Abstract
Pituitary adenomas usually develop temozolomide resistance, which could compromise the anticancer effects of temozolomide. Suppression of hypoxia-inducible factor 1α has been shown to sensitize glioblastoma cells to temozolomide treatment according to previous reports. However, whether and how the suppression of hypoxia-inducible factor 1α could sensitize pituitary adenomas to temozolomide treatment are still poorly understood. In the present study, using hypoxia-inducible factor 1α knockdown strategy, we demonstrated for the first time that hypoxia-inducible factor 1α knockdown could inhibit temozolomide-induced autophagy in rat pituitary adenoma GH3 cells and thus increase antitumor efficacy of temozolomide. Furthermore, we found hypoxia-inducible factor 1α knockdown could block autophagy process through neutralizing lysosomal pH value but not inhibiting autophagy induction. Finally, we found hypoxia-inducible factor 1α could regulate lysosomal pH value through regulating full length presenilin 1 expression, and exogenous reexpression of presenilin 1could restore lysosome acidic levels. Our data indicated hypoxia-inducible factor 1α knockdown could be a potential approach to improve the efficacy of temozolomide therapy for pituitary adenomas.
Introduction
Pituitary adenomas (PAs) are the second most common intracranial neoplasms and comprise 10% to 15% of all intracranial neoplasms. Although PAs are usually benign neoplasms, there are still approximately 35% of PAs that exhibit invasive or recurrent growth. 1 Pituitary tumors originated from highly specialized cell types expressing the respective pituitary polypeptide hormones and are divided into secreting and nonsecreting adenomas according to the origin of tumor cells. 2 Although surgery, radiotherapy, and pharmaceutical therapy are routinely employed, recurrence of these aggressive adenomas still result in high morbidity and mortality. 3
Temozolomide (TMZ) is an alkylating chemotherapeutic drug mainly used to treat glioblastoma multiforme. 4 However, in 2006, TMZ was proposed as a treatment reagent for pituitary carcinomas and aggressive adenomas. 5 Since then, TMZ has been used as the last-line treatment in aggressive PAs and has been found to be more efficient for ACTH- and GH-secreting adenomas than other types of PAs. 6 Although TMZ has shown a promising antitumor efficiency in certain PAs, only 60% of the published cases showed response to TMZ therapy, and a large number of invasive PAs failed to respond to TMZ treatment or even acquired resistance after treatment. 7 Therefore, development of new reagents to sensitize TMZ has become an urgent issue.
Previous study has implicated a role of hypoxia-inducible factor 1 (HIF-1) in modulating tumor response to therapies. 8 A large body of evidences have indicated that hypoxic cancer cells are more resistant to radiation or cytotoxic drugs, and this drug-resistant phenotype is closely related to the HIF-1 activity. 9 Therefore, inhibiting HIF-1 could sensitize hypoxic cancer cells to radiation or cytotoxic drugs and lead to a more profound antitumor efficacy. 10 However, the mechanisms underlying how HIF-1 involved in tumor drug resistance were poorly understood. Some recent studies have found autophagy could help cancer cells to escape from radiation or cytotoxic drug-induced apoptosis and thus help cancer cells to develop drug resistance. 11 –13 Considering hypoxia also enhances autophagy, 14 in the present study, we wanted to investigate (1) whether inhibiting HIF-1α could sensitize GH3 cells to TMZ treatment; (2) whether HIF-1α could reduce the effect of TMZ through autophagy; and (3) how HIF-1α regulates the autophagy process.
Materials and Methods
Plasmids and Reagents
mKeima Red, eGFP-LAMP1, presenilin 1 (PSEN-1), and TRPML-YPF constructs were purchased from OriGene (Rockville, Maryland). The oligonucleotides against rats’ HIF-1α (5′-AAGCATTTCTCTCATTTCCTCATGG-3′) were synthesized, annealed, and ligated into pSUPER vector (XbaI/EcoRI sites). The PSEN-1 shRNA was commercially purchased from Santa Cruz (Dallas, Texas, US). Temozolomide (100 μmol/L), Rapamycin (RA; 20 μmol/L), 3-methyladenine (3-MA; 10 mmol/L), chloroquine (CQ; 20 μmol/L), Bafilomycin (Baf; 50 nmol/L), and 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) were purchased from Sigma (Allentown, Pennsylvania, US). Antibody used here were mouse monoclonal HIF-1α, and β-actin antibodies were purchased from Sigma. Rabbit polyclonal antibody against p62, LC3, β-actin, and Beclin-1 were purchased from Cell signaling Technology (Danvers, Massachusetts, US). C-terminal (loop) of PSEN1 antibody was bought from Millipore (Billerica, Massachusetts, US). Lysotracker DND99 was purchased from Invitrogen (Carlsbad, California, US).
Cell culture and Transfection
The rat PA cells (GH3; Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, Shanghai, China) were maintained in Rosewell Park Memorial Institute-1640 medium, supplemented with 10% fetal bovine serum (Gibco [Grand Island, New York, US]), 100 U/mL penicillin, and 100 μg/mL streptomycin (Gibco) at 37°C, 5% CO2, and 95% air. For transfections, cells were transiently transfected with overexpression constructs (1 μg) or shRNA constructs (2 μg) with Lipofectamine 2000 (Invitrogen). After 48 hours (overexpression) or 72 hours (shRNA) incubation, cells were collected for protein/messenger RNA (mRNA) isolation or live imaging.
Western Blotting
Protein samples (20 μg total protein per lane) were subjected to 10% or 15% sodium dodecyl sulfate polyacrylamide gel electrophoresis, and the proteins were then electrophoretically transferred to a polyvinylidene fluoride membrane blocked by 5% BSA for 1 hour at room temperature and then incubated with related antibodies overnight at 4°C. Secondary antibody was incubated for 1 hour at room temperature. The blots were developed using the ECL system (Pierce [Rockford, Illinois, US]). Antibodies and dilutions used in this study were as follows: HIF-1α, 1:500; β-actin, 1:5000; LC3, 1:2000; Beclin-1, 1:3000; p62, 1:3000; and HRP-Goat anti mouse secondary antibody and HRP-Goat anti rabbit secondary antibody, 1:5000.
RNA Extraction and Reverse Transcriptional Polymerase Chain Reaction
The total RNA from the cell samples was extracted with Trizol (Invitrogen, Carlsbad, California) and reverse transcribed to complementary DNA using an oligo(dT)12 primer and Superscript II (Invitrogen). The mRNA levels of PSEN-1 and β-actin were measured by conventional polymerase chain reaction (PCR). Primers specific for the above-mentioned genes are listed as follows: PSEN-1: F 5′-- GGTCCACTTCGTATGCTGGT-3′ R 5′- GCTGTTGCTGAGGCTTTACC-3′ β-actin: F 5′- TGTACGCCTCTGGCCGTACC-3′ R 5′- CCACGTCACACTTCATGATGG-3′.
Polymerase chain reaction was performed in a 25 μL reaction mixture, and the PCR condition is 95°C 5 minutes, 95°C 30 seconds, 60°C 1 minute, and 72°C 2 minutes, 30 cycles. The relative mRNA expression levels were normalized to β-actin.
Image Capture and Quantification
Live cell images for GH3 cells with different constructs and treatments were captured by a single photon confocol microscopy (60× 1.0 NA objective, Olympus [Bridgeport, Connecticut, US]). Colocalization ratio of different vesicles or the number of single vesicles was quantified by ImageJ software (downloaded from NIH) with different plug-ins. A 2-tailed, 2-sample, unequal variance Student t test was used to calculate the P values. Error bars represent the standard error of the mean (SEM) of the samples.
Statistical Analysis
The data showed in figure were obtained from at least 3 independent experiments. Quantitative data were expressed as the means ± SEM. One-way analysis of variance or Student t test were used to analyze the difference between the means of test samples and controls, and P < .05 was considered to indicate a statistically significant result (*P < .05 and **P < .01).
Results
Elevation in HIF-1α Enhanced Autophagy and Thus Protected GH3 Cells From TMZ-Induced Apoptosis
Here we found after TMZ treatment that HIF-1α level elevated notably in GH3 cells, meanwhile HIF-1α shRNA could efficiently inhibit this elevation (Figure 1A and B), which meant HIF-1α level could be upregulated by TMZ treatment. Furthermore, MTT results showed that although 100 μmol/L TMZ treatment alone could reduce roughly 25% of cell viability, combined treatment with the autophagy inhibitor 3-MA (prevent autophagy induction) could significantly increase this rate to around 60% (Figure 1c). This result indicated blocking autophagy could actually sensitize antitumor effect of TMZ on GH3 cells. Interestingly, MTT results also showed HIF-1α shRNA could sensitize TMZ efficiency to the same extent as 3-MA, and this similarity indicated HIF-1α shRNA may also work through autophagy. To further investigate whether HIF-1α was involved in autophagy process, we did Western blot for a couple of autophagy markers including LC3, p62/SQSTM1, and Beclin-1 (Figure 1D). Results here showed that after TMZ treatment, LC3 II–LC3 I (lower/upper bands) ratio increased almost twice than control (Figure 1E), while p62 level reduced to nearly 40% of control (Figure 1F). These results demonstrated TMZ treatment could induce autophagy in GH3 cells. Consistent with previous reports, Beclin-1 level barely changed after TMZ treatment, which meant TMZ may not induce autophagy through Beclin-1 15 (Figure 1G). Finally, we found both 3-MA and HIF-1α shRNA could increase p62 protein level, which meant both 3-MA and HIF-1α shRNA could block autophagy in GH3 cells. However, to our surprise, when comparing LC3 II–LC3I ratio of TMZ alone, TMZ with 3-MA, and TMZ with HIF-1α shRNA groups, we found TMZ with HIF-1α shRNA group showed a strongest LC3 II–LC3 I ratio in all the groups, which indicated HIF-1α shRNA may not block autophagy at the initial stage as 3-MA did but more likely to function at final degradation stage of autophagy without influencing autophagy induction.

Autophagy could be induced after temozolomide (TMZ) treatment to protect GH3 cells from apoptosis through elevation in hypoxia-inducible factor 1α (HIF-1α). (A) GH3 cells were treated with TMZ together with 3-methyladenine (3-MA) or HIF-1α shRNA, and Western blots were used to detecte HIF-1α level. (B) Quantification for (A), normalized with actin. (C) GH3 cells were treated with TMZ together with 3-MA or HIF-1a shRNA, and 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay was performed to detect cell viability. (D) GH3 cells were treated with TMZ together with 3-MA or HIF-1α shRNA. LC3, Beclin-1, and p62 protein levels were detected by Western blot. (E-G) Quantification of LC3 II/I, p62, and Beclin-1 level intensity, respectively, normalized by actin. Data were represented as Mean ± standard error of the mean (SEM), n = 3. *P < .05; **P < .01; ***P < .001.
Hypoxia Inducible Factor 1 Regulated Autophagy Through Modulating Lysosomal Acidification
We employed autophagy inducer RA to treat GH3 cells with or without HIF-1α shRNA and found that although HIF-1α shRNA could not block RA-induced Beclin-1 increase, it could efficiently inhibit RA-induced p62 degradation (Figure 2A, C, and D), which is generally considered as autophagy dependent. Furthermore, we found HIF-1α shRNA combined with RA treatment resulted in the highest ratio of LC3 II–LC3 I within all the groups (Figure 2B). These results indicated that the inhibition effect of HIF-1α shRNA on autophagy was RA independent and probably occurred at the autolysosome stage. To further investigate this issue, we transfected mCherry-eGFP-LC3 tandem construct into GH3 cells with or without HIF-1α shRNA and RA. Normally, mCherry-eGFP-LC3 tandem construct shows “Cherry+” and “GFP+” in autophagysomes but only “Cherry+” in autolysosomes as the acidic environment of autolysosome/lysosome (usually around pH4.6-5.0) can easily quench the GFP fluorescence inside it (eGFP pKa value is around 5.0-5.5). To our surprise, the results here showed HIF-1α shRNA could significantly increase autophagosome number and the ratio of autophagosome–autolysosome, which meant either there was a fusion issue that autophagosome cannot merge into lysosome, or the acidic extent of autolysosome was impaired so that GFP fluorescence from tandem construct cannot be quenched normally (Figure 2F). To further investigate this issue, we transfected mKeima Red construct into GH3 cells. mKeima Red construct has a nearly 590 nm excitation and 620 nm emission wavelength only in acidic environment (pH <5), which was used here to indicate acidic vesicles in the GH3 cells. Results shown in Figure 2G and 2H indicated after HIF-1α shRNA treatment that the number of acidic vesicles decreased dramatically. Considering under physiological conditions most of the acidic vesicles inside the cell should be lysosomes, we cotransfected mKeima Red together with LAMP1-GFP (GFP here was attached in the C-terminal of LAMP1, which is finally located outside the lysosome, so the GFP fluorescence will not be quenched by lysosome) and found although under control or RA condition, most acidic vesicles (mKeima Red+) were colocalized with LAMP1-GFP, which meant lysosomes still had correct pH value, and after HIF-1α shRNA treatment, the LAMP1-positive vesicles showed no acidification anymore, which meant lysosomal pH was neutralized after HIF-1α knocking down (Figure 2I and J).

Hypoxia-inducible factor 1α (HIF-1α) regulated autophagy through modulating lysosome acidification. (A) GH3 cells were treated with Rapamycin (RA) or HIF-1α shRNA, LC3, and Beclin-1, and p62 protein levels were detected by Western blots. (B-D) Quantification of LC3 II/I, p62, and Beclin-1 level, respectively, normalized by actin. (E) GH3 cells were transfected with eGFP-mCherry-LC3 tandem construct under indicated conditions. (F) Numbers of autophagosome (GFP+ RFP+) and autolysosome (GFP− RFP+) were counted under each conditions. (G) GH3 cells were transfected with mKeima-Red and GFP-tagged control shRNA or HIF-1α shRNA; scale bar 10 μm. (H) Numbers of acidic vesicles (Red+) were counted under each indicated conditions. (I) GH3 cells were transfected with mKeima-Red and GFP-LAMP1; scale bar 20 μm. (J) Colocalization of acidic vesicles (Red) with lysosomes (GFP) were calculated. Data were represented as mean ± standard error of the mean (SEM); for Westerns, n = 3, for images, n = 10. *P < .05; **P < .01; ***P < .001.
Regulation of HIF-1α on Lysosome pH was Gamma Secretase-Independent and Relied on PSEN1 Expression
Next, we tried to find out the detailed mechanisms underlying how HIF-1α regulated lysosomal acidification. The PSEN1 is a key component of γ-secretase complex, which is important for many basic cellular processes such as Notch signal transduction. It has been reported previously that full-length PSEN1 could bind v-ATPase V0a1 subunit and conduct the latter one to translocate onto lysosomal membrane and thus maintain an appropriate acidic environment inside lysosome. We wondered whether HIF-1α could modulate lysosomal acidification through regulating PSEN1 expression. Figure 3 showed HIF-1α overexpression could both upregulate PSEN1 mRNA and protein level, while HIF-1α shRNA could decrease PSEN1 protein level (Figure 3 A-D). Furthermore, as PSEN1 is a component of γ-secretase complex, we tried to find out whether it affects lysosome in a γ-secretase-dependent or γ-secretase-independent way. We employed DAPT, a γ-secretase permanent inhibitor 16 , as the test group and lysosome degradation inhibitors Chloroquine (CQ; direct neutralize lysosome pH) and Bafilomycin (Baf; inhibit v-ATPase activity) as positive control. Results showed that while CQ and Baf could both efficiently increase LC ratio and p62 amount, DAPT did not have a significant influence on these 2 markers (Figure 3E-G). We also counted the acidic vesicle number under different conditions using lysotracker DND99 Invitrogen (Carlsbad, California, US), which could stain acidic vesicles inside the cell, and found that while both CQ and Baf could dramatically eliminate acidic vesicles, DAPT treatment showed a similar pattern as DMSO control. These results indicated PSEN1 regulated lysosomal acidification in a γ-secretase-independent way (Figure 3H and I). Finally, as mKeima and lysotracker are indeed recognizing acidic vesicle instead of lysosome, we developed TRPML-YFP-Keima tandem construct. The TRPML is a lysosomal-specific membrane cation channel protein, and Keima tag on its C-terminal could show the pH level inside lysosome. By calculating acidic TRPML+ vesicles (colocalized yellow vesicles/ green vesicles), we found both HIF-1α shRNA and PSEN1 shRNA but not DAPT could increase lysosomal pH, which confirmed that both HIF-1α and PSEN-1 could regulate lysosomal pH value (Figure 3J and K).

Hypoxia-inducible factor 1α (HIF-1α) regulation on lysosome pH is gamma-secretase independent and relies on presenilin 1 (PSEN1) expression. (A) GH3 cells were transfected with HIF-1α overexpression construct, and mRNA of PSEN1 was determined by polymerase chain reaction (PCR). (B) Quantification of (A) normalized by actin. (C) GH3 cells were tranfected with HIF-1α overexpression or HIF-1α shRNA, and C-terminal of PSEN1 was determined by Western blots. (D) quantification of (C) normalized with actin. (E) GH3 cells treated with DAPT, CQ, or Baf, and LC3 and p62 protein level were detected by Western blot. (F-G) Quantification of LC3 II/I and p62 level, respectively, normalized by actin. (H) GH3 cells transfected with GFP-LAMP1 and treated with lysoTracker Red for 5 minutes under each indicated conditions. (I) Acidic vesicle (Red) numbers was counted. (J) GH3 cells were transfected with TRPML-YFP-Keima tandem construct under each indicated conditions. (K) Percentages of acidic TRPML-YFP vesicle was calculated (yellow vesicles (Red+, Green+)/ Green+ vesicles). Data were represent as mean ± standard error of the mean (SEM), n = 3. *P < .05; **P < .01; ***P < .001. Scale bar 20 μm.
Presenilin 1 Could Attenuate HIF-1α shRNA-Enhanced Cell Death Under TMZ Treatment by Restoring Autophagy
As we already confirmed HIF-1α could regulate PSEN1 expression, we finally wanted to ask whether HIF-1α actually regulated lysosomal acidification and autophagy process through PSEN1. We administered TMZ to GH3 cells together with HIF-1α shRNA and PSEN1 overexpression construct and detected p62, the autophagy degradation marker. Data here were consistent with our previous results that HIF-1α shRNA could block p62 degradation and resulted in accumulation of p62 inside the cell, however PSEN1 reexpression could totally reverse this phenotype (Figure 4A and B). Furthermore, by using MTT assay, we found reexpression of PSEN1 could attenuate HIF-1α shRNA-induced cell death after TMZ treatment (Figure 4c). Finally, by counting acidic vesicle number or its colocalization with LAMP1-GFP vesicles, we found the neutralized lysosomal H+ concentration (pH value) by HIF-1α knockdown could be restored by PSEN1 reexpression (Figure 4D-G).
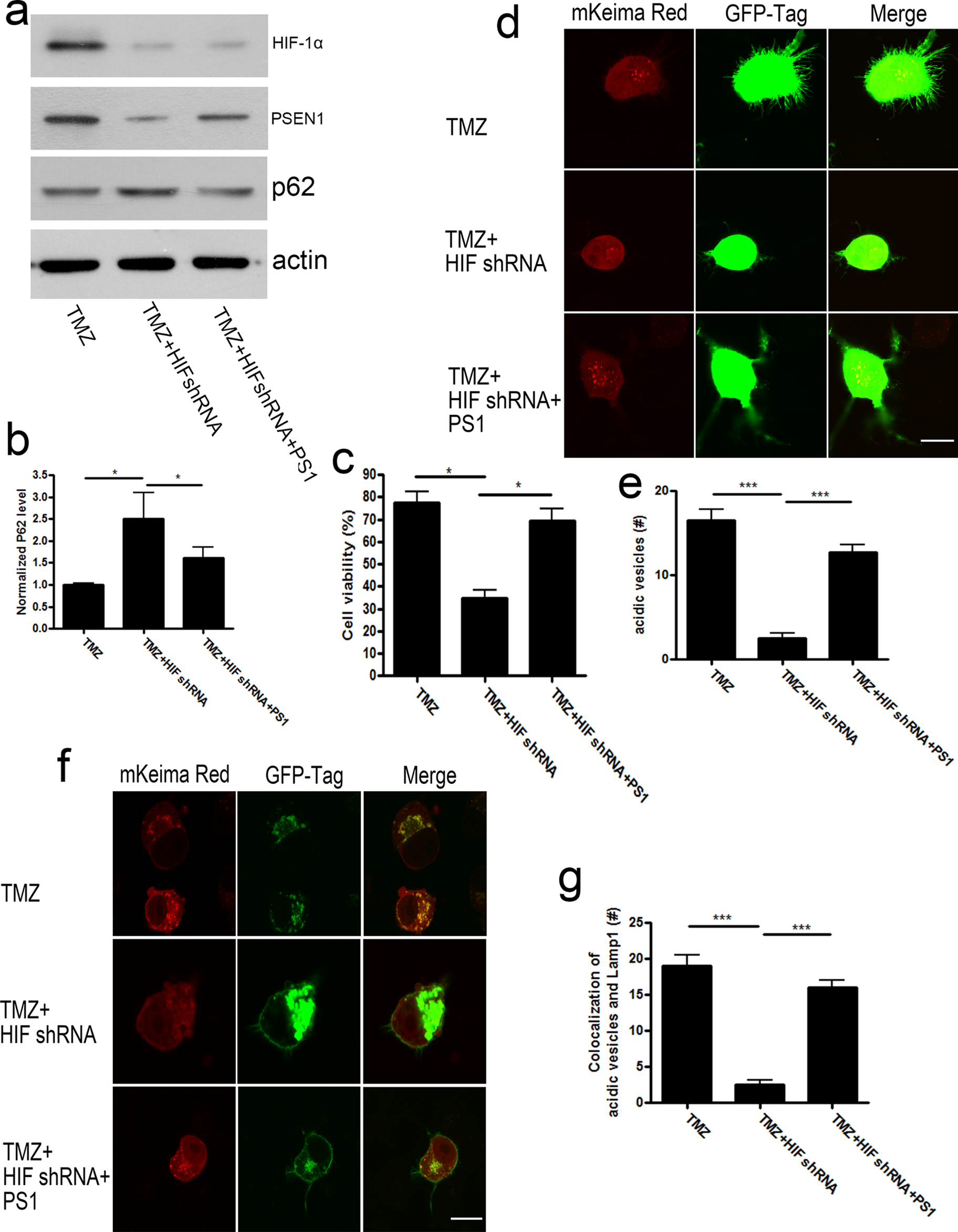
Presenilin 1 (PSEN1) could attenuate hypoxia-inducible factor 1α (HIF-1α) shRNA-induced cell death under temozolomide (TMZ) treatment by facilitating autophagy. (A) GH3 cells treated with TMZ were transfected with HIF-1α shRNA along or together with PSEN1 overexpression construct. P62 level was determined by Western blots. (B) Quantification of (A) normalized with actin. (C) GH3 cells treated with TMZ were transfected with HIF-1α shRNA along or together with PSEN1 overexpression construct. 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay was performed. (D) GH3 cells were transfected with mKeima Red with GFP-tagged control shRNA or HIF-1α shRNA. (E) numbers of acidic vesicles (Red+) were counted under each conditions. (F) GH3 cells were transfected with mKeima-Red and GFP-LAMP1. (G) colocalization of acidic vesicles (Red) with lysosomes (GFP) were calculated. Data were represented as mean ± standard error of the mean (SEM). For Westerns n = 3 and for images n = 10. *P < .05; **P < .01; ***P < .001. Scale bar 20 μm.
Discussion
It has long been considered that autophagy plays a role in both cancer genesis and antitumor treatment, but its detailed effects are still in debate. Although some previous reports have found activated autophagy may induce cancer cell death, many other studies have also suggested cancer cell may escape drug-induced apoptosis through activating autophagy. Autophagy itself is an important way for cells to degrade larger organelles to recycle amino acids when facing nutrition-deficient microenvironments as well as eliminate impaired organelles to limit damages spreading inside the cell. 17 Some recent reports showed CQ, a lysosomal pH neutralization reagent, could sensitize glioma cells to TMZ treatment through blocking autophagy process, which indicated activation of autophagy could resist antitumor effects of TMZ in glioma cells. 18 In the present study, we found that when using 3-MA to block autophagy initialization, cytotoxicity effects of TMZ on rat PAs GH3 cells could be enhanced significantly, which indicated autophagy is a way used by GH3 cells to escape TMZ-conducted anticancer effects.
Ratio of LC3 II–LC3I (lower/upper bands) is commonly used to evaluate autophagy levels. During autophagy, LC3-I is converted to LC3-II through lipidation by an ubiquitin-like system. So this ratio would increase when autophagy was induced or lysosome degradation was blocked; on the other hand, it would decrease when autophagy induction was blocked. Protein p62/ SQSTM1 is a receptor for cargo destined to be degraded by autophagy; when autophagy is induced, the p62 protein localizes to the autophagosomes and will be subsequently degraded in lysosome. Conversely, when autophagy is inhibited, the p62 protein would then accumulate in the autophagosome. Thus, p62 protein amount could indicate function of lysosome degradation system. 19 In the present study, we used LC3 II–LC3 I ratio and p62 protein level to evaluate autophagic activity. We found TMZ could increase LC3 II–LC3 I ratio but decrease P62 level, which strongly indicated TMZ could induce autophagy in GH3 cells.
As solid tumors usually develop in environments with a limited oxygen supply, HIF-1α, which is the center regulator of low oxygen-related gene expression and cell adaptation, has long been considered to be only involved in cancer genesis. 20 Until recently, studies revealed that HIF-1 also played a part in drug resistance through regulating O6-methylguanine-DNA-methyltransferase (MGMT) gene expression. MGMT is a DNA repair protein that could remove TMZ-induced DNA methylation in glioma cells and thus protect cells from TMZ-induced apoptosis. 21 In the present study, we found HIF-1 could also induce drug resistance through regulating lysosomal acidification. Knockdown of HIF-1α resulted in neutralized lysosomal pH and blockade of autolysosome contents degradation. Although failed fusion of autophagosomes and lysosome could also block autolysosome degradation, the present data showed colocalaization of LC3+ and LAMP1+ vesicles were not changed after HIF-1α knock down, which suggested fusion of autophagosomes with autolysosomes was unlikely to be influenced. Finally, considering shRNA usually have “off-target” effects, we employed HIF-1α inhibitor 2-Methoxyestradiol to carry on experiments parallel with HIF-1α shRNA and got similar results (data not shown).
Presenilin 1 is a key component of γ-secretase complex. As a core component of γ-secretase, the relation between hypoxia and PSEN1 has been studied thoroughly in Alzheimer disease and in embryo development stage. In the embryo development stage, γ-secretase could cleave Notch protein, release the intracellular domain of Notch into cell nucleus, and thus control Notch signaling pathways, which are important for embryo development and are tightly regulated by hypoxia condition. 22 In contrast, PSEN1 mutations could induce familiar inherited dementia through shifting APP cleavages, which generate toxic amyloid beta (Abeta) peptide and thus cause neuronal death. Reports showed hypoxia could increase toxic Abeta peptide generation by increasing γ-secretase activity 23 or by negatively regulating gene expression of major degradation enzyme Neprilysin (NEP) of Abeta. 24 It is believed in the aging process, PSEN1-conducted gain of function of γ-secretase complex is a key point for Alzheimer disease occurrence. However, until recently, people start to focus on γ-secretase-independent functions of PSEN1 . Lee et al 25 have found PSEN1 could directly bind to v-ATPase V0a subunit and regulate v-ATPase, a core H+ channel protein on lysosome, and translocated onto lysosome membrane, thus maintaining a normal acidic pH value of lysosomes interior. In the present study, we found HIF-1α could regulate PSEN1 gene expression, and after HIF-1α knockdown, the lysosomal pH neutralization could be rescued by exogenous reexpression of PSEN1, which is consistent with previous report that PSEN1 null cells showed impaired lysosomal acidification. 25 Furthermore, we also found PSEN1 affected lysosomal function in a γ-secretase-independent way, as γ-secretase inhibitor DAPT failed to show a similar effect as HIF-1α knockdown or lysosome inhibitor CQ. This result is also consistent with previous findings that full length but not cleaved PSEN1(which functions in a γ-secretase-dependent way) could regulate lysosomal pH value. 24,25
Unfortunately, in the present study, we failed to identify any potential HRE motifs in PSEN1 promoter region, so we couldn’t conclude whether HIF-1α regulated PSEN1 expression directly or indirectly. However, in a recent study, a research group has identified HIF-1α could negatively regulate NEP gene expression through competitively binding to NEP promoter region with AICD, and the latter one could upregulate NEP expression through epigenetic acetylation instead of direct transcriptional regulation. 24 Their work showed that HIF-1α may regulate downstream gene expression through epigenetic way, which may also happen in the present study, which means HIF-1α may regulate PSEN1 expression through promoting histone acetylation or inhibiting CpG gene methylation.
Overall, in the present study, we demonstrated TMZ could elevate HIF-1α protein level and induce autophagy which could help GH3 cells to escape from TMZ-induced apoptosis. Meanwhile, autophagy inhibitor 3-MA and HIF-1α knockdown could sensitize GH3 cells to cytotoxic effect of TMZ to a similar extent. Furthermore, we found HIF-1α knockdown could neutralize lysosomal acidic pH level and thus block autophagy process. Finally, we found HIF-1α knockdown could neutralize lysosomal pH through regulating PSEN1 expression, and exogenous reexpressing of PSEN1 after HIF-1α knockdown could restore lysosomal H+ concentration in a γ-secretase-independent way and thus recovered the autophagy and reduced TMZ’s effect. Taken together, our data found that HIF-1 could regulate autophagy process through regulation of lysosomal acidification and PSEN1 expression in GH3, which suggest HIF-1 knockdown could be a potential approach to improve the efficacy of TMZ therapy for PAs.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
