Abstract
The inward-rectifier potassium channel (Kir) 2.x family is an important family of ion channels in the context of human health. These potassium channels are involved in processes such as cardiac action potential, formation of skeletal muscle, bone development, vasodilation, and neuronal activity and are expressed centrally and peripherally. Given their importance, they are an attractive target for the development of tool compounds. The high homology between the members of the Kir family has made isoform selectivity challenging. In an effort to discover novel chemical matter related to this intriguing target, we performed a high-throughput screen utilizing compounds from the Vanderbilt Institute of Chemical Biology Discovery Collection. This screen of over 20,000 compounds resulted in 48 verified hits consisting of six novel chemical scaffolds. Of these hits, VU0523203 and VU0606851 were selected as promising starting points for initial medicinal chemistry optimization to improve potency and distribution, metabolism, and pharmacokinetic (DMPK) properties. These efforts resulted in the discovery of VU6073995, a compound with modest potency at Kir2.1 and improved DMPK properties compared with ML133.
Keywords
INTRODUCTION
The inward-rectifier potassium channels (Kir) are key players in the regulation of resting membrane potential, potassium secretion, and cell excitability. Kir channels are tetrameric complexes of subunits that assemble around a central water-filled pore that enables potassium ions to move down their electrochemical gradient and out of the cells. Each subunit contains two transmembrane domains, an extracellular, re-entrant pore-forming loop, and cytoplasmic domain comprising the amino- and carboxyl-termini. 1 The Kir channel family is made of 16 genes whose gene products can be subdivided into classical Kir channels (Kir2.x), G protein-gated channels (Kir3.x), ATP sensitive channels (Kir6.x/SURx), and K+ transport channels (Kir1.1, Kir4.x, Kir5.x, Kir7.x).2,3
The Kir2.x family of channels are expressed both peripherally and centrally in the brain, retina, heart, skeletal muscle, and endothelium/smooth muscle. 4 They are involved in processes such as cardiac action potential, formation of skeletal muscle, bone development, vasodilation, and neuronal activity.5–9 In particular, Kir2.1 has historically been acknowledged for its role in Andersen-Tawil syndrome, periodic paralysis, and short-QT syndrome.10–12 Recently, it has also been reported to potentially modulate neuropathic pain. 13 Their wide-reaching influence on body processes make the Kir2.x family of channels attractive targets for the development of tool compounds and eventual drugs. Ideal tool compounds would also possess favorable distribution, metabolism, and pharmacokinetic (DMPK) properties enabling effective in vivo studies.
Tool compounds to investigate Kir2.1 are highly desirable given its role in human disease. 14 Previous efforts have included the repurposing of existing drug molecules (tamoxifen and chloroquine), as well as high-throughput screening (HTS) approaches.1,15–17 While these compounds have achieved potencies as low as sub-micromolar, they lack selectivity against other members of the Kir2.x family and other Kir channels. According to previous reports, ML133 (N-(4-methoxybenzyl)−1-(naphthalen-1-yl)methanamine) possesses a Kir2.1 IC50 in manual patch clamp (MPC) ranging from 1.51 to 1.8 µM at pH 7.4 and 290 nM at pH 8.5.1,18 While it exhibits minimal selectivity within the Kir2.x family, it possesses selectivity against Kir1.1, Kir4.1, and Kir7.1. 1 These limitations provide a new opportunity for HTS efforts to uncover novel chemical matter for Kir2.1 inhibition. Toward this goal, we performed a high-throughput screen utilizing compounds from the Vanderbilt Institute of Chemical Biology (VICB) Discovery Collection.
MATERIALS AND METHODS
Thallium (Tl+) Flux Assay
The Tl+ flux assays were performed with modifications tailored for the Kir2.1 cell line, following the methodology described previously. 19 Briefly, stable T-REx-HEK-293 cells expressing human Kir2.1 were cultured in Dulbecco's Modified Eagle Medium (DMEM) growth medium (Cat#11965-092, Gibco, Billings, MT), supplemented with 10% heat-inactivated fetal bovine serum (FBS) (Catalog #16140, Gibco), 100 U/mL of penicillin-streptomycin, 5 μg/mL blasticidin S, and 250 μg/mL hygromycin. The day before the experiment, cells that reached approximately 90% confluency were resuspended in plating media, which contained the same components as growth media, with dialyzed FBS (Catalog #26400, Gibco) replacing the heat-inactivated FBS. Tetracycline was added to the plating media to achieve a final concentration of 1 μg/mL, promoting hKir2.1 expression. Following this, cells were counted and plated at a density of 15,000 cells per well in black-walled, amine- or poly-D-lysine (PDL)-coated 384-well plates, with 20 μL of the cell suspension per well (Corning, Corning, NY). On the experimental day, the media was removed from the cell plate and replaced with 20 μL per well of Tl+-sensitive dye loading solution, containing 2.5 μg/mL Thallos-AM (Cat# 11000-050, ION Biosciences, San Marcos, TX), in assay buffer. This was accomplished using a Multidrop Combi dispenser (Thermo Fisher Scientific, Waltham, MA). The plates containing the dye solution were incubated at room temperature for 1 h. After incubation, the cell plate was washed with Hanks’ balanced salt solution (HBSS) assay buffer containing 20 mM N-2-hydroxyethylpiperazine-N-2-ethane sulfonic acid (HEPES) using a BioTek ELx 405 TS plate washer (Agilent, Santa Clara, CA), leaving 20 μL of assay buffer in each well. The cell plate was then subjected to imaging on a Panoptic kinetic imaging plate reader (WaveFront Biosciences, Franklin, TN) at a frequency of 1 Hz, with excitation at 482/35 nm and emission at 536/40 nm for 10 s. Following the initial imaging, 20 μL per well of HBSS assay buffer containing test compounds at twice their final concentrations was added, and imaging continued for an additional 8 min. Finally, 10 μL per well of chloride-free Tl+ stimulus solution was introduced (Cat# 11000-050, ION Biosciences), resulting in a final concentration of 0.25 mM Tl2SO4 in the cell plate post-stimulation. Images were collected for an additional 2 min. See Table 1 for additional details.
Protocols for High-Throughput Thallium (Tl+) Flux Assay
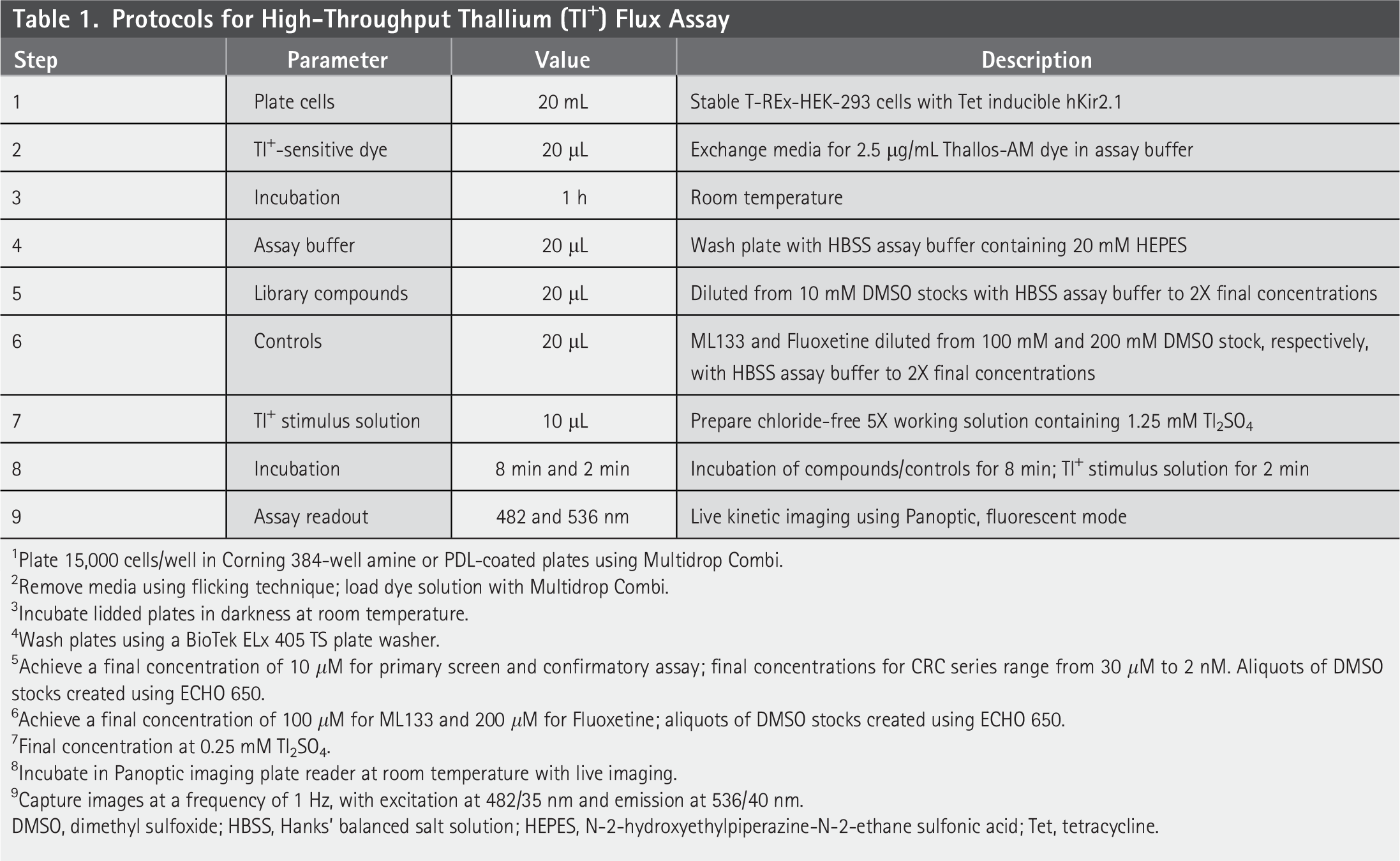
1Plate 15,000 cells/well in Corning 384-well amine or PDL-coated plates using Multidrop Combi.
2Remove media using flicking technique; load dye solution with Multidrop Combi.
3Incubate lidded plates in darkness at room temperature.
4Wash plates using a BioTek ELx 405 TS plate washer.
5Achieve a final concentration of 10 μM for primary screen and confirmatory assay; final concentrations for CRC series range from 30 μM to 2 nM. Aliquots of DMSO stocks created using ECHO 650.
6Achieve a final concentration of 100 μM for ML133 and 200 μM for Fluoxetine; aliquots of DMSO stocks created using ECHO 650.
7Final concentration at 0.25 mM Tl2SO4.
8Incubate in Panoptic imaging plate reader at room temperature with live imaging.
9Capture images at a frequency of 1 Hz, with excitation at 482/35 nm and emission at 536/40 nm.
DMSO, dimethyl sulfoxide; HBSS, Hanks’ balanced salt solution; HEPES, N-2-hydroxyethylpiperazine-N-2-ethane sulfonic acid; Tet, tetracycline.
Primary Screen, Confirmation, and Concentration-Response Curve Evaluation
Test compounds from the VICB Discovery Collection were screened against human Kir2.1 in a high-throughput Tl+ flux assay in singlicate (n = 1) at a fixed concentration of 10 μM. To account for baseline variability, the fluorescent signal for each well was normalized by dividing it by the average fluorescent amplitude from the first 5 s, resulting in a static ratio (F/F0) for each well. This normalization controlled for variations due to Thallos-AM loading and cell density. The initial slopes of the normalized data were calculated for each well between 2 and 8 s following thallium stimulation and were normalized to the response percentage of the maximal inhibitor control (100 μM ML133) for each 384-well plate. A checkerboard assay conducted prior to the primary screening confirmed a significant separation between positive and negative signals, yielding a Z’ factor of 0.79 (see Supplementary Figure S1). During the primary screening, the Z’ factor was determined for each plate by analyzing negative control wells (dimethyl sulfoxide [DMSO], n = 46) and positive control wells (ML133, n = 16) located in columns 1, 2, 23, and 24. Library compounds were tested in columns 3 through 22. In total, 63 plates were used for the primary screen, resulting in an average Z’ factor of 0.75. Compounds were classified as hits if they satisfied both Z-score and Robust Z-score criteria (i.e., meeting thresholds of 3 standard deviations from the mean and 3 mean absolute deviations from the median) and displayed no activity prior to thallium addition (e.g., no fluorescent tags). These hits underwent retesting in a confirmatory Tl+ flux assay at 10 μM in duplicate (n = 2). Selected confirmed hits were further evaluated in triplicate (n = 3) through a 10-point concentration series ranging from 30 μM to 2 nM, with a dilution factor of 1:3. The final DMSO concentration of 0.3% (v/v) will remain constant across all compound concentrations. The initial slopes of Tl+-induced fluorescence changes were calculated and fitted with a four-parameter logistic equation to derive potency and efficacy values. Efficacy measurements were relative to the maximally effective concentration (100 μM) of the control inhibitor compound ML133. All data were analyzed using our in-house software, Waveguide.
Patch Clamp Electrophysiology
Whole-cell patch-clamp electrophysiology was used to record Kir2.1 currents from a stably transfected monoclonal T-REx-HEK-293 cell line with a tetracycline-inducible promoter expressing human Kir2.1, as described previously cultured in a 5% CO2 incubator at 37°C. 20 Kir2.1 expression was induced by overnight treatment with 1 µg/mL tetracycline. Cells were dissociated using 0.25% trypsin-EDTA and seeded onto 12 mm circular coverslips (Electron Microscopy Sciences, PA, USA) for same-day whole-cell patch-clamp recordings. Kir2.1 currents were recorded at room temperature using an AxoPatch 200B amplifier with Clampex 10.7 software (Molecular Devices, USA). The voltage-clamp protocol consisted of a 1-s step to −120 mV from a −80 mV holding potential, followed by a 2-s ramp to +40 mV, repeated every 10 s. Data were sampled at 5 kHz and filtered at 1 kHz. Patch pipettes, pulled from 1.5 mm OD thin-walled capillaries (Warner Instruments, MA, USA) using a Sutter P-1000 puller (Sutter Instrument, CA, USA), had resistances of 2–3 MΩ when filled with intracellular solution containing (mM): 135 KCl, 10 HEPES, 1 Ethyleneglycol- bis(β-aminoethyl)-N,N,Nʹ,Nʹ-tetraacetic Acid (EGTA), 2 MgCl2, 2 Na2ATP (pH 7.35, adjusted with KOH). The extracellular solution contained (mM): 135 NaCl, 5 KCl, 2 CaCl2, 1 MgCl2, 5 glucose, 10 HEPES, 10 sucrose (pH 7.35, adjusted with NaOH). After establishing stable baseline Kir2.1 currents, cells were perfused with compounds at concentrations of 0.03–30 µM until response saturation (within minutes). The osmolarity of the internal and external solutions were 285 mOsm (adjusted with sucrose) and 300 mOsm, respectively. The pH of both solutions was 7.35. Recordings concluded with a 2 mM Ba2+ solution to isolate Ba2+-sensitive currents, with compound responses expressed as fractions of this current. Current amplitudes at the −120 mV step were analyzed using Clampfit 11.2 (Molecular Devices). IC50 values were calculated by fitting the Hill equation to concentration-response curves (CRCs) using variable-slope nonlinear regression in GraphPad Prism 10.4 (GraphPad Software).
Hepatic Microsomal Intrinsic Clearance
Human and rat hepatic microsomes (0.5 mg/mL) and 1 µM test compound were incubated in 100 mM potassium phosphate pH 7.4 buffer with 3 mM MgCl2 at 37° C with constant shaking. After a 5 min preincubation, the reaction was initiated by addition of NADPH (1 mM). At selected time intervals (0, 3, 7, 15, 25, and 45 min), aliquots were taken and subsequently placed into a 96-well plate containing cold acetonitrile with internal standard (50 nM carbamazepine). Plates were then centrifuged at 3,000 RCF (4°C) for 10 min, and the supernatant was transferred to a separate 96-well plate and diluted 1:1 with water for liquid-chromatography mass spectrometry (LC/MS/MS) analysis. The in vitro half-life (t1/2, min), intrinsic clearance (CLint, mL/min/kg), and subsequent predicted hepatic clearance (CLhep, mL/min/kg) were determined using Equations 1–3:

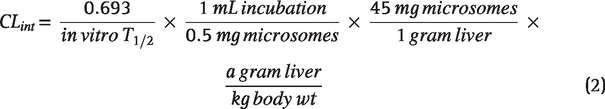

Plasma Protein Binding
The protein binding of each compound was determined in rat or human plasma via equilibrium dialysis employing HTDialysis membrane plates. Plasma was added to the 96-well plate containing test compound and mixed thoroughly for a final concentration of 5 μM. Subsequently, an aliquot of the plasma-compound mixture was transferred to the cis chamber (red) of the RED plate, with a phosphate buffer (25 mM, pH 7.4) in the trans chamber. The RED plate was sealed and incubated for 6 h at 37° C with shaking (120 rpm). At completion, aliquots from each chamber were transferred to a new 96-well plate and were diluted 1:1 with either plasma (trans) or buffer (cis), at which time ice-cold acetonitrile containing internal standard (50 nM carbamazepine) (3 volumes) was added to extract the matrices. The plate was centrifuged (3,000 RCF, 10 min) and supernatants transferred and diluted 1:1 (supernatant: water) into a new 96-well plate, which was then sealed in preparation for LC/MS/MS analysis. Each compound was assayed in triplicate within the same 96-well plate. Fraction unbound was determined using Equation 4.

Equation 4. Determination of fraction unbound.
Compounds
Compounds used for HTS screening were obtained from the VICB Discovery Collection. Compounds for CRC reconfirmation were purchased from Life Chemicals, Inc. except for 4-chloro-N-(3-(3,5-dimethyl-1H-pyrazol-1-yl)−3-oxopropyl)benzenesulfonamide (VU0507577), which was purchased from ChemDiv, Inc., and 2-ethyl-1-((5-ethyl-2-methoxyphenyl)sulfonyl)−4-methyl-1H-imidazole (VU0606851), which was synthesized at the Warren Center for Neuroscience Drug Discovery (WCNDD). All the purchased compounds were assigned Vanderbilt-specific ID numbers for purposes of chemical inventory. The compounds highlighted in the medicinal chemistry section that were not part of the original HTS screen were synthesized at the WCNDD. Compounds were determined to have >95% purity by LCMS analysis at 215 nm, and structures were confirmed by 1H nuclear magnetic resonance (NMR) analysis. Low-resolution mass spectra were obtained on a Waters QDa (Performance) single quadrupole mass spectrometer (SQ MS) with eletrospray ionization (ESI) source. MS parameters were as follows: cone voltage: 15 V, capillary voltage: 0.8 kV, probe temperature: 600° C. Samples were introduced via an Acquity I-Class PLUS ultra-performance liquid chromatography (UPLC) comprised of a binary solvent manager (BSM), fixed-loop sample manager (FL-SM), column heater (CH-A), and photodiode array (PDA). UV absorption was generally observed at 215 nm and 254 nm; 4 nm bandwidth. Column: Phenomenex EVO C18, 1.0 × 50 mm, 1.7 um. Column temperature: 55° C. Flow rate: 0.4 mL/min. Default gradient: 5% to 95% CH3CN (0.05% TFA) in H2O (0.05% TFA) over 1.4 min, hold at 95% CH3CN for 0.1 min.
RESULTS AND DISCUSSION
The HTS consisted of 20,157 compounds from the VICB Discovery Collection, tested at a 10 µM single-point concentration in the thallium (Tl+) flux assay as shown in Figure 1. The labels on the y-axis are adjusted for the primary screen plot. Each point on the plot represents an individual well from each 384 well plate, where normalization calculations are based off the DMSO response (n = 42 per plate), representing 0% or no change with respect to those plate controls, and 100% inhibition represented by the mean response of 14 wells of ML133 per plate. Within these control populations per plate, the individual wells represented by each point on the graph, do respond above and below the mean calculation of 100%. Each well value is a single point from each 384 well screening plate scaled with respect to the mean control response on the plate.
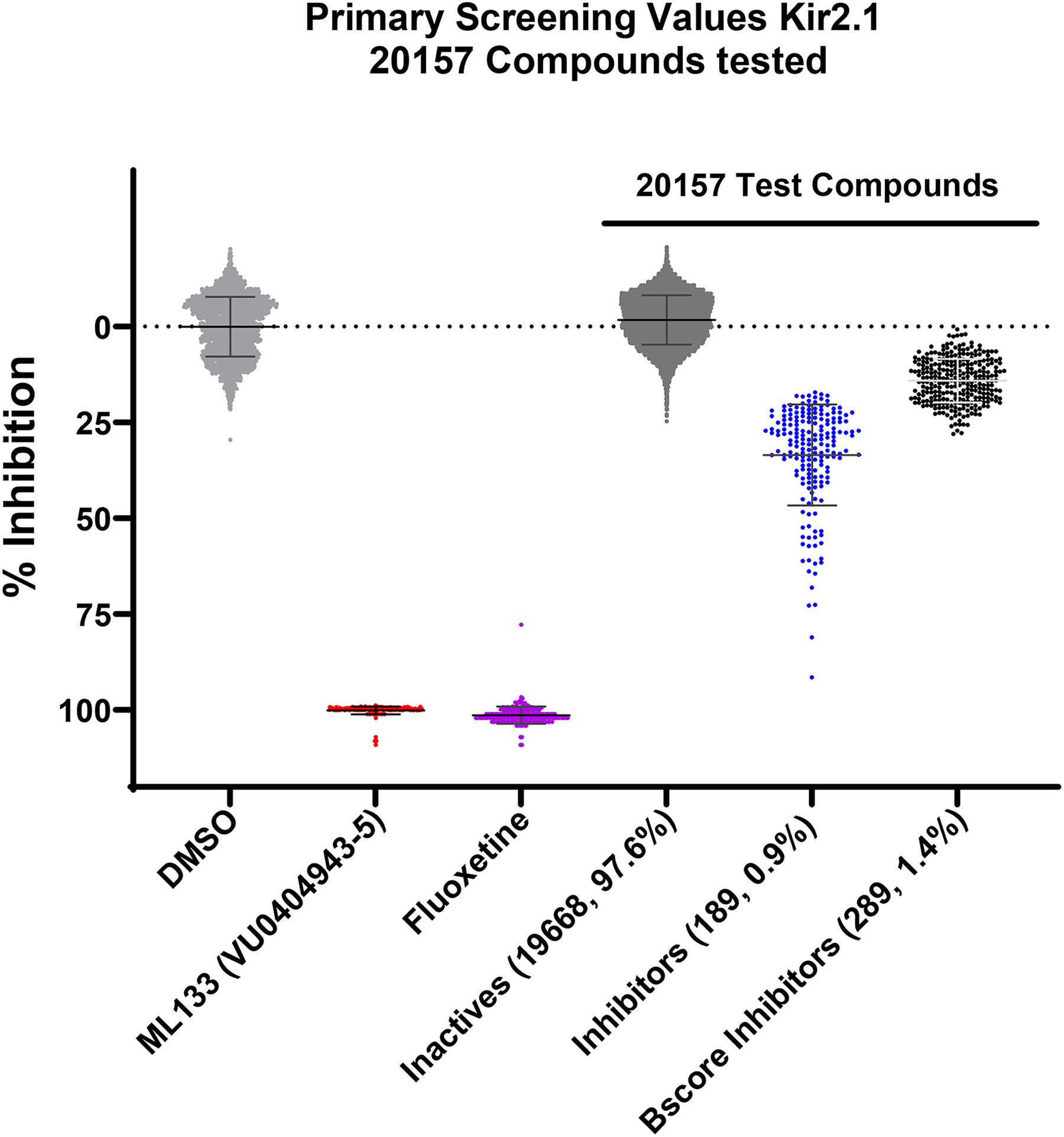
Violin plot of primary Tl+ flux screen.
Compounds that exhibited a Zscore < −3 were selected as Kir2.1 inhibitors with an inhibition range from 18.0% to 91.4%. These 189 compounds also had a Bscore <−5 on all but 4 compounds. A reconfirmation screen at 10 µM, led to 123 compounds retesting positive as Kir2.1 inhibitors. Compounds that exhibited >30% inhibition in the second screen were then selected for a 10-point concentration-response curve (CRC) in the thallium flux assay to determine IC50 values. A total of 50 compounds were tested, affording IC50 values ranging from 6.24 µM to >30 µM. These compounds were reconfirmed in an additional 10-point CRC. A total of 48 compounds were re-verified as hits with 5 compounds displaying IC50 values of < 10 µM, 22 compounds displaying IC50 values between 10–20 µM, and 9 compounds with IC50 values between 20–30 µM. From the 50 compounds that were retested, 12 displayed either an IC50 >30 µM or a poorly shaped curve and were no longer considered hits. Figure 2 provides a summary of this workflow.
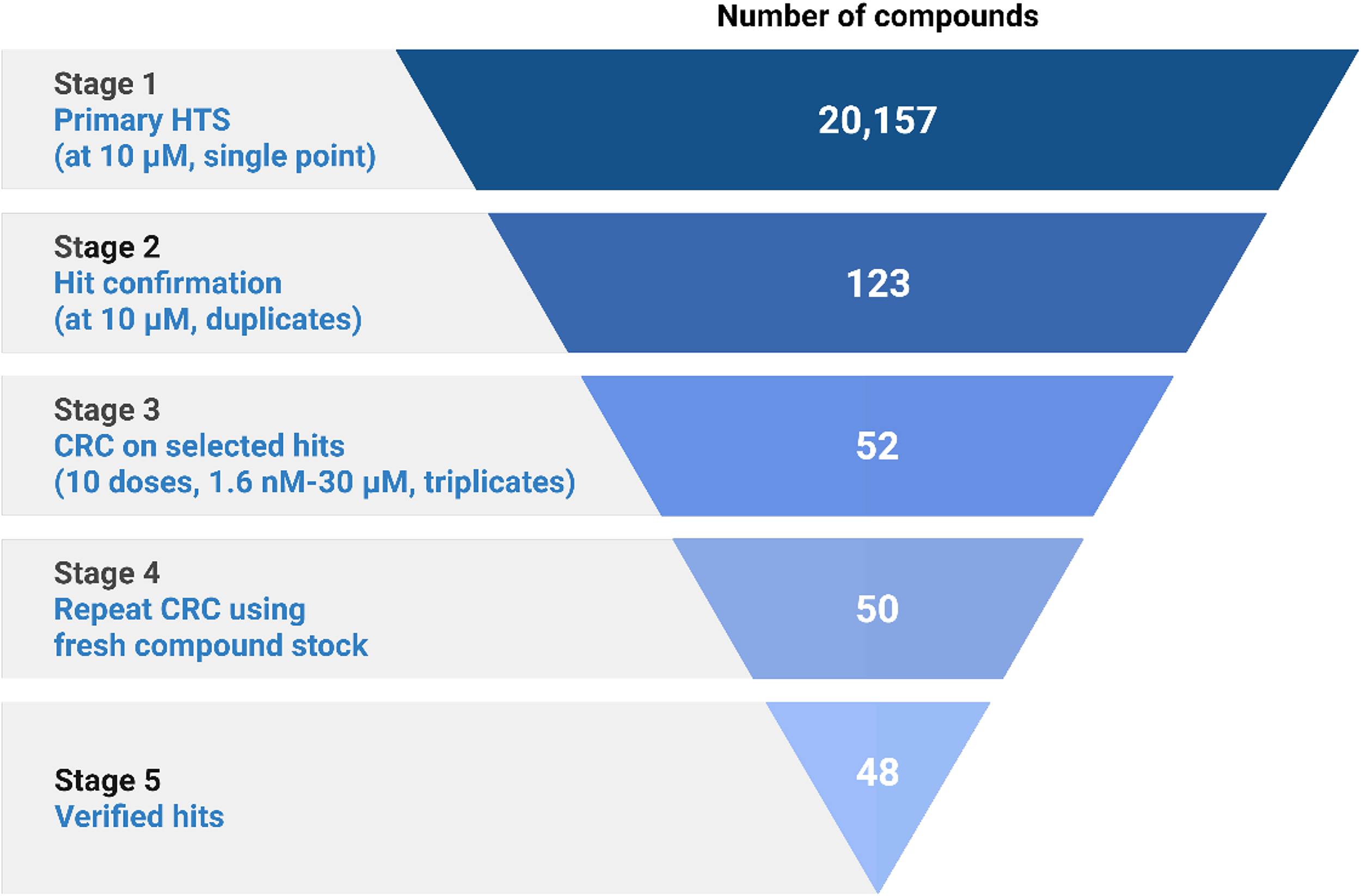
Summary of high-throughput screening (HTS) campaign using thallium (Tl+) flux assay. Figure 2 was generated with Biorender.com.
The 48 verified hits were then categorized into different chemotypes (Figure 3). The first chemotype consisted of aryl sulfonamides containing 14 hits displaying IC50 values varying from 12.3 µM to > 30 µM. The amide-linker aryl class contained 18 hits with IC50 values in the 7.92 µM to > 30 µM range. The terminal aryl class contained 4 hits with IC50 values ranging from 19.9 µM to > 30 µM. The aryl central core class contained 8 compounds displaying IC50 values from 6.24 µM to > 30 µM. The propargyl sulfonamide class contained 2 compounds displaying IC50 values from 9.69 µM to 15.6 µM. Finally, 2 compounds, VU0510678 (IC50 = 18.7 µM) and VU0493926 (IC50 = 20.1 µM), did not fall into any of these chemotypes, and were labeled as “miscellaneous.”
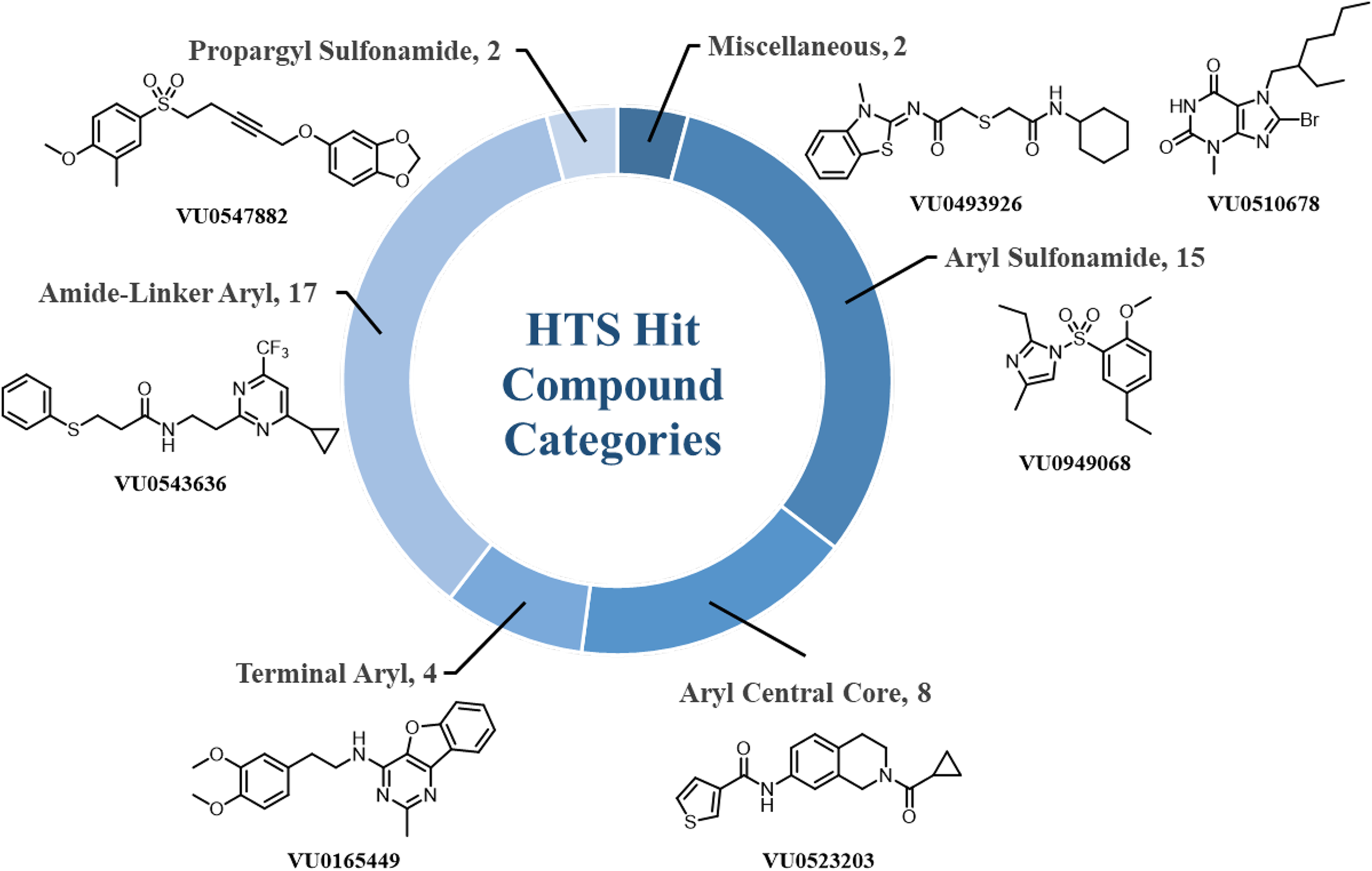
Classification of HTS hits into categories based on chemotypes.
From the previously mentioned classes of compounds, the aryl central core, aryl sulfonamide, and propargyl sulfonamide chemotypes were selected as the starting points for structure-activity relationship (SAR) exploration due to factors such as average potency of hits, preliminary clearance data, and synthetic accessibility for rapid library production. From this effort, compounds VU6073995, VU6074621, and VU6074628 were selected for additional profiling alongside their HTS hit counterparts. In the Tl+ assay, most of the compounds in Table 2 were comparable with ML133 (Tl+ IC50 = 10.1 µM, 101%), while some of the compounds showed enhanced potency such as VU6073995 (Tl+ IC50 = 5.49 µM, 103%) and VU6074621 (Tl+ IC50 = 2.62 µM, 98%). 18 The representative hit expansion data are depicted in Table 2. Following these results, those compounds were then tested in MPC at pH = 7.4. Interestingly, in MPC, VU6074621 exhibited a nearly 1,000-fold loss in potency (Tl+ IC50 = 2.62 µM vs. MPC IC50 = 2370 µM), while the MPC potency of VU6073995 was comparable with Tl+ results (Tl+ IC50 = 5.49 µM vs. MPC IC50 = 5.71 µM). We speculate that the large discrepancy observed for VU6074621 reflects its voltage dependency of block. In thallium assays, the resting membrane potential will be close to the calculated Nernst potential for potassium, which is approximately −80 mV under the assay conditions. In MPC experiments; however, the voltage-gradient acting on Kir2.1 is −120 mV. ML133 and many other Kir channel inhibitors are pore blockers that can be “knocked out” of the pore by inwardly directed potassium ions. 22 We suspect that the potency of VU6074621 from MPC experiments is underestimated due to the voltage-dependent unblocking at −120 mV. In contrast, ML133 shows an improvement of about 10-fold in potency in MPC at pH = 7.4. (Tl+ IC50 = 10.1 µM vs. MPC IC50 = 1.51–1.8 µM).1,18 This difference between values could potentially be due to pH dependent inhibition sometimes observed for ion channel blockers. Following MPC analysis, compounds of interest were assessed for DMPK properties.
IC50 of Selected Compound in Tl+ and MPC Assays. a
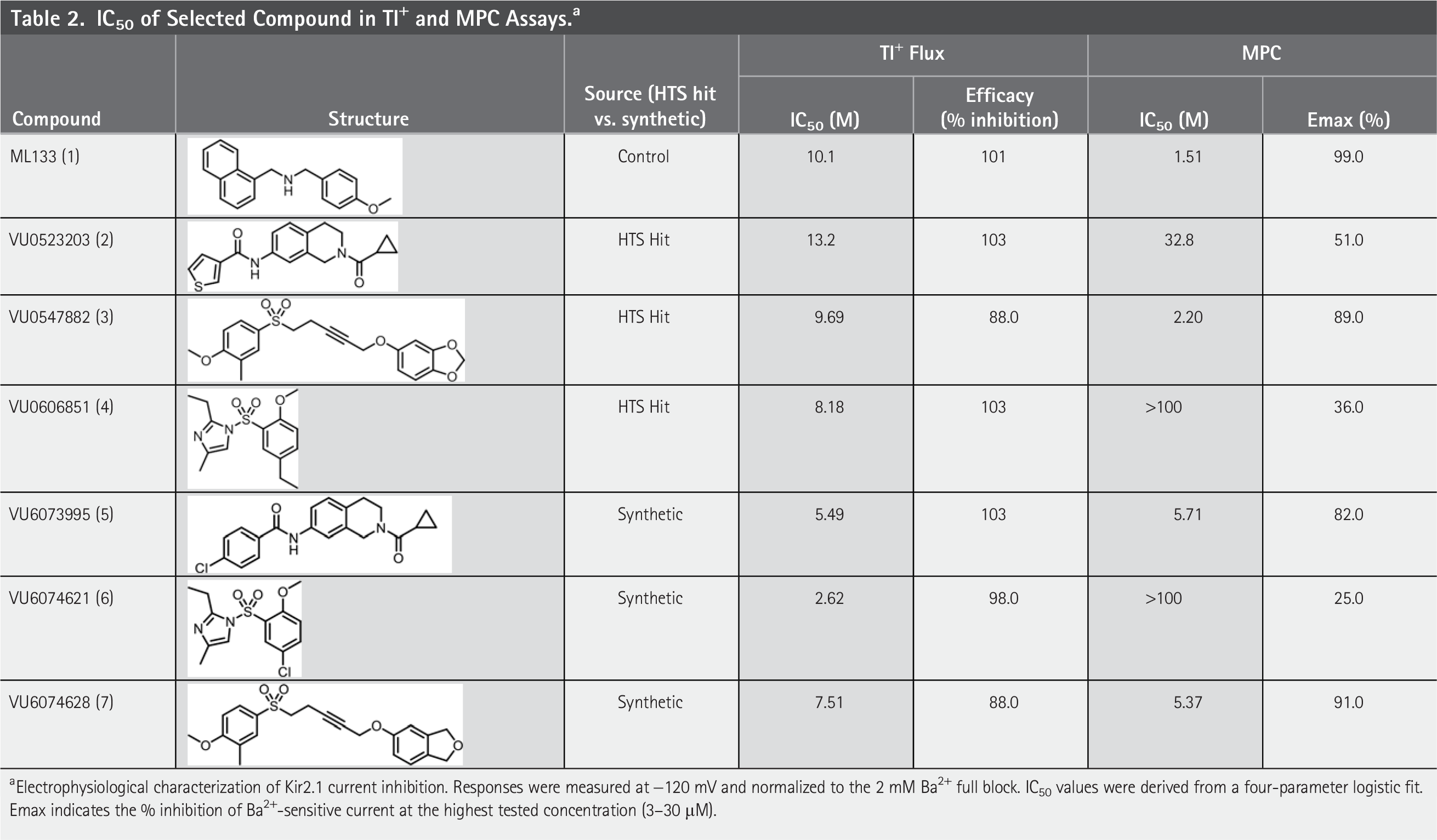
Electrophysiological characterization of Kir2.1 current inhibition. Responses were measured at −120 mV and normalized to the 2 mM Ba2+ full block. IC50 values were derived from a four-parameter logistic fit. Emax indicates the % inhibition of Ba2+-sensitive current at the highest tested concentration (3–30 µM).
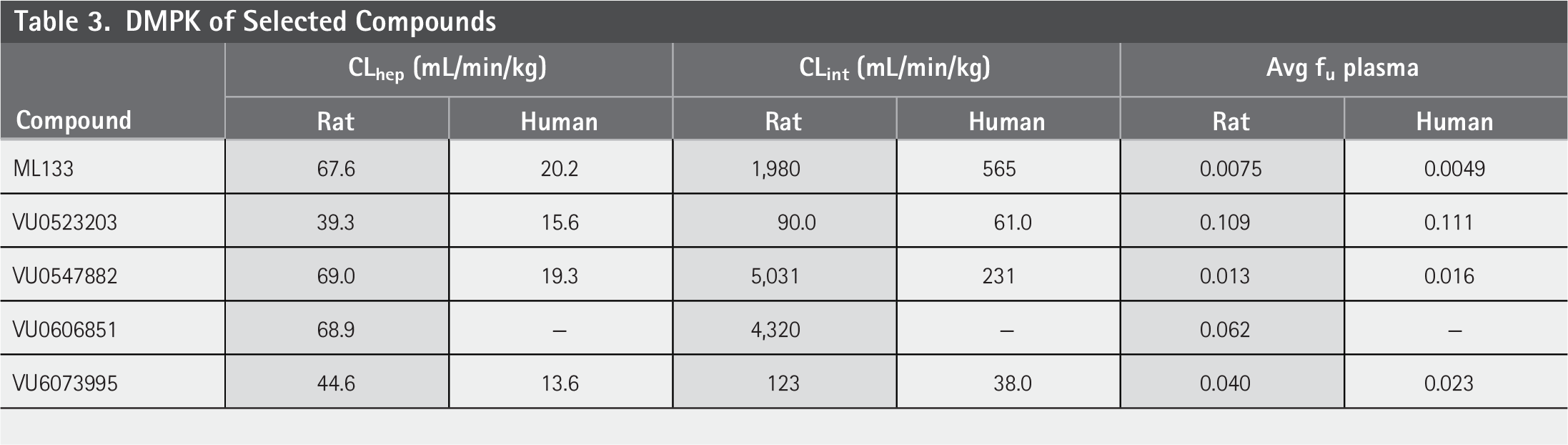
Compound ML133 possesses high in vitro rat and human clearance (predicted hepatic and intrinsic), while also being highly bound (Table 3). VU0523203, presented an improvement for in vitro rat and human clearance (predicted hepatic and intrinsic) and plasma protein binding. Next, VU0547882 and VU0606851 displayed comparable clearance profiles with ML133. VU6073995 presents an overall profile comparable with VU0523203 while being more plasma bound. The improved DMPK properties of VU0523203 and VU6073995 demonstrate the potential of the aryl central core scaffold as a promising starting point for additional medicinal chemistry efforts focused on potency and selectivity.
DMPK of Selected Compounds
CONCLUSIONS
In summary, a successful HTS campaign was conducted with the aim of uncovering novel chemical matter in the context of Kir2.1 inhibitors. To this end 20,157 compounds from VICB’s Discovery Collection were screened, affording 48 verified hits after CRC confirmation. These compounds were classified under 6 different chemotype categories, with the most promising being the aryl central core class. Compounds belonging to this class, VU0523203 and VU6073995, both possess modest potency against Kir2.1 and improved DMPK profiles compared with the initial lead compound ML133. We hope that these novel leads can provide starting points for the next generation of Kir2.1 tool compounds and eventual drug molecules that may address any of the diseases this ion channel is associated with.
AUTHORS’ CONTRIBUTIONS
J.L.B., P.d.A.H., M.A., and Y.N. performed synthetic chemistry and compound characterization. R.M.L. performed manual patch clamp experiments and R.M.L. and J.S.D. performed data analysis. L.L. and E.L.D. performed thallium flux experiments and L.L., E.L.D., and J.A.B. performed data analysis. S.C. and S.S. performed PK experiments and bioanalysis. O.B. analyzed PK experimental data. C.W.L. and M.A. oversaw experimental design and conceived the study, and J.L.B. and P.d.A.H. wrote the article with final approval from all authors. J.LB. and P.d.A.H. contributed equally.
Footnotes
ACKNOWLEDGMENTS
The authors thank the William K. Warren Family and Foundation for endowing the WCNDD and support of our programs.
FUNDING INFORMATION
No external funding was received for this article.
DISCLOSURE STATEMENT
The authors declare no competing financial or conflict of interest.
