Abstract
Cysteine peptidases (CPs) are phylogenetically ubiquitous enzymes that can be classified into clans of evolutionarily independent proteins based on the structural organization of the active site. In mammals, two of the major clans represented in the genome are: the CA clan, whose members share a structure and evolutionary history with papain; and the CD clan, which includes the legumains and caspases. This review focuses on the properties of these enzymes, with an emphasis on their potential roles in the oral cavity. The human genome encodes at least (but possibly no more than) 11 distinct enzymes, called cathepsins, that are members of the papain family C1A. Ten of these are present in rodents, which also carry additional genes encoding other cathepsins and cathepsin-like proteins. Human cathepsins are best known from the ubiquitously expressed lysosomal cathepsins B, H, and L, and dipeptidyl peptidase I (DPP I), which until recently were considered to mediate primarily “housekeeping” functions in the cell. However, mutations in DPP I have now been shown to underlie Papillon-Lefèvre syndrome and pre-pubertal periodontitis. Other cathepsins are involved in tissue-specific functions such as bone remodeling, but relatively little is known about the functions of several recently discovered enzymes. Collectively, CPs participate in multiple host systems that are active in health and in disease. They are involved in tissue remodeling and turnover of the extracellular matrix, immune system function, and modulation and alteration of cell function. Intracellularly, CPs function in diverse processes including normal protein turnover, antigen and proprotein processing, and apoptosis. Extracellularly, they can contribute directly to the degradation of foreign proteins and the extracellular matrix. However, CPs can also participate in proteolytic cascades that amplify the degradative capacity, potentially leading to pathological damage, and facilitating the penetration of tissues by cancer cells. We know relatively little regarding the role of human CPs in the oral cavity in health or disease. Most studies to date have focused on the potential use of the lysosomal enzymes as markers for periodontal disease activity. Human saliva contains high levels of cystatins, which are potent CP inhibitors. Although these proteins are presumed to serve a protective function, their in vivo targets are unknown, and it remains to be discovered whether they serve to control any human CP activity.
(I) Introduction
As evidenced by the deleterious effects of salivary insufficiency, saliva is important in maintaining oral health, and it is largely taken for granted that much of this protective activity is mediated by the variety of different salivary proteins (reviewed in Schenkels et al., 1995). The salivary cystatin proteins (reviewed in Bobek and Levine, 1992; Henskens et al., 1996) are abundant salivary (and tear) inhibitors of CPs, a phylogenetically ubiquitous and diverse class of peptidase. Thus, it is generally assumed that one function of salivary cystatins in vivo is to provide protection in the oral cavity by inhibiting CPs. However, the in vivo target(s) for salivary cystatins has yet to be identified, and to date, no disease has been reported that is associated with a defect in a salivary cystatin gene. Therefore, the biological function of salivary cystatins can be inferred only from circumstantial data. What are potential sources of exposure of oral and nasopharyngeal tissues to CPs, and the consequences that could warrant a protective mechanism?
The number of identified mammalian CPs has grown considerably in the past few years. The purpose of this review is to summarize the salient features of these enzymes, concentrating on known (or potential) functions, and to indicate relevance to oral tissues. In addition, the potential for interactions with other peptidases (and their inhibitors) to form proteolytic networks will be examined. It is important to note that while some endogenous CPs have been shown to be involved in processes such as inflammation, antigen presentation, bone remodeling, and cancer, in only a few cases has their role been examined in oral tissues. Further, there are several CPs whose function remains to be established, but whose properties warrant investigation of potential oral sources. Exogenous sources of CPs (e.g., from pathogens) and the cystatins themselves will be the subjects of a separate review.
(A) Overview of peptidases
Enzymatic cleavage of peptide bonds is fundamental to almost every aspect of life, and peptidases represent about 2% of all gene products. Examples can be found in digestion, blood coagulation and fibrinolysis, processing of preproproteins such as collagen, immune function, development, and apoptosis (reviewed in Twining, 1994). Not surprisingly, peptidase genes are found in the genomes of all cellular organisms (and several types of virus), and there are arrays of proteolytic enzymes distributed in cellular and tissue compartments. Peptide bond scission proceeds via a nucleophilic attack on the carbonyl carbon, followed by a general acid-base hydrolysis. The general term peptidase is preferred for enzymes that catalyze this reaction, although protease and proteinase are commonly found in the literature. Peptidases are generally grouped into five major types (cysteine, serine, threonine, aspartate, and metalloproteinase) according to the mechanism used to generate the nucleophile in the active site (reviewed in Barrett et al., 1998). In the CPs, an activated cysteine residue is used as the nucleophile and a histidine residue as the proton donor. In some enzymes, a third residue serves to orient the His residue.
The CPs comprise a complex set of enzymes (reviewed in Rawlings and Barrett, 1994; Kirschke et al., 1995; Chapman et al., 1997; Barrett et al., 1998). CPs of various physical and biochemical types are found in all kingdoms of organisms, indicating that they are among the most ancient proteins. Phylogenetic analysis is proving to be a powerful tool for elucidating the relationships between and among the large numbers of CP-like protein sequences that have been identified thus far, and phylogeny now provides the foundation for classifying the CPs (and other peptidases) (Rawlings and Barrett, 1993, 1994; see Barrett et al., 1998, for a detailed compilation of peptidases using this scheme, and detailed descriptions of their enzymatic properties). However, in the literature, descriptions of the different groups of CPs and their relationships to each other are complicated by differences in usage of the hierarchical phylogenetic terms class, superfamily, family, and subfamily. For example, the latter three terms have all been applied to papain-like cysteine peptidases (Karrer et al., 1993; Rawlings and Barrett, 1993; Berti and Storer, 1995). Used in a phylogenetic context (that is, where evolutionary relationships are considered), these terms implicitly mean a grouping based on the position of a node on a phylogenetic tree that links all members of the group, and reflects the degree of divergence of that group from other groups. Strictly, a family of proteins (or matching domains of chimeric proteins) should be a monophyletic group—that is, the members share a most recent common ancestor that is not an ancestor of one or more proteins not included in the group. The term subfamily is often used to denote clear divisions within a family, such as functional differences in proteins of otherwise similar sequences. The term superfamily is often used to denote groups of proteins that have relatively low (but detectable) sequence similarity to each other, but which have structural and functional properties consistent with a common evolutionary origin. In the system of Rawlings and Barrett, the term family is used to denote recognizable groups such as the papain-like enzymes. In a family, every member has a statistically significant relationship to at least one other member of the family (at least in the sequence of residues involved in catalytic activity), which implies evolution from a common ancestral peptidase. Deep divergences within the phylogenetic tree generally warrant the designation of the main branches as subfamilies. The term clan denotes groups of families for which there are indications of evolutionary relationships (e.g., active site arrangements, common three-dimensional structures) but which lack statistically significant similarities in sequence. CP clans are designated CA, CD, etc., and families by C+number (e.g., C12).
(B) Overview of CPs
From sequence and structural comparisons, it is clear that several types of CPs had independent origins, and there are at least six distinct clans consisting of 43 families, of which over half are from viruses (Rawlings and Barrett, 2000). Most and perhaps all clans represent convergent evolution, since in the majority of cases the enzymes have been shown to have a distinctly different organization of catalytic residues. Enzymes in clan CA (papain-related) have the catalytic residues in the order Cys....His....Asn/Asp, while enzymes in clan CD (legumain-related) have an active site catalytic dyad with a His-Gly-spacer-Ala-Cys motif (Chen JM et al., 1998). In the oral cavity and the nasopharynx, the substantial majority of CPs encountered—whether from the host, from viruses, bacteria, parasitic protozoa or helminths, or the diet—are from these two clans.
CPs are best known from members of the CA clan that were purified by classic biochemical techniques: the plant enzyme papain and the related mammalian lysosomal cathepsins B, H, L, and DPP I. (It should be noted that the term cathepsin is a general term for a peptidase, especially lysosomal, with an acidic pH optimum that is involved in protein degradation, and is not restricted to peptidases of any one catalytic type. For example, cathepsin D is an aspartic peptidase.) More recently, molecular cloning has been used to identify other related mammalian CP cathepsins—primarily human—and now the human genome is known to contain at least 11 related, but distinct, CP cathepsins: B, F, H, K, L, O, S, V, X, DPP I, and lymphopain. At this time, a search of the near-complete human genome database does not reveal any additional functional genes (data not shown), and thus these cathepsins may represent the entire human complement of enzymes most closely related to papain. Most of the 11 CP cathepsins evolved relatively early and are present in all mammals. The literature concerning these proteins has been greatly complicated by the use of the same letter to designate different proteins, and different letters to designate the same protein. Some of the various designations, and presently accepted or recommended ones, are summarized in the Table, together with the human chromosomal locations. Amino acid identity between and among these mammalian cathepsins is high in the vicinity of the active site, but overall levels of similarity generally range from 20 to 60% (Wiederanders et al., 1992). However, the recently evolved cathepsin V shares 78% similarity with cathepsin L (see below). Other cathepsin activities have been described in the literature, but their relationship to the 11 listed above is uncertain (Kirschke et al., 1995). In addition to other, evolutionarily independent, types of CPs in the human genome (e.g., clan CD enzymes), there are other CPs, such as cancer procoagulant, that remain to be characterized.
(C) Regulation of CPs and other peptidases
It is axiomatic that peptidases represent potentially dangerous enzyme activities that must be subject to strict control and containment within appropriate compartments, and that a failure in these controls could lead to pathology. Several mechanisms are used to regulate peptidase activity, in addition to transcriptional and post-transcriptional controls (Twining, 1994; Chapman et al., 1997). Most enzymes are synthesized as inactive zymogens that must be activated by proteolytic cleavage. This may be autocatalytic under specific conditions, such as low pH. Release of peptidases from a cell is generally a controlled process. The CPs are readily inactivated by oxidation of the active site cysteine and require a reducing environment for full activity. Many human CPs are unstable at neutral pH and require an acidic pH for full activity. Once activated, enzyme activity can be lost by degradation. A major control governing peptidase activity is the presence of protein inhibitors that bind tightly to the enzyme, blocking substrate binding. It must be emphasized that, in many basic tissue reactions (e.g., inflammation), CPs probably do not function in isolation. Rather, they are components of complex networks constituting representatives of different types of peptidase and their respective inhibitors. Within these networks, cross-activation of zymogens and cross-inactivation of inhibitors can provide an amplification of an initial perturbation.
Numerous low-molecular-weight CP inhibitors have been developed. One of the most widely used is E-64 [L-trans-epoxysuccinyl-leucylamino(4-guanidino)butane], a potent irreversible inhibitor of many (but not all) CPs that forms a thioether with the active site cysteine (Barrett et al., 1982). The vinylsulfone N-morpholinurea-leucine-homophenylalanine-vinylsulfone-phenyl (LHVS) is a potent inhibitor of cathepsin S, and quite effective against cathepsin F (Shi et al., 2000).
(II) Clan CA
The CA clan is the largest characterized to date: It consists of 25 families, of which many are viral. Three are represented in prokaryotes, and four in eukaryotes (including protozoa, yeast, plants, and animals) and in some cases prokaryotes (Rawlings and Barrett, 2000). This clan includes the ancient papain-related enzymes found in bacteria, protozoan and metazoan organisms (family C1), and the calpains (C2). Several viral CPs also have papain-like structures (e.g., C28, foot-and-mouth disease virus leader peptidase). Many members of clan CA can be inhibited by the cystatins, and also by the synthetic inhibitor E-64.
(A) Evolution of clan CA enzymes
The papain-related enzyme family C1 is grouped into two subfamilies: C1A, comprised of “papain-like” enzymes (in a general sense; i.e., statistically similar in sequence and related in structure to papain), and C1B, consisting of intracellular bleomycin hydrolases and related bacterial aminopeptidases. These subfamilies display significant similarity only in the vicinity of the active-site residues, and represent the earliest evolutionary divergence in the C1 enzymes (Berti and Storer, 1995). Clan CA enzymes arose in bacteria in the Archean(4000 to 2500 millennia ago) and papain-related CPs are phylogenetically ubiquitous (reviewed in Tort et al., 1999).
Several phylogenetic analyses of the C1A enzymes have been reported, often in conjunction with structural considerations, although not all of these studies have included a statistical analysis of branch support (e.g., Wiederanders et al., 1992; Berti and Storer, 1995; Santamaria et al., 1999; Tort et al., 1999; Wex et al., 1999; Rawlings and Barrett, 2000). An alignment of vertebrate cathepsins and related proteins and two representative plant enzymes is shown in Fig. 1, and a phylogenetic tree based on this alignment in Fig. 2. Each branch on the tree represents either a duplication of a common ancestor (leading to paralogous genes) or speciation (leading to orthologs). The various trees are generally consistent, regardless of the methodologies used, and the C1A family of “papain-like” enzymes can be divided into two major, ancient groups (subfamilies), which have been referred to as Branch A and Branch B (Tort et al., 1999). These branches arose by duplication of a common ancestor over 2700 Ma, before the eukaryote-prokaryote divergence. A few other enzymes from parasitic and free-living protozoa and metazoa, as well as the recently described human cathepsin O, do not strongly localize to either branch (although they appear to group weakly with Branch B), and the timing of their divergence from the ancestors of the two major groups is uncertain.
Branch A divides into three strongly supported major groups: the cathepsins B- and X-like, and DPP I groups. Their order of origin is uncertain. Cathepsin B-like enzymes have been found in plants, the primitive protozoan Giardia as well as trypanosomatids, nematodes, trematodes, arthropods, and vertebrates; DPP I has been found in trematodes and vertebrates, and cathepsin X-like enzymes in nematodes and vertebrates. Thus, DPP I and cathepsin X likely evolved by duplication of an ancient cathepsin B-like ancestral gene. Branch B is more complex. Analysis of sequences from a wide phylogenetic range identified four major groups (subfamilies), plus two minor groups of enzymes from Dictyostelium, certain parasitic and free-living protozoa, and a nematode (Berti and Storer, 1995). All groups appear to have evolved early in the history of Branch B, reflecting a rapid initial duplication and divergence. The major groups are the cruzipain (cruzain)-like (corresponding to the Ddis1 group of Berti and Storer, 1995), cathepsin L-like, papain-like, and cathepsin H/aleurain-like enzymes. Cruzipain-like enzymes are found in plants, Dictyostelium, free-living and parasitic protozoa, nematodes, trematodes, and vertebrates (cathepsin F and lymphopain), while cathepsin H and aleurain-like (e.g., aleurain, orizain gamma, maize cysteine proteinase 2) enzymes have thus far been found in vertebrates and plants. Papain-like enzymes (in the strictest sense of the term, i.e., a monophyletic group, e.g., papain, chymopapain, caricain, stem bromelain, actinidain, vignain, orizain a) are restricted to plants, while cathepsin L-like enzymes are restricted to nematodes, trematodes, cestodes, arthropods, and vertebrates. Since they are represented in plants and eukaryotes, the cruzipain and the cathepsin H/aleurain groups must have evolved prior to the plant-animal divergence around 1600 Ma, while the cathepsin L-like group must have arisen before the divergence of nematodes from the lineage leading to arthropods and chordates, around 1200 Ma. The L-like cathepsins S and K have evolved more recently, and may be vertebrate-specific, pre-dating the evolution of bony fish over 450 Ma.
The mammalian cathepsin L-like CPs (cathepsins L, V, K, and S and the rodent M, J/P, Q, R, CLRP subgroup) form a distinct, well-supported (99% bootstrap) group (Fig. 2). Cathepsin F and lymphopain are also related to each other, although with weaker support (65% in this analysis). Significantly, gene pairs with a later common ancestor have tended to remain together on the chromosome (see Table). Thus, cathepsin pairs F and lymphopain co-localize to 11q13.1-13.3, K and S to 1q21, and L and V to 9q21-22. Members of each pair also have intron/exon organizations that are similar to each other. Other cathepsins map to unique positions in the human genome and tend to have unique intron/exon arrangements. Cathepsins L and V show 78% sequence similarity, consistent with a relatively recent gene duplication of an ancestral cathepsin L gene (Brömme et al., 1999; Itoh et al., 1999). They are sufficiently similar as to suggest that cathepsin V evolved after the mammalian radiation: Efforts to find a mouse ortholog of cathepsin V have failed, and thus the distribution of cathepsin V in mammals may be limited (Brömme et al., 1999). In general, vertebrate cathepsins representing phylogenetically ancient types (cathepsins B, X, L, H, O, F, and DPP I) are ubiquitously expressed in mammalian tissues, although often at quite different levels in different sites (see below). It should be noted that this does not preclude specific functions in certain tissues, as has been shown for cathepsins B, L, and DPP I (see below). However, within Branch B, other cathepsins have evolved tissue-specific patterns of expression (see below), primarily associated with cells of the immune system.
(B) General properties of Papainrelated (family C1A) CPs
(1) Structure and activity
The three-dimensional structures of several papain-related enzymes have been determined (Kirschke et al., 1995; reviewed in Turk et al., 1997, 1998; McGrath, 1999). With the exception of DPP I , which is oligomeric, all are relatively small monomers of 20 to 35 kDa. Some (e.g., cathepsins B, H, and L) undergo internal cleavage to produce two-chain forms, and many enzymes are glycosylated. The different proteins are similar in the number and position of α-helices and β-pleated sheets, and most relative insertions and deletions between the proteins occur in the loops and turns linking these elements, consistent with a common ancestry. The molecules are bi-lobed. The two domains are designated L and R, and the L domain consists of the majority of the N-terminal half of the protein, while the R domain consists of the C-terminal half of the molecule and the most N-terminal residues. The catalytic site is located in a cleft between the lobes: the catalytic Cys on the L domain, the His opposite on the R domain. Typically, the structure is stabilized by three disulfide bonds.
In contrast to many other peptidases, in no case has a CP cathepsin been shown to have a single specific substrate, although they do differ considerably in their preferred cleavage site. Polypeptide substrates bind along the cleft in an extended conformation. Binding sites for substrate residues N-terminal to the cleaved peptide bond are designated as S1, S2...etc.; those C-terminal are designated as S1′, S2′...(where S1 and S1′ are proximal to the cleaved bond). Similarly, P1, P2...; P1′, P2′......, etc., are used to designate the corresponding substrate residues (Barrett, 1994). Only the S2 subsite is a real pocket: the other sites are shallow indentations on the surface of the enzyme. Most enzymes favor a hydrophobic residue at the P2 site, whose side chain projects down into a hydrophobic S2 pocket, but papain-like CPs vary widely in their accommodation of an aromatic residue in this position. Some (e.g., cathepsin B, cruzipain) will accept an arginine at this position by forming a salt bridge. Interactions with the substrates P3 and P2′ residues involve the side chains, while P2, P1, and P1′ involve both main and side-chain contacts (reviewed in Turk et al., 1998; McGrath, 1999). Most enzymes are endopeptidases, but cathepsin B has strong carboxypeptidase activity, whereas cathepsin H has strong aminopeptidase and limited endopeptidase activity (reviewed in Kirschke et al., 1995). Cathepsin B has an 18-residue insertion proximal to the active-site cleft that forms an occluding loop. This restricts access to potential substrates in the prime sites and helps provide an anchor for the C-terminal carboxyl group. Flexibility in the loop facilitates endopeptidase activity (reviewed in Mort and Buttle, 1997; McGrath, 1999). In Cathepsin H, an 8-residue segment of the proregion, the minichain, remains attached via a disulfide bond and restricts access to the non-primed sites (reviewed in Turk et al., 1997; McGrath, 1999). The plant aminopeptidases aleurain and oryzain share sequence similarity with cathepsin H, and presumably a similar structure-function relationship (Kirschke et al., 1995; reviewed in Turk et al., 1997, 1998). The cystatins are CP inhibitors that can place N-terminal residues along the active site in the same orientation as a substrate, thereby occupying the S2 subsite, and also place hairpin loop regions in the active site (Turk et al., 1997). Just as the CP loops can interfere with the binding of polypeptide substrates, they can also interfere with the binding of cystatins. For example, cystatin C inhibits cathepsin B less well than cathepsin L. There are also proteins with homology to papain that lack peptidase activity due to substitutions in the active site (e.g., the testins, see below).
The overall conservation of cathepsin CP structure is maintained in both the ancient A and B branches. However, sensitivity to inhibition by cystatins is not uniformly distributed. In particular, cathepsin B is only poorly inhibited, while cathepsin X is resistant. Taking into account the ancient origins of DPP I, and the more recent mammalian origin of salivary cystatins (Dickinson, manuscript in preparation), it seems more probable that the target(s) for salivary cystatins is a member of branch B.
(2) The proregion
Typically, Clan CA enzymes are synthesized as inactive preproenzymes with a signal peptide and a multifunctional N-terminal proregion. The C1A subfamily proregion varies between 38 and 251 residues, depending on the enzyme and, to a smaller extent, on the species. Branch B-type CPs have a proregion around 60-100 residues in length. Cathepsin X is unusual in having a very short proregion (38 or 41 residues, depending on the prepeptide cleavage site), while DPP I has a long 206 (human)-residue prosegment due to an N-terminal extension. Alignments of the proregions of the 11 known human cathepsins, novel rodent cathepsins, and those of papain and aleurain are shown in Fig. 3. It can be seen that the definitive human Branch B-type CPs (cathepsins L, V, S, K, and H), as well as the rodent placental cathepsins (M, P and Q), have related proregions with similarities to those of papain and aleurain (Fig. 3A). The rodent proteins CTLA-2 and the testins also have homology to the Branch B proregions. As noted above, the origins of the related cathepsins F and lymphopain, and the more distantly related cathepsin O, are not well-resolved with respect to Branches A and B. However, cathepsin F and lymphopain show clear similarity to other Branch B enzymes in their proregions, and they have quite good matches to three conserved motifs within the Branch B-type prodomain (see below), consistent with the closer relationship to Branch B suggested by the tree constructed with the use of the mature enzyme sequences (Fig. 2). The cathepsin O proregion has a modest overall similarity, and partial matches to the three motifs. In contrast, the proregions of Branch A-type CPs (cathepsins B, X, and DPP I) are distinct from those of Branch B (Fig. 3), and lack matches to the conserved motifs. The proregion of cathepsin X shows some similarity to that of cathepsin B (Fig. 3B), while that of DPP I appears to be unrelated to other CPs (Fig. 1C). This fits with the results summarized above from phylogenetic analyses of larger numbers of enzymes from diverse species that identify these two distinct major branches. Despite the low sequence similarity of the proregions of cathepsins B, L, and K, analysis of their crystal structures shows that they have a common fold (Coulombe et al., 1996; LaLonde et al., 1999). Most likely this represents high divergence of ancestrally related sequences, but convergent evolution of ancestrally unrelated domains cannot be excluded at present.
The proregion is multifunctional. That of cathepsin L is essential for proper folding of the enzyme, and the proregions of other enzymes, e.g., aleurain, cannot substitute for it (Tao et al., 1994; Vernet et al., 1995). Many mammalian cathepsins are predominantly lysosomal. Newly synthesized CPs are routed via the signal peptide to the endoplasmic reticulum and then to the lysosomes (vacuoles in plants) or for secretion (reviewed in Rawlings and Barrett, 1994; Mort and Buttle, 1997). Mammalian lysosomal CPs have one or more potential N-glycosylation sites that can be located in either the proregion and/or the mature enzyme. Phosphorylation of certain mannose residues and binding to mannose 6-phosphate receptors in the Golgi results in targeting to the lysosome. However, there is evidence for mannose 6-phosphate independent transport (reviewed in von Figura and Hasilik, 1986). A 9-residue sequence in the proregion of cathepsin L was shown to mediate pH-dependent, mannose 6-phosphate-independent association with microsomal membranes (McIntyre et al., 1994). This motif has similarity to the yeast vacuolar-sorting sequences. An homologous motif in the protozoal cruzipain (cruzain) was subsequently shown to be necessary and sufficient for lysosomal targeting of green fluorescent protein in trypanosomes (Huete-Perez et al., 1999), suggesting that this motif represents an ancient lysosomal targeting system. This Branch B-type proregion peptide lysosome targeting motif is absent from the cathepsin B proregion (Fig. 3).
The N-terminal propeptide segment has been shown to be a potent and relatively specific inhibitor that serves to maintain the precursor in an inactive state until cleaved (Carmona et al., 1996; reviewed in Turk et al., 1997). The proregions bind to the active site in a linear, extended conformation, but in the reverse orientation to normal substrates. This binding is distinct to the interaction of CPs with cystatins, and likely provides a combination of a good fit to the site with resistance to proteolysis. The inhibitory activities of the cathepsin L and B proregions are sharply pH-dependent; for cathepsin L, the Ki is less than 0.5 nM at a pH above 4.3, but rises to 3.0 nM at pH 4.0, consistent with a low pH-dependent autoactivation of the enzyme (Fox et al., 1992; Carmona et al., 1996). Construction of cathepsin L proregion peptides containing various N- and C-terminal deletions allowed the inhibitory domain to be localized to a stretch of 30 residues located just following the peptide lysosome targeting motif (Carmona et al., 1996). Deletion of this domain caused a more than 200-fold increase in the Ki value. This domain contains the so-called ERFNIN motif: the conserved propeptide sequence E X3 R X2 (I/V) F/W X2 N X3 I X3 N previously identified in Branch B enzymes, but not other members of clan CA, from a phylogenetically diverse group of organisms (Karrer et al., 1993) (see Fig. 3). It is located within an alpha helix, and the conserved residues are in contact with the surface of the enzyme. Interestingly, this motif is also found in the mouse cytotoxic T-lymphocyte antigen-2 (CTLA-2) α and β gene products (Karrer et al., 1993; see Fig. 3A). These proteins show significant similarity to the L-type proregions (Denizot et al., 1989), and the CTLA-2β protein has been shown to be a good inhibitor of cathepsin L (Ki = 24 nM), H (IC50 = 67 nM), and papain (Ki = 25 nM), but not cathepsin B (Delaria et al., 1994). It is likely that the CTLA-2 genes evolved by duplication of an ancestral L-type cathepsin gene and subsequent deletion of the enzyme-coding region. However, it should be noted that CTLA-2β is a relatively non-specific inhibitor and can exist as a dimer or tetramer. Thus, it may be functionally distinct from the L-type proregions. The species distribution of CTLA-2-like proteins has not been explored. Inhibitory regions of the 56-amino-acid rat cathepsin B proregion have also been examined (Chen et al., 1996). Two regions were identified that caused 150- and 625-fold increases in the Ki. Alanine scanning identified W-24p and C-42p (rat procathepsin B numbering) as the most important residues within these regions.
In general, the N-terminal propeptide must be removed proteolytically for activation (but see below). pH-dependent auto-catalysis is believed to proceed in trans (reviewed in Turk et al., 2000). For the lysosomal cathepsins, the acidic conditions of the lysosome promote autocatalytic cleavage and dissociation of the propeptide (Mason and Massey, 1992; Ishidoh et al., 1998). Removal of the proregion can also occur by cleavage by other peptidases. However, the propeptide also serves to stabilize the enzyme against inactivation by denaturation under neutral or alkaline conditions (Mason et al., 1987; Yamaguchi et al., 1990). A conserved GXNXFXD motif (the GNFD motif) that is involved in both autoactivation and appropriate folding of the enzyme (Vernet et al., 1995) was identified within L-type but not other clan CA enzymes. Site-directed mutagenesis of the conserved residues and expression in yeast (which does not process the wild-type propapain) indicated that the negative charge of this region is involved in triggering processing at low pH. It will be of interest to examine the function of a highly conserved, negatively charged region that follows the GNFD motif (Fig. 3A). Processing of procathepsin L can be considerably enhanced by polyanions such as dextran sulfate and glycosaminoglycans (GAGs) (Mason and Massey, 1992; Ishidoh and Kominami, 1995). There is no information on whether polyanions interact with the proregion.
The propeptides of the clan CA enzymes generally share no structural similarity with the cystatins. It is therefore interesting that a region of the N-terminal propeptide extension of cathepsin F has been shown to display similarity to a cystatin domain (Nagler et al., 1999a). The level of sequence similarity is weak (it is not detected by a BLAST search of Genbank at http://www.ncbi.nlm.nih.gov), but molecular modeling predicted that the propeptide would have a structure similar to that of chicken egg white cystatin. Consistent with this conclusion, searches of a non-redundant peptide database with cystatin sequences revealed proteins in Japanese flounder, Drosophila, and Caenorhabditis elegans that contain an N-terminal domain with homology to cystatins and a C-terminal domain with significant homology to mammalian cathepsin F (Fig. 4). Thus, there appears to be an ancient CP lineage within the papain-related subfamily in which the propeptide contains a cystatin-like inhibitory domain. Whether this cystatin domain is a functional inhibitor remains to be established.
(C) Biochemical properties, expression, and normal functions of mammalian CPs of subfamily C1A
(1) Widely expressed cathepsins
(a) Cathepsins B, H, and L
(i) Properties and tissue distribution
Early studies of cathepsins B, H, and L unequivocally localized these peptidases to the lysosomes of cells, to which they are all targeted in mammals by the addition of mannose-6-phosphate. The concentrations of cathepsins B and L in lysosomes of cultured cells can be as high as 1 mM (Xing et al., 1998). In contrast to most members of the papain-like subfamily (C1A), which commonly have a neutral pH optimum, these lysosomal peptidases have an acidic pH optimum (around pH 6.0, depending in part on the substrate), and their activity would be maximal in the acidic environment of the lysosome (reviewed in Barrett and Kirschke, 1981). Cathepsin L will degrade almost any protein, while cathepsins B and H are more limited in their degradative abilities. Cathepsins L and H are efficiently inhibited by chicken egg-white cystatin and human cystatin C, while cathepsin B is less efficiently inhibited.
Cathepsin B, H, and L proteins and activities (reviewed in Barrett and Kirschke, 1981; Howie et al., 1985; Kirschke et al., 1995; Xing et al., 1998) and mRNAs (Qian et al., 1989; Söderström et al., 1999) have been detected in all tissues and cells examined. Consistent with this ubiquitous pattern of expression, these genes lack the TATA box motifs normally found in highly regulated genes but frequently absent from constitutively expressed genes (Ishidoh et al., 1989a,b; Qian et al., 1991). However, there is significant variation in the levels of these enzymes and their ratios in different tissues and cells (e.g., Qian et al., 1989; Gong et al., 1993; Katunuma et al., 1993; Söderström et al., 1999). Cathepsin B is the most abundant and widely expressed cathepsin and is found at high levels in macrophages. At the mRNA level, the highest levels are found in non-skeletal tissues. Cathepsin B levels in skeletal tissues are not greatly lower than those of non-skeletal tissues, while cathepsin H skeletal tissue mRNA levels are very low. Cathepsin L levels are generally higher in tissues that turn over more rapidly, such as the liver and ovary, and in phagocytic cells such as stimulated macrophages. In the rat, high levels of cathepsin B and L mRNA are found in the kidney.
In addition to transcriptional regulation, there is evidence that cathepsin levels may be governed by post-transcriptional processing and differences in translation rates of alternative transcripts (Chauhan et al., 1993; Gong et al., 1993; Yan et al., 1998). The level of expression of cathepsin L in fibroblasts is increased by several growth factors (e.g., epidermal growth factor, fibroblast growth factor, and platelet-derived growth factor), phorbol esters, and by oncogene-mediated transformation in vitro (reviewed in Ishidoh and Kominami, 1998). Cathepsin L has also been shown to be induced in granulosa cells of growing follicles by follicle-stimulating hormone, and in pre-ovulatory follicles in response to leuteinizing hormone in a progesterone receptor-dependent manner (Robker et al., 2000). Cathepsin B levels may be associated with cell differentiation (reviewed in Yan et al., 1998). Cathepsin B was not detected immunohistochemically in normal minor salivary glands (Steinfeld et al., 2000). However, minor glands in organotypic culture expressed significant levels of cathepsin B, primarily in the ducts, and these levels were substantially increased by treatment with prolactin.
(ii) Functions of cathepsins B, H, and L
Until recently, the ubiquitous lysosomal distribution of cathepsins B, H, and L has led them to be considered primarily “housekeeping” enzymes essential to the normal protein turnover of cells. Consistent with this view, broad CP inhibitors block up to 40% of cellular protein turnover (Shaw and Dean, 1980; reviewed in Barrett and Kirschke, 1981). However, regulation by growth factors and variation in expression levels imply duties beyond those of “housekeeping” and raise the possibility of tissue-specific functions. A powerful approach to the study of function in vivo is the generation of homozygous null mutants through the generation of transgenic knockout mice. Surprisingly, homozygous cathepsin-B-deficient mice have an apparently normal phenotype (Deussing et al., 1998). This suggests functional redundancy but raises the question of why a redundant gene has been so conserved throughout vertebrate evolution. Cathepsin-L-deficient mice have periodic shedding of fur and abnormal skin morphology but are otherwise viable (Nakagawa et al., 1998). Significantly, these mice also have a defect in major histocompatibility complex (MHC) class-II-mediated antigen presentation. In antigen-presenting cells (APCs), extracellular foreign proteins are internalized via endocytosis or phagocytosis and degraded to peptides in the endocytic pathway. Major histocompatibility complex (MHC) class II molecules bind derived antigenic peptides and present them on the cell surface to CD4+ T-helper cells. Intracellular trafficking of the MHC class II molecules and binding of antigen are regulated processes (reviewed in Wolf and Ploegh, 1995). In the endoplasmic reticulum, class II α and β chains form a heterodimer, and three αβ heterodimers associate with an invariant (Ii) chain trimer. This nonamer is then transported to the Golgi apparatus and sorted to the endocytic pathway by a signal in the Ii chain cytoplasmic domain, preventing it from entering the constitutive secretory pathway. In the endocytic compartment, the MHC class II molecules can encounter the foreign peptides. However, the Ii chain binds to the peptide-binding domain, blocking this interaction until it is removed by sequential proteolytic cleavage. The Iip10 fragment is the smallest that retains the N-terminal endosome-targeting sequence and a C-terminal extension in the peptide-binding groove. Further cleavage of the Iip10 fragment causes dissociation of the nonamer and release of αβ heterodimers bound to the CLIP fragment of the Ii chain, which occupies the peptide-binding site until the heterodimer interacts with another class-II-like chaperone molecule (HLA-DM in humans). This causes release of CLIP and allows peptide binding to occur. If the Ii chain is not cleaved, the nonamers can be targeted to the lysosome by the Ii chain cytoplasmic tail and degraded. Thus, cleavage of Iip10 is an important regulatory step, and the sequence and timing of Ii cleavage events likely determine the antigenic peptides presented. With a transgenic mouse knockout, cathepsin L has been shown to be essential for the degradation of the invariant (Ii) chain and cleavage of Iip10 to produce the CLIP fragment during MHC class-II-restricted antigen presentation in cortical thymic epithelial cells, but not in bone-marrow-derived antigen-presenting cells, which instead use cathepsin S (Nakagawa et al., 1998). Interestingly, the p41 form of the invariant chain contains a 64-amino-acid fragment with a thyroglobulin type 1 domain (Lenarcic et al., 1997) that binds and inhibits cathepsin L (Ki 1.7 pM), but not cathepsin S (Guncar et al., 1999). It may therefore be involved in regulation of Ii degradation, and in production of antigenic epitopes in endosomes (Fineschi et al., 1996).
Although lysosomal peptidases, including CPs, are undoubtedly involved in peptide antigen processing, the exact role of individual enzymes remains equivocal (Villadangos and Ploegh, 2000). The use of inhibitors “specific” for individual cathepsins has provided evidence for a role for cathepsins B (a CP) and D (an aspartyl peptidase) in antigen processing both in vitro and in vivo. For example, treatment of a mouse T-cell clonal line with a cathepsin B inhibitor suppressed processing of an ovalbumin antigenic epitope, and treatment of mice immunized with ovalbumin with this inhibitor suppressed the Th2 response and IgE production (Katunuma et al., 1998). Similarly, treatment of mice experimentally infected with Leishmania major with a cathepsin B inhibitor causes a switch in the immune response from Th2 to Th1, possibly reflecting a change in antigen processing (Maekawa et al., 1998). However, the use of inhibitors in these studies is complicated by the potential lack of complete specificity, and by the fact that the various cathepsins are involved in transprocessing of each other (e.g., Ishidoh et al., 1999). Cathepsin-B-deficient mice show no evidence for a role of cathepsin B in MHC class-II-mediated antigen presentation (including ovalbumin), indicating either that cathepsin B is not involved in this process, or that there is redundancy in the proteolytic system (Deussing et al., 1998).
Cathepsin L and, to a lesser extent, cathepsin B have been implicated in normal tissue-remodeling events. Hormonal regulation of cathepsin L levels in the granulosa cells of follicles suggests that it may be involved in the degradation of the follicle wall that leads to release of the mature oocyte (Robker et al., 2000). Cathepsin B mRNA levels rise in the apoptotic lumenal epithelial cells of regressing prostate and mammary glands, consistent with a role in degradation of the basement membrane, an early event in cell death (Guenette et al., 1994). Cathepsin CPs have been implicated in various stages of embryogenesis. The supply of amino acids to the developing mouse embryo prior to development of the chorioallentoic placenta is mediated by proteolysis of proteins in the visceral yolk sac, and levels of active cathepsin L are relatively high in this tissue at this time, in comparison both with later times and with the placenta (prior to parturition), as well as with cathepsin B (Sol-Church et al., 1999b). During implantation of the embryo, the embryonic trophoblasts invade the uterine stroma in a controlled manner, degrading the extracellular matrix (ECM). The endometrial connective tissue cells respond with the decidual reaction, which involves an enlargement of the cells and remodeling of the ECM. This provides a barrier to uncontrolled trophoblast invasion, and facilitates formation of an immunologically privileged site. As the placenta forms, decidual cells adjacent to the embryo undergo apoptosis and are phagocytized by the trophoblasts. The mouse placenta expresses substantially higher levels of cathepsin L mRNA relative to tissues such as the liver and kidney, and these levels are at their highest during implantation, suggesting a possible role in this process (Hamilton et al., 1991; Sol-Church et al., 1999b). The placenta also secretes procathepsin L, which may have proteolytic activity under certain circumstances, as well as other activities (see below). Injection of higher doses of E-64 into pregnant mice during the period of blastocyst attachment leads to a complete failure of implantation. Lower doses result in stunted embryos and a reduced decidual reaction (Afonso et al., 1997). These results suggest that CPs are essential for normal embryo development and decidualization of the uterus. Previously, it was suggested that cathepsins B and L were important in these processes (Afonso et al., 1997). However, the subsequent construction of cathepsin-B- and L-deficient mice (see above), which appear to grow and develop normally during gestation, makes this possibility seem less likely, although there could easily be redundancy in the enzyme systems. The recent discovery of placental-specific CPs (see below) might lead to clarification of these issues in the future. Placental cathepsin L mRNA levels also rise prior to parturition, possibly related to the degeneration of tissue around the placenta in preparation for birth (Hamilton et al., 1991). The role of cathepsin CPs in human implantation is unknown.
Thus far, discussion of the functions of cathepsins B, H, and L has primarily been confined to a lysosomal or endosomal location: the degradation of proteins trafficking through the endosomal system. However, it is also now clear that cathepsins B, H, and L are not purely lysosomal, and that they can be released from cells under various circumstances (see below). In the presence of thiol compounds, cathepsin B is active in the pH range of 5-6, while cathepsin L is active at pH 3-6.5, and cathepsin H has an optimum of 6.5-6.8 (Kirschke et al., 1995). In these pH ranges, cathepsins B and L and, to a lesser extent, cathepsin H can degrade a variety of components of the extracellular matrix, such as proteoglycans, laminin, and collagens II, IX, and XI (Maciewicz et al., 1990a; Buck et al., 1992; reviewed in Kirschke et al., 1995). Cathepsin L is a potent elastase at the optimal pH (5.5), where it is almost as active as pancreatic elastase, and significantly more than neutrophil elastase (both serine peptidases) (Chapman et al., 1994). In contrast, cathepsin B is 100-fold less active than cathepsin L against this substrate (Mason et al., 1986). Cathepsins B, H, and L are unstable at neutral pH, and are irreversibly inactivated above pH 7 (Barrett and Kirschke, 1981). Cathepsin L has a half-life of only about 1 minute at pH 7.2 and 37°C (Wang B et al., 1998), while cathepsin B is about 15-fold more stable (Turk et al., 1995). The rate of auto-degradation of cathepsin B at neutral pH is reduced in the presence of alternative substrates (Buck et al., 1992). Such instability would be expected to limit the extracellular degradative activity of these enzymes severely. Further, the concentration of cystatin C in vivo is sufficiently high to provide rapid and effective inhibition of cathepsin L and cathepsin B (even though the latter is less-well-inhibited by this cystatin), provided it remains in molar excess (Turk et al., 1995). However, various conditions can arise to enhance the stability of these enzymes (see below), and in contrast to the active enzymes, the proenzymes (which can also be released (see below)) are stable at neutral pH, as is a complex of mature cathepsin B and the proregion.
(iii) Cathepsins B, H, and L in the oral cavity
As lysosomal enzymes, cathepsins B, H, and L likely function in normal protein turnover of intracellular and endocytosed proteins in oral as in other tissues. Cathepsin B has been immunolocalized to granular duct cells in the rat submandibular gland and co-localized with renin in secretory granules, suggesting a role in processing secreted proteins (Sano et al., 1993). The role of these cathepsins—either intracellular or extracellular—in normal remodeling of oral tissues has not been addressed to any great extent. Cathepsins B and L have been localized to gingival fibroblasts, and this source may have a role to play in periodontal disease (discussed in detail below). Interestingly, phenytoin and cyclosporin A suppress the expression of cathepsin L (as well as of MMP-1 and TIMP-1), but not cathepsin B, in cultured gingival fibroblasts. Both these drugs induce gingival overgrowth, suggesting that some of this overgrowth is the result of impaired extracellular matrix degradation involving cathepsin L (Yamada et al., 2000).
It is axiomatic that the immune system is central to the maintenance of oral health, and the progression from gingivitis to periodontal disease. Therefore, the involvement of cathepsins B, H, and L in the function of the immune system described above also applies to the oral cavity. As lysosomal enzymes, they also function in phagocytosis and can be released extracellularly by immune cells, where they can be involved in remodeling (or damaging) the extracellular matrix and tissues as outlined above. However, these released cathepsins can also participate in more powerful proteolytic cascades. This area, and the number of studies which have examined the activities of cathepsins B, H, and L in gingival fluids with respect to periodontal disease, are discussed below. Their potential role in Sjögren's syndrome is also discussed in a separate section.
(b) Dipeptidyl peptidase I (DPP I)
(i) Properties and tissue distribution
Dipeptidyl peptidase I (DPP I) is the accepted nomenclature for an enzyme previously called cathepsin C, among other names (e.g., cathepsin J). DPP I is a Branch A enzyme most closely related to cathepsin B, and is likely to be phylogenetically widely distributed. It is a lysosomal CP with a pH optimum of 5-6 that primarily cleaves dipeptides from the N-terminus of polypeptides, although it also has endopeptidase activity (Kirschke et al., 1995). It does not cleave substrates with N-terminal Arg, Lys, or Pro, or Pro in the penultimate position. It has some distinct differences from other lysosomal CPs: It has a long 206 (human)-residue prosegment that has an N-terminal extension relative to the papain-related CPs, it forms oligomers of around 200 kDa, and it requires halide ion to be maximally active. The enzyme is inhibited by stefins A and B and chicken cystatin, but only weakly by E-64, and is unstable at > pH 7.5 (Nikawa et al., 1992; Dolenc et al., 1996).
In the mouse, Western blot analysis demonstrated DPP I in the majority of tissues examined (Pham et al., 1997): The highest levels were found in the spleen, lung, liver, and small and large intestines, while very low levels were found in the heart and brain. Comparable results were found for the mRNA distribution (Rao et al., 1997). DPP I is also present in various immune cells, including neutrophils, lymphocytes, and macrophages, and treatment of lymphocytes with interleukin-2 (IL-2) was shown to cause a significant increase in DPP I mRNA levels (Rao et al., 1997).
(ii) Functions of DPP I and its role in pre-pubertal periodontitis and Papillon-Lefèvre and Haim-Munk syndromes
In mammals, multiple functions have been ascribed to DPP I. It is thought to have a role in general protein degradation and turnover. More specific functions have been suggested, such as activation of platelet factor XIII. Recently, DPP I was shown to be required for the activation of granzymes—serine peptidases important in cytotoxic lymphocyte granule-mediated apoptosis—and could be involved in activation of other serine peptidase zymogens such as neutrophil elastase (Pham and Ley, 1999).
Missense mutations in the DPP I gene, located at 11q14, have very recently been shown to be responsible for one recessive form of pre-pubertal periodontitis, a rapidly progressing, heritable form of the disease that affects the primary dentition (Hart et al., 2000). Two distinct autosomal-recessive palmoplantar keratoderma disorders, Papillon-Lefèvre syndrome and Haim-Munk syndrome, characterized by hyperkeratosis of specific epithelial areas, particularly the hands and feet, are also characterized by severe early-onset periodontitis, resulting in the loss of the primary and secondary dentition. Papillon-Lefèvre syndrome is usually first diagnosed by dentists. Both syndromes have now been shown to result from mutations in the DPP I gene (Hart et al., 1999, 2000; Toomes et al., 1999). Why loss of this widely distributed lysosomal enzyme should preferentially affect these tissues is unknown, although Chediak-Higashi syndrome, which also affects lysosomes, is also associated with immune dysfunction and severe early-onset periodontal disease (Tempel et al., 1972; Introne et al., 1999). The association of DPP I with these disorders illustrates that a ubiquitously expressed cathepsin can have tissue-specific functions, and need not be confined to a housekeeping function.
(c) Cathepsins O and X
(i) Properties and tissue distribution
Little is known about cathepsin O. It was originally cloned from breast tumor tissue by the polymerase chain-reaction (PCR) by means of primers directed to conserved CP sequences (Velasco et al., 1994). It has a predicted prodomain of a typical length (84 residues) but with only partial matches to the three consensus sequences discussed above (see Fig. 3A). Northern analysis demonstrated mRNA in all tissues, with the highest levels in the ovary, kidney, liver, and placenta and the lowest in the thymus and skeletal muscle. The native protein has not been purified, although a recombinant protein has been obtained by expression in E. coli. No enzymatic properties (pH profile, stability, inhibition) have been reported.
Another novel human cathepsin has been independently characterized by three groups, who used identification of novel ESTs in the database, followed by screening of cDNA libraries. It was initially designated as cathepsin X (Nagler and Menard, 1998), cathepsin Z (Santamaria et al., 1998a), and cathepsin P (Pungercar and Ivanovski, 2000). Phylogenetic analysis (see Fig. 2) indicates that a CP designated as cathepsin Y cloned from rat spleen (Sakamoto et al., 1999) is the rat ortholog of cathepsin X. Cathepsin X is unusual in having a very short proregion (38 or 41 residues, depending on the prepeptide cleavage site) that is even smaller than that of cathepsin B. It completely lacks the N-terminal region that contains the lysosomal targeting consensus and the ERNF/WNIN motif found in the cathepsin L group. The role of this short proregion in folding, inhibition, and stabilization at different pHs remains to be determined. However, it does contain a cysteine residue in a position similar to that of a cysteine in cathepsin B that has been shown to be important in inhibition by the proregion (Chen et al., 1996; see above). Two potential N-glycosylation sites are present in the mature protein that could serve to target it to the lysosome. Interestingly, the proregion also contains an RGD integrin-binding motif.
Recombinant human procathepsin X was obtained by expression in Pichia pastoris (Nagler et al., 1999b). Unlike other cathepsins, it did not activate auto-catalytically at low pH, but cathepsin L was found to convert the proenzyme efficiently to the active form. Cathepsin X was found to be a very good carboxypeptidase, with a pH optimum around 5.0, and a relatively poor endopeptidase. The 3D structure of human procathepsin X has been determined. A Cys residue in the proregion is covalently bound to the active-site Cys, and a 3-residue “mini-loop” insertion between the Gln of the oxyanion hole and the active-site cysteine (predicted by primary sequence alignment algorithms) partially occludes the S2′ subsite, providing an explanation for the carboxypeptidase activity (Sivaraman et al., 2000). It is not inhibited by human cystatin C.
Northern blot and RT-PCR analysis demonstrated ubiquitous expression of cathepsin X, although the levels varied considerably between tissues (Nagler and Menard, 1998; Santamaria et al., 1998a; Deussing et al., 2000; Pungercar et al., 2000). Ubiquitous expression in the mouse and human was consistent with the characterization of the promoter as housekeeping-type (Deussing et al., 2000). Cathepsin X was also highly expressed in a variety of cancer cell lines, and may therefore be up-regulated with malignant transformation (Santamaria et al., 1998a; Pungercar et al., 2000). Cathepsin X was immunolocalized in human hepatocytes and Kupffer cells, and in the epithelial cells of distal tubules (Pungercar et al., 2000). It showed a diffuse, mostly peri-membranous distribution, in contrast to the punctated, granular distribution shown by cathepsin B, which was also primarily localized to the proximal tubules of the kidney. This suggests that cathepsin X may be localized to the membrane or the adjacent extracellular space. An examination of expression in oral tissues has not been reported.
(ii) Functions
The physiological functions of cathepsin X are unknown. The rat enzyme was initially identified based on its ability to produce bradykinin-potentiating peptide from plasma (Sakamoto et al., 1999). In equimolar amounts, this peptide increases the activity of bradykinin seven-fold, and in two-fold excess, by 23-fold. The precursor protein for this peptide is unknown. Bradykinin has been shown to synergize with IL-1 or TNFα to stimulate IL-6 production by human gingival fibroblasts (Modéer et al., 1998). Therefore, cathepsin X activity could contribute to the pathogenesis of periodontal disease by increasing the effect of this pro-inflammatory mediator.
(d) Cathepsin F
(i) Properties and tissue distribution
Cathepsin F was independently cloned by three groups either by using PCR and degenerate oligonucleotides directed to conserved CP regions or by identifying novel ESTs in the database (Wang B et al., 1998; Nagler et al., 1999a; Santamaria et al., 1999). The proregion is very large (251 residues), due to an N-terminal extension with an N-terminal region which has similarity to a cystatin domain, followed by a 50-residue flexible linker peptide (Nagler et al., 1999a). The following C-terminal segment of this proregion has overall similarity to the Branch B-like group, although it is most similar to lymphopain. Like lymphopain, it contains a peptide lysosome-targeting motif, followed by a partial match to the ERWNIN motif, ERFNAQ, consistent with these enzymes forming a phylogenetically distinct subgroup (Wex et al., 1999). The proprotein contains 5 potential N-glycosylation sites. Transient expression in Cos-7 cells localized the protein to vesicles, most likely lysosomes (Wang B et al., 1998).
Cathepsin F has been expressed in Pichia pastoris (Wang B et al., 1998). The enzyme autocatalytically activated at an acidic pH, and was shown to have a level of activity toward synthetic substrates similar to that of cathepsin L, with a broad pH optimum between 5.2 and 6.8. The catalytic efficiency (kcat/Km) was comparable with that of cathepsin L, which is the most active lysosomal CP cathepsin. Like cathepsins K, L, and S, cathepsin F prefers a bulky hydrophobic or aromatic residue at the P2 position. The enzyme was unstable at neutral pH, with a short half-life of about 2 minutes at pH 7.2 and 37°C. Cystatin C is an effective inhibitor of recombinant cathepsin F, with a Ki of 0.532 nM (Shi et al., 2000).
Northern blot analysis demonstrated that cathepsin F is ubiquitously expressed, although at different levels in each tissue (Wang B et al., 1998; Nagler et al., 1999a; Santamaria et al., 1999). The highest levels were found in the brain, heart, ovary, skeletal muscle, and testes. High-level expression in the brain is unusual for a CP cathepsin. Very low or undetectable levels were found in leukocytes and leukemia cell lines. Cathepsin F was found to be expressed in macrophages (although at lower levels than cathepsin S), but not in B-cells or dendritic cells (Shi et al., 2000).
(ii) Functions of cathepsin F
Cathepsin F is likely to be important in antigen presentation by macrophages, and in a normal Th-2 response. Degradation of Iip10 to CLIP involves cathepsin L in the thymic cortex (see above), and cathepsin S in dendritic and B-cells (see below). However, alveolar and peritoneal macrophages from cathepsin S-/- L-/- mice showed normal MHC class II peptide loading, indicating the involvement of one or more other peptidases in antigen presentation (Shi et al., 2000). Very efficient cleavage of Iip10 to CLIP by cathepsin F was demonstrated, and macrophages from cathepsin S-/- L-/- mice showed defective Iip10 processing and peptide loading in the presence, but not the absence, of 1 μM LHVS. High doses of this inhibitor will block the rise in IgE titers and eosinophilic infiltration seen in a mouse Th-2-driven model of pulmonary hypersensitivity (Riese et al., 1998).
(2) Tissue-specific cathepsins
(a) Cathepsin S
(i) Properties and tissue distribution
Cathepsin S protein was first purified from bovine lymph node and spleen, and then cDNA cloned from bovine spleen by RT-PCR (Wiederanders et al., 1991; reviewed in Kirschke et al., 1995; Kirschke and Wiederanders, 1994). Subsequently, human cathepsin S was independently cloned by two groups (Shi et al., 1992; Wiederanders et al., 1992). Phylogenetically, cathepsin S is most closely related to cathepsin K, and both map to 1q21 (Shi et al., 1994). It has a gene organization similar to that of cathepsin K (Wex et al., 1999) and the more distantly related cathepsin L, and also lacks TATA and CAAT boxes in the promoter. Cathepsin S has a proregion of 99 residues that includes a conserved peptide lysosome-targeting signal and a following ERWNIN motif. Like cathepsin K, it has a variant on the GNFD motif (GNLD). Recombinant procathepsin S will autocatalytically activate at pH 4.5 (Brömme et al., 1993). The mature protein is a single-chain polypeptide of 217 amino acids.
Cathepsin S has significant endopeptidase activity toward a wide range of proteins and synthetic substrates over a broad pH range of 5-8 (Brömme et al., 1993; reviewed in Kirschke and Wiederanders, 1994). Unlike cathepsin L (which prefers aromatic residues), it prefers branched hydrophobic residues in the S2 subsite. Importantly, cathepsin S will degrade several components of the extracellular matrix, including insoluble collagen and elastin. It is a potent elastinolytic enzyme at pH 4.5-5.5 (comparable with pancreatic elastase), but is unique among the CP cathepsins in retaining considerable activity, about 20-25% of maximal, at neutral pH (where it is comparable with leukocyte elastase in activity) (Shi et al., 1992; Chapman et al., 1997). At neutral pH, cathepsin S will degrade fibronectin, laminin, and myelin basic protein, and completely degrade proteoglycans (Petanceska et al., 1996; Liuzzo et al., 1999b). Further, unlike cathepsins B, H, and L, cathepsin S is stable at pH 7.0. No loss of activity occurred after 4 hours at 40°C at pH values up to 7.0, while these other cathepsins are inactivated. Only 10-12% of the activity was lost in 1 hour at pH 7.5 (Kirschke et al., 1989; Brömme et al., 1993). Cystatins and stefins are potent inhibitors of cathepsin S (Brömme et al., 1991; Turk et al., 1994).
Expression of cathepsin S has been examined in bovine tissues by Western blot analysis (Kirschke et al., 1989), and in human (Shi et al., 1994; Linnevers et al., 1997), rat (Petanceska and Devi, 1992), and mouse (Qian et al., 1989; Söderström et al., 1999) tissues by Northern blot analysis. Collectively, cathepsin S mRNA and protein were found in a wide range of tissues, but at significantly different levels in different tissues, and in a pattern quantitatively different from those of cathepsins B, H, and L. Of the tissues examined in these studies, some of the highest cathepsin S mRNA levels were found in the spleen. Human lung had the highest levels in the study by Linnevers et al. (1997), who also found relatively high levels of expression in peripheral blood lymphocytes and spleen and lymph nodes, moderate levels in the appendix and bone marrow, and low levels in the thymus. Mouse lung also had high levels, while the lowest levels were in the pancreas. High levels of the expected 1.4-kb mRNA were also found in rat ileum and brain, and the lowest in lung and heart (although an abundant 6-kb transcript was present in the lung). In bovine tissues, the highest protein level was found in the kidney. The extent to which the various species differences reflect genuine differences in expression, as opposed to methodological differences, is unclear.
For a limited number of tissues, cellular localization of cathepsin S expression has been performed by immunohistochemistry or in situ hybridization. In the rat, in situ hybridization localized expression in the spleen to cells in the marginal zone (Petanceska et al., 1996). This region contains macrophages and dendritic cells. In the human lung, immunohistochemistry localized cathepsin S protein to alveolar macrophages, but not airway or vascular lining cells (Shi et al., 1994). In the rat lung, in situ hybridization also localized cathepsin S mRNA to alveolar macrophages (Petanceska et al., 1996), while in the normal rat brain, low levels of cathepsin S mRNA localized to the microglial cells (Petanceska et al., 1994). Expression in the bovine kidney was localized by immunohistochemistry to the proximal tubule cells, which are also phagocytic (Kirschke et al., 1989). Thus, expression of cathepsin S is actually tissue-specific, and it is preferentially expressed in phagocytic/antigen-presenting cells. Consistent with this conclusion, high-level cathepsin S expression was demonstrated in CD14+ human monocytes, and very low level expression in CD19+ B-lymphocytes (Linnevers et al., 1997). Northern blot analysis of mouse and rat cell lines detected cathepsin S mRNA only in a monocytic leukemia and a microglial-like cell line (Petanceska et al., 1996). Various other cell lines, including fibroblasts, showed no expression. Similarly, cathepsin S mRNA was found in primary culture microglial cells, but not type 1 astrocytes or O2A progenitors. In contrast, cathepsins B and L were expressed at significant levels in all of these cell types. Further, cathepsin S was detected by Western blot in both early and mature mouse bone marrow dendritic cells (Pierre and Mellman, 1998), and by RT-PCR in cultured human retinal pigment epithelial cells, which are also phagocytic (Rakoczy et al., 1994). The moderate-low mRNA levels detected in multiple tissues by Northern analysis could be explained by the presence of phagocytic/antigen-presenting cells and blood retained in vessels. However, cathepsin S is not purely restricted to these cell types: It is also expressed in cultured rat thyroid cells (Petanceska and Devi, 1992) and, at least under certain circumstances, it can be expressed in smooth-muscle cells (see below). Nothing is known regarding cathepsin S levels in human oral tissues in health or disease.
Little is known about the regulation of cathepsin S expression. Like many housekeeping genes, the cathepsin S gene lacks recognizable TATA or CAAT boxes in the promoter region. However, it also contains only two Sp1 sites and multiple potential AP-1 sites, features that would be consistent with a regulated promoter driving expression in a limited range of tissues (Shi et al., 1994). In addition to regulation by cytokines, thyroid-stimulating hormone increases cathepsin S mRNA levels about four-fold in a rat thyroid cell line, raising the possibility that the enzyme is involved in thyroid hormone biosynthesis (Petanceska and Devi, 1992). Most significantly, cathepsin S activity has been shown to be regulated by intracellularly localized cystatin C levels in dendritic cells (Pierre and Mellman, 1998; see below).
(ii) Functions of cathepsin S and its role in antigen presentation
Cathepsin S appears to be multifunctional: intracellularly, as a processing enzyme during trafficking and secretion of proteins; and extracellularly, during response to damage and tissue remodeling. A key function for cathepsin S is in antigen processing and presentation, which would explain its prominent expression in phagocytic and antigen-presenting cells. Cathepsin S activity is distributed along the entire endocytic system in dendritic cells, consistent with its broad pH optimum (Driessen et al., 1999). Relatively specific inhibition of cathepsin S with low levels of a vinylsulfone inhibitor (also see cathepsin F, above) markedly impaired antigen presentation in vitro and in vivo, and caused a build-up of intermediate Ii breakdown products (Pierre and Mellman, 1998; Riese et al., 1998). In dendritic cells, antigen processing and presentation are uncoupled. Immature cells, such as those found in peripheral tissues, are highly endocytic and accumulate antigen, but they are inefficient at antigen presentation, in part because newly synthesized class II-Ii complexes are targeted to lysosomes and degraded. When dendritic cells migrate from the periphery to the lymphoid tissue and mature, the endocytic activity is reduced, and class II molecules are targeted to the surface. In bone-marrow-derived dendritic cells, the vinylsulfone inhibitor blocked Ii processing and arrested class II molecules in an immature dendritic cell-like subcellular distribution, with increased localization to lysosomes (Pierre and Mellman, 1998). However, it did not impair production of antigenic peptide. The switch in class II transport was associated with an increase in cathepsin S activity (but not protein), and, surprisingly, cystatin C (normally considered an extracellular protein) was found to be important in regulating intracellular cathepsin S. Dendritic cell maturation was accompanied by a significant decrease in intracellular cystatin C levels, and the localization changed: In immature cells, it co-localized with class II and Ii-containing endocytic structures; in mature cells, it localized to non-endocytic structures, while most of the class II molecules were found on the surface. Overexpression of cystatin C caused lysosomal class II accumulation in transfected cells. Similarly, in spleen dendritic cells of transgenic mice that are cathepsin S homozygous-null, the distribution of MHC class II molecules to late endocytic structures and, to a lesser extent, the lysosomes is enhanced (Driessen et al., 1999). These mice show defective Th1-type antibody responses, and are markedly impaired in antibody class switching to IgG2a and IgG3 (Shi et al., 1999). However, their IgE and pulmonary responses to antigen challenge are normal, indicating that cathepsin S is not required for some normal Th2-type responses. Cathepsin F has been shown to mediate Ii cleavage in macrophages (Shi et al., 2000; see above). Collectively, the effects on the immune response likely depend on the type of APC (and its complement of CPs) and the antigen itself.
Cathepsin S has been implicated in the production of the amyloid β peptide (Aβ) plaques and vascular deposits that accumulate in the brain in Alzheimer's disease (AD) and Down syndrome (DS). This peptide is formed by proteolytic processing of β-amyloid precursor protein (βAPP). By transient expression, cathepsin S, but not cathepsins B, L, or D, was shown to be able to produce amyloid β peptide (Aβ) (Munger et al., 1995). Cathepsin S protein levels have been examined in normal, DS, and AD brains (Lemere et al., 1995). In most normal individuals, no cathepsin S protein could be detected in hippocampal neurons or pyramidal cells, but protein was present in the leptomeningeal vessel smooth-muscle cells. In contrast, neuronal and glial immunostaining for cathepsin S was observed in the majority of AD and DS brains, in addition to the vascular staining seen in normal brains. The general distribution of cathepsin S paralleled sites of plaque formation.
Cathepsin S may also function in the intracellular degradation of phagocytized material. Non-dividing retinal pigment epithelial cells rapidly turn over their own structural components, and phagocytize and degrade large quantities of material shed from adjacent rods and cones. In cultured human retinal pigment epithelial cells, leupeptin (a general CP inhibitor), but not the aspartate peptidase inhibitor pepstatin, caused an increase in auto-fluorescent (incompletely degraded) material (Rakoczy et al., 1994). Cathepsin S expression in these cells was demonstrated by RT-PCR, although levels were too low to be detected by Northern blot analysis (Rakoczy et al., 1994). However, antisense cathepsin S oligonucleotides caused a pronounced increase in accumulation of auto-fluorescent material, and in the number of undigested phagosomes, while the sense oligonucleotide did not.
Cathepsin S likely also functions extracellularly in tissue remodeling. Under certain culture conditions in vitro, human monocyte-derived macrophages can develop a powerful ECM-degradative phenotype (Reddy et al., 1995). These cells express cathepsins B, L, and S, as well as elastinolytic matrix metalloproteinases (MMPs). However, only the CPs were detected extracellularly as processed, active enzymes, and inhibition of the CPs, but not the MMPs, almost completely blocked the elastinolytic activity. Consistent with these observations, cathepsin S mRNA was found in mature rat macrophages and microglial cells, but not in undifferentiated monocytes (Liuzzo et al., 1999a). Activators of macrophages negatively regulated the levels of mRNA, intracellular protein, and activity, but stimulated secretion of cathepsin S activity. Thus, activation of macrophages would cause extracellular delivery of a finite payload. Surprisingly, basic fibroblast growth factor and nerve growth factor—growth factors that are both present during inflammation but which were not known to activate macrophages or microglia—significantly increased the levels of cathepsin S mRNA (and also those of cathepsins B and L), and basic fibroblast growth factor also increased intra- and extracellular cathepsin S activity (Liuzzo et al., 1999b). In experimental entorhinal cortex lesions of the rat brain, which invoke a strong microglial response, a significant increase in cathepsin S mRNA was observed in the de-afferented dentate gyrus and the wound site, likely within activated microglia and blood-borne macrophages (Petanceska et al., 1996).
Remodeling of the ECM of blood vessels is important to the pathogenesis of atherosclerotic lesions. Negligible expression of cathepsin S (or K) was found in normal arteries by immunostaining (Sukhova et al., 1998). In contrast, in atheromas, cathepsins S and K were found to co-localize and were present in the expanding intima and the subjacent smooth-muscle cells, and in the abundant macrophages present in the lesion. Cathepsin S (although not cathepsin K) mRNA levels were increased three- to four-fold in atherosclerotic tissue, and plaque extracts had almost two-fold higher elastinolytic activity. Cultured vascular smooth-muscle cells did not contain immunoreactive cathepsin S, contained very low levels of cathepsin K, and had little or no elastinolytic activity. However, exposure to IL-1β, tumor necrosis factor-α (TNF-α), or interferon-γ (IFN-γ) (which are found in developing plaques) stimulated cathepsin S but not cathepsin K expression at both the protein and mRNA levels. Further, the stimulated cells were shown to secrete active enzyme. In parallel, the cells gained elastinolytic activity, the majority of which could be inhibited by the broad CP inhibitor E-64. The cervix undergoes extensive remodeling in preparation for parturition analogous to an inflammatory response, and pro-inflammatory cytokines are believed to participate in this process. TNF-α caused about a 100-fold and IL-1β a 34-fold increase in cathepsin S mRNA in cultured human cervical smooth-muscle cells (Watari et al., 1999). Procathepsin S, but not cathepsin S (37-kDa and 28-kDa proteins, respectively), was detected in the media of treated, but not untreated, cells. Other MMPs were also up-regulated by TNF-α. Thus, pro-inflammatory cytokines stimulate cathepsin S production and release by smooth-muscle cells of blood vessels and the cervix. Possibly also related to local remodeling of the ECM, cathepsin S expression was detected in 65% of soft-tissue sarcomas examined, and the expression of cathepsin S was correlated with local recurrence of the tumor, and moderately correlated with survival rate (Wurl et al., 1997).
(iii) Potential roles for cathepsin S in oral tissues and the oral cavity
Based on the above data, cathepsin S is a good candidate for a CP with a role in oral tissues in health and disease: It is expressed by antigen-presenting cells, including macrophages; it is up-regulated in vascular smooth-muscle cells in response to inflammation; it is relatively stable at pH 7; and it is a potent peptidase that can degrade extracellular matrix components and that is strongly inhibited by cystatins. Nothing is known regarding cathepsin S in oral tissues. Surprisingly, cathepsin S protein is readily detectable by immunohistochemistry on the cilia of conducting airway cells. Since it can function at neutral pH, this raises the possibility that cathepsin S has been captured onto the surface of the cilia to promote motility by degradation of other adherent proteins (Chapman et al., 1997). How much cathepsin S reaches the oral cavity or the pharynx, where it can interact with salivary cystatins, is unknown.
(b) Cathepsin K
(i) Properties and tissue distribution
A novel CP was independently cloned from rabbit osteoclasts and various human tissues by several groups around the same time. Thus, the same CP (now called cathepsin K) was variously called OC-2 (Tezuka et al., 1994), cathepsin K (Inaoka et al., 1995), cathepsin O (Shi et al., 1995), cathepsin O2 (Brömme and Okamoto, 1995), and cathepsin X (Li et al., 1995). Cathepsin K has a signal peptide and a typical 99-residue proregion with a peptide lysosome targeting signal and a following ERWNIN motif. The proprotein is about 35 kDa, and the mature enzyme is a monomeric protein with a calculated size of 23.5 kDa. There are three consensus N-glycosylation sites in the proprotein, although the sites in the mature protein region are unlikely to be used, due to flanking proline residues. Cathepsin K is most similar to cathepsins S and L, shares a similar gene structure, and maps adjacent to cathepsin S on 1q21 (Chapman et al., 1997; Gelb et al., 1997).
Like most other CP cathepsins, recombinant procathepsin K will autoactivate at pH 4 (D'Alessio et al., 1999). The enzyme activities of cathepsin K are similar to those of cathepsin S (Brömme et al., 1996). It prefers a bulky hydrophobic residue in the S2 subsite, although, unlike cathepsin S, it will accept the β-branched valine. The enzyme has a bell-shaped pH activity profile with a broad optimum between pH 5 and 8. At neutral pH, it is more stable than cathepsin L, but less so than cathepsin S. After 1 hour at pH 6.5, 37°C, cathepsin K retained only 49% of its activity, and was inactivated after 1 hour at pH 7.5, 37°C. Cathepsin K is the most potent mammalian elastase described, and it is also a powerful collagenase and gelatinase (Brömme et al., 1996; reviewed in Chapman et al., 1997). Even at neutral pH, its elastinolytic activity is 2.4 times greater than that of pancreatic elastase. It will extensively degrade collagen at a pH between 5 and 6, but much less so at pH 7, while gelatin is totally degraded within 30 minutes, even at neutral pH. The stability of the enzyme at neutral pH is greatly increased by chondroitin-4-sulfate (found in cartilage), and the collagenolytic activity is also stimulated by this GAG at acidic, but not neutral, pH, suggesting a novel regulatory mechanism (Li et al., 2000). Similar effects were not observed with cathepsin L. Cathepsin K will also degrade non-collagenous matrix proteins such as aggrecan, osteopontin, and osteonectin, even at neutral pH (Bossard et al., 1996; Gowen et al., 1999; Li et al., 2000). It is inhibited by cystatin and E-64 (Guay et al., 1999).
Like the cathepsin S gene, and many housekeeping genes, the promoter region of the cathepsin K gene lacks a TATA box. However, it also contains two AP-1 sites, and as with cathepsin S, high-level expression of cathepsin K is tissue-restricted. It is highly expressed in rabbit osteoclasts, but not osteoblasts or chondrocytes (Tezuka et al., 1994). Much lower levels of expression were detected by Northern analysis in other tissues, including alveolar macrophages. Cathepsin K mRNA expression in adult mouse tissues has been examined with RT-PCR (Lazner et al., 1999). Expression was detected in most tissues examined, with high levels in bone, brain, heart, lung, and skeletal muscle, and generally low levels elsewhere. Qualitatively but not quantitatively similar results were found in Northern blot analysis of young mouse tissue, with high cathepsin K mRNA levels in bone and cartilage, and trace levels in the majority of other tissues, including the brain, heart, and lung (Söderström et al., 1999).
Northern blot analysis of human cathepsin K expression has provided partially contradictory results. In both adult (Inaoka et al., 1995) and fetal (Söderström et al., 1999) human tissues, very low levels of cathepsin K mRNA expression were found in the majority of human soft tissues examined. Söderström et al. (1999) detected moderate levels of expression in skin and ovary, and high levels in bone, under conditions where the probe showed no cross-hybridization to cathepsins B, H, L, or S. Brömme and Okamoto (1995) observed high levels in ovary and colon, and medium-low levels in most other soft tissues. No expression was detected in brain, liver, and peripheral leukocytes. In contrast, Drake et al. (1996) did not detect cathepsin K mRNA by Northern blot analysis or in situ hybridization, or protein by Western blot analysis in a similar panel of human tissues. Interestingly, low levels of cathepsin K mRNA were detected in normal arteries and cultured vascular smooth-muscle cells, while immunoreactive protein was not (Sukhova et al., 1998), suggesting post-transcription control of expression. These results also suggest that the low levels of cathepsin K mRNA detected in various tissues could be from blood vessels, although cross-hybridization of probes with more widely distributed cathepsins has not been fully excluded. Also, cathepsin K is expressed in activated macrophages (see below). Since the human tissues examined were obtained at autopsy, some of the differences may reflect different causes of death.
Extremely high levels of cathepsin K expression were found in osteoclastoma cells (Brömme and Okamoto, 1995; Inaoka et al., 1995; Drake et al., 1996). Further, high levels of mRNA and protein expression were found in normal human bone. High levels of cathepsin K were also found in chondroclasts at sites of cartilage remodeling in osteophyte and in articular cartilage from sites of rheumatoid arthritis (Drake et al., 1996; Li et al., 2000). Thus, the primary site of cathepsin K expression in normal human and rodent tissues is the osteoclast, and also chondroclasts actively engaged in cartilage resorption. Macrophages are present in large numbers in rheumatoid synovium, but cathepsin K is expressed in only a minority of CD68-positive cells (macrophages and monocytes, giant cells, and osteoclasts) (Drake et al., 1996; Hummel et al., 1998). Shi et al. (1995) also found no expression in alveolar macrophages or peripheral monocytes. However, under certain conditions, macrophages can assume a highly degradative phenotype (Reddy et al., 1995). Macrophages in human atheromas contain abundant immunoreactive cathepsin K, as well as cathepsin S (Sukhova et al., 1998). Another type of matrix-degrading cell is the giant cell (macrophage polykaryon) found in diseased synovial (e.g., osteoarthritic) and inflamed tissues. These cells may be formed by fusion of fully differentiated macrophages or of synoviocytes (Wilkinson et al., 1993). They express osteoclast markers at sites of bone remodeling, including high levels of cathepsin K, but unlike macrophages, they do not express cathepsins B, L, or S (Dodds et al., 1999). Cathepsin K may therefore be involved in digestion of bone and cartilage fragments that break off the joint surface in osteoarthritis. Thus, there are at least five matrix-degrading cell populations that express cathepsin K and which may be derived from the monocyte/macrophage lineage in the hematopoietic compartment.
The importance of cathepsin K in bone remodeling (reviewed in Lazner et al., 1999) is revealed by the human genetic disease pycnodysostosis, a rare autosomal-recessive trait characterized by osteosclerosis and fragility of the bones, and abnormal bone and tooth development. Osteoclast numbers are normal, but the region of demineralized bone around the osteoclast is increased, and the cells contain large numbers of cytoplasmic vacuoles containing bone collagen fibrils, indicating normal demineralization of bone, but inadequate degradation of the ECM. Linkage analysis localized the defect to the vicinity of cathepsins S and K on 1q21, and sequence analysis demonstrated concordance of the affected individuals in unrelated families with mutations in the cathepsin K gene that would alter and likely inactivate the enzyme (Gelb et al., 1996). As a result, turnover of the ECM would be reduced. This conclusion is confirmed in cathepsin K knockout mice, which show skeletal abnormalities marked by osteopetrosis and reduced bone marrow cellularity (Gowen et al., 1999). The osteopetrosis primarily affected those skeletal tissues that undergo rapid remodeling, such as long bones, while tissues such as the epiphyses, which turn over slowly, were relatively unaffected. Intact bone matrix could be observed directly adjacent to osteoclasts, which were present in normal numbers. These results indicate that osteoclast differentiation and bone demineralization are unaffected by the cathepsin K deficiency, but matrix degradation is incomplete. At the mRNA level, interferon-γ (IFN-γ) decreases, whereas IL-1α increases cathepsin K expression in osteclasts (Kamolmatyakul et al., 2001). Therefore, modulation of cathepsin K levels is likely to be one mechanism by which these cytokines affect bone metabolism.
The expression of cathepsin K mRNA has been examined during mouse embryogenesis by in situ hybridization (Dodds et al., 1998). No cathepsin K mRNA was detected in any tissues in 11- to 13-day-old fetuses. At this stage, no osteoclasts are present. In the 15- to 17-day-old fetus, after osteoclast formation, cathepsin K expression was observed in pre-osteoclasts and osteoclasts at sites of bone and cartilage remodeling, but not in any other cells or tissues, including chondrocytes, the liver, and kidney. In contrast, during human development, immunohistochemistry, in situ hybridization, and Northern blotting have showed cathepsin K to be expressed at significant levels in the epithelial cells of various tissues (Haeckel et al., 1999). It is expressed in respiratory and gastrointestinal mucosa, including bile duct epithelia and urothelia, usually in the surface layers. It is not clear to what extent these different observations reflect species differences versus methodological ones. Recently, cathepsin K has been shown to be expressed in normal adult bronchial epithelial cells (Buhling et al., 1999).
(ii) Potential functions of cathepsin K in oral tissues
Clearly, there is a role for cathepsin K in bone loss during progression of periodontal disease. However, the expressing cell populations (e.g., osteoclasts and macrophages) have not been examined. Odontoclasts, which are similar to osteoclasts, are believed to mediate the root resorption that occurs during orthodontic tooth movement. Consistent with this, cathepsin K mRNA was detected by in situ hybridization in rat odontoclasts during experimental tooth movement, and immunoelectron microscopy localized the protein to resorption lacunae in the tooth root on the pressure side (Domon et al., 1999; Tsuji et al., 2001). Cathepsin K was also detected in the pressure-side periodontal ligament fibroblasts. Cathepsin K may therefore have a central role in bone resorption during tooth movement. Examination of cathepsin K expression in oral epithelia has not been reported.
(c) Lymphopain
Lymphopain was originally identified as a unique human EST in the database and designated as cathepsin W (Linnevers et al., 1997). The same cathepsin was independently cloned by subtraction from a hematopoietic progenitor cell line and named lymphopain (Brown et al., 1998). Unlike cathepsin F, to which it is otherwise most closely related, lymphopain contains a typical length proregion with overall similarity to the cathepsin L-like proregions, although it is most similar to cathepsin F. Like cathepsin F, the proregion has a good match to the peptide lysosome targeting sequence, followed by a partial match to the ERFWNIN motif, ERFNAQ. Lymphopain does contain two potential N-glycosylation sites that could serve for lysosomal targeting. However, a preliminary study of its subcellular localization by transient expression in Cos-7 and HeLa cells indicated that, in contrast to the other human cathepsins, it localizes to the endoplasmic reticulum (Wex et al., 1998). Lymphopain contains a 20-residue insertion between the active-site His and Asn residues (see Fig. 1). This would localize on the opposite side of the molecule to the active site, and therefore would not be expected to affect enzyme activity directly. However, the enzyme also contains substitutions in several residues that would be expected to affect substrate binding, including several that are otherwise highly conserved in the human cathepsins. To date, no detailed characterization of the properties of the enzyme have been performed.
Northern blot analysis demonstrated high lymphopain mRNA levels in bone marrow, lymph node, spleen, lymph nodes, and PBLs, and no expression in most other tissues, including the lung and liver (Linnevers et al., 1997; Brown et al., 1998). No expression was found in various cancer cell lines, including HSG, a salivary gland carcinoma line. These results are consistent with specific expression in immune system cells and are in contrast to the broader tissue distribution of cathepsin S, primarily in resident macrophages. Northern analysis of normal lymphocyte populations and lymphocytic leukemias demonstrated that lymphopain is primarily expressed in the T-cell population, and at particularly high levels in the CD8+ cytotoxic T-cells. In contrast, cathepsin S is expressed at high levels in the CD14 monocyte cell population and at very low levels in B-cells. The majority of thymic T-cells are CD4+CD8+ immature thymocytes. Since expression of lymphopain in the thymus is low, high levels of expression in the T-cell population must be coupled to thymocyte selection. Since DPP I has recently been found to have a critical function in immune cells, lymphopain may also serve a similar function in cytotoxic T- and natural killer cells.
(d) Cathepsin V and other cathepsin L-like sequences
(i) Properties and tissue distribution
In addition to cathepsin L, humans and other mammals also carry a second gene encoding a cathepsin L-like protein. Using ESTs or degenerate-primer PCR products to screen human cDNA libraries, three groups independently identified the same CP, designated as cathepsin L2 (Santamaria et al., 1998b), cathepsin V (Adachi et al., 1998), and cathepsin U (Brömme et al., 1999). A partially active cathepsin V protein has been obtained by expression in E. coli (Santamaria et al., 1998b), and the proform has been expressed in Pichia pastoris and characterized (Brömme et al., 1999). The recombinant zymogen is autocatalytically activated at acidic pH. The enzyme has a pH activity profile similar to that of cathepsin L, with an optimum of pH 5.7. Unlike cathepsin L, cathepsin V is more stable at neutral pH, although much less so than cathepsin S, with only around 15% of activity remaining after 30 minutes at pH 7.0, 37°C. The S2 binding pocket accepts both aromatic and non-aromatic hydrophobic residues. In contrast, the closely related cathepsin L prefers an aromatic residue, while the related cathepsin S prefers a non-aromatic residue. Like cathepsin S, but unlike cathepsin L, collagen is a poor substrate for cathepsin V. Although closely related to the widely expressed cathepsin L, cathepsin V is expressed in a limited number of tissues, primarily the thymus, testes, and corneal epithelium (Adachi et al., 1998; Santamaria et al., 1998b; Brömme et al., 1999). It may be the most abundant peptidase in the corneal epithelium, but the protein is not found in tear fluid.
A human genomic DNA clone was isolated during efforts to clone cathepsin L that had 88% similarity to the cathepsin L coding sequence and a similar exon arrangement (Bryce et al., 1994). This gene, designated cathepsin L-like gene 1 (CTSLL1), and at least two other related genes (CTSLL2 and 3), were mapped to three different positions on chromosome 10q, and are thus distinct from cathepsins L and V. The three genes were estimated to have evolved during the early primate radiation. Evidence was found for transcription of CTSLL2 and 3 (but not CTSLL1) in a variety of tissues. Both CTSLL1 and 2 were shown to contain the same premature stop codon early in the reading frame, and thus would not produce a functional cathepsin. CTSLL3 did not contain this stop codon but was not fully sequenced. Therefore, it is not known whether it encodes a functional protein.
(ii) Functions
Thymus expression suggests that cathepsin V might be involved in some aspect of the immune system, as has been shown for cathepsins L and S. Interestingly, cathepsin V is expressed at significant levels in various cancer cells, but not in the corresponding normal tissue, and the expression pattern does not parallel that of cathepsin L (Santamaria et al., 1998b). These results suggest that cathepsin V may be involved in tumor processes, as has been suggested for other cathepsins (see below).
(e) Cathepsins J, P, CLRP, M, Q, R, and the testins
(i) Properties and tissue distribution
Several placenta-specific CPs have recently been cloned from rodents, starting from unique EST sequences (Sol-Church et al., 1999a, 2000b,c; Tisljar et al., 1999). The mouse CPs designated cathepsin P (Sol-Church et al., 1999a) and cathepsin J (Tisljar et al., 1999) differ by only a single residue insertion, and are most likely the same protein. Sequence similarity and phylogenetic analysis (see Figs. 1, 2 ) suggest that cathepsin P is the mouse ortholog of a rat placenta-specific CP designated cathepsin L-related protein (CLRP) (Conliffe et al., 1995), and cathepsin R (Sol-Church et al., 2000c) is the mouse ortholog of rat cathepsin Q (Sol-Church et al., 2000b). A third distinct mouse placenta-specific CP, designated cathepsin M (Sol-Church et al., 2000a), is closely related to these proteins, although its branch point is less certain (Fig. 2 and data not shown). These closely related CPs are in turn most closely related to the vertebrate cathepsin L/V branch (Fig. 2). Cathepsin J/P was shown to map to the same region of mouse chromosome 13 as cathepsin L, consistent with a duplication of a common ancestor of these genes (Tisljar et al., 1999). However, substitutions in the residues located in the S1 and S2 subsites of cathepsin M suggest that these placenta-specific CPs will cleave at different sites. It has been suggested that these CPs are involved in embryo implantation, placental function, or both. It is not yet known if there are any human orthologs of these genes, although the phylogenetic tree suggests that they evolved prior to the mammalian radiation, indicating that this is a distinct possibility.
Testins I and II (testin) were first identified as moderately abundant testosterone-inducible glycoproteins synthesized and secreted by rat Sertoli cells (Cheng et al., 1989). They are produced by differential processing of a common precursor protein, and testin II has a 3-residue N-terminal extension (Grima et al., 1995). Testin is clearly homologous to the cathepsin L-like CPs (Figs. 1, 2 ). However, the active-site cysteine residue has been changed to a serine. While proteolytic activity is still theoretically possible, no such activity has been detected, nor has any peptidase-inhibitory activity been found, although a comprehensive analysis of substrates has not been conducted (Cheng et al., 1993; Grima et al., 1995). No human orthologs of testin have been reported, and a search of the near-complete human genome database revealed no close matches other than cathepsin L, raising the possibility that testin is rodent-specific, despite the apparent early origin of the protein (Fig. 2).
Sertoli cells are connected by occluding junctions to form a blood-testis barrier, protecting the cells at various stages of sperm formation from the immune system, and providing support and nutrition to the developing sperm via specialized junctions. Sertoli cells are the major secretory cells of the testes, and are also the major phagocytic component of the seminiferous epithelium responsible for removal of the residual bodies. They also secrete fluid into the seminiferous tubules to transport sperm. Secreted testin is localized at the cell surface near the basal lamina of the seminiferous epithelium during most stages of spermatogenesis, but levels near the modified adherens junctions between Sertoli cells and heads of spermatids rise dramatically in association with the disruption of these junctions as the germ cells traverse through the tissue (Grima et al., 1997, 1998). Testin is also found at significant levels in the rat ovary, and at very low levels in the epididymis, kidney, and small intestine (Grima et al., 1995).
(ii) Functions
It has been suggested that the placental CPs are involved in embryo implantation, placental function, or both. The existence of primate orthologs, or expression in oral tissues, is unknown. The function of testin is unclear. Once secreted, it can become tightly bound to the cell membrane. Therefore, testin might be a structural component of specialized junctions, or it might be involved in recycling junctions that are disrupted during tissue remodeling. Expression levels in rat testes and kidneys during development are coincident with tissue remodeling. In the ovary, the levels change during the estrus cycle, with the highest levels coincident with the highest rate of junction turnover. Thus, high testin levels are generally associated with disassembly of certain cell-cell junctions (Grima et al., 1997, 1998). The lack of peptidase or inhibitory activity has been presumed to mean that testin does not mediate cell junction damage or provide tissue protection, and studies with agents that disrupt Sertoli-germ cell junctions indicate that it is unlikely that it functions to re-assemble the junctions. One suggested possibility is that it might be induced in response to cell wounding (Grima et al., 1998). Cell-cell junctions are essential to the function of several tissues in the oral cavity, but there have been no reports of testin in the oral cavity or developing salivary glands of rodents. Curiously, several cystatin-like proteins of unknown function (e.g., the testatins) are also expressed in the testes. The possibility of species-specific differences in CP genes (and the definite difference between rodents and humans in their salivary cystatins) raises a note of caution for the extrapolation of conclusions between species.
(III) Calpains
The calpains in family C2 (Carafoli and Molinari, 1998; reviewed in Ono et al., 1998) are intracellular Ca2+ regulated peptidases found in fungi and higher eukaryotes, but not as yet in bacteria or plants. Some calpains are ubiquitously expressed in tissues, while others are tissue-specific. Several lines of evidence suggest that calpain-catalyzed limited proteolysis is an important regulatory mechanism in several cellular functions, and that activation of calpain itself is coupled to intra- and extracellular signals. Cytoskeletal proteins, certain membrane proteins, and particular cytosolic proteins appear to be preferred substrates. Despite intensive study, the functions of the calpains remain ambiguous, although they have been implicated in various pathological processes. Recently, mutations in the muscle-specific calpain 3 gene (CAPN3) have been shown to underlie one of the limb-girdle muscular dystrophies (reviewed in Bushby, 1999). The calpains are specifically inhibited by calpastatin, an intracellular protein with no homology to the cystatins. Almost nothing is known regarding calpains in salivary glands, the oral epithelium, or teeth.
(IV) Mammalian Peptidases of Clan CD
The CD clan consists of clostripain (C11, from Clostridium histolyticum), the gingipains (C25, from P. gingivalis), and the legumain (asparaginyl endopeptidase) (C13) and caspase (C14) families (superfamilies according to some authors) (Barrett et al., 1998). These are all very specific endopeptidases, and based on similarities in the active-site motif (His-Gly-spacer-Ala-Cys), substrate specificities dominated by interactions with the S1 subsite, and potentially similar structures, it is likely that these four families share a very distant but common ancestry (Chen et al., 1998).
(A) The legumains
Legumains have strict cleavage specificity for peptide bonds with Asn in the P1 position. They were first identified in plants (where they appear to be involved in processing and mobilization of storage protein precursors during development and germination), but have since been found in mammals and parasites (reviewed in Barrett et al., 1998). Recently, the mammalian lysosomal legumain has been shown to be the principal enzyme involved in processing a microbial antigen for class II MHC presentation (Manoury et al., 1998). Curiously, legumain has been found in marrow plasma from normal donors, and has been shown to be a significant inhibitor of osteoclast formation and bone resorption, suggesting that it may have a physiological role as a local inhibitor of osteoclast formation and activity (Choi et al., 1999).
(B) The caspases
(i) Properties
The caspases (
(ii) Function of caspases in inflammation and apoptosis
The mammalian caspases can be subdivided into two families (subfamilies) based on sequence similarity. The interleukin-1β-converting enzyme (ICE, caspase-1) subfamily contains caspases 1, 4, 5, and 11-13 (caspases 11 and 12 are mouse enzymes with no known human equivalent); the second subfamily contains caspases 2, 3, and 6-10 (Thornberry and Lazebnik, 1998). Although the functions of all of these caspases have not been fully established, at least some members of the first subfamily act primarily in cytokine processing, while members of the second subfamily are intimately involved in apoptosis: a controlled disassembly of the cell. The role of caspases in these processes is complex: They can have redundant functions, or act in a cell- and stimulus-specific manner.
Interleukin-1β (IL-1β) is a potent pyrogen and pro-inflammatory cytokine primarily released by activated monocytes and macrophages. It is synthesized as an inactive proprotein that lacks a signal peptide. Processing to a heterodimer and secretion are co-temporal, and proteolysis appears to be mediated by membrane-associated ICE during membrane translocation. ICE (caspase-1) is the only identified caspase that can process pro-IL-1β, and macrophages from ICE-deficient mice do not release the mature protein on stimulation. Further, these mice are resistant to LPS-induced lethal endotoxemia (reviewed in Dinarello, 1998). Unlike caspase-1, caspase-11 is highly inducible by LPS. This enzyme physically interacts with pro-ICE in cells, and is essential for the activation of ICE, although probably not by direct processing (Wang S et al., 1998). In turn, cathepsin B, which could be released to the cytosol by oxidative stress or ischemia, or by perturbations in the lysosomal membrane induced by an acidic sphingomyelinase involved in inflammation signaling, has been implicated in the activation of procaspase-11 (Schotte et al., 1998). The functions of other members of this ICE subfamily are less clear.
Caspase-1 knockout mice are developmentally normal. In contrast, caspase-3, -8, and -9 knockout mice show severe developmental abnormalities and early lethality as a result of defective heart muscle development or the lack of apoptosis in the proliferative neuroepithelium, although multiple other tissues are also affected, such as craniofacial development (reviewed in Salvesen and Dixit, 1997; Nunez et al., 1998). Programmed cell death is therefore essential to the modification of cell populations during development. Apoptosis-involved caspases can be divided loosely into initiators and effectors (executioners) that are assembled into an activation cascade. A proapoptotic signal activates an initiator caspase, and, in turn, effector caspases become active. Effector caspases perform limited cleavage of a distinct subset of cellular proteins that fall into two broad categories: regulators of apoptosis, and housekeeping or structural proteins (reviewed in Cohen, 1997; Nunez et al., 1998; Thornberry and Lazebnik, 1998; Wolf and Green, 1999). In mammals, caspase-2 and caspases-8-10 serve as initiators, and caspases-3, -6, and -7 as effectors (reviewed in Nunez et al., 1998; Wolf and Green, 1999). Initiator but not effector caspases have long prodomains that contain modules which mediate protein-protein interactions between the caspases themselves and with regulatory adapter molecules. Two types of modules have been defined: death effector domains (DEDs), and caspase recruitment domains (CARDs). These modules allow for the assembly of proteins into oligomeric structures, and this aggregation induces autoprocessing and activation of the initiator caspases (reviewed in Nunez et al., 1998; Wolf and Green, 1999; Stennicke and Salvesen, 2000). Different initiator caspases mediate the response to different proapoptotic signals, and they are coupled to these signaling systems via adapter molecules containing DED or CARD modules.
Pathways of activation can be divided into intrinsic and extrinsic (reviewed in Stennicke and Salvesen, 2000). Extracellular apoptotic signaling ligands, such as FasL, bind to transmembrane cell-surface death receptors of the TNFR1 family that in turn recruit and oligomerize adapter proteins such as FADD, that in turn recruit and oligomerize initiator procaspases, leading to autoactivation. Cytochrome c is released by mitochondria in response to a variety of apoptotic stimuli, a process controlled in part by pro- and anti-apoptotic members of the Bcl-2 protein family. Cytochrome c binds the regulatory adapter protein Apaf-1 in the cytosol, which in the presence of adenine nucleotides can associate with procaspase-9. Oligomerization of Apaf-1 via CARD modules leads to procaspase-9 aggregation and activation. Activated caspases-8 or -9 can in turn activate an overlapping set of effector caspases that includes caspase-3, -6, and -7 (reviewed in Nunez et al., 1998).
Non-caspase peptidases can also activate caspases and apoptosis (reviewed in Wolf and Green, 1999; Stennicke and Salvesen, 2000). The best-characterized is the aspartate-specific serine peptidase granzyme B, a potent activator of procaspases-3 and -7 that is delivered into target cells by cytotoxic T-lymphocytes. Recently, cathepsin B has been shown to be involved in TNF-α-mediated hepatocyte apoptosis (Guicciardi et al., 2000). TNF-α binds to the TNFR-1 receptor, which oligomerizes. This can trigger apoptosis by receptor interaction with DED-containing proteins, subsequently leading to activation of caspase-2 and -8. However, TNF-α signaling can also activate acid sphingomyelinase in acidic vesicles, an enzyme implicated in generation of apoptosis effectors, and suggesting an effect on lysosomes. TNF-α was shown to promote translocation of cathepsin B from a vesicular compartment to the cytosol, perhaps mediated by activation of caspase-2 or -8. In a cell-free system, cathepsin B and cytosol were shown markedly to enhance the release of cytochrome c from mitochondria, and catB-/- mice were shown to be substantially resistant to TNF-α-induced liver injury. There is therefore a connection among pro-apoptotic signaling, lysosomes, and cathepsin B.
Calpains (family C2) have been shown to cleave caspase-8 and -9, and may be negative regulators of apoptosis by inactivating initiators (Chua et al., 2000). The only known endogenous mammalian caspase inhibitor is X-linked inhibitor of apoptosis protein (XIAP), a member of a family of metazoan proteins first identified in baculovirus that targets caspases-3 and -7 (reviewed in Nunez et al., 1998; Sun et al., 1999). Viruses, e.g., cowpox, also produce inhibitors that block apoptosis (reviewed in Cohen, 1997; Thornberry and Lazebnik, 1998).
(iii) Apoptosis and oral tissues
The role of apoptosis in craniofacial development was noted above (reviewed in Graham et al., 1996). TdT-mediated dUTP-biotin nick-end labeling (TUNEL), a marker for DNA fragmentation, has been used to examine apoptosis in other oral tissues. During formation of the periodontium, epithelial cells of Hertwig's root sheath trigger root dentin formation, and then most of these epithelial cells move away from the root surface to form the rests of Malassez. However, some of these cells remain for a time, perhaps to participate in cementum formation, before they disappear. In the developing rat first molar, apoptosis may contribute to this disappearance (Cerri et al., 2000). Apoptosis of ameloblasts in the enamel organ is prominent during tooth development, and limited numbers of odontoblasts appear to undergo apoptosis during dentinogenesis (see Bronckers et al., 2000, and references therein). In the continuously erupting mouse incisor, a high level of apoptosis occurs in the periodontal ligament cells just below the junctional epithelium. Recently, apoptosis has been implicated in the disappearance of osteoclasts from bone surfaces during force-induced tooth movement (Kobayashi et al., 2000). Apoptosis may also have a role to play in oral diseases. There is increasing evidence for a role for apoptosis in the loss of acinar cells in Sjögren's syndrome (reviewed in Patel and McHugh, 2000), and p53-mediated apoptosis may be involved in maintenance of immune homeostasis in chronically inflamed human gingiva by reducing the inflammatory cell population (Tonetti et al., 1998b). Interestingly, this study also showed significant TUNEL labeling of suprabasal cells of the junctional epithelium close to the bacterial plaque, suggesting that plaque may promote loss of cells by triggering apoptosis.
(V) Mechanisms of Exposure of Tissues to Host CPs and the Consequences
(i) Release of mature enzymes
The primary destination for the host CP cathepsins following their synthesis and processing through the ER and Golgi is the lysosome (see above). However, under certain circumstances, these lysosomal enzymes can be released from the cell. Further, there is evidence that the cathepsins can also be targeted for secretion more directly in certain cell types. Once released, these CPs can contribute to normal physiology and to pathology. It must be emphasized that our knowledge of these contributions is largely confined to cathepsins B, H, L, S, and K; even here, it is incomplete.
While neutrophils do not produce large amounts of CPs, the levels of cathepsins B, H, and L are high in macrophages, which also produce cathepsin S (Jochum et al., 1994; see above). Phagocytes can release lysosomal contents during normal phagocytosis if the phagosome doesn’t close before lysosome fusion. During frustrated phagocytosis, in which the phagocyte is unable to ingest a particle (such as a bacterial microcolony) due to steric hindrance, considerable quantities of active peptidases can be released into the local environment. Lysosomal contents can also be released extracellularly by neutrophils and macrophages in response to stimulation. Under certain conditions, human monocyte-derived macrophages can secrete substantial amounts of fully processed and active cathepsins B, L, and S, resulting in an enormous increase in their ability to degrade elastin (Reddy et al., 1995). In several chronic inflammatory diseases (e.g., tuberculosis), macrophage death is evident, with concomitant release of the cellular contents. Cathepsins L and S are potent elastases at their optimal pH (5.5). Several lines of evidence indicate that, at least in vitro, cathepsin S is largely responsible for the elastase activity of alveolar macrophages (even though these cells also produce macrophage elastase, a metallopeptidase), although its contribution in vivo remains to be established (Chapman et al., 1994). Thus, release of cathepsins S and L by phagocytosis, secretion, and death could contribute to tissue destruction by digestion of elastin and other components of the ECM. Osteoclasts adhere to and acidify the mineralized surface, thus providing a low pH environment into which they secrete lysosomal enzymes, including CPs (and in particular, cathepsin K), that mediate destruction of bone matrix proteins.
CPs may also be involved in monocyte migration, since type I cystatins (stefins A and B) blocked migration at physiologically relevant concentrations (Lah et al., 1993). A similar result was found for melanoma cell migration (Boike et al., 1992). Activation of urokinase-type plasminogen activator (u-PA) at the leading edge by membrane-localized CPs may be one mechanism involved. Although cathepsin B is 100-fold less active than cathepsin L against elastin (Mason et al., 1986), it can activate other enzymes such as u-PA and, in turn, collagenases (see below).
(ii) Release of proenzymes
In addition to the release or secretion of active, processed enzymes, procathepsins can also be secreted under various conditions, primarily related to tissue remodeling and invasion of ECM by cells. A variety of different normal cells secretes high levels of procathepsin L. A mouse thymic epithelial cell line was shown to secrete procathepsin L, and cathepsin L was localized immunohistochemically to the epithelial cells of the subcapsular layer and around the small vessels of the cortex (Kasai et al., 1993). Procathepsin L, but not cathepsin L itself, was found to promote proliferation of immature CD4-8- thymocytes in the presence of IL-1 and various mitogens. On its own, it had no activity, and therefore it apparently serves as a co-factor. Whether this has any functional relationship to the production of CTLA-2α and β by mouse-activated T-cells is unknown. Osteoclasts also secrete procathepsin L, which could potentially be activated in the acidic extracellular lacunae, and the secretion was up-regulated by parathyroid hormone, 1,25-dihydroxyvitamin D3, and tumor necrosis factor α (Tagami et al., 1994; Kakegawa et al., 1995; reviewed in Ishidoh and Kominami, 1998). Mouse placental tissue secretes procathepsin L in vitro, presumably released by the spongiotrophoblasts and phagocytic giant cell trophoblasts, which contain, in vivo, significant levels of immunoreactive cathepsin L protein, as well as mRNA (Hamilton et al., 1991). In contrast, other cell types contain little, if any, immunoreactive protein. Placental cathepsin L mRNA levels were found to decline from a peak around the time of implantation, in parallel with changes in c-ras Ha mRNA levels, and then to rise rapidly just before parturition. Sertoli cells secrete procathepsin L at specific stages of the cycle of the seminiferous epithelium and in primary culture, and levels of synthesis and secretion change considerably during the cycle (Erickson-Lawrence et al., 1991; O'Brien et al., 1993). The epididymis is an important site of sperm maturation where they acquire motility and the ability to fertilize. In the boar, procathepsin L mRNA is most abundant in the distal caput epididymis, and this region of the epididymis secretes high levels of procathepsin L into the luminal fluid, which may be involved in sperm maturation (Okamura et al., 1995). The level of PDGF mRNA is very high in the mid-caput epididymis (which precedes the distal caput epididymis), suggesting that PDGF may be responsible for the local expression and secretion of procathepsin L in the adjacent downstream tissue. Curiously, low-level expression of certain salivary cystatin genes in the human seminal vesicle (and prostate) has been detected by RNAse protection assay (data not shown). Secretion is not limited to procathepsin L. For example, procathepsin B is secreted by cultured human lung epithelial cells, and the proenzyme, which is relatively stable at neutral pH, can be converted to an active form by elastase (Burnett et al., 1995). Cathepsin B may therefore have a role to play in inflammatory airway disease. As noted above, pro-inflammatory cytokines stimulate procathepsin S production and release by cultured human cervical smooth-muscle cells.
Many cancer cells secrete functional procathepsin L. One of the first discoveries was the observation that Kirsten-virus-transformed mouse 3T3 fibroblasts secrete an abundant 39-kDa protein, originally called MEP (major excreted protein), that proved to be procathepsin L (Mason et al., 1987). Human non-small-cell but not small-cell lung cancer cell lines also secrete procathepsin L into the media (Heidtmann et al., 1993). Secretion of procathepsin L (and other proenzymes) by cancer cells may reflect a dysregulation of intracellular trafficking, with “excess” enzymes defaulting to the secretory pathway. However, as described above, normal cells will also secrete procathepsins, raising the possibility that cancer cells more specifically up-regulate this trafficking.
Why would normal tissues release an inactive proenzyme that requires acidic conditions for autoactivation, and what advantage, if any, would cancer cells gain? Procathepsin L is stable at near-neutral pH (Yamaguchi et al., 1990), and under certain conditions it may have proteolytic activity without processing (although this is somewhat controversial and may be limited to low-molecular-weight substrates). Further, it can be taken up by other cells and added to their endosomal proteolytic pool. Alternatively, it can be activated extracellularly. In the presence of certain glycosaminoglycans, reducing agents, and mildly acidic conditions (pH 5.5), procathepsin L will degrade extracellular matrix proteins such as fibronectin and laminin. In the presence of excess substrate, this degradation apparently occurs without conversion of the procathepsin L to lower-molecular-weight active forms lacking the pro-region, and the pattern of degradation fragments differs from that obtained with processed cathepsin L (Ishidoh and Kominami, 1995). However, it cannot be fully excluded that cleavage in these experiments is mediated by low levels of processed cathepsin L. Under these conditions in the absence of excess substrate, procathepsin L will undergo autocatalytic processing to a 31-kDa active form in a multistep process (Ishidoh and Kominami, 1994, 1995). In the presence or absence of GAGs, no autoactivation occurs above pH 6.5 (Mason et al., 1987; Ishidoh and Kominami, 1995). Breast cancer cells have been shown to have a high capacity to reduce the pH of their local environment (Montcourrier et al., 1997), which could therefore lead to degradation of extracellular matrix during metastasis by either mechanism. Chicken cystatin will block the low-pH activation of procathepsin L in vitro (Heidtmann et al., 1993), but it is not known if cystatins will interfere with activation in the presence of GAGs under mildly acidic conditions. Secreted cathepsin L can reduce opsonization, facilitating tumor survival, but paradoxically, it can also potentially inhibit tumor growth through the production of endostatin, an inhibitor of angiogenesis (see below).
Secreted procathepsin L can be taken up by cells with mannose 6-phosphate receptors, although the protein appears to be a poor ligand for the cation-independent mannose 6-phosphate receptor (Lazzarino and Gabel, 1990). Mouse spermatocytes and spermatids will take up mannose 6-phosphate-containing glycoproteins, including procathepsin L, secreted by Sertoli cells, in a receptor-mediated process that is inhibited by mannose 6-phosphate (O'Brien et al., 1993). Human alveolar macrophages will take up significant amounts of the 43-kDa human procathepsin L from serum, process it to the 25-kDa active form, and traffick it to the vacuolar compartment (Reilly et al., 1990). The myelomonocytic cell line THP-1 behaves similarly after being induced to differentiate with phorbol myristate acetate. However, in these cases, uptake does not appear to be dependent on mannose 6-phosphate groups (or mannose or fucose groups), since it is not blocked by an excess of this agent, nor is it apparently the result of non-specific pinocytosis. CP cathepsin trafficking into and out of cells is still incompletely understood, and likely involves different receptors and pathways, with different contributions depending on the type of cell. Further, procathepsin L has been reported to stimulate DNA synthesis in various cell lines, and the stimulation requires the proregion and the mannose 6-phosphate moiety (reviewed in Ishidoh and Kominami, 1998). Analysis of these data suggests that extracellular procathepsin L may have functions distinct from serving as a peptidase precursor.
(iii) Control of released enzymes
The acidic pH optimum of the lysosomal cysteine peptidases may serve to minimize their activity following “accidental” release of processed enzymes into normal tissue by cell death or autocatalytic zymogen activation. Extracellular cathepsin activity is also regulated by their lability at neutral pH. Cathepsin L has the lowest stability above pH 7.0 (see above). Thus, within an acidic environment (such as the extracellular acidified lacunae of osteoclasts or an inflammatory infiltrate), the cathepsins would be relatively stable and active: Enzymes that escape from this environment would be diluted into near-neutral pH serum, and would be reduced in activity and destabilized. In the case of cathepsin L, this would rapidly and substantially reduce activity. However, cathepsin S differs from cathepsin L by retaining 25% of its maximal activity at neutral pH, and the residual activity is comparable with that of neutrophil elastase itself. Inactivation of cathepsin K at neutral pH also occurs relatively slowly (see above). Therefore, the availability of excess CP inhibitors would be anticipated to be important in containing cathepsins S and K, both of which are inhibited by cystatins.
(VI) The Proteolytic Cascade: Interactions between Enzymes and Their Inhibitors
It is clear that regulation of host peptidase activity, and the balance between peptidases and their inhibitors, is vital to the maintenance of health and to ensuring an adequate but appropriate response to disease. Tissues and fluids contain a large number of protein peptidase inhibitors that inactivate serine peptidases (the serine peptidase inhibitors, or serpins; and the Kazal, Kunitz, and leukoproteinase types of inhibitors), CPs (members of the cystatin superfamily), and metallopeptidases (e.g., the tissue inhibitors, TIMPs) (reviewed in Travis and Salvesen, 1983; Twining, 1994; Roberts et al., 1995). There are no known specific inhibitors of aspartate peptidases, but it is generally assumed that α2-macroglobulin (a broad-spectrum peptidase inhibitor) serves this purpose. About 10% of the human plasma proteins are inhibitors, mainly serpins. Some serpins will also inhibit certain CPs, which may reflect the similarities in the catalytic mechanisms and active sites of the enzymes. However, members of the cystatin superfamily are by far the most prevalent inhibitors of cysteine peptidase activity. Usually, peptidase inhibitors are present in significant molar excess over peptidases. The cystatins are particularly abundant in saliva and tears. In general, once a peptidase-inhibitor complex is formed, it can bind to receptors on cells such as fibroblasts, hepatocytes, and macrophages, and be endocytosed and degraded. Relatively little is known about CP clearance.
The serum glycoproteins α2-macroglobulin and α1-proteinase inhibitor (also called α1-antitrypsin, α1-PI) are abundant peptidase inhibitors. α2-macroglobulin forms complexes with peptidases of all catalytic types. Cleavage of a “bait region” triggers a conformational change that leads to the peptidase being entrapped within the inhibitor, and thus sequestered from high-molecular-weight substrates. α1-proteinase inhibitor will inhibit nearly all serine peptidases that have been examined, including neutrophil elastase (for which it is the main serum inhibitor) and cathepsin G. α1-antichymotrypsin is an acute-phase protein: Its level in plasma increases dramatically in response to a variety of traumas. It is a specific inhibitor of chymotrypsin-like serine peptidases, such as cathepsin G and proteinase 3 (for which it is a major serum inhibitor) and the mast cell chymases. Proteins with specific CP inhibitory activity belong to one of three families, designated types 1, 2, and 3, in the cystatin superfamily (reviewed in Bobek and Levine, 1992; Abrahamson, 1994; Henskens et al., 1996; Dickinson, in preparation). Type I cystatins, also called stefins, are ca. 100 residues in length, and lack disulfide bonds and carbohydrate. They are usually intracellular, and are widely distributed in human tissues, particularly epithelia. The type 2 cystatins are typically 115-120 residues in size and contain 2 disulfide bonds. They are primarily secreted, and certain type 2 cystatins (the salivary cystatins) are found at very high levels in human saliva, and at lower levels in tears and seminal plasma. Cystatin C is a ubiquitously expressed protein found in all bodily fluids, and is thought to provide a general CP inhibitory activity. The salivary cystatins are clustered with cystatin C at Chr 20p11.2 (Dickinson et al., 1994; Thiesse et al., 1994). The type 3 cystatins comprise the H- (high-molecular-weight, ca. 120 kDa) and L- (low-molecular-weight, ca. 68 kDa) kininogens (originally called α1- and α2-cysteine proteinase inhibitors). These are plasma proteins that contain 3 tandem type-2 cystatin-like domains, and are also the precursor peptides for vasoactive kinins. These inhibitors form technically reversible, but very tight, complexes with CPs, and block the active site. The recently discovered equistatins are evolutionarily distinct CP inhibitors that have been proposed to be members of a new superfamily of CP inhibitors, the thyroglobulin type-1 domain inhibitors (Lenarcic et al., 1997). More recently, it has become evident that the divisions between the different types of inhibitors are not as sharp as was once thought. For example, the human squamous cell carcinoma antigens (SCCA1 and SCCA2) have 92% identity: SCCA2 is a potent serpin, but SCCA1 is a potent CP inhibitor and lacks activity against serine peptidases (Schick et al., 1998).
Under certain circumstances, the balance of peptidase and peptidase inhibitors can be pathologically disturbed (Lah et al., 1993; reviewed in Travis and Bangalore, 1993). Peptidases elaborated by micro-organisms, such as Porphyromonas gingivalis, provide potential triggers (reviewed in Potempa et al., 2000). Various peptidases can cleave and activate zymogens of other enzyme types, and cleave and inactivate heterologous inhibitors. Once in excess, some enzymes can activate their own zymogens and inactivate their own inhibitors. The opportunities for complex interactions between the various types of peptidases and their inhibitors, leading to loss of inhibitor activity and increase in peptidase activity, have parallels with the positive feedback amplification of the coagulation cascade. The increase in proteolytic activity resulting from this enzyme cascade has been called the proteolytic burst (Lah et al., 1993). Not only would the capacity to degrade tissue be greatly increased, but also peptide fragments produced are potent chemoattractants that would recruit additional phagocytes and the lysosomal enzymes they carry. CPs have the capacity to participate in the proteolytic burst (and their inhibitors, the cystatins, in its control). Thus, by participating in an amplification, the consequences of CP activity could be far greater than expected from the amount of enzyme present or the stability. A few examples will be presented to illustrate these points.
Neutrophils (polymorphonuclear leukocytes, PMNs) are specialized phagocytic cells that provide the first line of defense against infection and foreign particles. Chemoattractants generated by bacteria (e.g., formyl-Met-Leu-Phe), the complement system (e.g., C5a, the most potent protein neutrophil chemoattractant), other phagocytes (e.g., IL-8), opsonization, and damaged tissues result in an influx of neutrophils to the site of infection and provoke their activation. These activated cells proceed to attempt to destroy the pathogen with an arsenal of weaponry (reviewed in Sandborg and Smolen, 1988; Weiss, 1989; Miyasaki, 1991). Their importance is illustrated in individuals with defects in neutrophil function or neutropenia. They are at risk for a variety of infections, including periodontal disease (reviewed in Daniel and Van Dyke, 1996). However, these weapons are not “smart”, and the surrounding tissues can suffer considerable collateral damage. Indeed, much of the damage caused by an infection is likely to be mediated by the host response and ensuing proteolytic burst, rather than by the pathogen itself, and neutrophils are considered to be the primary mediators of connective tissue destruction seen in chronic inflammatory disorders such as emphysema and periodontal disease, as well as acute trauma- or infection-induced inflammation leading to multiple organ failure (reviewed in Weiss, 1989; Lah et al., 1993; Doring, 1994; Jochum et al., 1994; Travis et al., 1994; see below). For example, serum levels of released peptidases were found to be related to the severity of trauma- or infection-induced inflammation and to the risk of multiple organ failure (Jochum et al., 1994).
The primary mechanism by which neutrophils remove foreign agents is phagocytosis followed by intracellular degradation within phagosomes. This degradation is mediated in part by a diverse collection of peptidases. In the neutrophil, these include the azurophil granule serine peptidases elastase, cathepsin G, and proteinase 3, and the aspartate peptidase cathepsin D, and the metallopeptidases collagenase (MMP8) and gelatinase in other granules (Sandborg and Smolen, 1988; Twining, 1994). The serine peptidase concentration in neutrophils is a considerable 3 μM (Travis et al., 1994), and in a healthy individual an estimated 2 g of elastase and 1.5 g of cathepsin G are released each day (Travis and Bangalore, 1993). Neutrophil elastase is a powerful and indiscriminate peptidase that will cleave many polypeptides and proteoglycans, including proteins such as collagen and elastin that are otherwise relatively resistant to proteolysis (see the relevant chapters in Barrett et al., 1998, for details). Cathepsin G is also a potent activator of neutrophil collagenase (Knäuper et al., 1990). Both elastase and cathepsin G have been shown to bind strongly to neutrophils under physiological conditions. They remain fully active, and partially resistant to inhibition by protein inhibitors. Not only can they damage tissue directly, but these peptidases can also cause the proteolytic activation of peptidase zymogens of other systems, such as the coagulation cascade and fibrinolytic system. Elastase will activate procathepsin B secreted by lung epithelial cells (Burnett et al., 1995).
Monocytes and other cells of the immune system must penetrate the ECM of basement membranes and connective tissues, and fibrin barriers, to localize to a source of chemotactic signals, such as an infection. Similarly, cancer cells must penetrate ECM barriers in metastasis. Proteolytic cascades involving CPs have been implicated in these events. Cathepsins B and L (and plasmin) are likely catalytic activators of pro-collagenases (MMPs) and pro-urokinase-type plasminogen activator (u-PA) (Eeckhout and Vaes, 1977; Kobayashi et al., 1991; Goretzki et al., 1992). Cathepsins B and L can efficiently convert soluble and receptor-bound forms of single-chain urokinase-type plasminogen activator (sc-uPA, or pro-uPA) to the two-chain, active form, and conversion by cathepsin L is relatively efficient even at pH 7 (Kobayashi et al., 1991; Goretzki et al., 1992). uPA associates with cells via a uPA receptor, and activity can be localized at the leading edge of migration. u-PA will activate plasmin, which is in turn a powerful activator of many MMPs, which are major effectors of matrix degradation and collagen breakdown (reviewed in Birkedal-Hansen et al., 1993; Murphy and Gavrilovic, 1999). Plasmin itself will directly degrade several matrix components and fibrin.
An imbalance in peptidase activities can lead to the inactivation of α1-PI and α1-antichymotrypsin, thus allowing for an increase in serine peptidase activities that can, in turn, degrade inhibitors of other classes of peptidases. The importance of α1-PI is exemplified by the destruction of the lungs in emphysema. The normal lung contains large numbers of neutrophils. Individuals with an hereditary deficiency in α1-PI are strongly predisposed toward development of emphysema, and it is thought that this disease results from an imbalance in the neutrophil elastase:α1-PI ratio, leading to elastin degradation (reviewed in Evans and Pryor, 1994). Given the large number of neutrophils in the gingival crevice, it might be anticipated that hereditary deficiency in α1-PI would predispose to periodontal disease. Young to middle-aged individuals of the ZZ phenotype with reduced serum α1-PI levels were found to have deeper pockets in comparison with controls matched for the amount of dental plaque, consistent with earlier studies that suggested an association between genotype and periodontal disease (Fokkema et al., 1998). However, the relationship between α1-PI genotype and risk of periodontal disease remains to be fully characterized in a larger study. α1-PI is normally present in excess in serum and very rapidly inactivates elastase (reviewed in Weiss, 1989). However, it is cleaved in the reactive site region and inactivated by catalytic amounts of cathepsin L (Johnson et al., 1986). Further, reactive oxygen species produced by neutrophils, and those present in cigarette smoke, are potent inactivators of α1-PI, although the importance of this mechanism in vivo has been questioned (reviewed in Weiss, 1989; Simon, 1993; Travis and Bangalore, 1993; Evans and Pryor, 1994). Cigarette smoking is an identified risk factor for periodontal disease, although the mechanism of involvement remains to be established (Genco, 1996). When elastase is in molar excess, it can inactivate α1-PI (reviewed in Doring, 1994). Elastase will also inactivate α1-antichymotrypsin, and the resulting modified protein is a potent neutrophil chemoattractant (Potempa et al., 1991). Further, neutrophil elastase degrades the protein inhibitors of the clotting and fibrinolytic proteolytic cascades, and will degrade TIMPs, thereby perturbing the balance of tissue MMPs and their inhibitors (Okada et al., 1988). The leukocyte collagenase MMP-8, which could be activated by either cathepsins B or G, will cleave the reactive loop and inactivate many serpins, and is highly effective against α1-PI (reviewed in Simon, 1993). MMP-8 is strongly inhibited by doxycycline, and it has been suggested that this collagenase is one target for this antibiotic that makes it of use in treating inflammatory disorders, such as periodontal disease (Ashley, 1999). Elastase will remove an N-terminal decapeptide from cystatin C, greatly reducing its inhibitory activity against cathepsins B and L (Abrahamson et al., 1991). Therefore, once active, elastase will degrade a major specific inhibitor of CPs. The aspartic peptidase cathepsin D will also inactivate cystatin C, and an inhibitory domain of kininogen, in an acidic environment (Lenarcic et al., 1991). Cathepsin L itself (but not cathepsins B or H) can cleave cystatin C and produce a protein with an 11-residue N-terminal truncation that is a much less effective inhibitor of cathepsin CPs (Popovic et al., 1999).
These examples indicate that the activity of a given peptidase in each class is held in check by the presence of inhibitors which are often specific for that class. However, once the balance is perturbed for any one peptidase, allowing excess activity, that peptidase can potentially initiate a proteolytic cascade by activating proenzymes in other classes, and inactivating their inhibitors. In turn, these enzymes can now activate latent forms of the initial trigger, and inactivate its inhibitors, leading to positive feedback and a burst of proteolytic activity. Collectively, released host peptidases have the capacity to cause extensive damage to cells and tissues unless otherwise contained, especially in conjunction with the simultaneous release of reactive oxygen species. Although the proportion of tissue damage that might be directly attributable to host CPs is not clear, and may even be relatively minor in comparison with that caused directly by other types of peptidase, CPs can, as just illustrated, contribute to an amplification of the proteolytic burst by several mechanisms. However, it is also likely that there is redundancy in the proteolytic burst components. Thus, in a given disease population, the same outcome could result from moderately different pathways, and correlations between peptidase levels (and their inhibitors) and disease activity, which could be useful diagnostically, may not be as statistically distinct as we might hope.
(VII) Interaction of Host CPs with the Immune System
In addition to the direct degradation of tissues, peptidases and peptidase-inhibitor complexes produced in a proteolytic burst also interact with the immune system, influencing immune cell behavior, including infiltration. Directly and indirectly, host CPs can modulate the immune system by several mechanisms.
Vertebrates can respond to exposure to foreign material (such as a parasite) with two immune defense systems. Innate (natural) immunity is the first line of defense. It is an otherwise non-specific response mediated in part by myeloid phagocytic cells that can recognize the material as foreign, and then proceed to attempt to engulf the material, which then becomes contained within an intracellular phagosome. Fusion of this vacuole with lysosomes allows enzymes, such as cathepsins, to begin digestion of the material. Phagocytes can also produce highly toxic reactive oxygen and nitrogen intermediates that can kill pathogens. The reticulo-endothelial system consists of various fixed phagocytes distributed in tissues. The mobile phagocytes consist primarily of polymorphonuclear leukocytes (PMNs, granulocytes), subdivided into the neutrophils (about 70% of total blood leukocytes), the eosinophils (2-5%), and the basophils (0.5%). Circulating monocytes (4-10% of leukocytes) can be stimulated to migrate into tissues in response to chemotactic signals, where they can differentiate into phagocytic tissue macrophages that express CPs such as cathepsins B and S. Macrophages also produce cytokines (e.g., TNF and IL-8) that recruit other inflammatory cells, particularly neutrophils. Recruitment can be indirect, through induction of chemokine expression and adhesion molecules in endothelial cells, gingiva, or osteoblasts (reviewed in Graves, 1999). Cytokines, especially TNF, are also important mediators of inflammation: a series of vascular events intended to serve as an additional defense mechanism. If phagocytic cells are unable to clear foreign material within a short period, vasoactive amines are released by mast cells in response to cytokines and complement fragments. These lead to vasodilation and increased blood flow to the area, increased capillary permeability, and pain. The vascular changes cause localized edema and redness, and allow for an enhanced influx of phagocytic cells in response to chemotactic signals. Neutrophils arrive within 30-60 minutes, macrophages within 5-6 hours, and if the signals still persist, lymphocytes will begin collecting. Kinins are peptides released locally by cleavage of plasma kininogens. They are hypotensive, increase vascular permeability, contract smooth muscle, and induce fever and pain. Although the best-understood mechanism for release of kinins is the plasma contact activation system involved in blood clotting, it has recently been shown that a CP on the surface of endothelial cells may be important in kinin release (Rojkjaer and Schmaier, 1999). Cathepsin X may potentiate bradykinin activity (see above). In addition to vascular changes, the clotting mechanism is locally activated, which serves to trap particles. Fibrin degradation products are also potent chemotactic agents.
The immune systems also interact with the complement system in several ways. Complement is a system of 14 components comprising two proteolytic cascades, the alternative and the classical pathways, that in turn can activate a common pathway which leads to the assembly of a membrane attack complex. This can kill cells non-specifically by disrupting membrane function. The alternative pathway can be activated non-specifically by activating surfaces, such as the LPS on bacteria. Spontaneous proteolytic cleavage of C3 releases C3b, which is stabilized by microbial surfaces. In association with other proteins, this leads to the formation of surface-localized alternative pathway C3 convertase, which cleaves C3 to C3a and C3b. C3b and other proteins form the alternative pathway C5 convertase. Conversion of C5 to C5a and C5b initiates assembly of the membrane attack complex. Microbial-surface bound C3b (and iC3b produced by Factor I cleavage) binds receptors on the surfaces of neutrophils and macrophages, promoting the adherence of these phagocytic cells to the organism, a process called opsonization. Fragments C3a, C4a, and C5a are anaphylatoxins; they trigger the release of soluble inflammatory mediators such as histamines from basophils and mast cells. C5a is also a potent chemoattractant for neutrophils and monocytes that stimulates directed migration. CP activity may be required for certain chemotactic responses. CP inhibitors, including E-64, were shown to reduce the chemotactic response of human neutrophils to C5a, but had no effect on the response to IL-8 or f-Met-Leu-Phe (Barna and Kew, 1995). Chicken egg white cystatin had a weak but statistically significant inhibitory effect. Consistent with this, human cystatin C is a powerful inhibitor of the chemotactic responses of PMNs to complement-derived factors (Leung-Tack et al., 1990). Procathepsin L released by cancer cells and localized at the cell surface has been implicated in reducing the ability of the immune system to destroy cancer cells by interfering with C3 function through cleavage, thereby reducing both opsonization and complement-mediated lysis (Jean et al., 1995; Frade et al., 1998). Elastase (increased in the proteolytic burst) also down-regulates neutrophils by impairing opsonization due to cleavage of C3 and the Fc region of immunoglobulins, and cleavage of the complement receptor. Neutrophil elastase-treated immune complexes fail to stimulate the oxidative burst (reviewed in Doring, 1994). Another consequence of elevated pro-inflammatory cytokines in the serum is the acute phase response by hepatocytes (reviewed in Moshage, 1997). The liver is induced to synthesize and secrete a variety of proteins, including peptidase inhibitors, and reduce the synthesis of certain other proteins, such as albumin. This response is thought to augment the immune response and protect tissues, although its exact function is unknown.
Macrophages (and other antigen-presenting cells, APCs) are also important in processing foreign proteins for presentation to the second defense system, adaptive (acquired) immunity, which provides a highly specific response to a challenge. The potential roles of CP cathepsins and legumain in antigen production were described above. Foreign protein antigens are processed by APCs, and peptide epitopes are presented on the cell surface as epitope-MHC complexes. These are recognized by CD4+ T-helper cells, which are stimulated by the IL-1 released by activated macrophages. Activation of T-helper cells leads to proliferation and differentiation. The acquired immune response has two arms: humoral immunity, which is based on antibodies, and cellular immunity, which involves interactions between cell surfaces. Which arm is predominantly activated depends on a complex series of interactions among the pathogen, T-helper subsets, other immune cells, and their products (reviewed in Finkelman and Urban, 1992; Okada and Murakami, 1998). There are two major T-helper subsets, Th1 and Th2, that differ in the cytokines they produce, and corresponding Th1 and Th2 responses to an antigen that differ in their effector cells. As a crude generalization, a Th1 response protects the host against intracellular pathogens (and some parasites), and a Th2 response against extracellular pathogens and toxins.
The balance between these responses has been implicated in progression of periodontal disease (reviewed in Okada and Murakami, 1998). The effects of pharmacological inhibition of cathepsin B, which can alter the response to an antigen in an animal model (although the exact mechanism is somewhat uncertain, given the absence of effect in a knockout mouse), and the expression of type 2 cystatins in antigen-presenting cells (e.g., Ni et al., 1998; Hansen et al., 2000) raise the possibility that host CPs and their endogenous inhibitors may have a role in modulating the Th1/Th2 response. Consistent with this, endocytic uptake of procathepsin L via mannose 6-phosphate receptors along with pigeon cytochrome c antigen was shown to lead to inhibition of antigen presentation, probably by degradation of the antigen in acidic endosomes to non-stimulatory fragments (McCoy et al., 1988). However, procathepsin L did not affect the presentation of an antigenic determinant derived from lambda repressor, most likely due to lack of cleavage sites in the antigenic region. Further, various pathogens appear to use CPs to influence the host immune response (Dickinson, manuscript in preparation). The recent availability of transgenic knockout mice lacking various CPs provides a powerful tool for the future elucidation of the contributions of these enzymes to the immune response.
Other peptidases, corresponding peptidase-inhibitor complexes, and inhibitor cleavage products are produced in a proteolytic burst with the involvement of CPs (see above), and these can also interact with the immune system. For example, α-1-antichymotrypsin-cathepsin G complexes were found to stimulate a nearly five-fold increase in IL-6 production by human lung fibroblasts (Kurdowska and Travis, 1990). α-1-antichymotrypsin can be inactivated by neutrophil elastase, and in this form will increase IL-6 production by fibroblasts. Il-6 in turn induces synthesis of acute phase proteins (which include several peptidase inhibitors, including α-1-antichymotrypsin). α-1-proteinase inhibitor that has been inactivated by macrophage elastase (a metallopeptidase) cleavage is a potent neutrophil chemotactic factor, as is the complex of neutrophil elastase and α-1-PI (Banda et al., 1988a,b). These would contribute to further recruitment of neutrophils to a site of inflammation. Levels of serine peptidase-inhibitor complexes, such as α-1-antiproteinase-neutrophil elastase and α-1-antichymotrypsin-cathepsin G, are controlled, in part, by multifunctional receptors on fibroblasts, hepatocytes, and other cells (Poller et al., 1995). These receptors facilitate the uptake and subsequent degradation of the complexes.
(VIII) CPs and Sjögren's Syndrome
Host CPs may be involved in the production of autoantibodies. Salivary and lacrimal gland acinar cells express MHC class II molecules, and these molecules appear to be up-regulated in response to a lymphocytic infiltrate. They are therefore potential APCs. In cultured rabbit lacrimal acinar cells, MHC class II molecules, the ribonuclear protein La/SSB autoantigen, and cathepsins B and D were shown to co-localize to membrane-bound compartments involved in trafficking to pre-lysosomes, secretory vesicles, and the basolateral membrane (Yang et al., 1999a). Further, acute cholinergic stimulation of cultured rabbit lacrimal gland acinar cells with carbachol caused changes in the distribution of procathepsin B, with a two- to three-fold increase in a basolateral membrane fraction, where it might be involved in antigen generation (Yang et al., 1999b). These observations support the hypothesis that MHC class-II-positive acinar cells, and endogenous CPs, may be involved in autoantigen processing and presentation to CD4+ T-cells, leading to autoantibody production, and to the development of Sjögren's syndrome. With the use of a substrate that can be hydrolyzed by papain-like enzymes (as well as by caspases), elevated levels of CP activity were found in the submandibular glands and saliva of non-obese diabetic (NOD) mice, which are a model for Sjögren's syndrome (Robinson et al., 1997). Mammalian legumain, which has been shown to be important in microbial antigen-processing (see above), has not been examined in salivary glands. It would be of interest to examine the expression profiles of CPs (especially cathepsins F and S) during the progression of Sjögren's syndrome.
(IX) CPs and Periodontal Disease
Even in health, there is a significant flux of PMNs into the oral cavity from the gingival crevice, promoted by chemotactic cytokines such as IL-8, produced by the junctional epithelium, and this flux can increase considerably with gingivitis and periodontal disease (Cimasoni, 1983; Tonetti et al., 1998a). A proportion of PMNs die and undergo lysis before being cleared by swallowing, and therefore the oral cavity is continuously exposed to host proteolytic enzymes. Periodontitis is characterized by gingival inflammation and the loss of the connective tissue supporting the teeth, including the periodontal ligaments and surrounding alveolar bone (Academy Reports, 1999). An acute inflammatory phase prompted by the expanded microflora of the pocket brings large numbers of neutrophils into the junctional and sulcular epithelium to form the “leukocyte wall” (Miyasaki, 1991). A subsequent chronic inflammation recruits monocytes, macrophages, and lymphocytes to the epithelia. Periodontal ligaments are destroyed, and osteoclast activity in alveolar bone is increased, resulting in loss of mineralized tissue. Immune function in the gingiva in health and disease is in part the result of interactions of cells with antigens, cytokines, and chemokines in complex patterns of production and response that remain to be fully elucidated, but which do not fall clearly into a Th1- or Th2-type response (reviewed in Wilson et al., 1996; Okada and Murakami, 1998; Graves, 1999). Organisms such as P. gingivalis have been identified as potential periodontopathogens, although the infection is distinctly polymicrobial (Dahlén, 1993; Genco, 1996).
Several representatives of the four major catalytic types of peptidases have been reported in inflamed gingiva and crevicular fluid, derived from the inflammatory cells, fibroblasts, and epithelial cells of the pocket (Lah et al., 1993). In the crevicular fluid of periodontal patients, the concentration of neutrophil peptidases can rise to nearly 300 μM (Travis et al., 1994). Thus, there is ample opportunity for peptidases, including CPs, to affect the gingival tissues and the intricate cross-talk of immune signaling systems. Further, alterations of the local cytokine cocktail could potentially alter the local synthesis and secretion of CPs, although our understanding of CP regulation is insufficiently developed to make predictions as to the consequences. Efforts have been made to determine if there is a correlation between levels of host peptidases in gingival tissues and crevicular fluid and the severity of periodontal disease. This would presumably indicate the extent of underlying host inflammatory activity and tissue damage, and might therefore have diagnostic value (reviewed in Lah et al., 1993; Zahradnik, 1998). These studies have focused primarily on the predominant neutrophil lysosomal enzymes elastase, cathepsin D, and CP cathepsins B, H, and L, with the idea that these would be important enzymes mediating tissue destruction, together with a select few inhibitors. As outlined above, this is likely to be an oversimplification of the system. Such studies also face several technical difficulties. Peptidase levels need to be correlated with disease activity, which is not a trivial task. It is hard to establish a baseline for activity in crevicular fluid, since only very small amounts can be collected from healthy individuals, and there are difficulties in measuring these volumes accurately. Trauma induced during sampling can lead to excessive contamination with blood components. The use of glass capillaries to collect GCF is likely to be more traumatic than the use of paper strips. Further complications are the concomitant release of peptidase inhibitors when tissue is homogenized, which can interfere with activity-based assays, and the presence of active and inactive forms of enzymes (both of which may react with test antibodies). It should also be noted, when reports are compared, that enzyme amounts can be reported in different ways.
Several studies have examined CP activity in gingival crevicular fluid (GCF), following an early report that cathepsin B activity could be detected in GCF from gingivitis patients, although the exact periodontal status of the collection sites was not specified (Eisenhauer et al., 1983). These studies have not been entirely consistent, no doubt due in part to the issues raised in the preceding paragraph. Kennett et al. (1997a) collected GCF from chronic periodontitis patients using Whatman No. 1 paper, and then eluted enzymes into detergent-free buffer by agitation at 4°C. A large proportion of total enzyme activity (99% of elastase and 87% of cathepsin B) was lost by centrifugation of the sample, which would remove cell-bound enzyme. Sonicating the eluate had little effect, but addition of detergent (0.1% Triton X-100) to the eluate after removal of the paper strip was found to increase the total activity in the sample greatly, possibly due to effective release of bound enzymes from cells. This was not complete, however, since centrifugation still caused a partial but significant loss in activity. The activity retained by the paper was not studied. Kunimatsu et al. (1990) collected GCF from chronic adult periodontitis patients and experimental gingivitis subjects for a fixed time on paper strips, and recovered enzymes by sonication in buffer. Selective synthetic substrates were used to determine the total enzyme activity of cathepsins B, H, and L per unit of collection time. The total collected activity of each enzyme was found to be correlated with the volume of GCF collected per unit of time (which was in turn correlated with the severity of disease). Thus, increased total amounts of enzymes were found at a site as tissue destruction increased. The total cathepsin L activity collected per unit of time showed a weak correlation with probing depth, but cathepsins B and H activities did not. The specific activity of each enzyme (i.e., units of enzyme activity/mg protein) was found to be negatively correlated with the volume collected/unit of time. One explanation for this observation is that as GCF flow rate (and disease severity) increases, more serum protein (and inhibitors) enter the pocket per unit of time. If this increase exceeds any increase in CPs released into the GCF, then the specific activity (units of CP activity/mg GCF protein) would decrease. In experimental gingivitis, no significant activity of either cathepsin B, H, or L was detected during the 21-day period tested. Lah et al. (1986) examined cathepsins L- and D-like activities in gingival fluid collected by glass capillary from a limited number of patients with periodontal disease, as indicated by pocket depth. Cellular material was removed by centrifugation, and activities in the supernatant were determined with the use of a protein substrate under conditions designed to be optimal for these enzymes. Proteolytic activity (in units of activity/unit volume) was found to increase with probing depth.
Cox, Eley, and co-workers (Eley and Cox 1991, 1992a,b,c, 1996; Cox and Eley, 1992; Kennett et al., 1994, 1997a, b) performed a series of studies aimed at determining the relationship among levels of elastase, cathepsin B/L, and other peptidases in GCF and disease status. GCF was collected by paper strips, followed by elution into detergent-containing buffer, but was not centrifuged. The synthetic CP substrate used did not discriminate between cathepsins B and L. Activities were expressed as both total enzyme activity units collected per unit of time and enzyme concentration (activity units/unit volume). In a cross-sectional study, GCF was collected from untreated chronic periodontitis patients (Eley and Cox, 1992a). Both the total enzyme activities collected per unit of time and enzyme concentrations of cathepsins B/L and elastase were found to be significantly correlated with clinical parameters of disease status. With total activities, cathepsins B/L and probing depth showed the best correlations, and good diagnostic specificity and sensitivity. Total enzyme activities and enzyme concentrations of elastase and cathepsins B/L were also shown to correlate with bone loss, although the relationship was strongest for total activity in inter- and intra-patient comparisons (Eley and Cox, 1992b). When enzyme levels were compared before and after periodontal treatment by scaling and root planing, there were reductions in all clinical parameters and levels of these peptidases, and there was a significant positive correlation with post-treatment levels and clinical parameters, although again the relationship was stronger for total levels rather than concentrations (Cox and Eley, 1992). Further statistically significant reductions in enzyme levels and concentrations were achieved following periodontal surgery (Eley and Cox, 1992c). Total enzyme levels fell by about 35-70% after scaling and planing, and by a further 70-90% after surgery, often to the point of being below detection limits. Importantly, a two-year longitudinal study of patients who had received periodontal treatment by scaling and planing demonstrated significantly higher cathepsin B levels (both total and concentration) in GCF from sites that subsequently showed rapid attachment loss, in comparison with paired control sites (Eley and Cox, 1996). Mean cathepsin B levels (both total and concentration) were significantly higher at rapid and gradual attachment loss sites than non-attachment loss sites in intra-patient comparisons.
Although these studies support a correspondence between CP activity and periodontal disease, it is perhaps surprising that a better correspondence is found between disease severity and total enzyme activity collected than with enzyme concentration. GCF flow rate increases with disease severity, as does the flux of granulocytes and macrophages into the pocket. These cells are responsible for much of the enzyme activity in GCF. If the amount of enzyme delivered to a site and flow rate increase roughly in proportion (albeit independently), then the enzyme concentration will also be roughly constant, even though much more enzyme activity would be delivered to the pocket per unit of time due to the higher flow rate.
Peptidase levels in gingival tissue have also been examined. The activities of cathepsins B, L, and D in homogenates of human gingival tissue from patients with gingivitis and moderate and severe periodontitis were found to be increased in the three groups in parallel with the severity of disease. However, the correlation coefficient with cathepsin D was much higher than for the CPs (Lah et al., 1985). Consistent with these observations, the levels of activities of cathepsins B and L were found to be significantly higher in gingival tissue from chronic periodontitis patients in comparison with levels in healthy gingiva (Eley and Cox, 1991). The levels of cathepsin B activity were greater than those of cathepsin L. In contrast to the results obtained from GCF, the mean patient enzyme activity did not correlate with clinical parameters, including probing depth. In inflamed gingiva from periodontitis patients, cathepsin B was localized by both histochemical and immunohistochemical techniques to CD68+ monocyte/macrophage cells, gingival fibroblasts, and possibly Langerhans cells (Kennett et al., 1994). Fibroblasts adjacent to the epithelium stained the most intensely. A diffuse staining was also observed over the epithelia, but it was not clear if this was due to intra- or extracellular enzyme. The staining pattern within macrophages and fibroblasts was consistent with a lysosomal distribution. This was confirmed for cathepsin B by ultrastructural localization with gold-conjugated antibody. However, both types of cells also showed surface labeling, raising the possibility of membrane-localized extracellular enzyme (Kennett et al., 1997b). Consistent with this, labeling was also observed on adjacent extracellular collagen fibers, indicating that these cells release cathepsin B. Cathepsins B, L, and D were found immunohistochemically in gingival fibroblasts and macrophage-like cells in gingival tissues from periodontitis patients, and a similar distribution for cathepsin D and L mRNAs was found by in situ hybridization (Trabandt et al., 1995). Again, the highest levels localized to fibroblasts adjacent to the epithelium. Expression of cathepsins B, L, and D mRNAs in gingival fibroblasts was confirmed by Northern blot analysis. Further, mRNA levels in early passaged cells from inflamed gingiva were significantly higher than those in identically cultured cells from non-inflamed gingiva. These studies suggest that gingival fibroblasts could be a significant source of CPs. An interesting possibility that remains to be examined is that they secrete procathepsins which are taken up by macrophages (see above). Consistent with these observations on gingival tissue, CD28+ monocyte/macrophages, which make up 10-20% of the cells in GCF, were shown by immunocytochemistry to contain cathepsin B (Kennett et al., 1997a). This study also demonstrated that most CD15+ granulocytes contained immunreactive elastase, although only a small proportion contained active enzyme as determined by cytochemistry. This may be due to the elastase being present in these cells as an immunoreactive but enzymatically inactive precursor. The discovery, since much of this work was published, of several additional Branch B cathepsins raises the obvious concern of antibody, substrate, and inhibitor cross-reactivity and -sensitivity.
u-PA, which can be activated efficiently by cathepsin B (see above), is present in gingival crevicular fluid at a significantly greater concentration than in plasma, as is tissue plasminogen activator. u-PA mRNA has been shown to be present in gingival tissues, primarily in cells of the junctional and sulcular epithelium—key sites in periodontal disease—although u-PA mRNA expression was limited to single cells, and tissue-type plasminogen activator appeared to be the predominant plasminogen activator in these tissues (Kinnby et al., 1999). However, as noted by the authors, the tissue samples were obtained from patients undergoing surgical therapy after a period of oral hygiene, and this distribution of activators may reflect a healing process. Tissue plasminogen activator itself can activate procathepsin B (Dalet-Fumeron et al., 1996). Collectively, GCF contains several identified host enzymes, often at significant levels, that could collaborate in a proteolytic burst. Cathepsin S largely accounts for the elastase activity of macrophages (Chapman et al., 1994) and would be expected to be important in the destruction of periodontal tissues, but there are no reports examining its activity in gingival crevicular fluid. Cathepsin K is likely to be a major mediator of alveolar bone loss. No studies of cathepsin K levels in periodontal lesions have been undertaken. It should be noted that in the studies cited above, the synthetic substrates used were not specific for cathepsins B, H, or L. Further, enzymes complexed with α2-macroglobulin can still be active with small substrates, leading to overestimates of the active enzyme available for digestion of polypeptides.
α2-macroglobulin protein levels (determined by ELISA) and cystatin activity (measured as papain-inhibitory activity) were both detected in GCF from chronic periodontitis patients (Chen et al., 1998). The cystatin activity could have been derived from serum, cellular secretions, or lysed cells. The total inhibitor levels were found to fall in parallel with those of cathepsin B activity after treatment. However, the correlations between inhibitor concentrations and clinical parameters were weak. In contrast, when cross-immunoelectrophoresis was used, an inverse relationship between total α2-macroglobulin levels and pocket depth and bone loss was observed (Skaleric et al., 1986). α2-macroglobulin in GCF would almost certainly be derived from plasma. Cystatin C levels measured by ELISA were found to be negatively correlated with probing depth (Skaleric et al., 1989). Cystatin C, and the salivary cystatins S and SN, could not be detected by immunoblotting in crevicular fluid samples from periodontitis patients (Blankenvoorde et al., 1997). However, cystatin A, normally considered to be an intracellular protein, could be detected in GCF and saliva, presumably released from epithelial cells and neutrophils. The concentration of the major plasma inhibitor of elastase, α1-PI, in GCF from healthy sites is comparable with that of plasma (Lee et al., 1997). The total level was found to rise significantly in GCF from gingivitis and periodontal disease sites, most likely as a result of local synthesis by macrophages. No degradation fragments were detected in GCF from healthy site. However, consistent with the notion of a proteolytic cascade outlined above, in samples from periodontally involved sites, a fraction of the α1-PI was found to be degraded. This degradation was most likely mediated by MMP-8 from PMNs. MMP-8 can itself be activated by cathepsin B.
(X) CPs in the Oral Cavity
PMNs entering the oral cavity remain functional for some time prior to being cleared by swallowing. These cells can release their lysosomal contents into the oral cavity when stimulated, or upon lysis. Our knowledge of other sources of CPs in the oral cavity is extremely limited. Human epidermal keratinocytes have been reported to secrete cathepsin B (Katz and Taichman, 1999), but oral keratinocytes have not been examined. When immunolabeling was used, cathepsins B and H, but not L, were identified in the granules of normal rat junctional epithelial cells; however, they were found only in the occasional cell in oral sulcular and oral epithelia (Yamaza et al., 1997). Junctional epithelial cells were shown to take up horseradish peroxidase by fluid-phase endocytosis, suggesting that they can take up and degrade foreign material from the gingival sulcus, and that this process is mediated in part by endosomal CPs. It is not known if these cells can release CPs.
We understand little about the consequences, if any, of the release of CPs (or other peptidases) into the oral cavity. An obvious likely result is proteolytic cleavage of salivary proteins. For example, released elastase has the potential to cleave acidic proline-rich proteins (PRPs), reducing their binding to hydroxyapatite (Boackle et al., 1999). The proteolytic processing of basic proline-rich proteins in parotid saliva has been shown to differ between individuals who have experienced dental caries and those who have remained caries-free (Ayad et al., 2000), and presumably continued proteolytic cleavage in the oral cavity could affect the protective activity of these proteins. It is possible that any effects of CPs released in the oral cavity would be greatly minimized by the presence of salivary cystatins. Is this why we have these proteins at such high levels in saliva?
(XI) CPs and Tooth Development and Movement
The role of apoptosis in tooth development and movement was discussed above. Cathepsin B has been identified in ameloblasts by immunocytochemistry, and may be targeted to the ruffled border, suggesting a role in enamel maturation (Al Kawas et al., 1996). Similarly, it has been localized to the lysosomes and vacuoles of odontoclasts in human deciduous teeth (Sasaki and Ueno-Matsuda, 1992). Cathepsin K has not been examined in tooth development, eruption, or shedding of deciduous teeth. Given the importance of cathepsin K in bone remodeling, it would be anticipated that it would be important in tooth movement. Analysis of cathepsin K levels during experimental tooth movement in the rat indicated that the numbers of cathepsin-K-positive osteoclasts do indeed change in response to pressure (Ohba et al., 2000). Initially, positive osteoclast numbers were found to increase significantly on the pressure side and then, after 3-4 days, on both the tension and pressure sides. The numbers declined to substantially lower levels after 7-12 days, and few positive osteoclasts were observed in untreated sites.
(XII) CPs and Cancer
Metastasis of tumor cells requires proteolysis of the extracellular matrix for penetration. Several studies have found a significant correlation between the production of peptidases (including CPs) by tumor cells and invasiveness and prognosis (Kolkhorst et al., 1998; reviewed in Kos and Lah, 1998; Koblinski et al., 2000; Lah et al., 2000a,b). Increased cathepsin B levels have been observed in a broad range of tumors, and the role of cathepsin B in cancer and its prognostic value have been the subject of recent extensive reviews (Lah and Kos, 1998; Yan et al., 1998; Koblinski et al., 2000). Key aspects will be summarized here. Oncogenic transformation results in increased levels of expression of cathepsin B (and L), and a relocalization to the plasma membrane adjacent to the underlying basement membrane. Cathepsin B also associates with the external cell surface, and increased levels of cathepsin B can be observed at the invasive edges of different human tumors (e.g., colorectal cancer; Hirai et al., 1999). Cathepsin B concentration in breast tumor cytosol is a significant indicator of prognosis for recurrence and survival, while cathepsin L levels are not (Lah et al., 2000a). In contrast, immunohistochemical localization of cathepsin B in tumor cells was of only borderline significance as a prognostic indicator of survival (Lah et al., 2000b). This discrepancy may be due to the various other non-tumor types of cells (e.g., sprouting endothelial cells) represented in homogenized samples that also express cathepsin B at increased levels. The data are conflicting regarding the prognostic value of cathepsin L levels in cancer (see Lah et al., 2000b, and references therein).
Experimental manipulation of cathepsin CP levels supports a role in cancer. The murine squamous carcinoma cell line SCC-VII secretes procathepsin B, and the invasiveness of these cells in an in vitro assay could be significantly inhibited by an E-64 derivative relatively specific for cathepsin B, or by transfection with human cystatin C (Coulibaly et al., 1999). Further, cells stably transfected with the human procathepsin B gene secreted increased amounts of enzyme and showed an increased invasiveness that could be reversed by the inhibitor. As noted above, tumors can establish mildly acidic conditions, and many tumor cells also secrete significant amounts of procathepsin L. Under these pH conditions, procathepsin L itself is active in the presence of GAGs, and can also be converted to the mature form. Overexpression of procathepsin L in melanoma cells increased their tumorigenicity and switched their phenotype from non-metastatic to highly metastatic (Frade et al., 1998). Procathepsin L will cleave human C3, and transfected clones showed up to 60% resistance to complement-mediated lysis. Conversely, treatment of a procathepsin-L-expressing melanoma cell line with anti-CP Fab fragments inhibited C3 cleavage, and increased the complement susceptibility of the cells by 60% (Jean et al., 1995). Suppression of cathepsin L expression by antisense expression resulted in decreased tumorigenicity of two myeloma cell lines (Kirschke et al., 2000). Thus, in addition to matrix degradation, another function of secreted pro-cathepsin L in tumor cells might be evasion of immune surveillance. However, cathepsin L may also contribute to reducing tumor growth. It will cleave collagen XVIII, a component of vascular and epithelial basement membranes, releasing a peptide fragment called endostatin (Felbor et al., 2000). Endostatin inhibits angiogenesis, which would slow tumor growth. The chicken ortholog of cathepsin K is significantly up-regulated by v-Jun, but not other oncogenes, in chicken embryo fibroblasts, suggesting a specific regulation by this oncogene (Hadman et al., 1996). Cathepsin K is significantly up-regulated in osteoclastomas (see above).
It might be anticipated that alterations in the balance between CPs and cystatins, such as cystatin C or stefins, could alter the susceptibility to cancer and tumorigenicity. Increases in CP activity and decreases in inhibitor levels would be predicted to be deleterious. However, the collective data are complex (reviewed in Kos and Lah, 1998). Depending upon the tumor and the individual, CP inhibitor levels may be increased, decreased, or unchanged. In some cases, lower levels of inhibitor, and in others higher levels, correlate with a poorer prognosis. These contradictions may in part reflect the fact that the inhibitors can be present in tissues as free, complexed, and processed forms of lower activity. In turn, different results may reflect the use of different techniques of sample preparation and assay (e.g., immunohistochemistry versus RT-PCR). In addition, their synthesis by tumor cells themselves may be up-regulated. It seems plausible that it is the net imbalance in the components of proteolytic cascades (enzymes and inhibitors) that affects tissue invasion, and that multiple combinations can be effective. Significantly, transgenic mice that are homozygous null for cystatin C are otherwise normal, and show a decreased metastatic spread of melanoma cells (Huh et al., 1999).
Analysis of the collective data indicates that regulation of CP expression and trafficking is perturbed in cancer cells, and that CPs play a complex, but not necessarily obligatory, role in proteolytic cascades involved in the interaction with neighboring cells, tumor invasion, and metastasis. As described above, an individual peptidase might be a primary effector directly involved in tissue breakdown, or it might be a catalytic participant in a proteolytic cascade. Tumors can contain elevated extracellular levels of active CPs that can activate u-PA efficiently, leading in turn to activation of plasmin and MMPs. Tumor uPA activity is strongly correlated with prognosis in a variety of cancers, such as breast cancer (e.g., Foekens et al., 2000, and references therein).
Very little is known regarding oral cancer, CPs, and their inhibitors. A human oral squamous cancer cell line (BHY) derived from a non-metastatic cancer was reported to express procathepsin L and pro-MMP7 (Kawamata et al., 1997). This cell line was able to invade mandibular bone in nude mice, but did not metastasize. Another cell line (HN) derived from a metastatic squamous cell cancer did not secrete procathepsin L, and did not invade mandibular bone, although it was metastatic. These results suggest that cathepsin L might be important to bone invasion by oral squamous cancer cells.
(XIII) CPs and Arthritis
Rheumatoid arthritis (RA) is characterized by chronic autoimmune inflammation of the synovium of the joints, leading to progressive damage to the cartilage and bone, and loss of function. In RA, the synovial membrane proliferates to form a pannus that destroys adjacent bone and cartilage. Fibroblast-like synoviocytes and activated macrophages that accumulate at the leading edge of the pannus are considered to be major effector cells of joint damage (Bresnihan, 1999). CPs and MMPs are thought to be important in this process. Cathepsin B and, to a lesser extent, cathepsin L (both most likely derived from macrophages and fibroblasts) were found in the synovial fluid and membrane in patients with rheumatoid arthritis, even at very early stages, while levels were very low in normal synovium (Cunnane et al., 1999; Hansen et al., 2000; Ikeda et al., 2000). There is evidence of a post-transcriptional up-regulation of these enzymes in RA (Keyszer et al., 1998). Cathepsin K mRNA is also found in synovial fibroblasts, and is up-regulated in RA (Hummel et al., 1998). Up to 90% of cells in RA lymphocytic infiltrates, but only 10-30% of the cells in a normal synovium, were found to express cathepsin K mRNA. Cathepsin K is a highly likely candidate for an enzyme mediating destruction of bone and cartilage (see above), while cathepsins B and L could be involved in a proteolytic cascade, such as pro-uPA activation.
Osteoarthritis (OA) is characterized by degeneration of the cartilage of joints. It can probably affect any vertebrate. High levels of cathepsin K are found in the giant cells of the synovium (Dodds et al., 1999). Cultured synoviocytes have been shown to secrete pro-cathepsin L (Maciewicz et al., 1990b). Although osteoarthritis can affect the temporomandibular joint (TMJ), there has been little investigation of CPs in this tissue in health or disease. Significant levels of cathepsins B and L were found immunocytochemically in macrophage-like cells of the synovial lining of the normal rat temporomandibular joint (Kiyoshima et al., 1993, 1994).
(XIV) Summary and Conclusions
CPs are potent proteolytic enzymes that are widely distributed in mammalian tissues. Many of the major enzymes are members of the CA (papain-related) and CD (legumain-related) clans. In the oral cavity and surrounding structures, potential endogenous sources of CPs include tissue and immune cells. However, expression of the majority of CPs has not been examined in oral tissues. With respect to inhibition by salivary cystatins, our knowledge of CP inhibition profiles is limited. Based on inhibition by cystatin C (the closest relative of the salivary cystatins), one or more Branch B enzymes would be predicted to be the most likely targets (although DPP I cannot be excluded).
Aside from a housekeeping lysosome function in the metabolic turnover of proteins, CPs participate in and affect multiple host systems in both health and disease. These can be divided into three broad, interrelated categories: tissue remodeling and turnover of the extracellular matrix; immune system function; and modulation and alteration of cell function. CPs can participate in these systems intracellularly and extracellularly. Intracellularly, they function in processes as diverse as normal protein turnover, antigen processing, and apoptosis. Extracellularly, CPs can mediate turnover of the extracellular matrix by direct proteolysis, or by amplification of a proteolytic cascade (e.g., activation of u-PA). Very little is known regarding trafficking and release of CPs from oral tissues. In normal tissues, CPs can contribute to tissue remodeling, including bone (e.g., cathepsin K). CPs are intimately involved in immune function. They are released in phagocytosis, and can cleave cytokines and complement to produce chemoattractants. There is mounting evidence that CPs have a role in modulating the immune system, and the switch between Th1 and Th2 responses. This can determine whether the response is protective (in terms of clearing the pathogen) or deleterious (in terms of damage to host tissues by the immune system). Significantly, cystatin C, a CP inhibitor previously thought to act extracellularly, has been shown to regulate the intracellular activity of cathepsin S during antigen presentation in bone-marrow-derived dendritic cells. Salivary cystatins are closely related to cystatin C, and the salivary glands are potential antigen-presenting cells. Could salivary cystatins actually function intracellularly in the control of antigen presentation? For example, they might function to suppress generation of novel epitopes from salivary proteins—and a potential autoimmune response—due to co-trafficking with CPs. CPs (and proCPs) also appear to have some signaling functions distinct from direct proteolytic cleavage, and can modulate cell behavior. CPs can modulate cell function by cleavage of surface receptors, and there are indications that they can affect activities such as DNA synthesis. However, our understanding regarding these “non-classic” properties of CPs is in its infancy. Collectively, CP function is much more than simple proteolytic breakdown, and CPs participate in complex regulated networks of protein-protein interaction consisting of other peptidases and inhibitors, as well as substrates.
In disease, CP activities can be viewed as a perturbation of these normal networks. Host CPs can participate in abnormal remodeling, tissue damage in inflammatory responses, and facilitate tumor metastasis. Any relationship among cathepsins, cystatins, and oral cancer is unknown. Any role CPs might play in tooth eruption or movement or in salivary gland remodeling in normal development or disease has received little attention. Only cathepsins B, H, and L have been studied in any detail with respect to periodontal disease, but their roles in mediating tissue damage in the pocket remain to be defined, as does any effect within the mouth following their exit from the pocket. The interaction of these cathepsins with salivary cystatins, and its significance, is somewhat ambiguous and needs clarification. The contributions of other cathepsins, such as cathepsin S, to periodontal disease are only just being explored. Significantly, defects in DPP I lead to severe early-onset periodontal disease, although the mechanism has not been established. It will be interesting to examine whether mutations in other CP genes, such as lymphopain, are associated with cases of early-onset periodontal disease not linked to CTSC. The consequences of oral exposure to CPs have not been examined in any detail, although degradation of salivary proteins is an obvious likelihood. Clearly, we still have much to learn about the biology of CPs and their inhibitors in relation to the oral cavity.
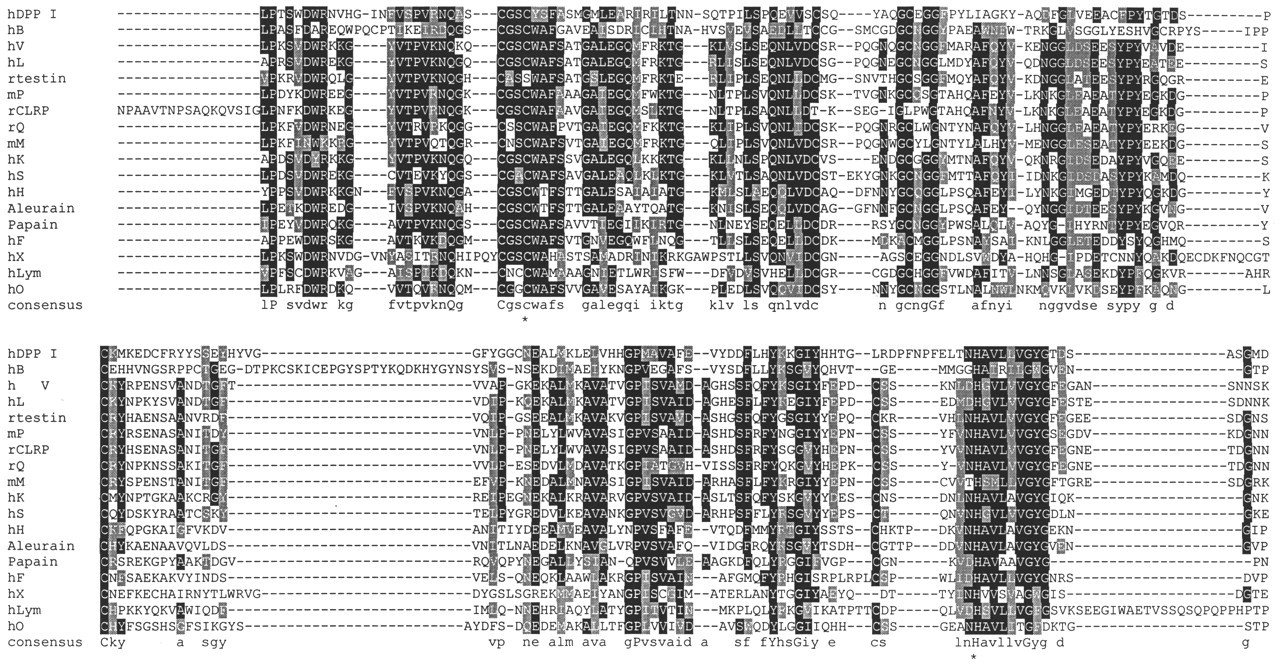
Protein sequence alignments of mature regions of known human cathepsins, other rodent cathepsins and related proteins, and plant papain and aleurain. Although only human and selected sequences are shown, the original dataset used for generating the alignments contained 43 vertebrate and two plant sequences, identified by a BLAST search of the GenBank non-redundant database (Altschul et al., 1990). The sequence abbreviations and GenBank entries for the sequences shown are: hDPP I, human dipeptidyl peptidase I, P53634; hB, human cathepsin B, NP_001899; hV, human cathepsin V, AAC23593; hL, human cathepsin L, NP_001903; rtestin, rat testin, P15242; mP, mouse cathepsin P, NP_036137; rCLRP, rat cathepsin L-related protein, I58002; rQ, rat cathepsin Q, AAF01247; mM, mouse cathepsin M, AAF68224; hK, human cathepsin K, P43235; hS, human cathepsin S, P25774; hH, human cathepsin H, P09668; Aleurain, plant, P05167; Papain, plant, P00784; hF, human cathepsin F, NP_003784; hX, human cathepsin X, NP_001327; hLym, human lymphopain, P56202; and hO, human cathepsin O, NP_00135. Propeptide cleavage sites were obtained from the GenBank entries, or from preliminary alignments. Alignments of the predicted mature proteins were generated based on the default settings of ClustalX (Thompson et al., 1997). Introduced gaps are shown as “-”. The alignment is in good agreement with that of other published ones, including those based on structure. To derive consensus sequences, we processed alignment files using the public domain software BOXSHADE (written by K. Hofmann and M. Baron, www.ch.embnet.org/software/BOX_form.html). Predominant identical residues (> 50%) at a position are shown with a black background, predominant similar residues on a grey background. The majority consensus residue is shown under each alignment. An uppercase letter shows a residue conserved in all aligned sequences. An * under the consensus sequence denotes the active-site cysteine and histidine residues conserved in all functional peptidases.
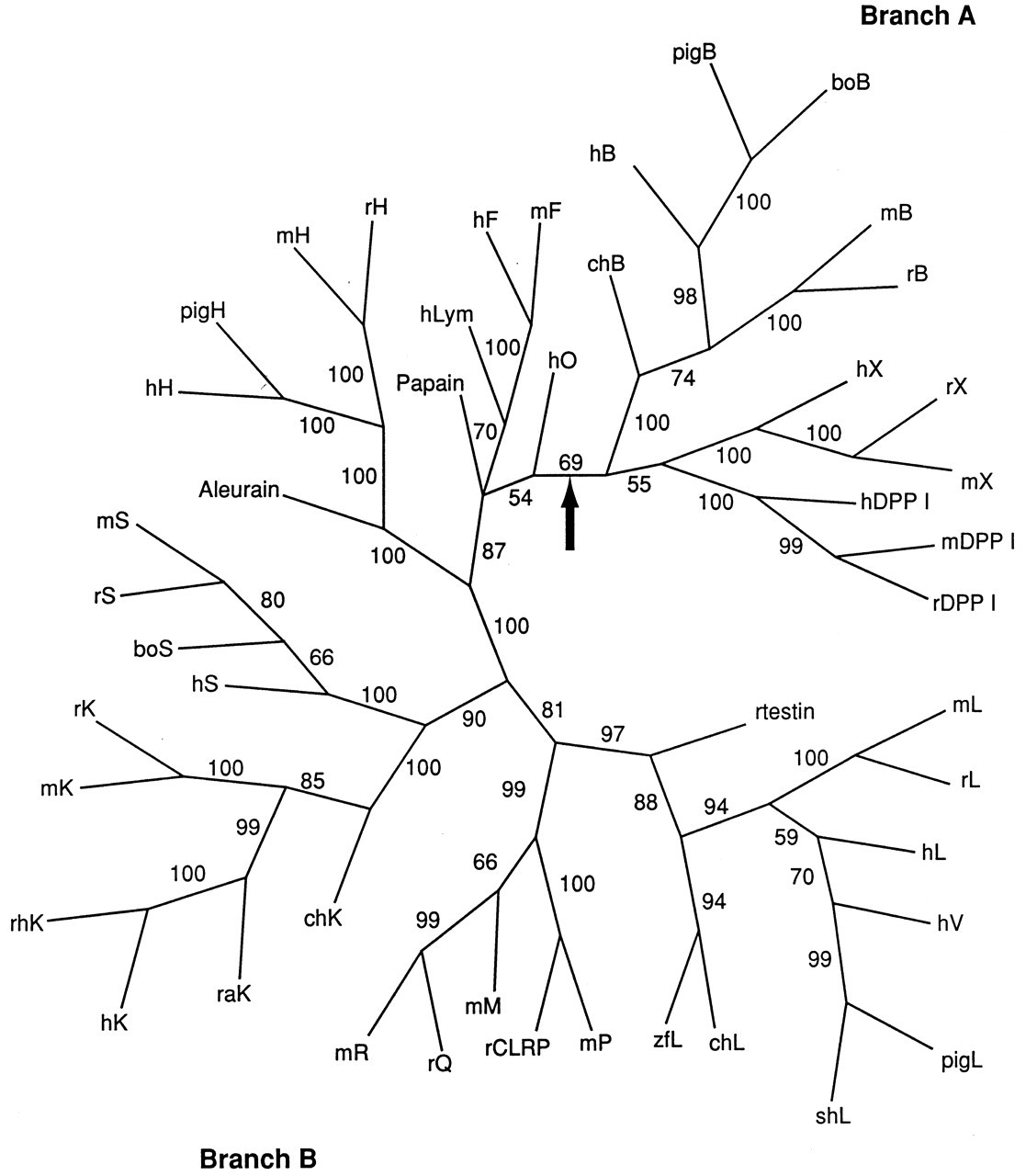
Phylogenetic tree of vertebrate and select plant CPs. The full alignment of CPs described in the legend to Fig. 1 was used, except that bovine cathepsin L was excluded, due to the incomplete C-terminal sequence. The PAUP 4b4a software package (Swofford, 2000) was used to search for trees by means of the distance optimality criterion and default parameters, and starting trees obtained by neighbor-joining. The tree shown is an unrooted 50% majority rule consensus tree obtained by the bootstrap method (500 replicates) with heuristic search. Values shown adjacent to branches are percentage support for a branch. The large arrow indicates the position of the presumptive root of the tree: CPs on the group of branches to the right of this point comprise Branch A, those to the left Branch B (see text for explanation of Branches A and B). The abbreviations used for species are: bo, bovine; ch, chicken; h, human; m, mouse; pig, pig; r, rat; ra, rabbit; rh, rhesus; sh, sheep; and zf, zebrafish.

Proregion alignments of known human cathepsins and selected rodent and plant proteins. Proregion alignments (excluding signal peptides and the N-terminal extension of cathepsin F) are shown in
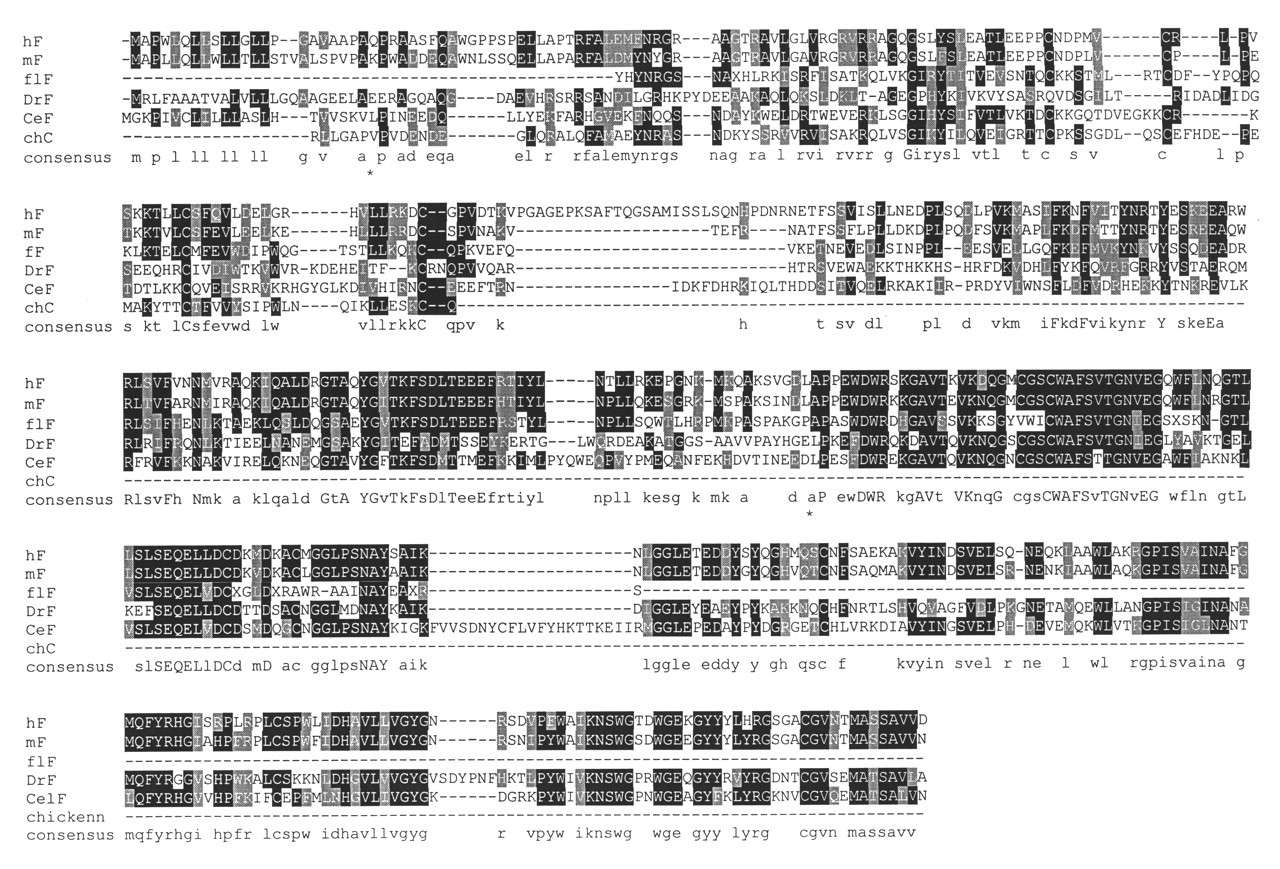
Cathepsin F-like and chicken cystatin C protein alignment. Consensus residues are indicated as described in Fig. 1. Residues identical in all six sequences are denoted by an uppercase letter in the consensus sequence. Presumptive N-terminal residues produced by cleavage of the leader peptide and the proregion are denoted by an * under the consensus sequence. The GenBank entries for the sequences shown are: hF, human cathepsin F, NP_003784; mF, mouse cathepsin F, AAF13147; flF, Japanese flounder Paralichthys olivaceus, AU050404 (partial peptide sequence derived from +1 frame of mRNA sequence); DrF, Drosophila melanogaster CG12163 gene product, AAF52055; CeF, Caenorhabditis elegans cathepsin F-like hypothetical protein F41E6.6, AAB65956; and chC, chicken egg white cystatin, P01038. The alignment was produced by separately aligning chicken cystatin to flounder cathepsin F, and human, mouse, Drosophila, and C. elegans cathepsin F proteins to each other by means of ClustalX. These two profiles were then aligned, and this alignment was manually adjusted by means of an alignment of all five CPs. The alignment of chicken cystatin with the cathepsin proregions agrees quite well with an alignment based on threading reported by Nagler et al. (1999a).
Footnotes
Acknowledgements
The author thanks Dr. R. Menard for providing information regarding the cathepsin-F-cystatin similarity search, Drs. Baldev Singh, Jill Lewis, and Keith Volkmann for critical reading of the manuscript, anonymous reviewers for valuable criticism, and Sandy Abbott and Karen Willis for assistance with manuscript preparation. This work was supported by the Medical College of Georgia School of Dentistry.
