Abstract
Background
Diabetic foot infections (DFIs) are the most complex and challenging pathological state, characterized by high complexity of management, morbidity, and mortality. Empirical antimicrobial administration is needed before the identification of the causative organisms. Microbiological profile of the diabetic foot patients is the basis of the antibiotic choice for clinicians.
Methods
76 patients visiting the Diabetic Foot Center of our Hospital for having ulcers with Wagner 3 to 5 were retrospectively surveyed for the bacterium infected. Antimicrobial susceptibility testing (AST) was implemented and the results were interpreted on the basis of the Clinical and Laboratory Standards Institute (CLSI) guidelines.
Results
A total of 92 isolates were obtained, of which 47 strains of gram-negative bacilli (GNB) accounted for 51.1%, 41 strains of gram-positive cocci (GPC) accounted for 44.6%, and 4 strains of fungi accounted for 4.3%. Species of Enterobacteriaceae and Staphylococcus are the two predominant strains. The top 4 most frequently isolated strains were Staphylococcus aureus (22.8%), enterococcus faecalis (9.8%), Pseudomonas aeruginosa (9.8%), Escherichia coli (5.4%). The rate of resistance to methicillin was 33.3% for S. aureus. The rate of extended-spectrum β-lactamases (ESBLs) production among E. coli was 60%. 50% of Klebsiella pneumoniae were resistant to carbapenems. P. aeruginosa showed high sensitivity to commonly used antibiotics. There was one isolate of Acinetobacter spp. resistant to most antimicrobials except tigecycline and aminoglycosides.
Conclusion
Enterobacteriaceae and Staphylococcus are the two predominant species. The resistance to antibiotics of S. aureus and P. aeruginosa showed optimistic results. However, the high prevalence of CRKP imposed a challenging issue for clinical physicians.
Keywords
Introduction
Diabetic foot (DF), often complicated with neuropathy, peripheral arterial diseases, foot deformities, is the main cause that leads to the disability and death of the patients with diabetes. The annual death rate of DF patients reached to 11%, and the amputated ones 22%. 1 Diabetic foot infections (DFIs) are the most frequent complications that require hospitalization and call for huge medical expenditure. Almost 40 to 70% ulcers have been infected when the patients come to the doctors.2,3 Data showed that 25% Mild superficial ulcers deteriorated and progressed to severe deep infections that often resulted in amputations. 4
When treating patients with acute DFIs, the administration of empirical antibiotics is needed for the delayed identification of the causative pathogens. Therefore, it is crucial to obtain the bacterial profile of DFIs patients with different severity of infection in various areas. In this study, we aimed to determine the microbiological and antibiotic resistance profiles of the hospitalized patients in the diabetic foot interdisciplinary center, in order to provide basis for our empirical antibiotic selection.
Materials and Methods
Bacterial Strains
A total of 76 patients with DFIs were included in this retrospective study. All patients with DFIs were admitted to our Hospital between July 1, 2016 to June 30, 2020. The diagnosis of DFIs was based on the presence of local or systemic signs and symptoms of inflammation according to the IDSA/IWGDF guidelines. All selected patients had ulcers with Wagner's grade 3 to 5. The specimens for culture from patients with DFIs were obtained by tissue biopsy. Culture specimens were obtained at the time of admission. The area surrounding the wound was cleaned with povidone iodine solution. After the surface of the wound had been washed vigorously by saline, there was debridement of superficial exudates. Specimens were then obtained by scrapping the base of the ulcer or the deep portion of the wound edge with a sterile curette after cleaning the base of ulcer with a sterile swab stick. None of the specimens was obtained as superficial swabs. All specimens were transported by sterile containers and taken to the microbiology laboratory in 30 min. Microbiological studies were carried out in the “Infectious Diseases and Clinical Microbiology Laboratory” within the hospital.
Antimicrobial Susceptibility Testing
Antimicrobial susceptibility testing (AST) was carried out at the microbiology laboratory department. The results were interpreted on the basis of the guidelines published by the Clinical and Laboratory Standards Institute (CLSI). The anaerobic culture was not performed due to lack of standard procedures for handling anaerobic samples.
Results
Distribution of Clinical Isolates
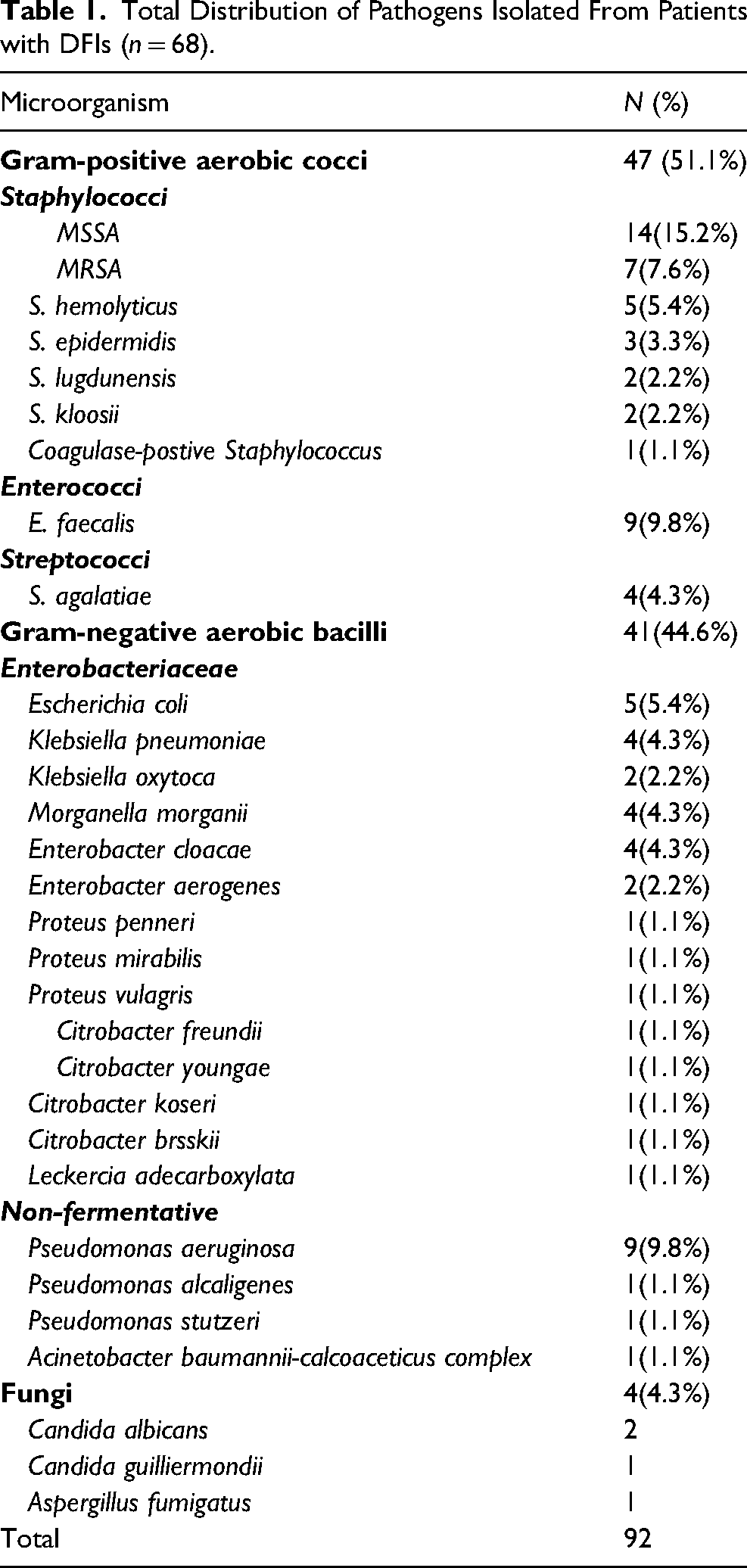
A total of 92 isolates were collected, of which 41 strains of gram-negative bacilli (GNB) accounted for 44.6%, 47 strains of gram-positive cocci (GPC) accounted for 51.1%, and 4 strains of fungi accounted for 4.3% (Table 1). The top 4 most frequently isolated strains were Staphylococcus aureus (22.8%), enterococcus faecalis (9.8%), Pseudomonas aeruginosa (9.8%) and Escherichia coli (5.4%). For GPCs, the top 4 most frequent isolates were staphylococcus aureus (44.7%), enterococcus faecalis (19.1%), staphylococcus hemolyticus (10.6%) and streptococcus agalactiae (8.5%). Among GNBs, Enterobacteriaceae ssp. accounted for 70.7% and non-fermentative bacteria accounted for 29.3%. The top 5 most common isolates were Pseudomonas aeruginosa (22.0%), Escherichia coli (12.2%), Klebsiella pneumoniae (9.8%), Morganella morganii spp morganii (9.8%) and Enterobacter cloacae (9.8%).
Total Distribution of Pathogens Isolated From Patients with DFIs (n = 68).
Bacterial Resistance in Gram-Positive Cocci (Table 2)
Staphylococcus aureus
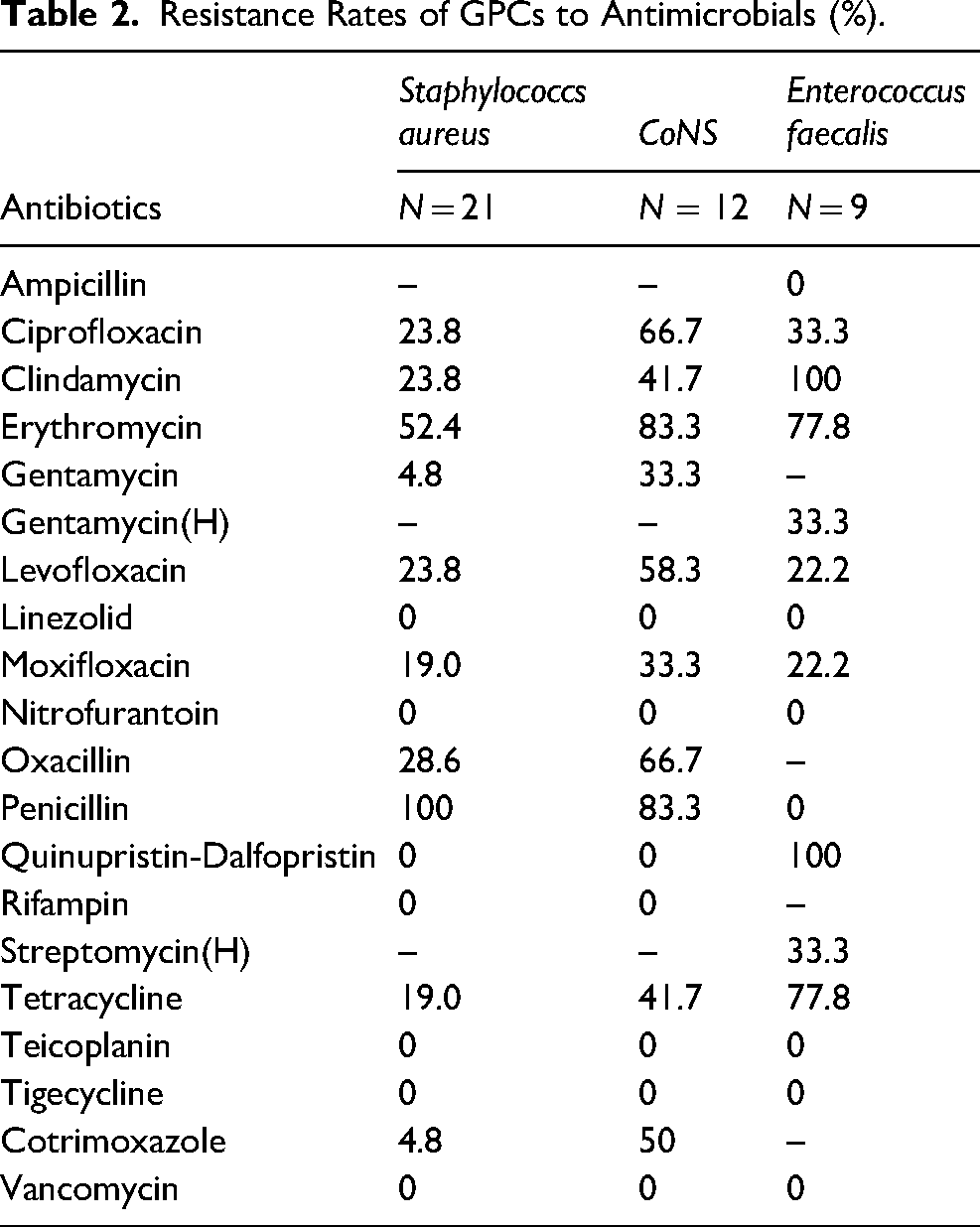
Of all strains, Methicillin-resistant staphylococcus aureus (MRSA) reached 7.6%. The resistance rates of MRSA to Quinolones, clindamycin, erythromycin and tetracycline are obviously higher than that of Methicillin-sensitive staphylococcus aureus (MSSA). No staphylococcus species were found resistant to linezolid, teicoplanin, vancomycin and tigecycline. High sensitivity to Gentamycin, rifampin, trimethoprim-sulfamethoxazole, nitrofurantoin, Quinupristin-Dalfopristin was determined for S. aureus.
Resistance Rates of GPCs to Antimicrobials (%).
Enterococcus faecalis
The resistance rates of E. faecalis to clindamycin, erythromycin and tetracycline were relatively higher than other antibiotics. All strains were showed susceptible to penicillin, vancomycin, linezolid and tigecycline.
Coagulase-Negative Staphylococci
Totally, 9 strains of methicillin-resistant coagulase-negative staphylococcus (MR-CoNS) including 7 isolates of S. haemolyticus and 2 isolate of S. epidermidis were identified from the culture, accounting for 66.7% in CoNS. The resistance rates of CoNS to commonly used antimicrobials are obviously higher than S. aureus.
Bacterial Resistance in Gram-Negative bacilli (Table 3)
Escherichia coli
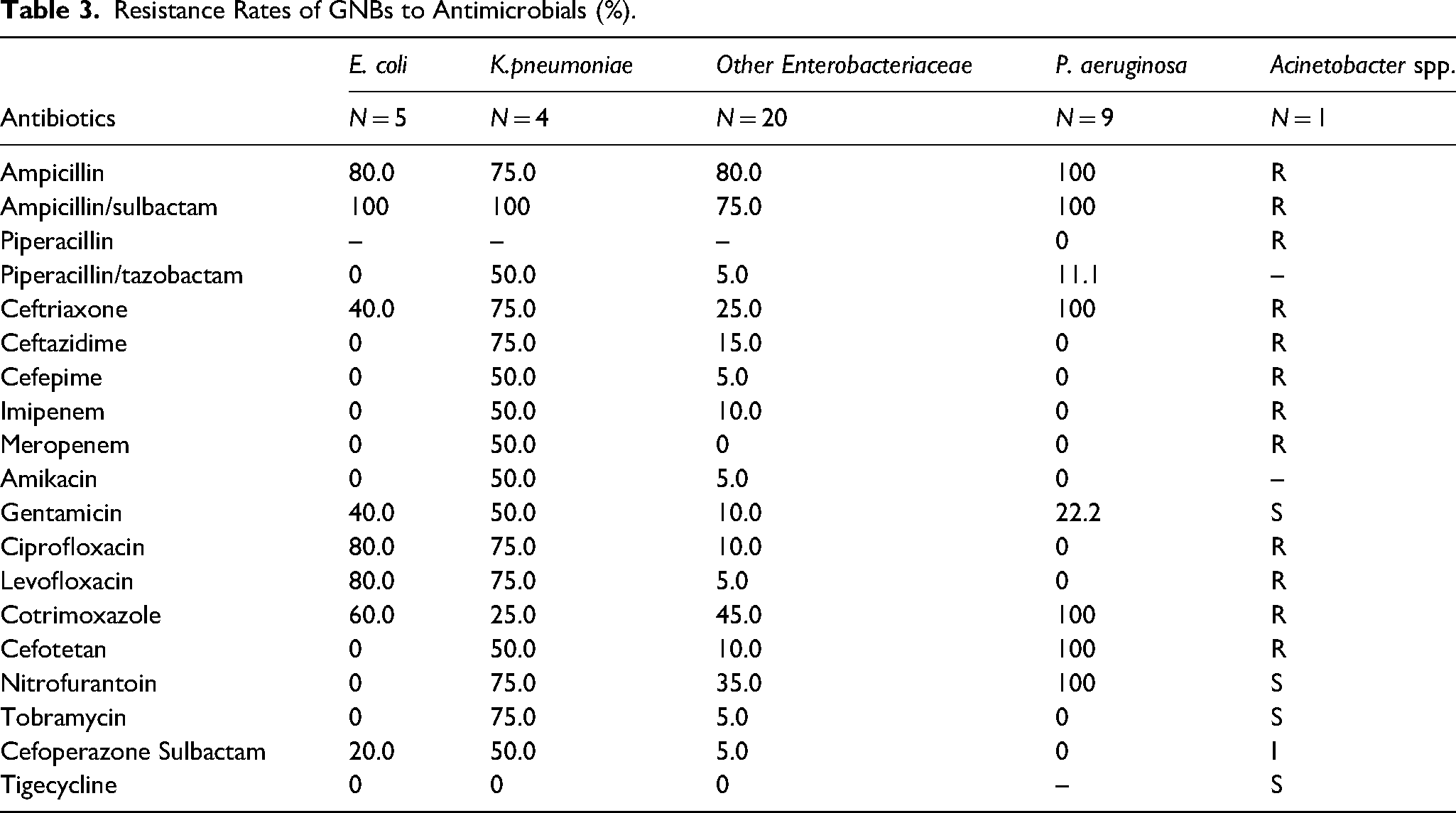
For 5 strains of E. coli, 3 isolates produced extended-spectrum β-lactamases (ESBLs). The resistance rates of E. coli to Ampicillin-sulbactam, ciprofloxacin, levofloxacin were higher than 80%. The resistance rates to ceftriaxone and gentamicin were both 40.0%. Total isolates were susceptible to amikacin, ceftazidime, piperacillin-tazobactam and carbapenems.
Resistance Rates of GNBs to Antimicrobials (%).
Klebsiella pneumoniae
In general, the resistance rates of K. pneumoniae to most antimicrobials were quite high. 50% of K. pneumoniae were resistant to carbapenems. These carbapenem-resistant K. pneumoniae (CRKP) isolates were also resistant to other β-lactam/β-lactamase inhibitor. The resistance rates to amikacin, gentamicin and cefotetan were 50%. The tigecycline was the only agent that CRKP was sensitive to.
Pseudomonas aeruginosa
The resistance rates of pseudomonas aeruginosa to ampicillin, ampicillin/sulbactam, cefotetan, nitrofurantoin, ceftriaxone, cotrimoxazole were 100%. All of the strains were non-resistant to cefoperazone/sulbactam, piperacillin, ceftazidime, cefepime, imipenem, meropenem, amikacin, tobramycin, ciprofloxacin, levofloxacin. The piperacillin/tazobactam and gentamicin resistance rates were 11.1% and 22.2%, respectively.
Acinetobacter species
There was one strain of acinetobacter species obtained, named acinetobacter baumannii-calcoaceticus complex, which was multi-drug resistant. It was resistant to all kinds of quinolones and β-lactam/β-lactamase inhibitors, including carbapenems. It was sensitive to gentamycin, tobramycin and tigecycline intermediate to cefoperazone/sulbactam.
Discussion
We obtained specimens partly by swabbing due to the retrospective nature of the study. However, all wound swabs were collected after effective debridement and wound cleaning with saline.
In our study, species of Enterobacteriaceae were the predominant isolates, and the staphylococcus spp. ranked second. Previous investigations showed that GPCs were the most common strains, especially in mild infections. 5 Our observations consistent with that.
For GPCs, staphylococcus aureus was the top 1 isolate. MRSA accounted for 7.6% of all the isolates. This is close to the result of a retrospective multi-center survey in the Beijing area. 6 A meta-analysis showed that the prevalence of MRSA for infections of 10 994 diabetic foot patients, from 41 data sets was 16.78%, which is much higher than our result. 7 According to the China Antimicrobial Surveillance Network (CHINET), the prevalence of MRSA has markedly decreased from 69.0% in 2005 to 35.3% in 2017. 8 The implementation of infection control measures such as hand hygiene and antimicrobial stewardship measures in clinical practice in China may contributed to the decreasing trends of MRSA prevalence. Furthermore, the relative low isolation rate of MRSA might be attributed to the absence of Wagner 1 to 2 ulcers which tend to be infected by GPCs.
Furthermore, the prevalence rate of MR-CoNS was considerably high in CoNS. Seng et al. 9 reported that MR-CoNS isolated from community and hospital environments accounted for 20.5% and 70.1%, respectively. S. haemolyticus was the predominant specie. All MR-CoNS strains carried mecA gene. The treatment of MR-CoNS is challenging for clinicians due to the high productive rate of biofilms which protect the pathogens from antimicrobials and host immune system.
Among Enterobacteriaceae, ESBLs were detected in the E. coli and Klebsiella species. Though the production rate of ESBLs was low in K. pneumoniae, its resistance rate to carbapenems were relatively high which means these CRKP strains were resistant to most commonly used antibiotics. It has raised a serious concern that the prevalence of CRKP showed a markedly increase in recent 10 years in CHINET. The resistance rate to carbapenems climbed from 3% in 2005 to 20.9% in 2017. 8 A nationwide survey in China covering a total of 1105 carbapenem-resistant Enterobacteriaceae (CRE) strains (including 668 CRKP strains) collected from 25 provinces confirmed that production of carbapenemase was the major mechanism for carbapenem resistance. It is noted that blakpc-2 was the most common carbapenemase gene harbored in by CRKP. 10
P. aeruginosa was the main isolate in the non-fermentative bacteria, which presented a relatively low resistance profile. Data from CHINET indicated that resistance rates of P. aeruginosa to amikacin, ceftazidime, ciprofloxacin, piperacillin/tazobactam, imipenem showed decreasing trends from 2005 to 2017. 8 Carbapenem-resistant P. aeruginosa was not found in our samples.
There was only one isolate of Acinetobacter species obtained, which was resistant to a majority of antimicrobials. Generally, the resistance rates to commonly used antimicrobials exhibited increasing trends except for amikacin. Our Acinetobacter sample were sensitive to gentamycin and tobramycin, both aminoglycosides. However, aminoglycosides are bacterial inhibitors, the capacity of which to kill microorganisms is not strong enough. There is another antimicrobial needed to enhance bacterial effect. Based on the AST result, tigecycline was the last choice.
Conclusion
This study presented a comprehensive microbiological profile of infected diabetic foot ulcers in hospitalized patients. Enterobacteriaceae and Staphylococcus are the two predominant species. The resistance to antibiotics of S. aurue and P. aeruginosa showed optimistic results. However, the high prevalence of CRKP imposed a challenging issue for clinical physicians.
Footnotes
Author Contributions
Yaping Xie: Funding acquisition, Writing-original draft. Ying Xu: Data curation, Investigation. Kuang Chen: Project administration, Writing-review & editing. Can Chen: Data curation, Software. Jiao Huang: Formal analysis, Methodology. Qiong Chen: Formal analysis, Methodology. Pengfei Shi: Conceptualization, Supervision, Writing-review & editing.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship and/or publication of this article.
Ethical Approval
Not applicable, because this article does not contain any studies with human or animal subjects.
