antigenic features but react differently when tested against the sera of patients affected by Behçet’s disease, suggesting cross-reactivity of host and pathogen HSP60s. Mycobacterial HSP65 sensitizes γδT-cells from patients (Haregewoin et al., 1989; O’Brien et al., 1989) with Behçet’s disease (Hamzaoui et al., 1994). Autoimmunity and antigenic cross-reactivity of bacterial and human HSPs may be involved in the progression of Behçet’s syndrome (Lehner et al., 1991; Pervin et al., 1993; Yoshikawa et al., 1996; Tanaka et al., 1999). Schett et al.(1997) reported that saliva from patients with gingivitis contains higher levels of anti-mycobacterial GroEL (HSP65) antibodies (mostly the IgA isotype) than does the saliva of healthy people and patients with periodontitis. Schett et al.(1997) also analyzed epitope specificity to recombinant mycobacterial GroEL (HSP65) and discovered six distinct binding sequences, two of which (171–195 and 491–515) are highly reactive. The tetrapeptide KDLL found in mycobacterial GroEL protein is also present in human proteins such as lactoferrin, transferrin, and MHC class II molecules (Aguas et al., 1990). The primary structure of human HSP60 is similar to the 545- to 555-amino-acid region of human cytokeratin, a characteristic that may be at the origin of autoimmune reactions (Jones et al., 1993). Hotokezaka et al.(1994) compared the amino acid sequences of P. gingivalis GroES and GroEL with that of human collagen. Seven glycine-rich homologous regions were found in P. gingivalis GroEL and human type 1 and type III collagen. Once again, these similarities may be involved in immunologic reactions leading to tissue destruction. Polyclonal antibodies raised against C. albicans receptors for laminin and fibrinogen cross-react with ubiquitin, which plays a role in protein degradation and stress resistance (Sepulveda et al., 1996). The ubiquitin-like epitopes in these receptors may allow ubiquitin to play a role in modulating the activity of the receptors and in interactions between C. albicans and host structures.
The 61- to 65-amino-acid sequence (EKRAA) in A. actinomycetemcomitans DnaJ corresponds to a highly conserved region represented by QKRAA in E. coli (Yoshida et al., 1997). Interestingly, approximately 90% of patients with severe rheumatoid arthritis have HLA-DR haplotypes that contain the QKRAA pattern (Albani et al., 1995). The immune response to A. actinomycetemcomitans DnaJ may thus contribute to periodontal diseases (Yoshida et al., 1997). SDS-PAGE/Western immunoblotting and enzyme-linked immunosorbent assay (ELISA) analyses recently performed in our laboratory showed that a polyclonal antibody directed against purified A. actinomycetemcomitans GroEL cross-reacted with human fibronectin but not with other human proteins, including type IV collagen (Yoshioka et al., 2004) (Fig. 4). On the other hand, a polyclonal antibody directed against E. coli GroEL and a monoclonal antibody directed against human HSP60 did not cross-react with fibronectin. A comparative analysis of the amino acid sequences of A. actinomycetemcomitans GroEL and fibronectin revealed the presence of eight instances of four-amino-acid sequence homology. Six of these tetrapeptide sequences are also shared with E. coli GroEL and three with human HSP60, suggesting that the remaining two tetrapeptides, GQLI and TGLE, may be associated with the epitope that the anti-A. actinomycetemcomitans GroEL antibody specifically recognizes. This molecular mimicry between A. actinomycetemcomitans GroEL and fibronectin may lead to an autoimmune response and contribute to tissue destruction during periodontitis.
Cross-reactive immune responses cannot be explained solely by an association between HSPs and inflammatory conditions. The effects of HSPs on the innate immune system should thus not be overlooked. HSPs activate the immune inflammation network. C. rectus GroEL stimulates the production of both interleukin-6 (IL-6) and interleukin-8 (IL-8) by human gingival fibroblasts without affecting their viability (Hinode et al., 1998b). C. rectus GroEL can also stimulate the production of IL-6 by a confluent monolayer of human gingival epithelial cells and is cytotoxic when used at high concentrations (Tanabe et al., in press). T-cells need help or co-stimulation to be activated. The antigen must be presented by antigen-presenting cells (pAPCs) such as dendritic cells (Banchereau and Steinman, 1998). T-cells are present during periodontal disease and in gingivitis inflammatory infiltrates and require the antigen to be presented by APCs (Gemmell et al., 2002a). More particularly, P. gingivalis-specific T-cells produce cytokines, regardless of the APC population (Gemmell et al., 2002b). pAPCs interact with microbial components and can be activated by endogenous substances released from stressed tissues (Janeway, 1992; Matzinger, 1998). HSPs are potent molecules that signal tissue damage and cell stress to the immune system (Wallin et al., 2002). HSP60 seems to be the best activator of human monocytes and dendritic cells (Bethke et al., 2002). A model presented by Costa et al.(2002) to evaluate the role of HSP60 in the stimulation of innate immune cells by Chlamydia would also be useful in oral diseases.
Bacterial HSPs may have deleterious effects on host cells. Low concentrations of native A. actinomycetemcomitans GroEL promote epithelial cell proliferation, while high concentrations produce toxic effects (Goulhen et al., 1998). A. actinomycetemcomitans GroEL may also increase the proliferation of human skin keratinocytes (HaCaT cell line) through sustained activation of ERK1 and two mitogen-activated protein (MAP) kinases (Zhang et al., 2001). Recombinant human HSP60 has no effect on HaCaT cell proliferation or MAP kinase phosphorylation, indicating that the proliferation-promoting effect does not result from the conserved protein sequences but rather is unique to the bacterial GroEL protein (Zhang et al., 2001). This specific effect may be related to the composition and conformation of A. actinomycetemcomitans GroEL. Paju et al.(2000) reported that the potential to promote the proliferation of periodontal ligament epithelial cells is strongly correlated with high levels of A. actinomycetemcomitans GroEL in membrane fractions. Saline-extracted surface-associated A. actinomycetemcomitans material has strong osteolytic activity that is 10- to 100-fold greater than that of lipopolysaccharides or surface-associated material from other periodontopathogens, such as C. rectus, P. intermedia, and P. gingivalis (Wilson et al., 1985; Meghji et al., 1992, 1994; Kirby et al., 1995; Reddi et al., 1995). A. actinomycetemcomitans GroEL may be the active moiety, even though it constitutes only 2% of the dry mass of surface-associated material (Kirby et al., 1995).
(6) Conclusions
In recent years, several studies have reported the characterization of HSPs produced by oral micro-organisms. Like the HSPs involved in other diseases, these stress-induced proteins may contribute to the pathogenic process of oral infections. Although some evidence indicates that HSPs are expressed in vivo by periodontopathogens in diseased periodontal sites and tissues, additional studies are needed. In addition, the contribution of A. actinomycetemcomitans GroEL to the adherence and invasion of mammalian cells such as epithelial cells remains to be investigated. This is particularly relevant since Garduño et al.(1998) recently reported that an L. pneumophila cell-surface GroEL mediates adherence and invasion in a HeLa cell model. For the importance of periodontopathogenic HSPs as virulence factors to be demonstrated, individual HSP-negative mutants must be constructed. Studies carried out in our laboratory have indicated that it is very difficult to obtain such mutants by insertional mutagenesis. It may be more appropriate to construct temperature-sensitive mutants to test their ability to survive in vivo (Lathigra et al., 1991).
It would also be interesting to evaluate the importance of cross-reactive epitopes in the HSPs of oral pathogens by mapping specific regions of the HSPs and evaluating humoral responses in animal models. The immune response to a cross-reactive epitope in H. pylori HSP60 is lower in infected than in healthy patients (Yamaguchi et al., 2000). On the other hand, infected patients had a higher antibody response to H. pylori HSP60. This was not observed for E. coli HSP60, suggesting that the humoral response is not induced by other bacteria. Humoral responses to specific HSP epitopes might thus be different from responses to the whole molecule. Specific epitopes might also be associated with protection against H. pylori infections, given that pre-immunized mice are protected (Yamaguchi et al., 2000). Interestingly, our phylogenetic tree shows that this epitope is highly conserved in all the micro-organisms investigated, including A. actinomycetemcomitans (100% match), P. gingivalis (100% match), S. mutans (87% match), and B. forsythus (93% match).
HSPs are now considered as key elements in the pathogenesis of several infections, and they are emerging as important diagnostic markers (Macario, 1995). Mycobacterial GroEL (HSP65) has a high specificity as a marker for Behçet’s disease and may be of interest in the development of a diagnostic test (Hasan et al., 1996). Interestingly, four peptides derived from this HSP and identified by T-cell mapping of patients affected by Behçet’s disease cross-react with human HSP60 and may also be of interest in the development of a diagnostic test (Lehner, 1997). Since the saliva of patients with gingivitis contains more anti-mycobacterial GroEL antibodies (mostly IgA isotype) than does saliva from healthy subjects or patients with periodontitis, Schett et al.(1997) suggested measuring anti-GroEL antibodies in saliva as a way of diagnosing gingivitis. Lopatin et al.(1999b) suggested that tests be developed to evaluate the risk of periodontitis and that vaccines based on HSP epitopes be developed, since anti-HSP antibodies appear to be protective. However, such vaccines should be used with great caution, since they could lead to autoimmune diseases, as has been the case with Lyme disease. Indeed, Trollmo et al.(2001) reported that a cross-reaction between B. burgdorferi outer surface protein A and human LFA-1a(L) at the T-cell level leads to autoimmune disorders.
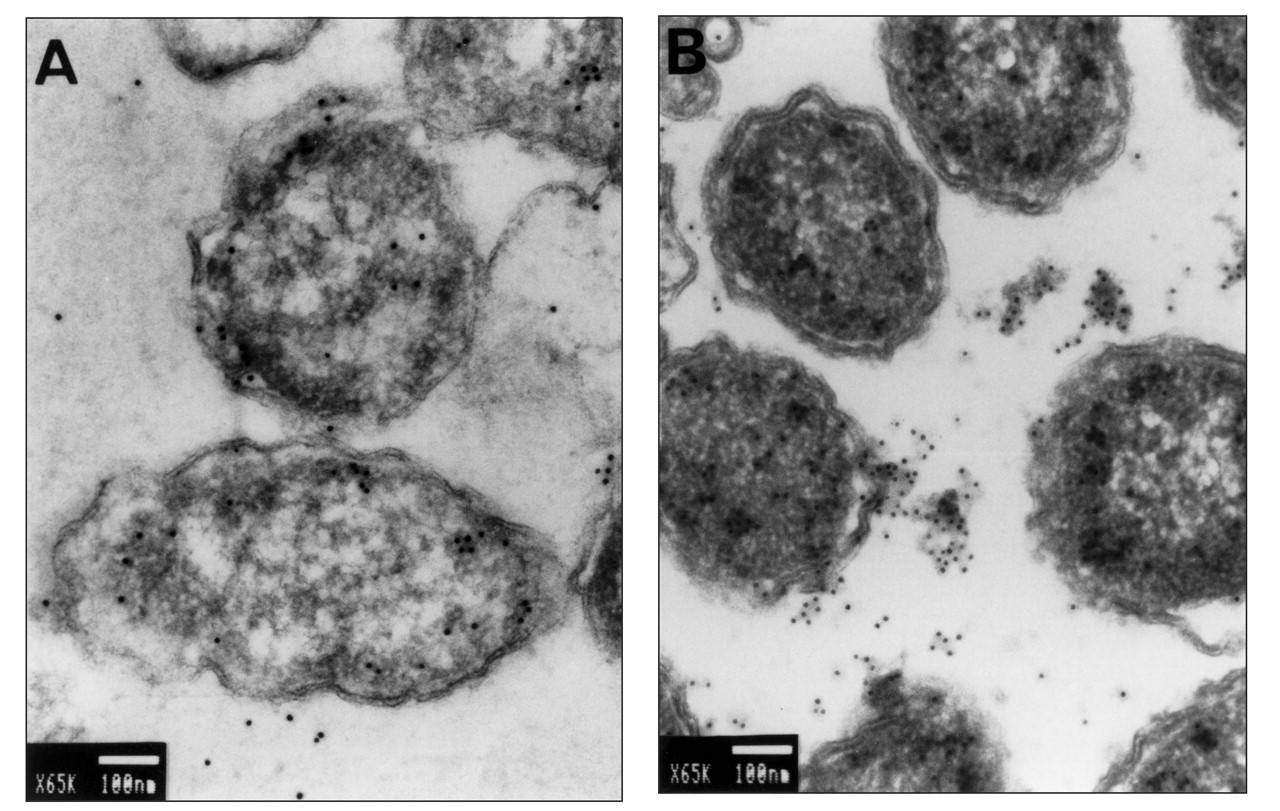
Immunolocalization by transmission electron microscopy of GroEL in ultrathin sections of A. actinomycetemcomitans ATCC 29522. Unstressed cells
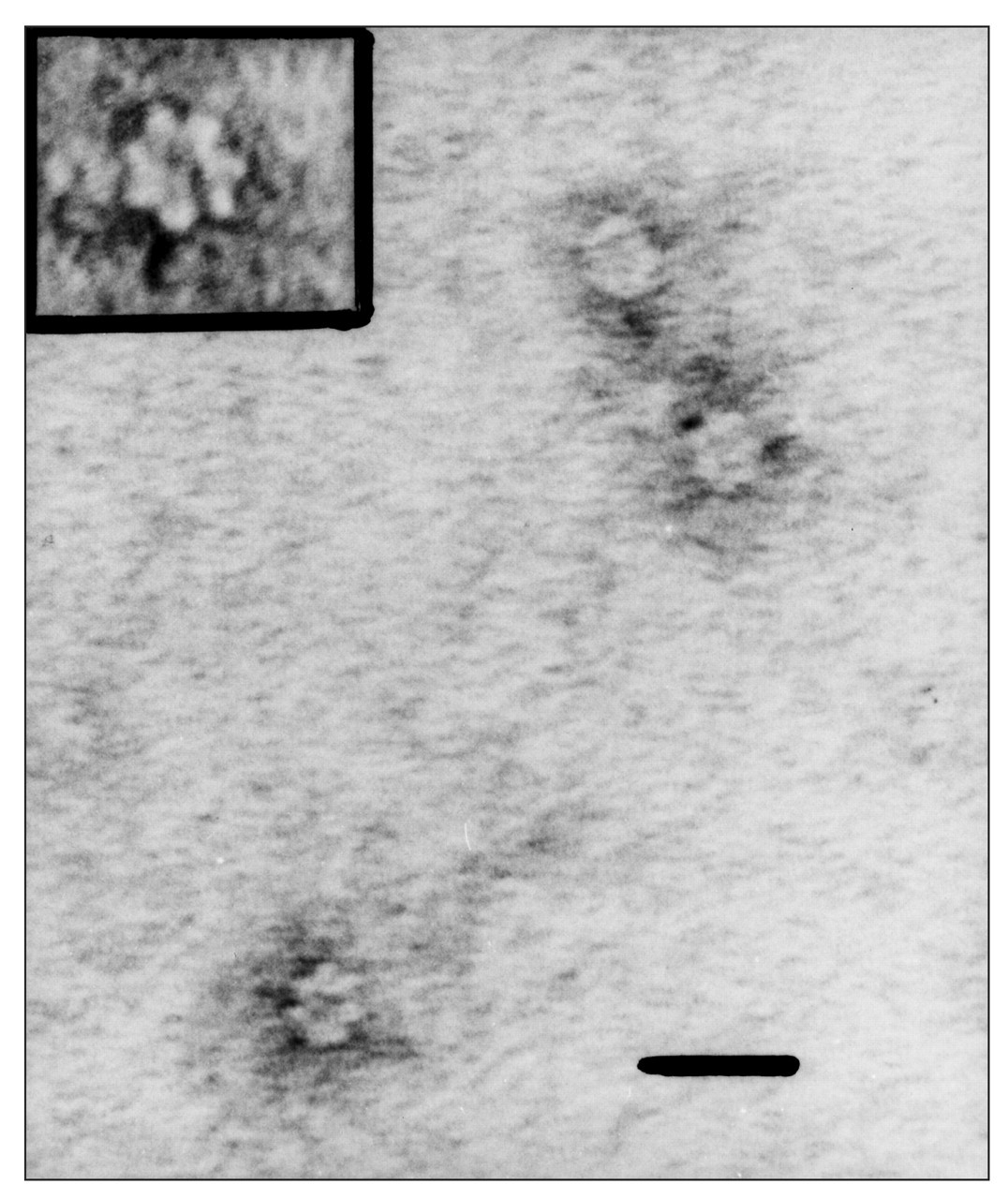
Transmission electron micrograph of the purified native A. actinomycetemcomitans ATCC 29522 GroEL stained with uranyl acetate. The heptameric ring can be seen in the enlargement. Bar = 20 nm.
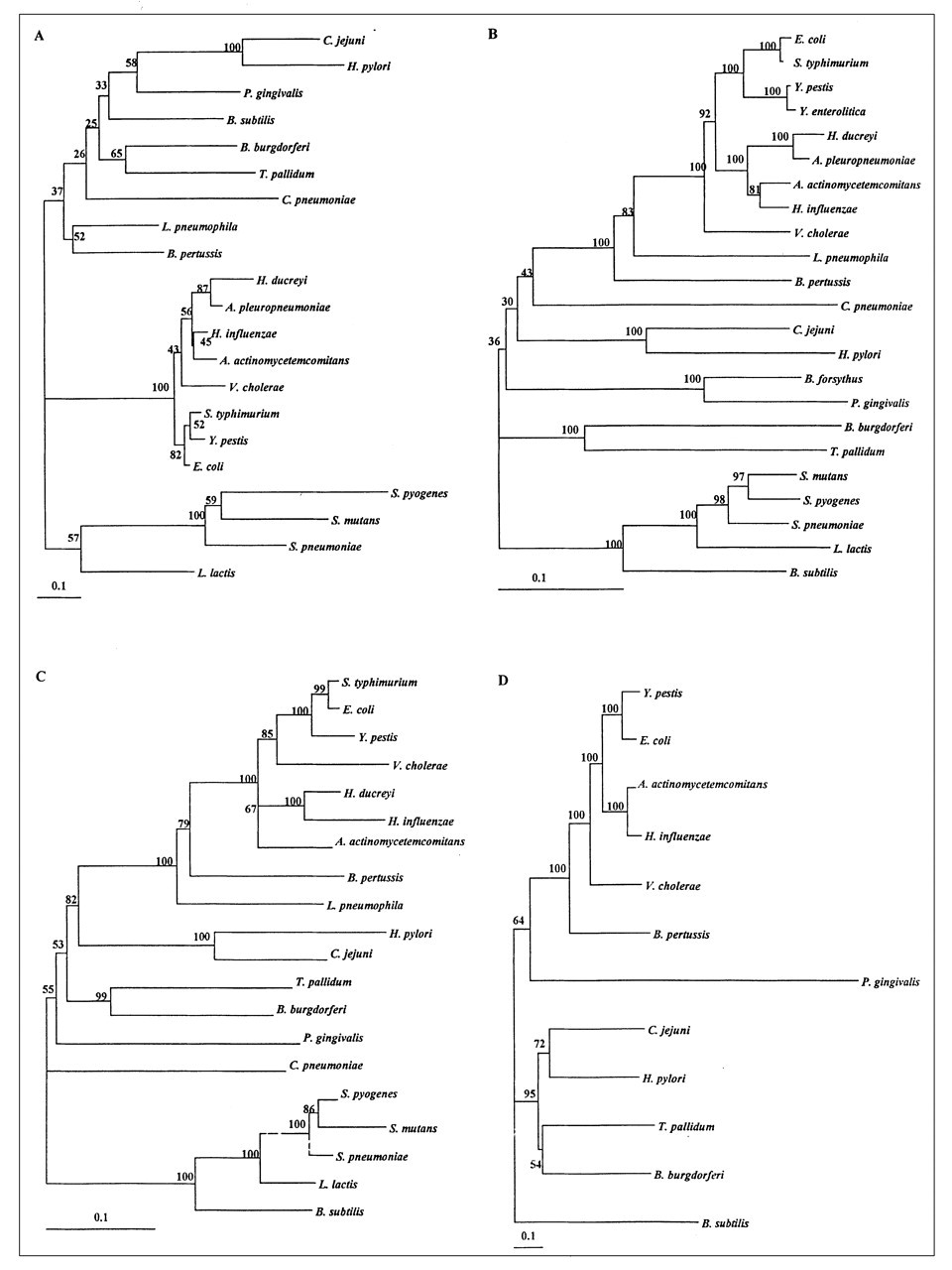
Phylograms of HSP10
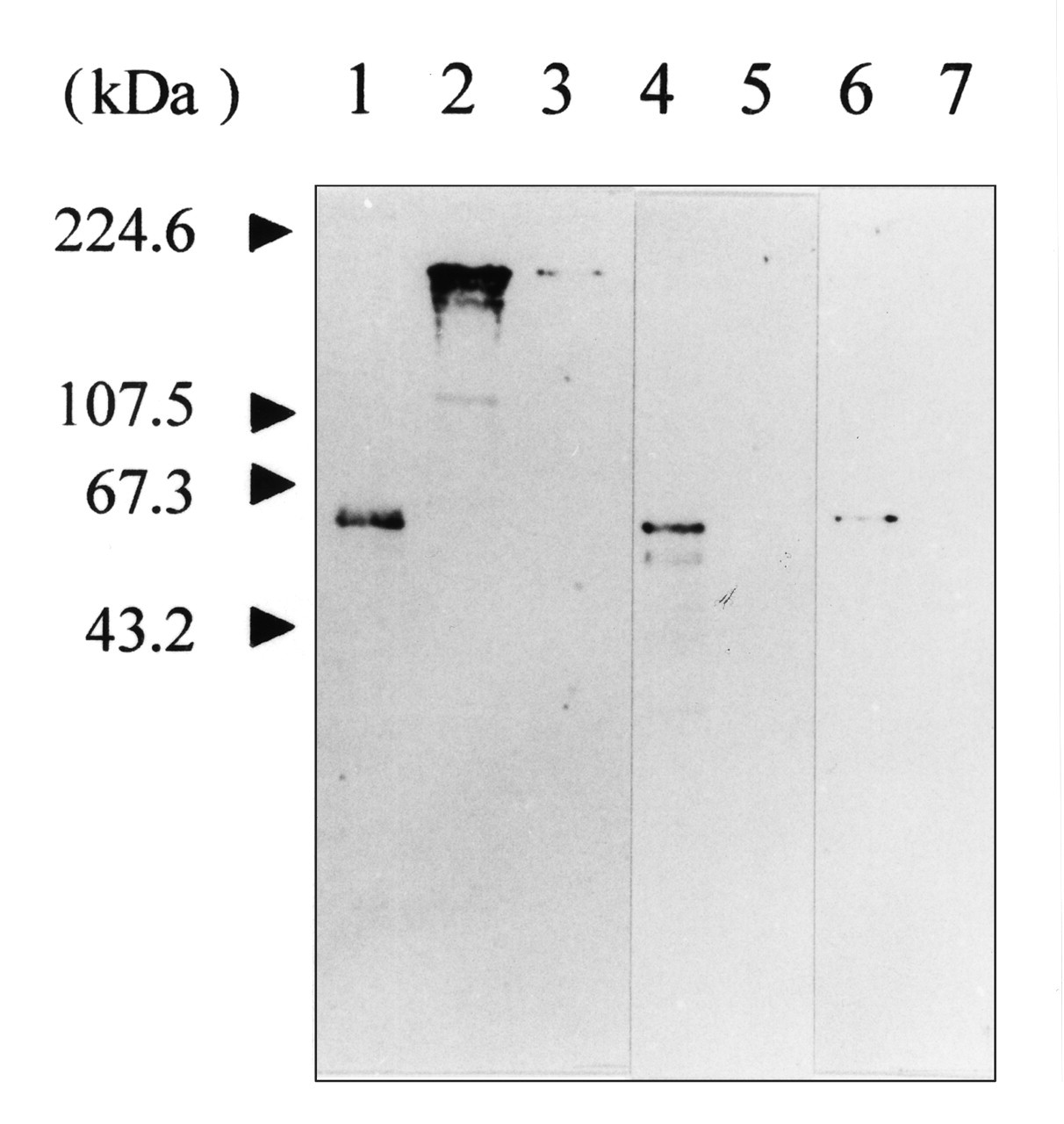
Immunoreactivity of human fibronectin with antibodies directed against bacterial GroEL and human HSP60.
Footnotes
Acknowledgements
We thank Dr. Dennis Lopatin and Dr. Fatiha Chandad for reading the manuscript and providing helpful suggestions and comments, and Dr. Claude Lemieux for his help with the phylogenetic analysis. F. Goulhen was a recipient of a scholarship from the Fonds pour la Formation de Chercheur et l’Aide à la Recherche (FCAR). The authors’ research is supported by grants from the Canadian Institutes of Health Research (CIHR) and the Fonds de la Recherche en Santé du Québec (FRSQ).
