Abstract
Hepatic and cardiac drug adverse effects are among the leading causes of attrition in drug development programs, in part due to predictive failures of current animal or in vitro models. Hepatocytes and cardiomyocytes differentiated from human induced pluripotent stem cells (iPSCs) hold promise for predicting clinical drug effects, given their human-specific properties and their ability to harbor genetically determined characteristics that underlie inter-individual variations in drug response. Currently, the fetal-like properties and heterogeneity of hepatocytes and cardiomyocytes differentiated from iPSCs make them physiologically different from their counterparts isolated from primary tissues and limit their use for predicting clinical drug effects. To address this hurdle, there have been ongoing advances in differentiation and maturation protocols to improve the quality and use of iPSC-differentiated lineages. Among these are in vitro hepatic and cardiac cellular microsystems that can further enhance the physiology of cultured cells, can be used to better predict drug adverse effects, and investigate drug metabolism, pharmacokinetics, and pharmacodynamics to facilitate successful drug development. In this article, we discuss how cellular microsystems can establish microenvironments for these applications and propose how they could be used for potentially controlling the differentiation of hepatocytes or cardiomyocytes. The physiological relevance of cells is enhanced in cellular microsystems by simulating properties of tissue microenvironments, such as structural dimensionality, media flow, microfluidic control of media composition, and co-cultures with interacting cell types. Recent studies demonstrated that these properties also affect iPSC differentiations and we further elaborate on how they could control differentiation efficiency in microengineered devices. In summary, we describe recent advances in the field of cellular microsystems that can control the differentiation and maturation of hepatocytes and cardiomyocytes for drug evaluation. We also propose how future research with iPSCs within engineered microenvironments could enable their differentiation for scalable evaluations of drug effects.
Impact statement
Cardiac and hepatic adverse drug effects are among the leading causes of attrition in preclinical and clinical drug development programs as well as marketing withdrawals. The insufficiency of animal testing models has led to considerable interest in the employment of cardiac and hepatic models using human-induced pluripotent stem cells (iPSCs) for drug toxicity testing. However, current batches of iPSC-derived cardiomyocytes and hepatocytes are variable and not matured as adult primary tissues, which limit their prediction of drug effects. This article discusses how the use of microfluidics can create microenvironments to better control differentiation protocols and increase the physiological relevance of iPSC-derived cardiomyocytes and hepatocytes. Development and standardization of technologies will enable evaluation of the potential value of cellular microsystems to improve the in vitro models used in drug development programs. Future steps in this field include controlled connections of organ systems to better recreate clinical metabolism and pharmacokinetics.
Introduction
The ability of cells differentiated from human induced pluripotent stem cells (iPSCs) to predict clinical drug effects is limited by the immaturity of their cellular function, variability between cells or differentiation batches, and the lack of physiologically relevant properties for several applications in modeling primary cells. In this article, we discuss the potential of microfluidic and microfabricated devices to recreate microenvironments in cellular systems that can address the hurdles of differentiating, maturing, and using iPSC-derived cardiomyocytes and hepatocytes. These lineages are of interest to the drug development field because cardiac and hepatic adverse effects are the leading causes for drug attrition. 1 Relevant in vitro research with these two lineages seeks to understand drug mechanisms of action and improve the predictivity of drug effects at the preclinical stage. Reliable cellular models with strong biological relevance are critical in drug development. To this end, microengineered cellular systems have been developed utilizing a range of strategies often tailored to respective cell lineages to create biomimetic microenvironments that have resulted in improved maturity and function of iPSC-derived cell lineages. 2 Key physiological elements of cellular microenvironments include the presence of multiple cell types and organ- or tissue-specific properties that stabilize and mature cell function. The fields of microfluidics, micrometrology, and microfabrication have enabled technologies that can control and define more physiological three-dimensional (3D) multicellular cultures as well as sense or manipulate their cellular function.3–5 In addition, engineered microfluidic connections of multi-organ systems, such as interconnected heart-liver systems, can further enhance in vitro models of drug response through direct flow of metabolites and soluble factors. Such approaches can be implemented to control and improve differentiations, cellular maturity, and overall physiological relevance.
Differentiation of iPSCs involves cellular populations with compositions that progress along stages of the differentiation processes,6–8 creating a naturally heterogeneous microenvironment with complex cellular interactions and, in 3D settings, these differentiating multicellular cultures have been demonstrated to develop spontaneously at the microscale. Although microengineered 3D cultures have been reported to improve differentiation quality,9–13 owing to difficulties in handling 3D cultures, the field currently favors monolayer differentiation approaches. This approach may evolve with the development of microfabricated devices designed to perform with higher robustness and reliability for handling multicellular 3D microenvironments and for phenotyping their function in higher throughput settings. Such devices offer an unprecedented opportunity to monitor, control, and study differentiation stages and to provide models for early-stage prediction of drug effects.
We review microengineered approaches to mature iPSC-derived hepatocytes and cardiomyocytes and examine how specific features of microfluidic cellular systems are used to mimic in vivo microenvironments for improved differentiation, maturation, and monitoring of iPSC-derived cells. For using these systems, we describe potential drug development applications for the employment of iPSC-derived cardiac and hepatic cellular models. The fundaments of reprogramming 14 and differentiating iPSCs,14,15 of microfabrication 16 and microfluidics, 17 of cardiac- or hepatic-specific function,15,18,19 and of in vitro drug evaluation assays17,20 have been reviewed in detail elsewhere. Concepts from these fields are referenced and summarized for discussing how microfabricated systems with iPSC-derived hepatic and cardiac cells can be developed and used for predicting drug adverse effects.
Maturity of iPSC-differentiated cells improves with enhanced extracellular physiology
The clinical predictivity of cellular models is highly dependent on the ability of cellular functions to mimic adult human physiology. However, the fetal-like profile of many iPSC-derived cardiac and hepatic cells has been a principal challenge for modeling human tissue.21,22 The definition of maturity differs by context and is dependent on mechanistic properties that establish specific cellular activities related to tissue function. Much current research seeks to addresses challenges with promoting and defining aspects of maturity, both for iPSC-derived cardiomyocytes23,24 and hepatocytes. 25 A variety of approaches using either or both biochemical and biophysical stimuli have been developed to enhance the maturity of iPSC-derived lineages, including extended time in culture after differentiation, activation of signaling pathways using chemical factors, 3D culture settings, electromechanical stimulation, and modulation of cell morphology.17,26–33 Cellular systems can integrate these features, often in combination, and be further microengineered to facilitate measurements of biological properties that define maturation. To this end, cellular systems can be microengineered to recreate tissue- or organ-specific conditions.34–36 Here we discuss approaches specific to applications with hepatocytes and cardiomyocytes.
Hepatocytes
Despite advances in hepatocyte differentiation methods, 37 the fetal-like profile of iPSC-hepatocytes remains a challenge for translational research.38,39 Hepatocyte maturity directly relates to cellular metabolic and transport functions, notably in lipid storage, glycogen synthesis, formation of biliary canaliculi, expression of cytochrome (CYP) P450 enzymes, other drug-metabolizing proteins, transporters and biomarkers.40–45 To address this, 3D culture formats,46–48extracellular microenvironments under fluid flow, 49 and co-culture with non-parenchymal cell types 50 have been demonstrated to enhance the maturity of iPSC-hepatocytes. Accordingly, future work should investigate the biological mechanisms underlying the maturation of these cells.
Data from studies involving microengineered systems with primary hepatocytes generally suggest that more stable cultures are achieved with 3D settings, fluid flow, and co-culture with other cells, which prolonged and stabilized hepatic function for several weeks.51,52Co-cultures with hepatocytes (as reviewed in depth by Cotovio and Fernandes 19 ) may involve derivation followed by mixing and loading of individual cell lineages (e.g. hepatic endoderm and endothelial cells 53 ) or differentiation to multiple cells types (e.g. hepatic endoderm and cholangiocytes 54 and/or Kupffer cells 55 ) within the same culture (see Figure 1(c)). Therefore, a combination of structural support and activation of signaling pathways from biomimetic microenvironments may lead to the maturation of iPSC-hepatocytes with 3D co-cultures under flow.56–58 Mechanistically, studies with 3D hepatocyte culture systems have demonstrated enhanced cell polarity, formation of bile canaliculi, and increased expression of cytochromes P450 (CYPs).11,59,60 These results in improved function of primary hepatocytes suggest that microfluidic systems can be developed to better understand the fundamentals of maturation and further optimize microenvironments for culturing iPSC-hepatocytes.
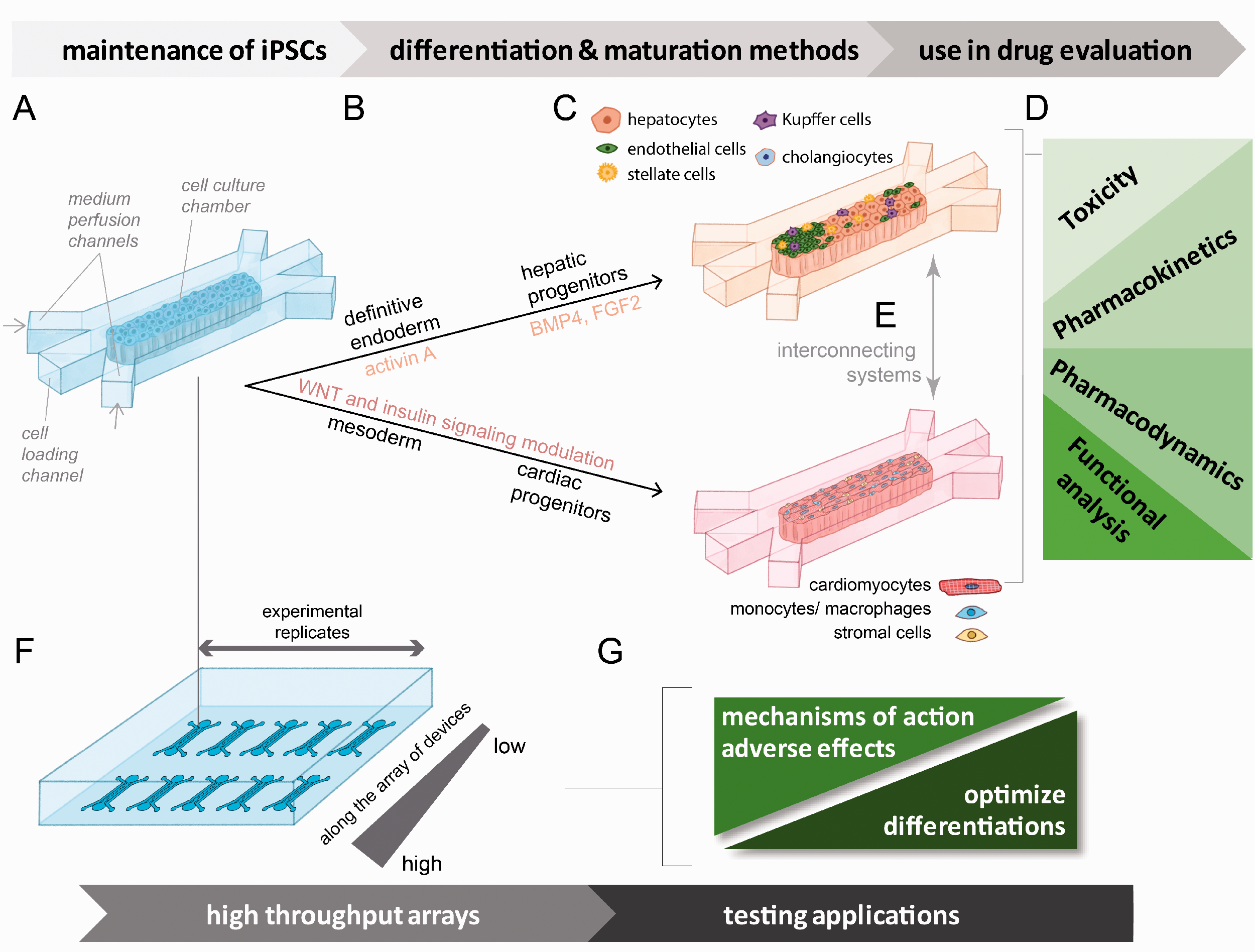
Schematic of a possible device array system for the differentiation of hiPSCs to hepatic or cardiac lineages with interconnecting capabilities and broad testing and analysis platform potential. (a) General schematic of a single device unit containing a chamber for initial hiPSC culture. Cells can be loaded through an engineered inlet and the culture can be maintained by media perfused through parallel channels. (b) Directed differentiation from hiPSCs to specified lineages (hepatic or cardiac) has potential to be performed at scale within the chambers. Media containing differentiation signals (growth factors, small molecules, media supplements) can be perfused through the channels. Widely established differentiation methods are indicated for cardiac and hepatic differentiation. (c) Co-cultures that support both cardiac and hepatic functional maturation can be created by loading mixtures of cell types into the chambers to establish heterogenous cultures. (d) Examples of functional endpoints that can be evaluated using these types of developed systems. (e) Multi-organ systems can be simulated through connecting systems loaded with different cell types. (f) Graphical representation of a device array designed for miniaturized higher throughput experimental capacity. Gradients of variable conditions can be established through microfluidic controls. (g) Examples of screening applications that can be performed on miniaturized higher throughput arrays of devices. These systems offer opportunities to screen variables that affect the efficiency and quality of differentiations and offer a platform for the screening evaluation of drugs affecting cell function. (A color version of this figure is available in the online journal.)
The next challenge to advance cellular hepatic systems is to engineer microfluidic capabilities using gradient combinations of different factors along arrays of miniaturized systems. These types of systems can be used to identify signaling factors and concentrations to optimize differentiation methods and cellular maturation, as well as to perform higher throughput functional assays (see Figure 1(d), (f), and (g)). In addition to maintaining multicellular cultures in 3D under flow, efforts have been undertaken to further emulate liver microarchitecture, either by attempting to separate bile from the cell culture medium 61 or by inducing cells to organize into acinar-like microtissues. 62 The potential to implement improvements such as these for understanding and controlling the maturation of iPSC-hepatocytes has not yet been comprehensively explored.
Cardiomyocytes
Electrophysiology, calcium handling, metabolism, contractility, and the organization and composition of subcellular structures and organelles are biological properties of cardiomyocyte function that affect the duration and magnitude of contraction for each heartbeat and therefore define cardiomyocyte maturity.17,21,63–66iPSC-cardiomyocytes differ from primary cardiomyocytes for each one of these functional modes, and cellular systems have been microengineered to enhance the maturity of cellular function, as extensively reviewed by several authors.51,67–70 Briefly, the microenvironment can be engineered to stimulate cells mechanically and electrically, expose cells to chemical factors dissolved in the cell culture medium or attached to the surface of materials, and induce cells to assume a level of structural organization and morphology that is observed in primary tissue. In addition to recreating the organization of primary tissue, 3D engineered cellular constructs, termed engineered heart tissues, have the advantage of facilitating the co-culture of different cell types (see Figure 1(c)), which has been demonstrated to enhance the maturity of iPSC-cardiomyocytes.18,71–73 Cellular organization and composition of the human heart tissue are complex and difficult to fully replicate in vitro. In addition to cardiomyocytes, various cell types exist in the heart in high numbers relative to cardiomyocytes, such as endothelial cells, smooth muscle cells, and different types of stromal fibroblasts organized within an intricate and well-defined three-dimensional network of extracellular matrix proteins. 74 Establishing in vitro co-culture systems generally involves a limited number of available cell types (e.g. iPSC-cardiomyocytes with fibroblasts or mesenchymal stem cells) at an experimentally defined ratio.
Soluble components with demonstrated enhancement of cellular maturity include factors secreted by other co-cultured cell types,75,76 metabolic conditioning with glucose-free media (including other more physiological carbon sources30,77,78), and the addition of combinations of small molecules such as tri-iodo-L-thyronine and dexamethasone.29,31 Modulation of gene expression can also increase maturity, as seen in published applications for arrhythmogenicity79,80 and calcium handling 81 studies, as well as overexpression of micro-ribonucleic acids. 82 Strategies involving soluble factors do not primarily involve an engineered microenvironment, although microfluidic formulators and perfusion of small extracellular volumes 83 can optimize the timing and concentration in the delivery of reagents to cells, either for modulating gene expression, hormonal treatment, or metabolic manipulation.
The mechanistic causes that drive cellular maturity of iPSC-cardiomyocytes are still poorly understood and may be difficult to dissect with platforms that combine several maturation factors.84,85 As reviewed by Schroer et al., 18 microdevices can focus on a specific microenvironment property. Therefore, functional effects of cell shape, cell-cell adhesions, specific properties of extracellular mechanics, and electrical stimulation can be individually evaluated. For example, Lundy et al. 86 discovered that the structural maturation of iPSC-cardiomyocytes, defined by the expression and organization of the contractile machinery and associated subcellular structures, was related to functional maturation. Consequent work used microengineered devices to further investigate these observations and confirmed the dependence of cardiomyocyte maturity on the subcellular organization of its contractile machinery.28,87–89 Overall, these studies showed that single cell maturity was enhanced with an aligned rectangular morphology enabling myofibril alignment and setting sarcomeres size within 1.9–2.2 µm. Similar relationships between these properties and functional maturity were further confirmed in multicellular systems.90,91 Other studies have focused on the importance of cell–cell adhesions for enhancing maturity.92–94 This dependence of maturity on contractile and associated structures is still poorly understood and exemplifies the additional work that is needed in the field to set cardiac maturity in vitro.
Microengineered platforms for culturing iPSC-cardiomyocytes have been developed to simultaneously address two main goals: (i) enhance the maturity and physiological significance of their functional modes (electrophysiology, electrical conduction, calcium signaling, contractility, structural organization, metabolism, chamber specificity, etc.) and (ii) enable measurement of parameters that define these functional modes.17,18,95 While fabricating devices, implementing extracellular factors and proteins and combinations of cell types that can recreate microenvironments to enhance cellular maturity, features have been created within systems to assay cardiac function. Soft lithography, micromolding, 3D printing, and hydrogel formation are the most common approaches for fabricating such devices.73,96–101 Decellularized matrices as biomimetic cues to enhance cell maturity also hold great potential for recreating physiological settings in vitro. 102 Incorporation of cells is dependent on the design of devices and can involve seeding them directly on micromolds followed by centrifugation to force cellular aggregation,97,99 loading them in culture media into microfluidic chambers under pressure, 98 or mixing them homogeneously in a hydrogel to be molded or printed into a specific shape.73,100,101 To assay cellular properties for testing drug effects on different mechanisms of cardiac function, devices can be engineered or combined with instrumentation to quantify parameters that mechanistically define electrophysiology, contractility, calcium cycling, electrical conduction, structural organization, and bioenergetics.17,99,103,104
Despite progress in the field, there remains a gap in functional modes between primary myocytes and iPSC-cardiomyocytes. Moving forward, more successful models with enhanced maturity 84 should be evaluated in multiple sites and translated by independent laboratories for prediction of clinical drug effects. Of note, significant efforts have been made in the last decade to miniaturize engineered tissues with iPSC-cardiomyocytes and multiplex the amount of functional modes that can be evaluated.97,98,105,106
Assaying functional maturity in microsystems
Microsystems can facilitate the analysis of cellular function in relation to maturity. With cardiomyocytes, functional analysis is often based on parameters that provide information about mechanisms that drive cardiac contractility, such as electrophysiology, calcium flow, force generation, and metrics of cellular mechanics and conduction. These methods have been extensively applied in two-dimensional (2D) cultures and reviewed in depth elsewhere, such as by Laurila et al. 107 To transition to engineered cellular systems, comparable methods can be integrated into devices to perform analyses with image-based techniques, force sensors, or other stimulating or recording features. 17 For example, engineered heart tissues108,109 have deformable microposts for measuring contraction force and rate. Similarly, 2D approaches for cardiac functional assays can be engineered into 3D systems, for example, electrical probes to pace cells or to record electrophysiology metrics, or optical windows for image-based assays. In addition to these methods, cardiac and hepatic assays can involve the detection of soluble factors excreted into the media, i.e. the secretome, 110 which may include indicators of cellular maturity, biomarkers, or drug metabolites that can be collected with automated microfluidic approaches.
Image-based assays can especially benefit from the use of stable lines of iPSCs with fluorescently labeled proteins. In contrast to other established methods such as immunostaining, gene expression analysis, and flow cytometry, cell lines expressing fluorescently tagged proteins offer advantages by enabling non-destructive and minimally invasive assays for tracking the status of cellular function. Deep learning software can be particularly advantageous when analyzing subtle changes in cellular phenotypes from cell imaging data, either by monitoring cultures of iPSCs and early-stage differentiation, 111 determining maturation, or assaying drug responses using experimental cultures. Such methods have recently been validated to determine the quality of iPSC cultures, 112 detect early differentiation changes, 111 and for late-stage analysis such as the categorization of drug responses using iPSC-cardiomyocytes. 113
To generate imaging data with live cells, many engineered reporter iPSC-lines are being developed to aid in the determination of cellular structure and function, including fluorescent reporter lines for structural features 114 and a GCaMP calcium sensor. 115 In addition to screening function, image-based methods can be developed to provide information concerning the purity of differentiations.
Cell-to-cell and batch-to-batch variability is another recognized challenge in the standardization of iPSC differentiations, and it can confound experimental results concerning their reproducibility. 116 To address this challenge, algorithms can be developed to screen for cellular variability from images acquired with microscopy to quantify variability and assess its experimental implications.
Towards contexts of use of iPSC-derived cardiac and hepatic systems for drug development
From a historical perspective in the evolution of the 3D cellular systems field, Simian and Bissel 117 discussed translational steps that may lead to the broad adoption of these technologies. Despite considerable advances in the field of 3D cellular systems during the past decade, a stronger commitment by drug development stakeholders will be necessary to validate and translate these technologies. Since cellular systems lack several components of the self-controllable and often poorly understood multi-factorial complexity of in vivo models, these systems are not expected to replace the employment of animals in drug development in their current state. These systems must be developed for well-defined regulatory contexts of use, such as assessing the effects of drugs on specific mechanisms.
Toxicity
The liver and the heart are among the most sensitive organs to drug toxicity. 118 Cardiac and hepatic toxicity-related side effects are a leading cause of attrition during drug development, post-marketing withdrawals, and black box warnings added to FDA-approved drug labels. 119 Therefore, increasing the predictivity of in vitro tools for clinical drug toxicity is a high priority for regulatory stakeholders. Overall, the contexts of use in predicting toxicity for cellular systems need to be defined and validated to enable confidence in their use by the pharmaceutical industry.17,57
Microengineered cellular systems can improve the predictive outcome of toxicity relative to traditional 2D cultures. For example, the presence of Kupffer cells in hepatic systems is essential to investigate inflammatory roles in drug toxicity, and traditional 2D platforms do not maintain such type of co-cultures to the same degree as 3D systems.52,120,121 A similar need has been identified to include macrophages or monocytes for predicting immune-related effects in cardiac systems.122,123 In addition, both 3D cardiac or hepatic systems can more robustly predict toxicity mechanisms that rely on prolonged drug exposures as their function systematically lasts longer and is more stable than cells in 2D platforms.41,108,124–127
As contexts of use relate to biological mechanisms, Weaver et al. 20 proposed a roadmap for implementing preclinical models of hepatotoxicity in which different mechanisms of drug-induced liver injury are comprehensively reviewed. Toxicity mechanisms in the liver can involve hepatocyte mitochondrial dysfunction, abnormal transporter activity, abnormal formation of lysosomes, formation of reactive oxygen species, damage of endoplasmic reticulum, and immune-induced injury or damage of nonparenchymal cells that support liver function.20,128 Similar publications also review the mechanistic causes of cardiotoxicity.129–133 Cardiotoxicity can lead to the dysfunction of cardiomyocyte physiological mechanisms via drug effects that directly target cardiomyocytes or indirectly affect the function of other cell types that regulate cardiomyocyte function.130,134,135
Liver metabolism that converts parent drugs into metabolites also accounts for a potential source of cardiotoxic effects that can only be predicted in vitro with interconnected heart-liver cellular systems.136–138 Commonly screened cardiotoxicity mechanisms relate to electrophysiology, calcium cycling, mitochondrial function/bioenergetics, structural tissue and subcellular properties, contractility, and cell death.129–133
In addition to assaying phenotypes and markers screening toxicity mechanisms, lists of compounds with known mechanistic drug effects are often used to validate drug development tools and serve as comprehensive controls. For example, the Comprehensive in vitro Proarrhythmia Assay initiative established a set of compounds with varied levels of proarrhythmic clinical risk to validate in vitro assays, 139 and similar lists have been proposed to screen for mitochondrial toxicity, 130 tyrosine-kinase inhibitor toxicity, 140 and contractility. 17 As iPSCs can be developed to model patient- or population-specific risk factors, they offer the potential to predict toxicity in cases where such factors play a role in drug mechanisms of action.141,142
Pharmacokinetics and pharmacodynamics
Drug toxicity depends on the level of exposure and availability of a drug. 143 Reichel and Lienau 143 reviewed how preclinical pharmacokinetic data enhance the success rate of drug discovery and referenced practical examples from the industry.144,145 A key concept in pharmacokinetic studies is drug clearance. 146 Human-specific organ function related to drug clearance is systematically determined during drug development, but it is usually estimated from indirect measurements and it is rarely determined directly with clinical testing, which could be potentially done with engineered cellular systems. 147
The liver plays a major role in the pharmacokinetics of the majority of drugs.147,148 In contrast, myocardial metabolism is mostly for energetic cellular requirements, although the possibility of drug transformation and clearance in the heart has been identified.149–151 At the mechanistic level, the expression and activity of enzymes and transporters tightly regulate drug clearance, 152 and the use of iPSC-differentiated cells for such studies must ensure the stability of the expression and activity of enzymes and transporters. 153 Related to the stability of cellular function, culturing hepatic cells in 3D platforms and under fluidic flow has been shown to stabilize transport and metabolism.41,52,154
Initial studies focusing on differentiation and maturation of iPSC-hepatocytes in 3D systems yielded promising results for obtaining functional and gene expression profiles that are closer to the activity of primary tissues.155–157 However, in addition to better investigating the mechanistic implications of 3D microenvironments in iPSC-hepatocyte differentiation, the potential employment of these systems with differentiated cells for drug clearance studies still requires elucidation. Once validated for this context, engineered systems with iPSC-differentiated cells will provide the opportunity of replicating patient-specific properties that relate to the expression of enzymes and transporters. 158
Differences in drug metabolism and transport have been identified between populations,159–161 which has strong implications in inter-population variations in drug pharmacokinetics that may be modeled in engineered cellular systems containing cells differentiated from iPSCs that represent the genetic background of specific populations. In addition to enabling population-specific studies, iPSC-hepatocytes can also harbor disease-causing mutations or conditions that affect hepatic drug clearance.148,162–164 Altered metabolic induction can occur in association with genetic determinants, as it has been reported to occur with rifampicin induction when P-glycoprotein is downregulated, or with CYP1A1 inducibility upon mutations in the aromatic hydrocarbon receptor, which activates CYP enzymes along with other transcription factors.165–167 However, induction of enzyme activity in iPSC-hepatocytes, such as CYP3A4, CYP1A2, and CYP2C9 activities, is low compared with primary hepatocytes. Engineered 3D systems have been reported to enhance the maturity of metabolic induction,32,168,169 and further research should develop this application with iPSC-hepatocytes.
To our knowledge, cardiac cellular systems containing iPSC-cardiomyocytes have not yet been tested for the evaluation of drug metabolism in cardiac tissue.149,151 However, another pharmacokinetic concept of major interest for using cardiac cellular systems is drug systemic concentration, 170 which can be replicated in vitro by setting physiological levels of drug exposure in cell culture medium with microfluidic formulators. 83 Future work should be developed in drug clearance and varied drug exposure levels with both cardiac and liver systems to evaluate how clinical pharmacokinetic data can be predicted.
The discipline of pharmacodynamics studies drug mechanisms of action on living systems, including drug-induced reactions at the cellular level, drug binding to biological receptors, and effects on the biochemistry and physiology of cellular functions. 171 As drug pharmacodynamics are intimately dependent on pharmacokinetics, which govern the time and magnitude of tissue exposure to a drug, pharmacokinetic and pharmacodynamic studies are often conducted simultaneously. 172
For engineered cellular systems, the use of microfluidic formulators that can expose cultures to well-tuned drug concentrations is a promising approach to replicate clinical drug pharmacokinetics in vitro. 83 Such high-resolution studies are necessary in drug development as there is often a considerable gap between the proposed mechanisms of action based on preclinical research and the clinical therapeutic targets of new drugs. 113
Bedside-to-bench approaches aim to use clinical data as a starting point for mechanistic investigations with in vitro methods of drug effects, as reviewed by Brackman and Giacomini 113 for the gene that encodes the breast cancer resistance protein. Similar reverse translational approaches could be applied to evaluate the predictivity of cellular systems based on known clinical effects of drugs. The main challenge for validating cellular systems, however, relates to replicating physiological mechanisms that correlate to potential drug effects (e.g. expression of receptors, involvement of multiple cell types) and discordance between results from these systems and clinical data can inform on their limitations for predicting drug effects.
Integrating iPSC-differentiated cells in more predictive clinical effect platforms would also be key for applications that relate to rare diseases and personalized and precision medicine. In the cardiac field, several mutations and combined genetic conditions are known to cause cardiomyopathies and other rare cardiac conditions. 173 In addition, drugs effects such as doxorubicin toxicity174,175 can also be related to the genetic background of patients and related gene expression profiles, which further illustrates the opportunity of using patient-specific cells.
Similar opportunities for addressing patient-specific drug effects also exist for the use of iPSC-differentiated cells in liver cellular systems. 176 In addition to recreating settings dependent on genetic backgrounds, cellular systems can be engineered to replicate microenvironments that enhance the physiological effect of drugs. For example, a variety of liver systems can recreate oxygen zonation by setting extracellular oxygen tension, which can be critical to detecting specific drug effects.57,177 Physical exercise and general conditioning of cardiovascular physiology can also play a strong role in drug cardiac effects178–182 and several cardiac microphysiological systems have the ability to stimulate cells electrically and mechanically to recreate physiological conditioning.73,127,183 Altogether, patient-specific conditions can be recreated with iPSC-derived lineages and the pathophysiology of cardiac or hepatic function can be emulated by engineering the extracellular microenvironment.
Features of microfluidic cellular systems for cardiomyocyte and hepatocyte differentiation
Despite the progress in the fields of microfluidics and microfabrication, use of 2D cultures remains vastly predominant in differentiations, and testing of the potential for cellular microsystems, non-invasive functional monitoring, and programmed media changes is in early stages of initiation.
The differentiation methods for deriving cardiomyocytes or hepatocytes from iPSCs are currently well-standardized in the field for 2D culture. Derivation of cardiomyocytes most often relies on carefully timed modulation of WNT signaling to induce mesoderm and specify the cardiomyocyte progenitors along with ongoing modulation of insulin signaling. 184 The pathways of hepatocyte differentiation are reproduced in vitro using Activin A to induce formation of definitive endoderm, followed by the addition of BMP4 and FGF2 to specify hepatic progenitors.185,186 Figure 1(b) represents these two differentiation pathways. Most current differentiation protocols have been established using 2D or monolayer culture systems. In this review, we seek to emphasize the value of incorporating 3D systems at early or initial stages of differentiation, an approach that has led to development of tissue platforms with iPSC-CMs.187,188 Transitioning differentiations from 2D systems to 3D devices involves challenges at many stages. Firstly, the process of seeding or loading iPSCs into 3D devices may require more rigorous dissociation or additional manipulation. Compromising cell quality through experimental handling is a known challenge for sensitive cells such as iPSCs, 189 although alternative approaches to reduce stress during cell loading into devices have been developed, such as transient on-chip vacuums 190 or inducing aspiration using degassed PDMS (polydimethylsiloxane). 191 Some additional challenges during cell differentiation may include the need to adjust the timing of signaling factor addition since stage progression may be paced differently due to spatial-temporal differences in a 3D format, 192 and addressing the retainment of dead cells within a device that may accumulate as a natural process of ongoing differentiation. 193
Engineered cellular microsystems, such as 3D organoids or microphysiological systems, seek to recreate elements of the in vivo microenvironment by facilitating cell–cell interactions and allowing self-organization of structural features. 194 Ronaldson-Bouchard et al. 84 observed that iPSC-cardiomyocytes co-cultured with stromal cells in 3D cultures and under electrical training yielded more mature modes of function when cells were seeded into these settings relatively early in differentiation (around Day 12). In a similar manner, microsystems used for deriving iPSC-hepatocytes have demonstrated promising results when integrating elements of 3D culture and microfluidic perfusion.49,195,196 Furthermore, most differentiation methods rely on soluble factors as differentiation cues. When culturing iPSCs for differentiation in microfluidic devices, as exemplified in Figure 1, there is potential to tune the microenvironment to either enhance or replace signaling factors based on cell–cell interactions and co-cultures of cell lineages. 197 The extracellular matrix (ECM) is also a critical aspect of both in vivo and in vitro cell function and biology and can be experimentally defined both in 2D and 3D culture settings, as well as in devices that facilitate differentiation and maturation stages in 3D. The ECM is known to modulate cell signaling and differentiation cues, for example, ECM-integrin signaling has roles both in early-stage differentiation of iPSCs as well as through later stages of cardiomyocyte differentiation and maturation. 198 ECM studies open the possibility of biomimetic approaches to stem cell differentiation methods, which could be controlled in microsystems designed to generate specific lineages. 199
Different designs have been applied for the development of microengineered liver and cardiac cellular systems, which include open and closed systems, static or dynamic flow, as well as systems that require different quantities of cells.19,57,200 Figure 1(a) represents a general closed microfluidic device, featuring an inlet for loading cells as well as microfluidic channels for media perfusion. Cells can be loaded into culture chambers or wells as a mono- or co-cultures, generally by dissociating and mixing lineages, when applicable, before resuspending in culture medium at the appropriate density. 19 Depending on the device design, cells then form attached layers, spheroids, or organoids in either open or chambered systems where they have exposure to perfused media in defined environments 57 For example, the open-chamber device designed to mimic the structure of a single acinus by Prodanov et al. 201 with iPSC-hepatocytes differs from the scaffolds developed by Domansky et al. 200 to drive the self-assembly of iPSC-hepatocytes into arrays of 3D microtissues. For overall reference, Cotovio and Fernandes 19 reviewed current methods for the formation of iPSC-derived liver organoids.
Engineering cellular microsystems for improving differentiations can offer many advantages, and these systems can be miniaturized for using low cell numbers within microfluidic chambers (see Figure 1(a)). In addition, image-based and other analytical non-destructive assays can examine the cellular secretome within the perfusate to provide quality control criteria for evaluating the progress of differentiations. As the field evolves toward leveraging microengineered platforms for iPSC differentiations, we will increase our understanding of how the microenvironment impacts differentiation. 202 In the rest of this section, we discuss how specific features of microfluidic cellular systems (programmed media changes and formulation, perfusion flow settings to maintain homeostasis, set specified cell-to-media ratio, and establishment of shear force) can be used to mimic in vivo microenvironments for the enhancement of differentiation outcomes and improvement of analysis methods.
Overall, programmable microfluidic devices with automated capabilities for media formulation and media change can considerably reduce human-induced error and variability and allow for higher standards of consistency during often extended and sensitive experimental procedures. In the context of differentiations, media changes are critical and rely on careful optimization and standardization of timing and factor concentrations. Culture media often needs to be changed during brief time windows of developmental competence to progress cell fate through developmental intermediates. Competent stages can last mere hours, which can create challenges in optimizing protocols and keeping differentiations standardized due to user-to-user variability. Media formulations are also critical and can contain highly sensitive concentrations of growth factors affected by lot number, aliquot age, and media temperature or storage conditions. For example, the WNT inhibitor CHIR99021 in cardiomyocyte differentiation protocols needs finely tuned optimization for different cell lines.203,204 Programmable microfluidic devices, such as the MultiWell MicroFormulator designed to deliver customized reagent mixtures over 96-well culture plates, 205 have the potential to assist in optimization of timing and formulation as well as standardization of procedures across users and sites. However, media components and small molecules used in differentiation can be temperature sensitive and the development and use of programmed automatic media handling and formulation should incorporate solutions to protect perishable solutions.
In traditional cell culture platforms, media remain static between changes, which can result in temporal gradients of nutrient and oxygen depletion or build-up of cellular waste products. In addition, over the course of differentiations, cells are exposed to exogenous signaling factors (small molecules or growth factors) that guide sensitive developmental decisions, and many of these factors can be unstable in culture conditions. 206 Microsystems featuring continuous media exchange can address these hurdles by providing a constantly refreshed soluble microenvironment. Continual flow without media recirculation offers the advantage of a consistent and defined soluble microenvironment to minimize culture-based variability for cells whose metabolic activity can affect their microenvironment. 207 In systems with static culture relying on manual maintenance, intermittent media changes can cause biological fluctuations in the microenvironment that significantly impact the homeostasis of cellular function. While continuous flow can provide constant renewal of microenvironmental conditions, in some applications it also allows the concentration of excreted factors to reach a natural equilibrium by avoiding media changes. For example, hepatocytes have been shown to establish steady states of glucose and lactate concentrations under conditions of continuous but slow media supply. 208 Additionally, microfluidic systems can be designed to create signaling gradients within a culture to spatially influence cell response or development. For example, Cui et al. 209 demonstrated regionalized formation of the primitive streak from pluripotent stem cells within a device using controlled microfluidics. This same approach could be applied to developmental models to optimize differentiation or maturation conditions, differentiate co-cultures within a device, or for downstream evaluation of various drug concentration gradients.
Maintaining a low ratio between cellular mass and media volume in culture systems can potentially increase assay sensitivity and therefore enable either quantification of low-level factors or detection of subtle changes in soluble markers. The use of devices designed with low cell-to-media ratios maintained by slow consistent media exchange allowing for high concentrations of secreted factor can enable studies that are dependent on such variables. By decreasing the cellular volume density of microfluidic extracellular environments, analysis of the secretome of cultured cells in the perfusate of microengineered devices 210 could enable a more detailed analysis of maturation stages from bioanalytical assays.
Research with primary hepatocytes in low-volume systems has indicated that secreted factors such as HGF and TGF-β1 play a role in maintaining functional hepatocyte identity in culture, 211 further indicating the need to understand the effects of stabilizing the microenvironment at a low cell-to-media ratio. This technique has not yet been explored in directed differentiation methods. These cases highlight the need for carefully designed strategies tailoring the extracellular environment to the biological properties of the research.
In addition to slowly exchanging cell culture media and maintaining the homeostasis of the microenvironment, microfluidics can be used to recreate the extracellular physiology of cells exposed to shear stress caused by fluid flow. 57 Although cardiomyocytes are not directly exposed to shear forces in the beating heart, hepatocytes are exposed to fluid flow in the liver. 212 Recreating this native property of shear force exposure in vitro for hepatocyte cultures has been shown to enhance mature properties of iPSC-hepatocytes 213 and could also be used to facilitate differentiations by increasing efficiency and functional properties. 214 Most liver-on-a-chip devices on the market use controlled microfluidic flow to enhance and maintain hepatocyte function.215–217 Further research should evaluate how microfluidic systems can benefit differentiations and how to optimize flow conditions for developmental intermediates.
Many early differentiation methods used cell aggregate methods such as embryoid body formats, either free-floating or through hanging drop formation. Transitioning to 2D methods (monolayer culture) offered the advantage of a defined and homogenous environment, but it can limit unknown cell–cell interactions and structural heterogeneity. Microfluidic settings are ideal for recreating 3D microenvironments for differentiating iPSCs. One major approach is the containment of cells within a device, providing an enclosure for aggregates that minimizes inadvertent loss of embryoid bodies, organoids, or spheroids during handling, testing of compounds, or media changes that can occur with open culture platforms.
The method of culturing spheroids with hanging drops has been used for both hepatocytes and cardiomyocytes.105,218,219 Despite their improvements over monolayer cultures, these methods often remain subject to low-throughput limitations and influences including media exposure, aggregate size, and undefined cell–cell interactions. Currently, the challenge is to integrate similar tissue-building methods in more robust and highly defined microenvironments. The use of microfluidics in such systems can control for variables such as media flow, volume, oxygen tension and can provide opportunities for connected multi-lineage 3D systems.
Future challenges and opportunities
As an increasing number of cellular systems become implemented in iPSC fields, the next steps include connecting organ systems to further replicate in vivo physiology. Integrating organ systems through connected devices provides unique opportunities for studying drug effects that rely on multi-organ interactions. Generally, organ lineages are cultured within respective devices, and these devices can be connected via microfluidic systems to interchange soluble factors for experimental analysis.201,220Cardiac-hepatic interactions are of interest in drug development due to the prevalence of drug metabolites produced in the liver that can affect cardiac function. 221 Development of connected systems reliably replicating such in vivo interactions can provide a platform for predicting drug adverse effects and allow for mechanistic studies of organ physiology. For these systems to reliably mimic in vivo interactions, connections between tissues need to be carefully designed and monitored to have appropriate flow parameters and pharmacokinetic properties. When iPSC-derived tissues are used, all lineages should be assayed for appropriate levels of functional maturity and robustness (Figure 1(b) and (g)).
As the field moves forward, there is increasing potential for fully integrated systems that can accommodate start-to-finish experiments, from initial seeding of iPSCs to monitoring of differentiations to end-point functional assays (Figure 1). Developing and integrating higher throughput, non-destructive, minimally invasive phenotyping analysis methods is of interest. Image-based machine learning platforms lend themselves to such higher throughput capacities in addition to bioanalytical methods. The next challenge is to incorporate different capabilities for maturing cells, while assaying their function, in miniaturized cellular systems. Given the high sensitivity of iPSCs to experimental handling, it can also be assumed that loading these cells in microfluidic devices without affecting their quality will be a constant challenge in the field.
In summary, we include a list of recommendations (Table 1) for the field to consider as research continues to integrate iPSC-derived lineages with increasingly sophisticated cellular systems toward the goal of predicting clinical drug effects.
Summary of recommendations.

iPSC: induced pluripotent stem cell.
Footnotes
Authors’ contributions
KD and AJSR contributed equally to this manuscript.
ACKNOWLEDGEMENTS
The authors acknowledge the members of the Division of Applied Regulatory Science and of the Office of Clinical Pharmacology at FDA for valuable discussions.
Disclaimer
This article reflects the views of the authors and should not be construed to represent the views or policies of the FDA.
Declaration OF CONFLICTING INTERESTS
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported via the Food and Drug Administration – Medical Countermeasures Initiative Intramural Challenge Grants funding.
