Abstract
Background:
In patients with metastatic colorectal cancer (mCRC), Quxie Cap-sule (QX)—a combination of conventional therapy (including chemotherapy, targeted therapy or supportive care)—has shown a significant overall survival benefit compared with placebo and might have the property of dual effects of antitumor and immunity enhancement, both mediated by the microbiome. In preclinical models, QX has also shown activity against colorectal cancer. This study aimed to describe how the aforementioned effects of QX look after when focusing on the patients in third or above line setting.
Methods:
A Simon’s Minimax two-stage phase II design was used in this study, which enrolled mCRC patients who progressed after second-line treatment. Patients received conventional therapy plus QX until disease progression or unacceptable toxicity. Before and after 1-month intervention, we collected patients’ stool samples for microbiome analysis by 16s rRNA sequencing approaches. And the microbiome analysis before and after 1-month intervention was done through bioinformation analysis platform.
Results:
Fifteen patients were enrolled and gut microbiome were analyzed from 7 of 10 patients that with PFS over 3.7 months. Microbiome community analysis on genus level showed that the proportion of Lachnospiraceae_UCG-001 (0.04% vs 1.06%, P = .02249) significantly increased after conventional therapy plus QX while the proportion of Alistipes (2.96% vs 1.35%, P = .03461), Flavonifractor (0.04% vs 0.02%, P = .02249), Bifidobacterium (6.11% vs 1.14%, P = .02249) and Butyricimonas (0.24% vs 0.11%, P = .03603) significantly decreased after intervention . LEfSe analysis showed that after intervention, samples were highly related with unclassified-f-lachnospiraceae, Eubacterium and Lachnospiraceae_UCG-001.
Conclusions:
Decrease of gut bacteria with potential roles in carcinogenesis of colorectal cancer and increase in the abundance of gut anticancer bacteria such as Lachnospiraceae may partly explain how conventional therapy combined with QX can influence carcinogenesis and tumor progression in colon cancer.
Trial registration:
Chinese Clinical Trial Registry (ChiCTR2100053874).
Keywords
Introduction
Colorectal cancer (CRC) is the third-leading cause of cancer-related death. 1 Among people diagnosed with mCRC, approximately 30% to 35% of patients survive beyond 3 years, and fewer than 20% beyond 5 years from diagnosis. 2 Multivariate regression analysis identified use of third line chemotherapy (regorafenib or trifluridine/tipiracil) as associated with better survival. 1 At present, the three-line standard monotherapy regimen recommended by the NCCN and CSCO Guidelines is: Fuquinitinib, Regofenib, and TAS-102, and the combination therapy as will as the treatment after the third-line are still being explored.3,4 However, doctors need to consider various factors including treatment goals and patient wishes when making treatment decisions for the patients with mCRC of the third or above line setting. Due to the rapid progression of multi-line treatment, multiple drug intolerance or economic factors in existing prescriptions, many patients will face the dilemma of no standard treatment available. Therefore, new and effective treatment options are urgently needed to address these problems. QX, a compound of 21 kinds of Traditional Chinese Medicine (TCM) herbs, has been used to treat patients with metastatic colorectal cancer (mCRC) and could suppress the growth of colon cancer.5,6 Pre-clinical and clinical studies have revealed that QX can benefit patients with mCRC and play a role by affecting intestinal microbes.5,7 Specifically, QX could increase the abundance of gut anticancer bacteria such as Actinobacteria as well as butyrate-producing bacteria such as Lachnospiraceae. However, what is the specific effect of QX on the gut microbiome in the third or above line setting patients for which it is effective? We hypothesized that the combination of QX can be used safely in the third or above line setting therapy and is well tolerated by patients. Further, we hypothesized that conventional therapy plus QX partly works by affecting the gut microbiota. To test our hypothesis, we designed a Simon’s Minimax two-stage phase II trial to evaluate the efficacy and safety of conventional therapy with QX in patients with mCRC who had received second-line therapy, at the same time, stool samples were collected before treatment and 1 month after treatment, and the changes of gut microbiome in patients with effective QX treatment (PFS > 3.7 months) were analyzed before and after treatment. This study, to our knowledge, is the first to research the specific effect of conventional therapy plus QX on the gut microbiome in the third or above line setting patients.
The first stage of Simon two-stage design of this one-arm trial has met the primary endpoint, and the interim analysis shows that conventional therapy plus QX may have encouraging efficacy. Twelve patients were evaluable for efficacy, The median PFS was 5.3 months (95% CI: 4.6-6.0 months) by investigator assessment. For all 12 patients enrolled for whom efficacy can be evaluated, investigator-assessed progression-free survival at 16 weeks was 66.7% (95% CI: 35.4-98) and 33.3% (2-64.6) at 24 weeks, 1-year overall survival was 83.3% (80% CI: 68-98.7). The results of these clinical trials are being submitted, and the second stage of the trial is under way. At the same time, we observed that conventional therapy combined with QX modulated the gut microbiome, so we report the results below.
Material and Methods
Trial Patients
Fifteen patients (≥18 years of age) with an Eastern Cooperative Oncology Group (ECOG) performance status of 0 to 1 and histologically confirmed unresectable, metastatic colorectal adenocarcinoma, as well as a treatment history of 2 or more regimens of standard chemotherapy. Eligible patients should have measurable disease by the Response Evaluation Criteria in Solid tumors (RECIST) version 1.1, 8 adequate bone marrow, hepatic, and renal functions were required by blood testing within the 7 days before enrollment. Moreover, patients should have a treatment history of at least 2 regimens of chemotherapy, and were refractory or intolerant to fluoropyrimidine, irinotecan, oxaliplatin, anti-VEGF therapy, and anti-EGFR therapy (for patients with wild-type KRAS). 7 A patient was deemed refractory to a chemotherapy if they had confirmed tumor progression based on imaging during or within 3 months after the last administration of the chemotherapy in a metastatic setting. 7 Intolerance to a chemotherapy was defined as the patient having experienced clinically significant adverse events from chemotherapy that precluded the re-administration of that same chemotherapy. 7 Key exclusion criteria included active infections, require immunosuppressive treatment due to an autoimmune disorder or history of organ transplantation, or have unresolved adverse events of grade 2 or higher (classified with the National Cancer Institute Common Terminology Criteria for Adverse Events [CTCAE] version 4.03) from previous treatment.2,9,10
All patients gave their written informed consent before their enrollment to the study.
Trial Design, Treatment, and Procedures
This was a Simon’s Minimax two-stage single-arm, open-label, non-randomized phase 2 trial in Xiyuan Hospital of China Academy of Chinese Medical Sciences between November 2021 and September 2025. Participants were recruited from 216 individuals who were willing to participate, but only 15 met eligibility criteria (Figure 1).

Trial profile.
QX capsule contains TCM herbs—Euodiae Fructus, Zingiberis Rhizoma, Poria, Aurantii Fructus Immaturus, Magnoliae Officinalis Cortex, Succinum, Cinnamomi Cortex, Coptidis Rhizoma, Pinelliae Rhizoma, Acori Tatarinowii Rhizoma, Aquilariae Lignum Resinatum, Radix aconiti, Hordei fructus germinatus, Citri Exocarpium Rubrum, Crotonis Semen Pulveratum, Gleditsiae Sinensis Fructus, Ginseng Radix Et Rhizoma, Arecae Semen, Coptis chinensis, Platycodonis Radix, and Aconiti Radix Cocta.2,3All herbs were made by the Pharmaceutical Center of Xiyuan Hospital under the quality control of the institution (production date1: August, 2021; expiration date1: August, 2023; production date2: March,2023; expiration date2: March, 2025). All capsules were stored in a storage room under constant temperature and humidity. 2
The patients received conventional therapy which based on the patient’s prior chemotherapy exposure and physician preference including chemotherapy, targeted therapy, supportive care and Chinese herbal medicine plus QX (each capsule of 0.2 g was orally administered at 50 mg/kg, twice daily, day 1-20, in a 30-day course) for at least 3 months.
Both conventional therapy and QX treatment cycles overlapping or not overlapping were allowed. Safety assessments were performed on days 30 ± 7 of cycle 1, and days 30 ± 7 of subsequent cycles. Adverse events were characterized using CTCAE, version 4.03. Disease assessment with cross-sectional imaging (CT or MRI) was performed at baseline and every 2 cycles of conventional therapy or QX treatment. Tumor assessments were performed using RECIST 1.1. Stool samples was collected on cycle 1 day 1 of QX (pre-treatment) and QX cycle 2 day1 ± 7 for microbiome analysis of patients with effective QX treatment (PFS > 3.7 months) by 16s rRNA sequencing approaches.
Stool samples were collected from 8 of the 10 patients that with PFS over 3.7 months, 11 of which one set of collected samples was not suitable for analysis.
Total microbial genomic DNA was extracted from 7 samples using the PF Mag-Bind Stool DNA Kit (Omega Bio-tek, Georgia, U.S.) according to manufacturer’s instructions. The quality and concentration of DNA were determined by 1.0% agarose gel electrophoresis and a NanoDrop® ND-2000 spectrophotometer (Thermo Scientific Inc., USA) and kept at −80°C prior to further use. The hypervariable region V3-V4 of the bacterial 16 seconds rRNA gene were amplified with primer pairs 338F (5′-ACTCCTACGGGAGGCAGCAG-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′) 11 by an ABI GeneAmp® 9700 PCR thermocycler (ABI, CA, USA). The PCR reaction mixture including 4 μL 5 × Fast Pfu buffer, 2 μL 2.5 mM dNTPs, 0.8 μL each primer (5 μM), 0.4 μL Fast Pfu polymerase, 10 ng of template DNA, and ddH2O to a final volume of 20 µL. PCR amplification cycling conditions were as follows: initial denaturation at 95°C for 3 minutes, followed by 27 cycles of denaturing at 95°C for 30 seconds, annealing at 55°C for 30 seconds and extension at 72°C for 45 seconds, and single extension at 72°C for 10 minutes, and end at 4°C. All samples were amplified in triplicate. The PCR product was extracted from 2% agarose gel and purified. Then quantified using Quantus™ Fluorometer (Promega, USA). Purified amplicons were pooled in equimolar amounts and paired-end sequenced on an Illumina PE300 platform/PE250 platform (Illumina, San Diego, USA) according to the standard protocols by Majorbio Bio-Pharm Technology Co. Ltd. (Shanghai, China). After demultiplexing, the resulting sequences were quality filtered with fastp (0.19.6) 12 and merged with FLASH (v1.2.11). 13 Then the high-quality sequences were de-noised using DADA2 14 (or Deblur) 15 plugin in the Qiime2 16 (version 2020.2) pipeline with recommended parameters, which obtains single nucleotide resolution based on error profiles within samples. DADA2 (or Deblur) denoised sequences are usually called amplicon sequence variants (ASVs). To minimize the effects of sequencing depth on alpha and beta diversity measure, the number of sequence from each sample was rarefied to 20 000, which still yielded an average Good’s coverage of 97.90%.
Trial Objectives
The objectives were to determine the effects of conventional therapy plus QX when administered in combination as determined by analysis of pre- and post-treatment stool samples for gut microbiome analysis.
Sample Size Estimation
Simon’s Minimax two-stage phase II design was used to calculate the sample size. 17 We considered the study drug to be inactive if DCR was ≤15%, and considered the study drug as active if DCR was ≥35%.18,19 Assuming the type I error of 0.025 and the type II error of 0.20, running by g PASS 15.0.5, the sample size and stopping rule were as follows: accrue 12 patients in stage I; if 2 or fewer of the 12 patients were nonprogressors, we would stop the study and conclude the study drug as inactive, otherwise proceed to stage II and accrue an additional 33 patients. After stage I interim analysis, the study will proceeded to stage II. In stage I, we recruited a total of 15 patients as considering the censoring rate of 20%.
Statistical Methods
This single arm phase 2 study evaluated the conventional therapy plus QX. The study is descriptive in nature with the goal to initially explore the efficacy of conventional therapy plus QX in mCRC patients with the third or above line setting to determine whether a larger sample size and higher level of evidence-based medicine evidence can be conducted. Bioinformatic analysis of the gut microbiota was carried out using the Majorbio Cloud platform (https://cloud.majorbio.com). Based on the ASVs information, rarefaction curves and alpha diversity indices including observed ASVs, Chao1 richness, Shannon index and Good’s coverage were calculated with Mothur v1.30.2. 20 The similarity among the microbial communities in different samples was determined by principal coordinate analysis (PCoA) based on Bray-curtis dissimilarity using Vegan v2.4.3 package. The PERMANOVA test was used to assess the percentage of variation explained by the treatment along with its statistical significance using Vegan v2.4.3 package. The linear discriminant analysis (LDA) effect size (LEfSe) 21 (http://huttenhower.sph.harvard.edu/LEfSe) was performed to identify the significantly abundant taxa (phylum to genera) of bacteria among the different groups (LDA score > 2, P < .05).
Results
Patients
A total of 15 patients were enrolled from 24 November 2021 to 5 January 2023. 3 of the 15 patients withdrawn consent—these 3 patients did not provide patient diaries and were not followed up. Summarizes the baseline characteristics of 12 followed up patients. Overall, the median age was 62 years (range 43-77) with 25% of patients ≥65 years of age. Most patients were male (75%). All patients had an ECOG performance score of 0 (75%) or 1 (25%). Eleven patients were left-sided colorectum cancer (91.7%). Eleven (91.7%) patients’ number of metastatic organs were more than 2.
Safety
The cutoff date for both safety and activity analyses was 30 September 2023, with a median follow-up of 16.9 months (IQR 10.5-21.1). Treatment was well tolerated with manageable toxicities. Treatment-related adverse events (TRAEs) are defined as adverse events attributed to, or likely attributed to, any of the study drugs. The most common grade 1 TRAEs included diarrhea (66.7%), dyspepsia (50%), abdominal pain (41.7%), GGT increased(8.3%) in Table 1. Grade 3 TRAEs were reported in 1 patient (8.7%) with occurrence of GGT increased, which was accompanied by grade 2 alkaline phosphatase increased and blood bilirubin increased, when the liver target lesions were enlarged and investigator determined disease progression.
Adverse Events During Study Treatment Period (according to CTCAE v4.03).

No grade 4 adverse events and treatment-related deaths occurred.
Treatment-Associated Changes in the Gut Microbiome Analysis
Alpha diversity
The Shannon/ACE curves based on the Shannon/ACE indexes showed that the sequencing data and depth and sample size were sufficient (Figure 2A). Alpha analysis on OTU level showed that Shannon/ACE indexes declined after conventional therapy with QX, however there were no statistically significant differences between before and after intervention (Figure 2B and Table 2).
Alpha Diversity Index.


(A) The rarefaction curve before and after QX. (B) Alpha analysis of Shannon/ACE indexes.
Beta diversity
For beta analysis, PCoA on OTU level did not show significant differences on principal component 1 (PC1) before and after intervention(P = .068) as shown in Figure 3 A1. Meanwhile, PCoA at the genus and phylum level show significant differences before and after intervention as shown in Figure 3 A2 and A3 (P = .031 and 0.013, respectively).
Alterations in the composition of gut microbiome
Through 16 seconds rRNA sequencing analysis of gut microbiome after intervention, it was found that there were 166 common genera among the groups at the genus level, and there were 57 distinct genera before treatment and 21 distinct genera after treatment (Figure 3B) .

(A) PCoA on OUT, Genus and Phylum show the differences on principal component. (A1) PCoA on OTU level. (A2) PCoA at the genus level. (A3) PCoA at the phylum level. (B) Specific species before and after intervention of the combination therapy of QX.
The relative proportions of dominant taxa at the phylum level were assessed by microbial taxon assignment in both groups before and after intervention. We observed considerable variability in gut microbiome across samples in each group (Figure 4). Five phyla were identified in each group. Firmicutes was the most predominant phylum, accounting for 46.44% and 77.07% of the OTUs before and after intervention, respectively. In addition, Bacteroidota (24.59% vs 13.00%), Proteobacteria (19.04% vs 8.52%), Actinobacteriota (9.29% vs 1.16%), and others (0.64% vs 0.25%) were declined after intervention. To compare the differences in gut microbiome between before and after intervention, Wilcoxon signed-rank test was performed for different classification levels. At the phylum level, Actinobacteriota (P < .05) were significantly lower abundance after intervention, meanwhile Bacteroidota, Proteobacteria, and others were also declined but there were no statistically significant differences between before and after intervention. And Firmicutes increased after intervention, however there were also no statistically significant differences.
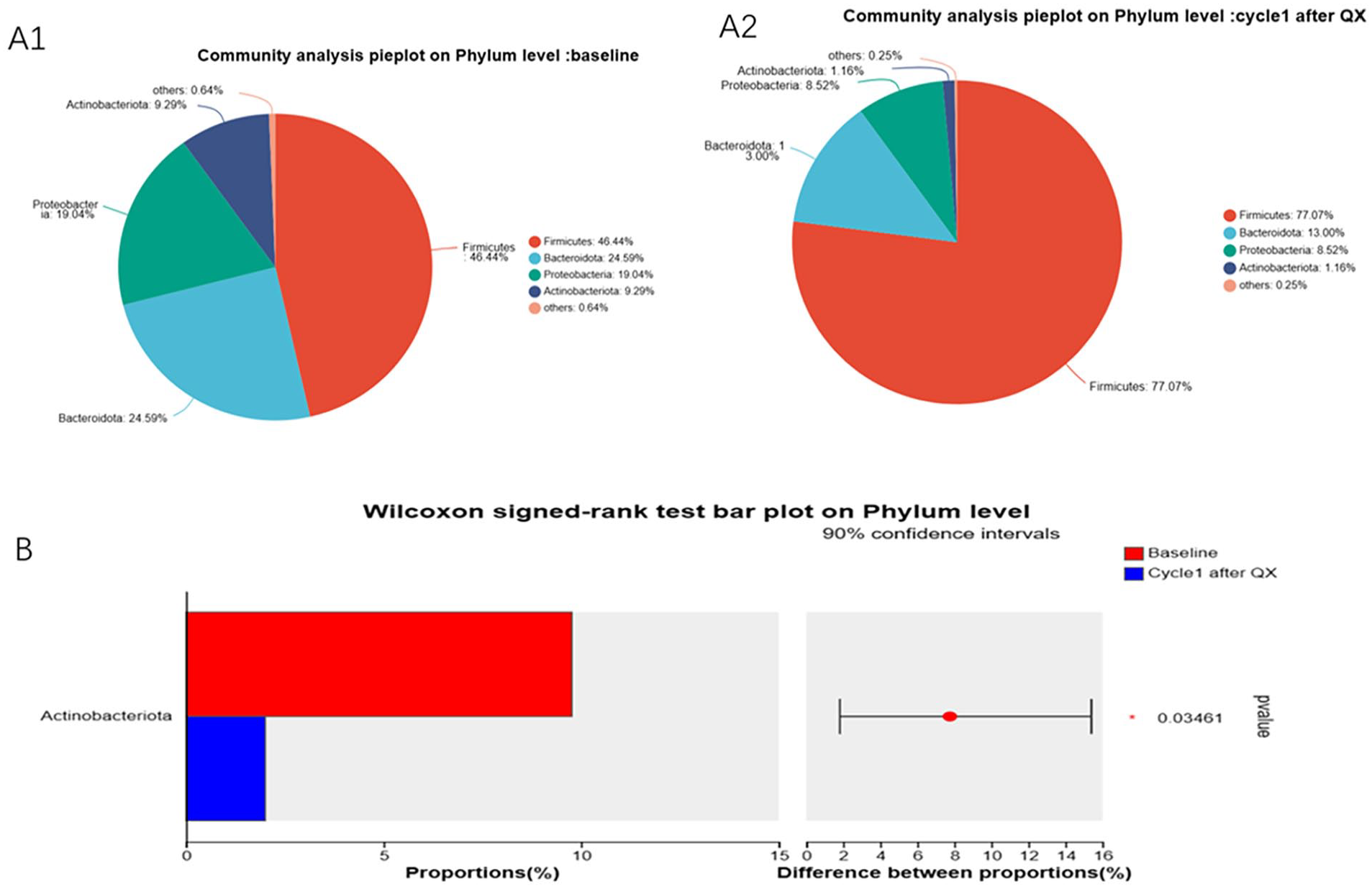
Difference of community proportions on at the phylum level before and after intervention. (A) The pie chart showed that Firmicutes were more dominant after QX intervention. (B) The histogram shows that the abundance of Actinobacteriota has increased significantly after intervention.
At the genus level, a total of 32 genera were found to be significantly different between before and after intervention. Of these discriminatory taxa, Bacteroides (12.46% vs 9.10%), Subdoligranulum (4.63% vs 13.28%), Blautia (4.47% vs 11.15%), Faecalibacterium (3.96% vs 10.06%) were the most predominant genera at the genus level. In addition, Bifidobacterium (P = .02249), Alistipes (P = .03461), Butyri-cimonas (P = .03603), and Flavonifractor (P = .02249), were found to be significantly lower abundance after intervention, Lachnospiraceae_UCG-001 (P = .02249) were significantly increased after intervention, respectively (Figure 5).

Difference of community proportions on at the genus level before and after intervention. (A) The bar chart shows 32 genera that were found to be significantly different between before and after intervention. (B) The heat map showed that the abundance of bifidobacterium decreased after treatment, which was similar to the results of previous studies, and then we conducted a species difference analysis. (C) The results of Circos map and heat map are similar, which support each other. (D) Species difference analysis shows that although Bifidobacterium, Flavonifractor and Butyricimonas are not the highest in abundance, there are significant differences before and after treatment.
Considering that this discriminant analysis did not distinguish the predominant taxon, LEfSe was used to generate a cladogram to identify the specific bacteria associated with intervention. We identified 33 discriminatory genera at the genus level as key discriminants. Only taxa with an LDA value > 2 are presented. Circles indicate phylogenetic levels, diameter and color of each circle represent its abundance and enterotype, respectively. Actinobacteriota prevailed before intervention, and after intervention, Lachnospiraceae were abundant (Figure 6).

LEfSe multi level species differences analysis before and after intervention. (A) Dots are marked for significant enrichment either before the intervention (n = 7) (red) or after the intervention (n = 7) (blue) at multi level species, while nondiscriminant taxon nodes are represented by yellow. Only taxa with an LDA value >2 are presented. (B) Bacterial histograms of unique biomarkers based on LEfSe (>2). The length of the bar chart represents the magnitude of the impact of significantly different genus.
The 16 second taxonomic lineages based on Silva database were converted into the taxonomic lineages of prokaryotes in KEGG database by Tax4Fun, and then KEGG functional annotation of 16S RNA gene sequences was performed. Tax4Fun analysis revealed differences in potential microbial functional signatures (Figure 7A). A decrease in the function of potential Human Diseases pathways was observed after intervention.
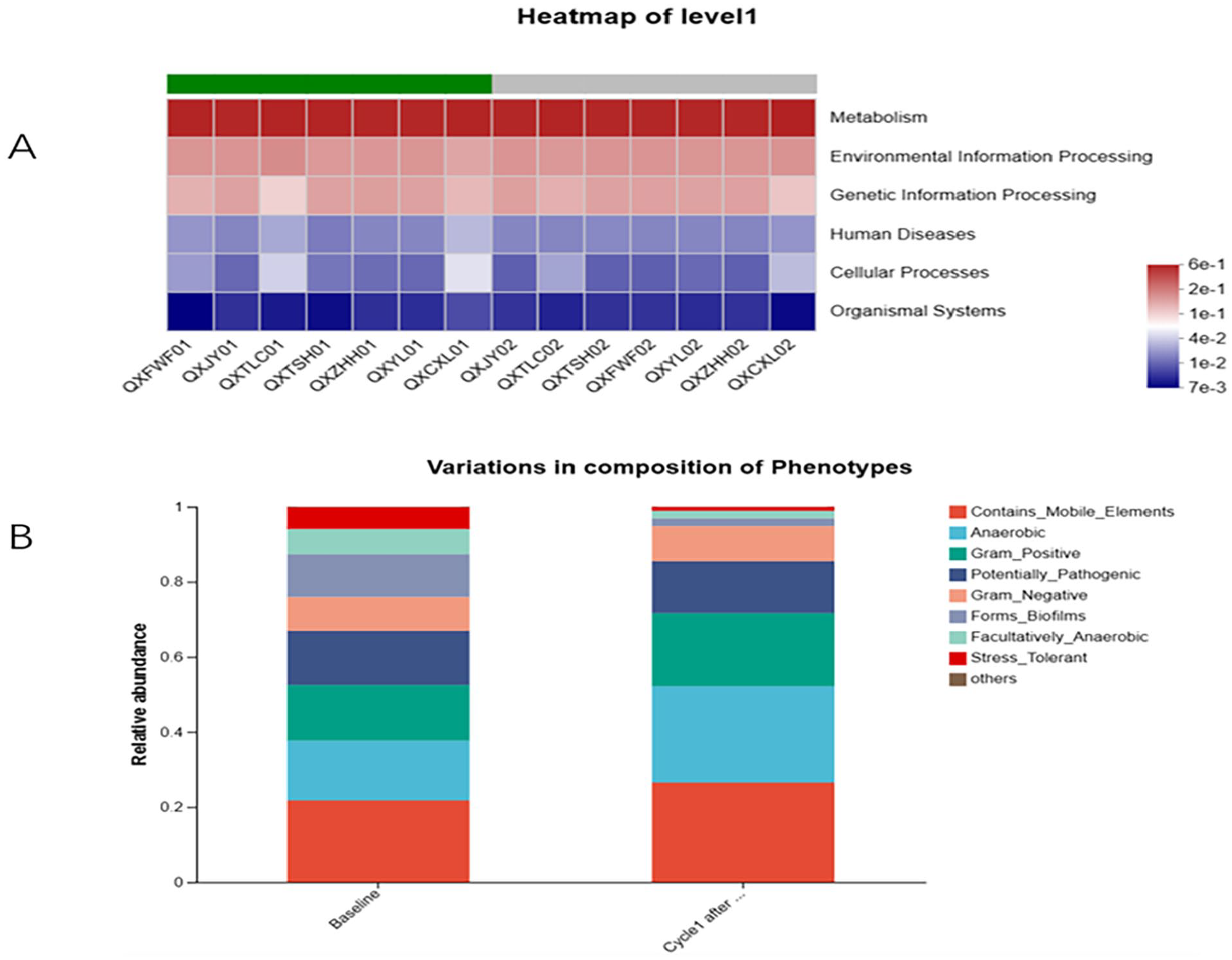
Tax4Fun analysis and BugBase prediction before and after intervention. (A) Six metabolism pathways enriched before and after intervention using Tax4Fun. The labels with 01 indicates pre-intervention samples and with 02 indicates post-intervention samples. (B) Via the BugBase potential prediction for phenotypic functions of bacteria before and after intervention, nine potential microbial phenotypes are included.
The effect of intervention on gut microbiome phenotype is based on BugBase predictive analysis, including Contains_Mobile_Elements, Anaerobic, Facultatively anaerobic, Gram_Positive, Potentially_Pathogenic, Gram_Negative, Forms_Biofilms, and Stress_Tolerant. According to the potential prediction of bacterial phenotype function before and after intervention by BugBase, the above 8 potential microbial phenotypes were found. Conventional treatment plus QX increased Anaerobic, Gram_Positive Gram_Negative and Contains_Mobile_Elements, while reducing Potentially_Patho-genic, Stress_Tolerant, facultatively anaerobic, and Forms_Biofilms (Figure 7B).
Discussion
Our trial met the primary endpoint, demonstrating trend of activity of conventional therapy plus QX, as shown by investigator assessed median progression-free survival (mPFS). mPFS exceeded the prespecified threshold determined based on the study of Fruquintinib monotherapy, which is considered to be one of the third-line standard treatments for mCRC. 19 To the best of our knowledge, this trial is the first phase 2 study to preliminarily and prospectively explore the effects of conventional therapy plus QX on gut microbiome in the third or above line setting in mCRC.
From the aspect of gut microbiome, the Shannon/ACE curves based on the Shannon/ACE indexes showed that the sequencing data and depth and sample size were sufficient for subsequent analysis. On the one hand, Alpha analysis on OTU level showed that Shannon/ACE indexes declined after intervention, which means a lower species richness with the gut microbiome of patients treated with intervention. Previous studies showed that the gut microbiome related to CRC was different compared with the gut microbiome of healthy individuals as controls, with a higher species richness and diversity.22,23After intervention, the lower species richness in patients with stable disease may be one of the reasons for the effectiveness of the conventional therapy plus QX. On the other hand, through 16 seconds rRNA sequencing analysis of gut microbiome, the microbiome of patients before and after intervention were different. At the phylum level, Firmicutes was the most predominant phylum and increased, while Bacteroidota, Proteobacteria and Actinobacteriota (P < .05) were declined after intervention. At the genus level, Bacteroides, Subdoligranulum, Blautia, Faecalibacterium were the most predominant genera, in addition to the decrease in the number of Bacteroides, the latter 3 were found to increase, although not significantly. Furthermore, Bifidobacterium, Alistipes, Butyricimonas and Flavo-nifractor were found to be significantly lower proportions, Lachnospiraceae was significantly increased proportion after intervention, respectively. Eckburg et al 24 found that Firmicutes, Bacteroidetes, Proteobacteria and Actinobacteria constituted the most abundant phyla in the human intestinal microbiota, Firmicutes and Bacteroidetes being predominant in adults. Compared with normal samples, the presence of Proteobacteria and Fusobacteria increased considerably in CRC samples; conversely, the abundance of Firmicutes and Spirochetes decreased markedly. 25 In this study, Actinobacteriota decreased significantly, which was inconsistent with the previous research conclusions of our team. The first possible reason is that the intervention in this study is conventional therapy plus QX, which might be affected by some drugs such as capecitabine. Second, all the patients in this study are treated with third-line or above, which are different from the first-line and second-line patients in the previous study. In addition, this study is a small sample size research so there is a possibility of bias. Blautia, Faecalibacterium, Lachnospiraceae (P < .05) are respectively considered to be probiotic species, anti-inflammatory commensal bacterium and a protective role against CRC, which the abundance of all 3 increased after intervention.26 -28 In addition, Flavonifractor plautii is an important bacterial species in mCRC, which the abundance decreased after intervention. 29
The abundance of Subdoligranulum which may drive systemic autoantibody generation and immune activation were increased in this study. 30 This phenomenon may be one of the reasons for the progression of target lesions, but whether it is the target of treatment or the mechanism of drug resistance remains to be further studied. 30 Similarly, Bifidobacterium and Butyricimonas, which are considered probiotics, decreased in abundance after intervention in this study. This phenomenon is partly consistent with some of the previous results of our team, which may be due to the combination of targeted therapy and chemotherapy in this study, and the decrease in abundance of Bifidobacterium may also explain the diarrhea symptoms of TRAEs.31 -33 In addition, the phenotype and function analysis of gut microbiome showed that potentially-pathogenic and Human Diseases were reduced after intervention, which may support the efficacy of conventional therapy plus QX. In summary, decrease of gut bacteria with potential roles in carcinogenesis of colorectal cancer and increase in the abundance of gut anticancer bacteria such as Lachnospiraceae may partly explain how conventional therapy combined with QX can influence carcinogenesis and tumor progression in colon cancer. This study is a phase 2 clinical study with a small sample size, and the sample size can be increased in the future to obtain a more accurate clinical mechanism of QX.
Study Limitations
The study did not include a concurrent randomized control group, which could have provided a better comparison of the effects of treatment regimens with QX on gut bacteria. In future studies, including a control group and clearly identify what specific drugs are used in conventional therapies will be important to accurately assess the impact of treatment regimens with QX on gut bacteria in mCRC patients with the third or above line setting.
Additionally, this study aimed to investigate the impact of QX along with conventional therapies—including other Chinese herbal medicine, chemotherapy, targeted therapy, and supportive care—on the gut bacteria of mCRC patients in the third or above line setting. It is important to acknowledge that there may be some confounding factors in this study that could have influenced our judgment of the specific relationships between gut bacteria, mCRC patients in the third or above line setting and conventional therapy combined with QX. Therefore, it is imperative to establish more stringent research protocols in the future in order to accurately examine the interaction between treatment regimens involving QX and the gut bacteria of mCRC patients in the third or above line setting.
Conclusion
The results of the first phase of our study provide valuable insights into the potential effects of conventional therapy combined with QX on gut bacteria in mCRC patients with the third or above line setting. The expansion of our study into the second stage will allow for a more comprehensive analysis of the data and a more robust assessment of the efficacy of treatment regimens with QX in this patient population.
Footnotes
Acknowledgements
We thank all patients and their families who supported and participated our study.We also thank Dr Lingyun Sun from China Academy of Chinese Medical Sciences for her guidance of the study, as well as technical support provided by Majorbio Bio-Pharm Technology Co. Ltd.
Contributors
Zhien Gu: manuscript drafting, patient recruitment, observation and statistical analysis. Lei Wang: patient recruitment, observation, statistical analysis and study design. Jiawei Zhai: patient recruitment and observation. Tong Zhang: study design and administration support. Yufei Yang: study design and administration support.
Data Accessibility Statement
Some data generated or used during the study are available from the corresponding author by request.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was funded by grants from National Natural Science Foundation of China (General program 2021, No. 82174461) and Hospital capability enhancement project of Xiyuan Hospital, CACMS. (NO. XYZX0201-22).
Ethics Approval and Consent to Participate
The study had been approved by the Ethics Committee of Xiyuan Hospital (2021XLA080-2) and all patients gave their written informed consent before their enrollment to the study.
Consent for Publication
Not applicable.
