Abstract
Apolipoprotein E (ApoE) gene polymorphisms are thought to be the most important genetic risk factor in the pathogenesis of late onset and sporadic Alzheimer’s disease (AD). Moreover, interleukin-1α (IL-1α) is found to be associated with the pathogenesis of AD. In this research, ∊2, ∊3, and ∊4 polymorphisms of ApoE gene and C889T polymorphism of IL-1α gene were genotyped in patients with AD and controls. Genotyping was performed by real-time polymerase chain reaction. ∊3/∊3 and ∊3/∊4 genotype frequencies were significantly higher in control and case groups, respectively. While ∊3 allele frequencies were significantly higher in the control group, ∊2 and ∊4 allele frequencies were significantly higher among the cases with AD. No difference was found between the groups according to C889T polymorphism of IL-1α. In conclusion, we demonstrated that there was a strong association between ApoE ∊4 allele and AD, while there was no relation with IL-1α C889T polymorphisms for this study.
Introduction
Alzheimer's disease (AD) is a multifactorial, progressive neurodegenerative condition responsible for the most common form of dementia. 1 The concept of preclinical AD postulates that AD lesions accumulate in the brain for years prior to the appearance of cognitive deficits or symptoms of dementia. 2 Late-onset Alzheimer's disease (LOAD), a type of AD and the most common form of dementia, is defined by the onset of symptoms after 60 years and has annual incidence rates increasing from 1% at 65 to 70 years to 6% to 8% at 85 years and older. 3 Late-onset Alzheimer's disease is multifactorial, that is, both genetic and nongenetic risk factors contributing etiologically. 4 Early-onset dementia conventionally includes patients with disease onset before 65 years of age. 5
The strongest known risk factor influencing the incidence of sporadic AD is the genotype for apolipoprotein E (ApoE).Apolipoprotein E, a 299-amino-acid protein, mediates the binding of lipid particles to lipoprotein receptors. 6 Apolipoprotein E has 3 major isoforms: ApoE2 (Cys 112, Cys 158), ApoE3 (Cys 112, Arg 158), and ApoE4 (Arg 112, Arg 158), which are the products of 3 ApoE allele (∊2, ∊3, and ∊4) at a single gene locus on chromosome 19q13.2. 7 Apolipoprotein E alleles ∊2, ∊3, and ∊4 give rise to 6 different phenotypes (∊2/2, ∊2/3, ∊2/4, ∊3/3, ∊3/4, and ∊4/4). 8 Apolipoprotein E ∊3 is the most common isoform (77%-78%) in the general population, while ApoE ∊2 is found in 7% to 8% and ApoE ∊4 in 14% to 16% of individuals. 9 A meta-analysis that included data from several ethnic groups showed that the ∊4 allele of the ApoE is a major susceptibility factor in the development of AD. 10 However, ∊4 allele has been reported to increase an individual’s risk of developing AD and decrease the age of disease onset, in proportion to the number of ∊4 alleles present. 10,11 Indeed, it is reported that the ∊4 allele of ApoE is the only repeatedly proven genetic risk factor for LOAD. 4 Inheritance of the ∊4 allele of ApoE increases the risk of AD, whereas inheritance of the ∊2 allele decreases the risk. 12 Numerous hypotheses have been proposed to explain ApoE affects deposition of amyloid plaques and formation of neurofibrillary tangles. 13 Still, it remains unclear how ApoE ∊4 accelerates and ApoE ∊2 retards AD pathology to influence cognitive decline. 14,15
In the brains of patients with AD, the microglial cells are activated and produce a large number of inflammatory mediators including cytokines such as interleukins (IL)-1A, IL-1B, IL-6, and tumor necrosis factor-α. 16 Interleukin 1 is a potent proinflammatory cytokine that is markedly overexpressed in the brains of patients with AD. 17 Overexpression of IL-1 by activated microglia across brain regions correlates with the pattern and distribution of amyloid plaques in AD. 18 A further role for IL-1 overexpression in neuronal dysfunction may arise from IL-1 effects on cholinergic systems, as the cholinergic decline characteristic of AD is thought to contribute to memory loss. 19 The IL-1 gene polymorphisms, especially IL-1α C889T polymorphism, have been suggested to be associated with AD risk and onset age. 20 Genetic variation in the IL-1α gene was reported to increase the risk of early-onset AD 21 or LOAD. 22
To determine whether ApoE and IL-1α polymorphisms are genetic risk factors for developing AD in studied group, we analyzed ∊2, ∊3, and ∊4 polymorphisms of ApoE gene and C889T polymorphism of IL-1α gene in patients with AD and healthy controls.
Methods
The study sample comprised 95 Turkish patients (mean age at the time of study was 73.5 years) who met NINCDS-ADRDA criteria (National Institute of Neurological and Communicative Diseases and Stroke/ Alzheimer's Disease and Related Disorders Association) for probable Alzheimer's Disease. Patients with AD were admitted to Afyon Kocatepe University, Faculty of Medicine, Department of Neurology and treated by Alzheimer's Association at City Nursing Home. Control participants were 60 unrelated individuals (mean age at the time of study was 74 years) randomly selected and these participants had no dementia history. All patients and control group underwent a comprehensive evaluation by a research physician and ancillary staff, including cognitive assessment, medical history, and pedigree analysis. All participants (or their responsible next of kin) provided written informed consent and were studied under a protocol approved by the Afyon Kocatepe University Medical Ethic Committee.
Sample Collection
About 2 mL aliquots of peripheral blood samples were collected from the participants and stored in EDTA-coated vacutainers. Genomic DNA was extracted from a 200-µL peripheral blood sample by using a High Pure Template Preparation (Roche Diagnostics, Indianapolis, Indiana) kit. Then, DNA amount and DNA purity were quantified for each DNA sample by Nanodrop ND-1000 spectrophotometer V3.7. DNA samples were stored at −20°C until use. Each genomic DNA sample was analyzed for ApoE ∊2, ∊3, and ∊4 polymorphisms and IL-1α C889T polymorphism. Apolipoprotein E and IL-1α genotyping were performed by real-time polymerase chain reaction (PCR) on a LightCycler Instrument.
APOE Genotyping
A 265-bp fragment of ApoE gene including codons 112 and 158 of polymorphic regions was amplified using the Roche LightCycler 1.5 -ApoE mutation determination kit (Roche Diagnostics, Mannheim, Germany). Amplicon was determined with fluorescence using specific probes that hybridize at the annealing phase of PCR cycle. After preparation of the master mixture (the mutation detection mixture 4 µL, reaction mixture 2 µL, and 12 µL sterile PCR-grade H2O), 18 µL of the reaction mixture and 2 µL of the isolated genomic DNA template or the control template were loaded into LightCycler capillaries. For negative control, sterile PCR-grade H2O was added instead of template. Polymerase chain reaction cycling conditions were 60 seconds at 95°C for DNA denaturation, followed by 45 cycles of PCR (0 second denaturation at 95°C, 10 seconds annealing at 60°C, and 10 seconds extension at 72°C). After the PCR, a melting curve analysis was performed by heating to 95°C for 30 seconds, followed by cooling to 42°C for 4 minutes to achieve maximum hybridization and then heating slowly at 0.1°C/s to 80°C. After the melting curve analysis, a final cooling was performed at 40°C for 30 seconds. The fluorescence signals recorded in the respective channels were then converted to melting peaks by plotting the negative derivative of the fluorescence with respect to the temperature (−dF/dT vs T). The resulting melting peaks in the different fluorescence channels allowed us to discriminate among the homozygous as well as the heterozygous genotypes. Tm values were obtained for each allele: for ∊2, these were 56°C and 57.5°C; for ∊3, 56°C and 66°C; and for ∊4, 62.5°C and 66°C, respectively.
Interleukin 1α C889T Genotyping
A 136-bp fragment of IL-1α including C889T polymorphic region was amplified using the IL-1α C-889T ToolSet for LightCycler kit (Genes-4U). Amplicon was determined with fluorescence using specific probes that hybridize at the annealing phase of PCR cycle. During PCR, the amplicon was detected using 2 specific hybridization probes, one labeled with fluorescein and the other with LightCycler Red 640 (LCRed640). The presence of a T-allele introduces a destabilizing mismatch that results in a decreased melting temperature. Primers and probes were designed and custom-made by Genes-4U AG, Switzerland. For mutation detection with the LightCycler, a 20 μl reaction mixture was performed. The reaction mixture contained 2.8 μL OligoTool (lyophilized oligos for PCR contains mutation detection and anchor probe, primers), 10.4 μL solvent, 2 μL LightCycler DNA Master HybProbe, 25 mmol/L MgCl2, and 4 µL genomic DNA. Polymerase chain reaction cycling conditions were 30 seconds at 95°C for DNA denaturation, followed by 50 cycles of PCR (5 seconds denaturation at 95°C, 10 seconds annealing at 55°C, 7 seconds extension at 72°C). After the PCR, a melting curve analysis was performed by heating to 95°C for 30 seconds, followed by cooling to 40°C for 60 seconds and gradual heating to 85°C at 0.2°C/s. After the melting curve analysis, a final cooling was performed at 40°C for 30 seconds. To analyze the melting curves, the corresponding melting peaks were calculated by plotting the first negative derivative of the fluorescence with respect to the temperature (−dF/dT vs T). During the melting curve analysis, the probes were dissociated from the target DNA. In case of a T-allele (mutant genotype), the fluorescein-labeled probe has a fully homologous sequence with the target DNA and was dissociated from the target DNA at a melting temperature of 62.1°C (±2.5°C). In case of a C-allele (wild-type genotype), 1 mismatch occurs and the melting temperature was decreased to 57.8°C (±2.5°C).
Statistical Analysis
Statistical analysis was performed using the SPSS 15.0 and MiniTab Programs. In patients and controls, allele frequencies related to ApoE alleles polymorphisms ∊2, ∊3, and ∊4 and IL-1α gene C889T polymorphisms were compared using 2 proportions test. In patient group, the relationship between ApoE gene ∊4 allele and IL-1α gene C889T genotypes was compared using χ 2 test. Comparison of ApoE ∊4 allele frequency between the 2 patient groups with or without a family history of dementia was made using χ 2 test.
Results
Genotype and Allele Frequencies of ∊2, ∊3, and ∊4 Polymorphisms of ApoE
A total of 3 homozygous (ApoE ∊2/∊2, ∊3/∊3, and ∊4/∊4) and 3 heterozygous (ApoE ∊2/∊3, ∊2/∊4, and ∊3/∊4) genotypes occur in humans. In this study, these polymorphisms were evaluated in the patients diagnosed with AD and in the control group without a history of dementia.
The distribution of ApoE genotype frequencies in AD case group was 0% for ∊2/∊2, 7.4% for ∊2/∊3, 2.1% for ∊2/∊4, 63.2% for ∊3/∊3, 21.1% for ∊3/∊4, and 6.3% for ∊4/∊4, and in control group the distribution of genotypes was 0% for ∊2/∊2, 1.7% for ∊2/∊3, 0% for ∊2/∊4, 93.3% for ∊3/∊3, 5.0% for ∊3/∊4, and 0% for ∊4/∊4. Figure 1 shows the distribution of the ApoE genotype frequencies in both control and AD case groups. While ∊3/∊3 genotype frequency was significantly higher in control, ∊3/∊4 and ∊4/∊4 genotype frequencies were significantly higher in AD case group. In our study, ∊2/∊2 homozygote was not detected in both control and the AD case group.
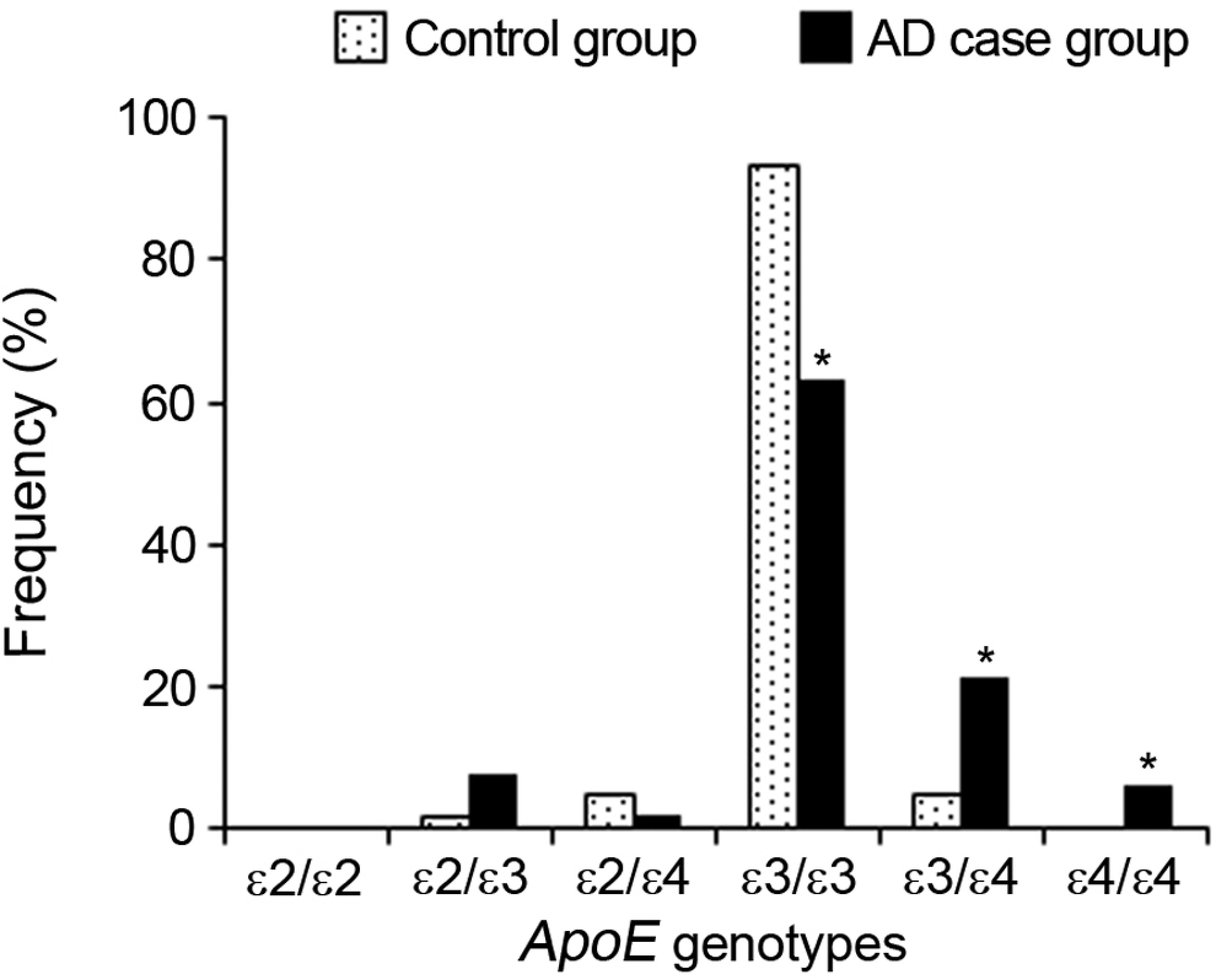
Distribution of the apolipoprotein E genotype frequencies in control and Alzheimer’s disease case groups.
The distribution of alleles in the control group was 0.8% for ∊2, 96.7% for ∊3, and 2.5% for ∊4, and in the AD case group, it was 4.7% for ∊2, 77.4% for ∊3, and 17.8% for ∊4. The frequency of the ∊3 allele in control group (96.7%) was significantly higher than in AD case group (77.4%; P < .05). In addition, the frequencies of the ∊2 (4.7%) and ∊4 (17.8%) alleles were significantly higher in the AD case group than in the controls (0.8% and 2.5%, respectively; P < .05). Figure 2 shows the distribution of the ApoE allele frequencies in both control and AD case groups.
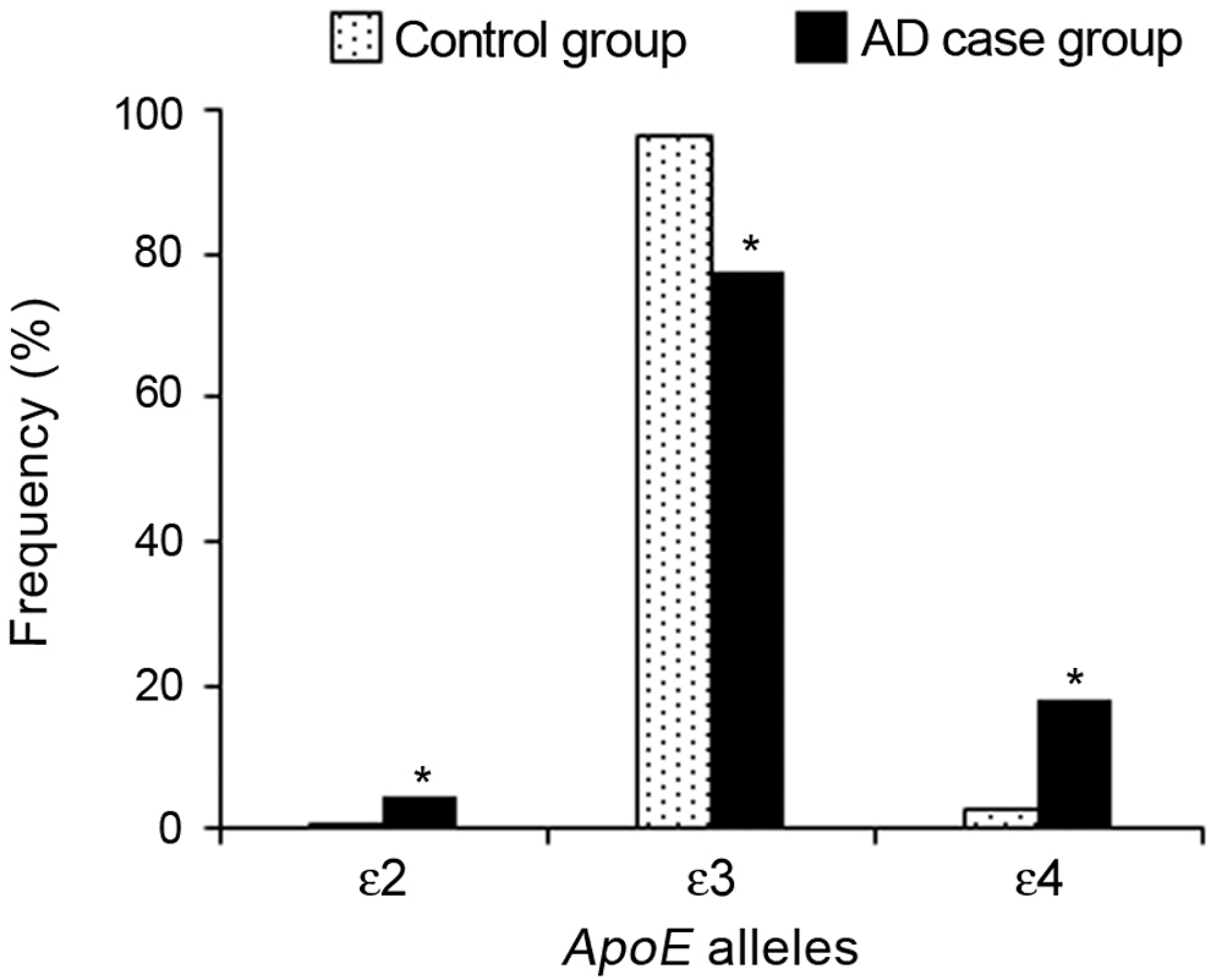
Distribution of the apolipoprotein E allele frequencies in control and Alzheimer’s disease case groups.
There is a significant difference between control and the AD case groups in the terms of ∊4 allele carrier (P < .05). Positive ∊4 allele frequency was higher (29.5%) in the AD case group than in the control group (5.0%). Figure 3 shows the distribution of the positive and negative ∊4 allele frequencies in both control and AD case groups.
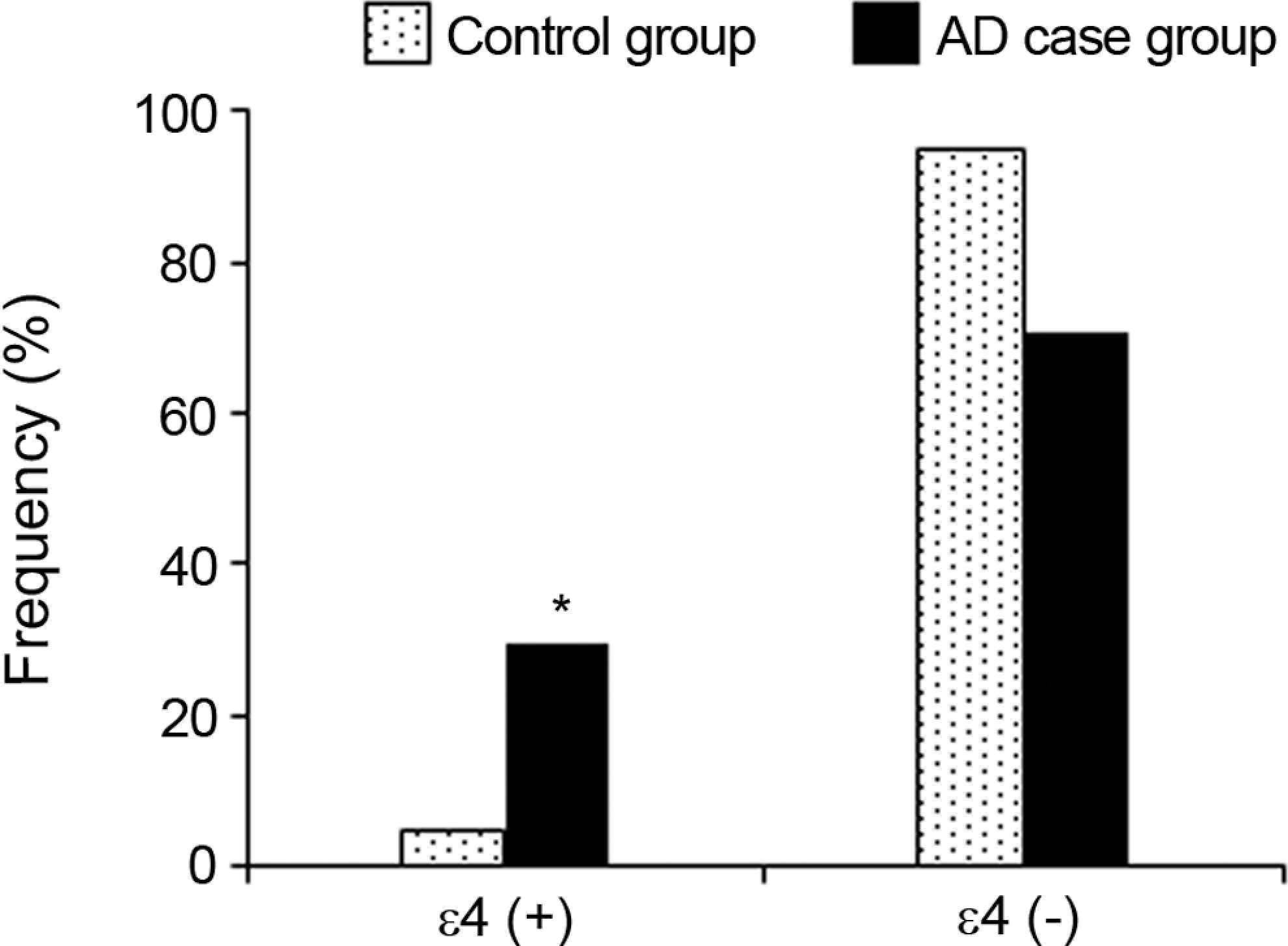
Distribution of the positive and negative ∊4 allele frequencies in control and Alzheimer’s disease case group.
In all, 25.2% of the AD case group has a dementia history in their family. In this group, ∊4 allele frequency was significantly higher than the AD case group, which has no family history of dementia (P < .05).
Genotype and Allele Frequencies of IL-1α Gene
The distribution of genotypes of IL-1α gene in AD case group was 54.7% for CC, 38.9% for CT, and 6.3% for TT; and in control group, the distribution of genotypes was 60% for CC, 37.3% for CT, and 1.7% for TT. The distribution of alleles in the AD case group was 74.2% for C allele and 25.8% for T allele; and in control group, it was 79.2% for C allele and 20.8% for T allele. Figure 4 shows the distribution of the IL-1α C889T genotype, and Table 1 shows the distribution of the IL-1α C889T allele frequencies in both control and AD samples. There were no significant differences in IL-1α genotype or allele frequencies comparing the AD cases and controls (P > .05).
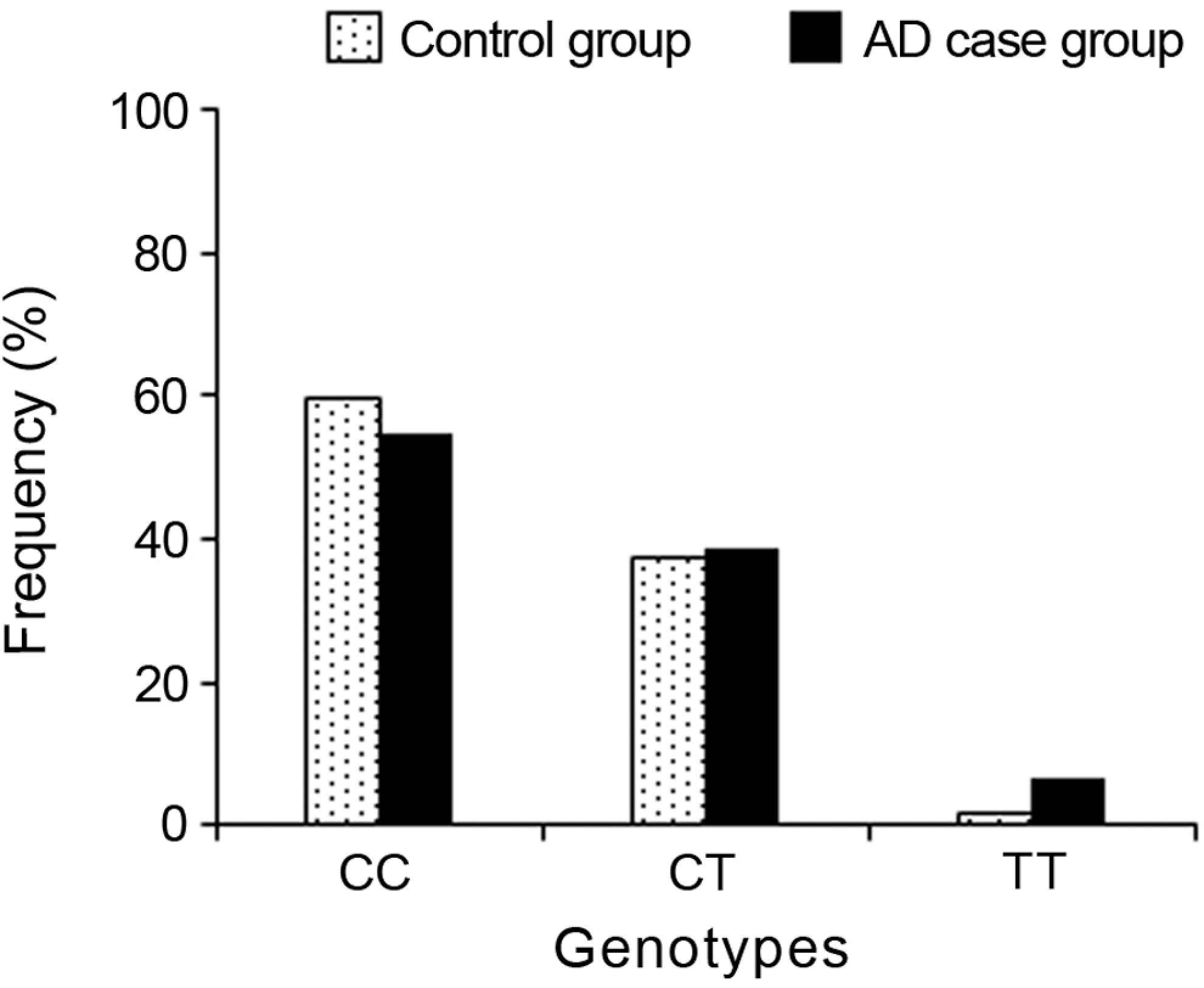
Distribution of the IL-1α genotype frequencies in control and Alzheimer’s disease case groups.
Distribution of IL-1α C889T allele frequencies in control and Alzheimer’s disease samples
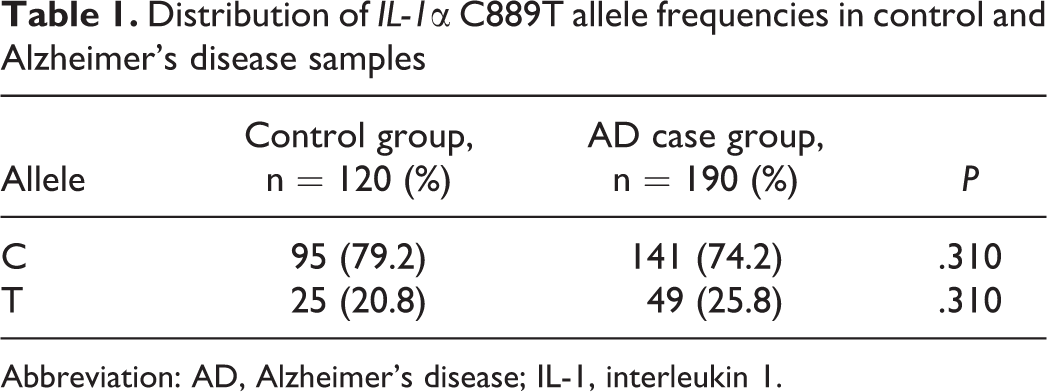

Abbreviation: AD, Alzheimer’s disease; IL-1, interleukin 1.
In addition to all these results, there was no significant difference (P > .05) between ∊4 allele carrier and noncarrier AD case group in the terms of IL-1α C889T genotype frequencies.
Discussion
We investigated the association of AD with ApoE and IL-1α gene polymorphisms. In our current analysis including 95 participants with AD, we found a significant relationship between ApoE ∊4 and AD in the studied group. There was a significant difference between the control and the AD case groups in the terms of ∊4 allele carrier (P < .05). As shown in Figure3, positive ∊4 allele frequency was higher in the AD case group than in the control group. In all, 25.2% of the AD case group has a dementia history in their family. In this group, ∊4 allele frequency was significantly higher than the AD case group, which has no family history of dementia (P < .05). The ApoE ∊4 allele is now known to be a major susceptibility factor associated with approximately 40% to 50% of sporadic and familial AD compared with 30% of the normal population. 23 The ∊4 allele associates with late-onset familial AD in several populations and in late-onset sporadic AD. 24 Beside, epidemiological studies demonstrated that ApoE ∊4 carriers have a higher risk of developing an early onset disease. 25 However, Ohara et al 26 reported that the ApoE ∊4 allele is a risk factor for AD in the Japanese population.
It is reported that inheritance of the ∊4 allele of ApoE increases the risk of AD, whereas inheritance of the ∊2 allele decreases the risk. 11 The ApoE ∊2 allele confers relative protection against AD, lowering the risk and delaying the age of onset of the disease. 27 The Apo E ∊2 allele seems to have a protective effect against AD. Furthermore, it is associated with longevity. 28 Taken together, contrary to this, not only ∊4 allele frequencies were significantly higher in the cases, but also ∊2 allele frequencies were significantly higher incases of our study. These findings are not in the range of the previous studies. We have found the ∊2 allele frequency strikingly higher in the AD case group. The reason for the discrepancy may be due to the regional differences or low ∊2 allele existence in the control group. Saunders and Roses 29 studied 497 healthy blood donors and estimated the ApoE allele frequencies in the general population. They reported that, because the ∊4 and ∊2 alleles are risk factors for atherothrombotic diseases, people bearing these alleles probably die at a younger age on average. Thus, they suggested that the ∊3 allele frequency in the population tends to increase with age, rendering comparisons less informative. Although we studied polymorphisms for AD, similarly ∊3 allele frequencies were significantly higher in control group. While ∊3/∊3 genotype frequency was significantly higher in control, ∊3/∊4 and ∊4/∊4 genotype frequencies were significantly higher in AD case group. Consistent with our findings, as the number of ApoE ∊4 alleles increases from 0 to 2, the risk of developing LOAD increases from 20% to 90%, and the mean age of onset decreases from 84 to 68 years. 11 A significantly increased risk of AD was observed in carriers of the ApoE ∊4 allele. 30 Although APOE e4 is a well-studied genetic risk factor for developing AD, in some regions most patients do not carry this genotype, and additional research is needed to understand both other genetic and environmental risk factors. 31 In our study, ∊2/∊2 homozygote was not detected in both control and the AD case groups. Similarly, in other studies, ∊2/∊2 homozygote was not detected in patients with AD. 32,33
Interleukin 1 gene polymorphisms, especially IL-1α C889T polymorphism, have been suggested to be associated with AD risk and onset age. 20 Qin et al 34 reported that the IL-1α is a candidate gene for AD susceptibility. The IL-1α −889C/T889C/T polymorphism may be a risk factor for AD in Caucasians. Genetic variation in the IL-1α gene was reported to increase the risk of early-onset AD 21 or LOAD. 22 On contrary to this, there were no significant differences (P > .05) between AD cases and controls in terms of IL-1α allele and genotype frequencies in our study. These findings are consistent with many studies in the literature. 20,35 –38 Also, it is suggested that the presence of the heterozygous IL-1α T allele cannot be used for distinguishing AD or vascular dementia from controls. 39 There was no significant difference (P > .05) between ∊4 allele carrier and noncarrier AD case groups in the terms of IL-1α C889T genotype frequencies. Similarly, Kuo et al 39 reported that the presence of the ApoE ∊4 allele did not influence the IL-1α genotype or allele distribution.
We have evaluated the existence of probable associations relating ApoE and IL-1α polymorphisms to AD risk in a studied group, which have also been observed in other ethnic groups. Our genetic association and functional data indicate that the existence of ApoE ∊4 allele is associated with a modest risk of AD in our study. The presence of ApoE ∊4 and homozygous state was associated especially with AD, confirming the results of other studies done before. The ∊4 allele frequency was significantly higher in AD case group that has a dementia history in their family than the AD case group that has no family history of dementia. In this regard, if there is a dementia history in the family, it is important to examine individuals in terms of carriage of the ∊4 allele.
When study results were evaluated together with the literature data, it was observed that many different results were presented about ApoE and IL-1α gene polymorphisms related to AD. Some of the possible reasons for these different results may be different ethnic groups, different numbers of patients, or regional differences. As a conclusion, these polymorphisms must be studied with more number of cases, with another probable candidate gene and different ethnic groups at the same time.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the financial support for the research, authorship, and/or publication of this article: supported by the Afyon Kocatepe University Scientific Research Projects Commission with a project numbered as 06.TIP.25.
