Abstract
The Clock gene, an indispensable component of the circadian clock, not only modulates circadian oscillations but also regulates organismal function. We examined whether silencing the expression of the human Clock gene in glioma cells influences cell growth and induces apoptosis after irradiation. Silencing the expression of Clock in a human glioma cell line (U87MG), but not in a control glioma cell line, resulted in increased apoptosis and cell cycle arrest. Moreover, silencing Clock expression altered the expression of apoptosis-related genes. The protein levels of c-Myc and Cyclin B1 were downregulated, but those of p53 were upregulated, in human Clock-silenced U87MG cells compared with control cells. Our results suggest that the circadian gene human Clock may play an important role in carcinogenesis by inhibiting apoptotic cell death via attenuating proapoptotic signaling.
Introduction
Glioma is the most common form of primary brain tumor. Astrocytoma (World Health Organization grades I and II) is a type of glioma that is derived from star-shaped astrocyte-like cells, and high-grade astrocytomas are referred to as anaplastic astrocytomas (grade III) or multiform glioblastomas (GBM; grade IV). Despite advances in surgery and radiotherapy, the prognosis of malignant glioma remains poor, as the median survival duration of patients with GBM is 5 to 8 months. 1,2
The prevention, screening, and treatment of this type of cancer are all problematic, emphasizing the need for the development of new diagnostic and therapeutic strategies. Epigenetic events are an important physiologic cellular function, and recent evidence has suggested that these events are critical mediators of the initiation and progression of cancer. 3,4 Recent studies have shown that the deregulated expression of rhythm genes due to epigenetic alterations is an early event in many human malignancies, including glioma.
The circadian rhythm is the daily oscillation of organismal processes that is regulated by an endogenous clock. The rhythm of the circadian clock is modulated by the interaction between positive and negative feedback loops that consist of critical clock regulators (including Clock and Bmal1), 2 cryptochromes (Cry1 and Cry2), and 3 period genes (mPer1, mPer2, and mPer3).
5,6
Epidemiological studies have shown that disruption of the normal circadian rhythm may increase the risk of developing various types of cancer, such as breast, prostate, colorectal, and endometrial cancer.
7
–12
Chen
13
In the present study, we aimed to understand the role of Clock expression in modulating the sensitivity of tumor cells to radiation and in promoting tumor growth and progression. We assessed the effect of silenced Clock expression on cell growth, apoptosis, and radiation sensitivity, and we found that Clock plays an important role in regulating both the growth and apoptosis of human glioma cells.
Materials and Methods
Cell Culture
The wild-type p53-carrying human glioma cell line U87MG was obtained from American Type Culture Collection and maintained in Dulbecco’s modified Eagle’s medium (HyClone, Logan, Utah) supplemented with 10% fetal bovine serum (HyClone) and antibiotics in a humidified atmosphere consisting of 95% air and 5% CO2 at 37°C. To evaluate cellular apoptosis, the cells were cultured on coverslips in 6-well plates until reaching 70% confluence.
Small interfering RNA Transfection
Small interfering RNAs (siRNAs) targeting Clock were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA) as a pool of 3 target-specific, 19- to 25-nt siRNAs. A scrambled-sequence negative control siRNA with no known homology to the human genome was used to control for the effects of transfection and was chemically synthesized by Integrated DNA Technologies (Coralville, Iowa, USA). The efficacy of this method and of the oligodeoxynucleotide sequences has been described previously. 14
We treated the cells with 100 nmol/L tissue plasminogen activator (TPA; Sigma, St. Louis, MO, USA) to induce endogenous human Clock (hclock) expression in glioma cells, and the Clock siRNA was used to block this expression. Following the addition of TPA and the oligonucleotides for 24 hours, the cells were grown in serum-containing medium and treated with 10 Gy of x-rays using a Varian 2100C/D linear accelerator (Varian, Palo Alto, CA, USA). We then analyzed the irradiation-induced apoptosis and DNA damage in the following 3 groups: (1) TPA alone, (2) TPA and scrambled oligonucleotides (TPA Rand), and (3) TPA and antisense oligonucleotides targeting hclock (TPA siClock).
Apoptosis Assay
A cell that is undergoing apoptosis displays nuclear condensation and DNA fragmentation, which can be detected via Hoechst 33342 staining and fluorescence microscopy.
Forty-eight hours after irradiation, cells grown on coverslips were fixed with 4% paraformaldehyde and stained with Hoechst 33342 (5 mg/mL) for 20 minutes at room temperature. The coverslips were then washed, mounted on glass slides, and stored at 4°C until quantification using a laser scanning confocal microscope (FluoView™ FV1000 Viewer, Olympus, Japan), with an excitation wavelength of 350 nm and an emission wavelength of 460 nm. Three coverslips were used per experimental group, and at least 200 cells in 4 random fields were counted for each coverslip.
Cell Cycle Analysis
Forty-eight hours after irradiation, the cells were washed with 4°C phosphate-buffered saline (PBS) and fixed in 70% ethanol at 4°C overnight. Then, the cells were treated with RNase A solution (500 U/mL) at 37°C for 15 minutes and stained with propidium iodide (50 μg/mL) at room temperature. The DNA content was determined via flow cytometry (Flow CytoMeter, Millipore) using CytoSoft 5.3 software (Millipore, Billerica, Massachusetts, US), and the percentages of cells in G1, S, or G2/M phase were determined using FCS Express software (version 3; Denovo).
DNA Damage Assay
DNA damage was assessed using the alkaline single-cell gel electrophoresis assay (comet assay). After the above-described irradiation treatments, the cells were washed in PBS and incubated in normal medium for 3 hours. Then, the cells were harvested, mixed with low-melting agarose, and fixed to slides. The cells were subsequently lysed, and the DNA was denatured using an alkaline (pH >14) solution at 4°C. Afterward, the slides were placed on a horizontal electrophoresis device that allowed damaged and/or broken DNA to migrate away from the nucleus. Next, the slides were stained with ethidium bromide, visualized via fluorescence microscopy, and analyzed using Komet 5 comet assay analysis software (Andor Technology, Belfast, UK), which quantitatively determines the extent of DNA damage in each sample according to the mean Olive tail moment, as described previously.
Western Blotting
Forty-eight hours after irradiation, the cells were lysed in cold radioimmunoprecipitation assay lysis buffer containing protease inhibitors, and the proteins were collected via centrifugation. The protein concentrations were then determined using the bicinchoninic acid protein assay (Pierce, Rockford, Illinois), and immunoblotting was performed using antibodies against the following proteins: Clock, p53, c-myc, Cyclin B1, Cyclin D, Mdm2, and β-actin. The immunoblots were developed using Supersignal WestPico chemiluminescence substrate (Pierce, Rockford, Illinois, USA) and were imaged and analyzed via photodensitometry using an imaging system (Bio-Rad, Hercules, California, USA).
Results
Effects of Clock on Cellular Apoptosis
A cell that is undergoing apoptosis displays nuclear condensation and DNA fragmentation, which can be detected via Hoechst 33342 staining and fluorescence microscopy. Forty-eight hours after irradiation, apoptotic cell damage was analyzed using Hoechst 33342 staining, as shown in Figure 1A. As illustrated in Figure 1B, many cells were transfected with Clock siRNA and then stained with Hoechst 33342 exhibited fragmented nuclei; the proportion of apoptotic cells in the Clock siRNA-treated group was significantly higher than that of the group treated with TPA alone (
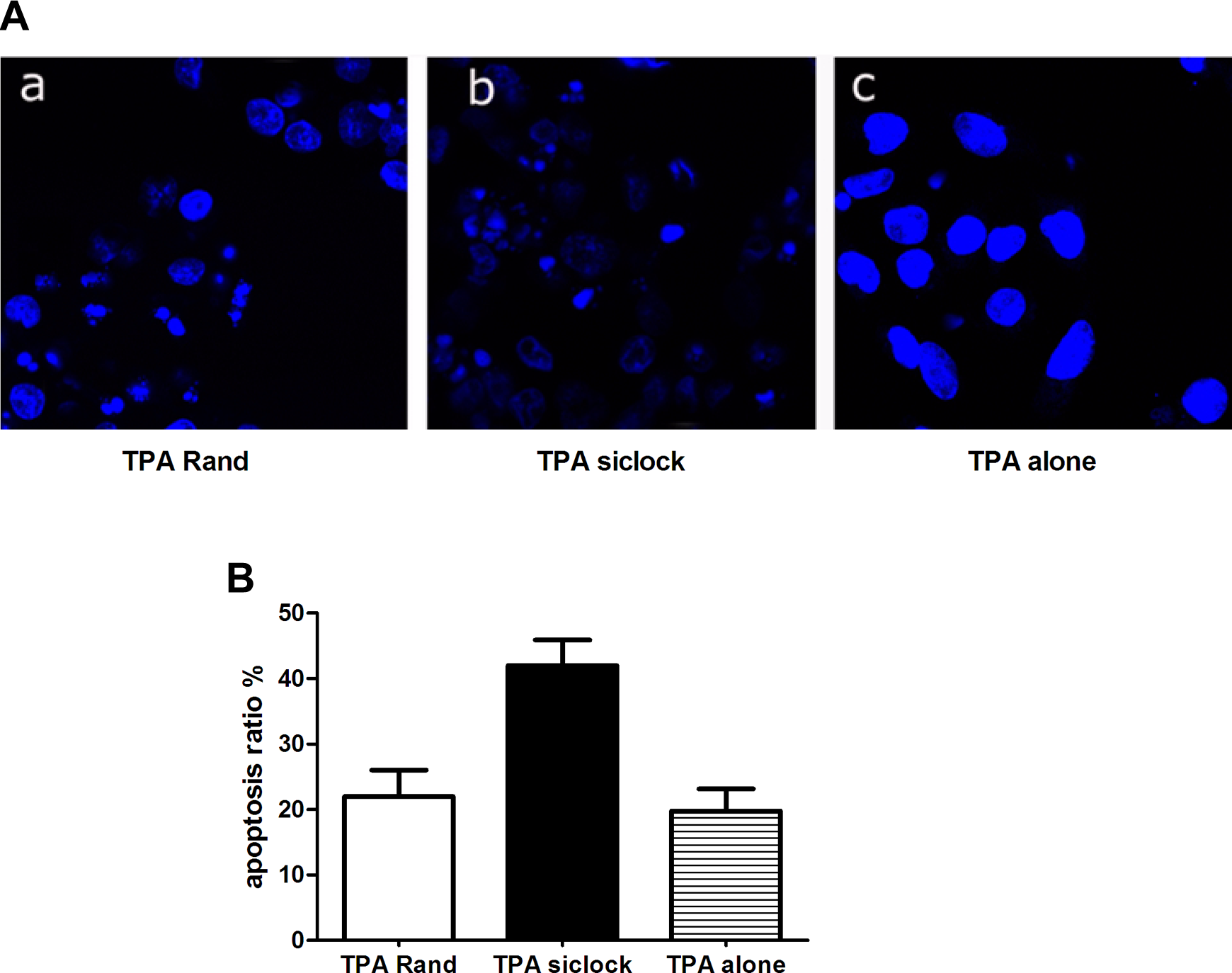
Following treatment with irradiation and staining with Hoechst 33342, cells were subjected to fluorescence microscopic analysis to determine the proportion of apoptotic cells. a, The TPA and scrambled oligonucleotides (TPA Rand), (b) TPA and antisense oligonucleotides targeting hclock (TPA siClock), and (c) TPA alone. TPA indicates tissue plasminogen activator.
Effects of Clock on DNA Repair Capacity
DNA damage was assessed using the alkaline single-cell gel electrophoresis assay (comet assay) and analyzed using the Komet 5 comet assay analysis software, which quantitatively determines the extent of DNA damage in each sample using the mean Olive tail moment calculation as described previously. The mean Olive tail moment was determined for 50 cells from each of the 3 treatment groups (targeted siRNA, negative control siRNA, and TPA alone; Figure 2). Longer “tails” for an individual cell correspond to a greater accumulation of damage, as the head material consists of intact DNA, and the tail material is composed of damaged DNA. The mean Olive tail moment was 0.722 ± 0.069 for the TPA alone group (1), 0.113 ± 0.034 for the targeted siRNA group (2), and 0.791 ± 0.152 for the negative control siRNA group (3). The targeted siRNA group displayed a significantly longer mean Olive tail moment (
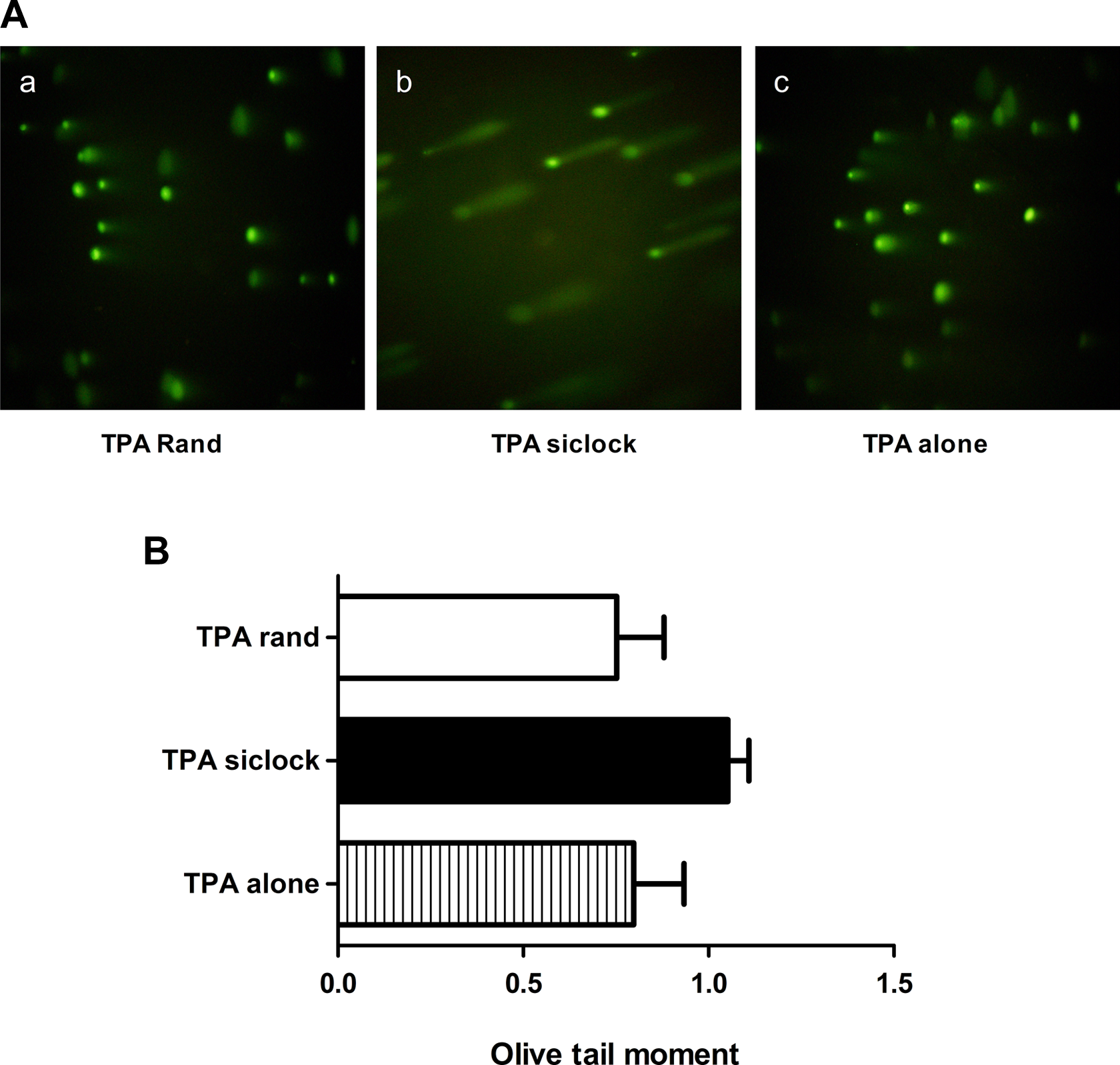
DNA damage repair capacity assessed via the comet assay. a, The TPA and scrambled oligonucleotides (TPA Rand), (b) TPA and antisense oligonucleotides targeting hclock (TPA siClock), and (c) TPA alone. TPA indicates tissue plasminogen activator.
Effects of siClock on Cell Cycle-Related Genes
U87MG cells were transiently transfected with either the negative control siRNA vector or the siClock vector. Forty-eight hours posttransfection, Western blot analysis was performed to evaluate the protein expression level of Clock. As shown in Figure 3, antisense oligonucleotide treatment resulted in a clear reduction in the expression of the Clock protein compared to the control treatment. Moreover, the protein expression of p53 was downregulated, and those of c-myc and Cyclin B1 were upregulated by siClock treatment (Figure 3A and B). However, the protein expression levels of Mdm2 and Cyclin D were not significantly altered by siClock treatment.

Relative expression of Clock, p53, c-myc, Cyclin B1, Cyclin D, and Mdm2 in glioma cells. A, The protein levels of Clock, p53, c-myc, Cyclin B1, Cyclin D, and Mdm2 were determined via Western blot analysis and were normalized to the level of β-actin using a Molecular Dynamics densitometer. B, The ratios of the Clock, p53, Cyclin B1, or c-myc signals to the β-actin signal in the control cells were arbitrarily set at 1.0. The error bars indicate the standard errors of the means (SEMs; n = 3). (1) The TPA and scrambled oligonucleotides (TPA Rand), (2) TPA alone, and (3) TPA and antisense oligonucleotides targeting hclock (TPA siClock). TPA indicates tissue plasminogen activator.
Effects of Clock on the Cell Cycle
The proportion of U87MG cells in G2/M phase in the siClock group after radiation treatment (10.95%) was significantly higher than that in the TPA alone (3.1%,

Effects of Clock on the cell cycle; cells were harvested, and DNA was stained with propidium iodide (PI) for flow cytometric analysis and cell phases were determined using the Watson pragmatic algorithm. A, The TPA and scrambled oligonucleotides (TPA Rand), (B) TPA alone, and (C) TPA and antisense oligonucleotides targeting hclock (TPA siClock). TPA indicates tissue plasminogen activator.
Discussion
In separate studies, the deregulated expression of Clock genes was commonly found in human gliomas, and the Clock gene was shown to play a role in glioma development by enhancing the growth and proliferation of cancer cells. 15 –17 In the present study, the effects of Clock were evaluated. As a core circadian gene, Clock not only maintains the circadian rhythm of cells but also sustains the physiologic cell cycle by modulating the expression of cell cycle-related genes such as P53, c-Myc, and Cyclin B1. In our study, silencing the expression of Clock induced a significant effect on cellular apoptosis, as demonstrated by Hoechst 33342 staining and Olive tail moment analyses. Alternatively, Clock overexpression in human glioma cells exerted an adverse regulatory effect on cell cycle-related genes. The circadian clock controls downstream events by regulating the expression of clock-controlled genes. Therefore, we postulated that certain genes that function in cell proliferation and tumor suppression act as clock-controlled genes and are deregulated in siClock-treated glioma cells. The clock-controlled genes that are involved in regulating the cell cycle and apoptosis include c-Myc and the tumor suppressor p53, as well as genes that encode caspases, cyclins, and transcription factors. 9,10,18,19
In general, the circadian clock and the cell cycle are 2 global regulatory systems that are present in most eukaryotic organisms. 20 Clock plays a role in a wide variety of physiological processes, including the circadian rhythm of cells and the maintenance of the cell cycle. It has been reported that 2% to 10% of all mammalian genes are clock controlled, 21 –24 and recent studies reported that approximately 7% of all clock-controlled genes that have been identified in rodents regulate either cell proliferation or apoptosis. 25 Cell cycle checkpoints are important growth arrest mechanisms that ensure the orderly progression of cell cycle events and that prevent aberrant mitosis in response to DNA damage. Therefore, cell cycle checkpoints are critical components of the mechanism by which the Clock gene affects the apoptosis of glioma cells.
Cyclin B1 is the key regulator of the G2/M phase of the cell cycle, and recent investigations have demonstrated that the level of Cyclin B1 expression is likely correlated with the malignant potential of human tumors. 26 –29 A high level of Cyclin B1 may accelerate the progression through G2/M, and the absence of Cyclin B1 prior to the G2/M transition prevents the cell from entering mitosis, resulting in cell arrest at G2. Our results indicate that the disturbance of Cyclin B1 by Clock may lead to cell cycle arrest at G2/M and strongly agree with those of earlier studies demonstrating that Clock genes regulate the G2/M transition during murine liver regeneration. 30 In addition, circadian genes regulate cell cycle- and apoptosis-related proteins in mouse bone marrow and tumors. 31 Therefore, it is conceivable that the Clock gene may control cell cycle progression via its transcriptional regulation of Cyclin B1. These results, together with our findings, support a role of the circadian clock in orchestrating mitotic events.
The tumor suppressor p53 is a transcription factor that is activated in response to DNA damage or oncogenic transformation. The loss or mutation of p53 in many cancers leads to impaired cell cycle regulation, genomic instability, and the inhibition of apoptosis. 32 –34 The protein levels of p53 in the Clock-silenced U87MG cells tended to be higher than those in the control cells, indicating that Clock reduces p53 expression, and this activity may at least partially contribute to the apoptosis of Clock-silenced cells.
When mammalian cells contain damaged DNA, p53 and the Rb family of transcriptional repressors act together to downregulate many genes that encode proteins required for the G2/M transition. The elimination of these essential cell cycle proteins helps to maintain the cells in G2 arrest. 34 Furthermore, p53 regulates Cdc2 and Cyclin B1 through transcriptional suppression. Moreover, previous studies indicated that the circadian regulation of Cyclin B1 appears to be indirect and to involve a p53-dependent mechanism, as p53 helps to block entry into mitosis and strengthens G2 arrest. 35
The circadian clock-controlled gene c-myc plays a key role in cell proliferation and apoptosis. The deregulation of c-myc has been associated with various cancers and with the hyperplastic growth of mammalian tissues. 36,37 Previous studies have also indicated that c-myc overexpression may facilitate the progression of cells through the cell cycle in the presence of genomic DNA damage. 37 In our study, we have focused on examining circadian clock-regulated gene expression to emphasize the fundamental role that the circadian clock plays in glioma cell apoptosis following irradiation. We found that the levels of c-myc were significantly decreased in irradiated siClock-transfected glioma cells and that c-myc overexpression induced genomic DNA damage and compromised p53 function, presumably via a reactive oxygen species-mediated mechanism. 38 Thus, silencing the expression of Clock partially promotes p53-mediated apoptosis in glioma cells.
Based on recent discoveries from Clock studies and our own results, we propose a model for the role of Clock in oncogenic transformation. In this model, the deregulation of Clock results in decreased intracellular levels of p53, leading to the derepression of c-myc and Cyclin B1. In contrast, overexpression of c-myc and Cyclin B1 causes genomic DNA damage and ultimately induces hyperplasia and tumor development.
In summary, our data establish a strong association between the circadian clock and the cell cycle, which may explain how disruption of the circadian clock may shift the cellular balance between proproliferative and antiproliferative genes, which increases the risk of cancer initiation and progression. Studies in this field over the next few years will likely reveal additional associations between the circadian clock and signal transduction pathways and will further increase our understanding of how these factors interact and, equally importantly, of how such interactions and subsequent cellular responses are exquisitely orchestrated.
Footnotes
Authors’ Note
Fan Wang and Caiyan Li contributed equally to this work.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Natural Science Foundation of HuBei Province of China (Grant 2014CFC1029).
