Abstract
We evaluated and compared the accuracy of 2 deformable image registration algorithms in 4-dimensional computed tomography images for patients with lung cancer. Ten patients with non-small cell lung cancer or small cell lung cancer were enrolled in this institutional review board-approved study. The displacement vector fields relative to a specific reference image were calculated by using the diffeomorphic demons (DD) algorithm and the finite element method (FEM)-based algorithm. The registration accuracy was evaluated by using normalized mutual information (NMI), the sum of squared intensity difference (SSD), modified Hausdorff distance (dH_M), and ratio of gross tumor volume (rGTV) difference between reference image and deformed phase image. We also compared the registration speed of the 2 algorithms. Of all patients, the FEM-based algorithm showed stronger ability in aligning 2 images than the DD algorithm. The means (±standard deviation) of NMI were 0.86 (±0.05) and 0.90 (±0.05) using the DD algorithm and the FEM-based algorithm, respectively. The means of SSD were 0.006 (±0.003) and 0.003 (±0.002) using the DD algorithm and the FEM-based algorithm, respectively. The means of dH_M were 0.04 (±0.02) and 0.03 (±0.03) using the DD algorithm and the FEM-based algorithm, respectively. The means of rGTV were 3.9% (±1.01%) and 2.9% (±1.1%) using the DD algorithm and the FEM-based algorithm, respectively. However, the FEM-based algorithm costs a longer time than the DD algorithm, with the average running time of 31.4 minutes compared to 21.9 minutes for all patients. The preliminary results showed that the FEM-based algorithm was more accurate than the DD algorithm while compromised with the registration speed.
Keywords
Introduction
Four-dimensional computed tomography (4D CT) has been widely used in radiation therapy to acquire time-resolved volumetric images 1 –3 and access patient-specific breathing motion for determining individual safety margins. 4 –8 Since the thoracic organs expand and contract nonuniformly with the breathing, deformable image registration (DIR) is required to extract the motion and anatomical information from 4D CT images.
Several different algorithms have been proposed and validated for nonrigid registration of 4D CT images. 9 –17 For example, Wang et al 9 proposed an accelerated “demons” algorithm and validated it on patients with prostate, head-and-neck, and lung cancer. An “active” force along with an adaptive force strength adjustment during the iterative process was introduced to accelerate the registration algorithm. The results showed that the accelerated demons algorithm had a significant potential for delineating and tracking doses in target and critical structures during computed tomography (CT)-guided radiotherapy. Wu et al 10 developed a registration method that could be used to create discontinuous vector fields at the boundaries of anatomic subregions and overcame the difficulties in most registration algorithms, which assumed that the deformation field was smooth and continuous everywhere. The authors indicated that the lung was allowed to slide almost independently along the chest wall, resulting in a discontinuous deformation vector field, which could not be generated using standard deformable registration methods. In their study, registration was performed independently on each subregion, with a boundary-matching penalty used to prevent gaps. Both B-spline and demons registration algorithms were used to implement and test this method. The approach was validated on 4D CT data sets of 4 patients with lung cancer for registration of the end-inhalation and end-exhalation volumes. The results showed that the registration achieved consistently improved accuracy for both the B-spline and the demons algorithms in both moving and less-moving subregions. Retzel et al 11 registered 4D CT data to a reference respiratory phase using a deformable registration based on B-splines. Anatomic landmarks were selected across 10 respiratory phases in 5 patients to assess registration performance. The average registration error for 5 landmarks for each of the patients was 2.1 mm. Velec et al 12 used a biomechanical model-based DIR algorithm, previously developed by Brock et al 13 to retrospectively deform each exhale CT to inhale CT. The deformation map was combined with exhale and inhale dose grids from the treatment planning system to accumulate dose over the breathing cycle. The diffeomorphic demons (DD) algorithm proposed by Vercauteren et al 14 has been recognized as a robust and reliable method. 15 This algorithm is based on the classic demons, 16 which is a well-established technique for nonrigid registration and computes the pixel displacements driven by the image intensity difference. Instead of performing the optimization on the displacement vector field (DVF) as in the classic demons algorithm, the DD algorithm combines the Lie group framework on diffeomorphisms and optimizes for the Lie group. Thus, this method ensures to generate invertible output transformation. Furthermore, most nonrigid registration methods are established by solving an energy function minimization problem or a partial differential equation. The finite element method (FEM) 17 is a powerful technique to solve these problems, which commonly divides the problem domain into elements and further solves the equations element-by-element. The advantage of FEM-based method is the ability of dividing the domain into nonuniform elements. Thus, it can reduce the computational time by applying small elements where accurate registration is required and large elements in areas of less importance.
The goal of our study is to evaluate and compare the accuracy of the DD- and FEM-based algorithms. This is a novel study because no investigation has been presented to report the comparison between the 2 algorithms. Ten 4D CT data sets of patients with lung cancer were included in this study. The CT image at phase T = 0% (CT00) was chosen as the reference image, while the CT images at other respiratory phases were treated as the floating images successively. To speedup the registration process, maintain robustness, and avoid local extremum, the reference and floating images were downsampled into 3 resolution levels, forming 2 multiresolution image pyramids, and the multiresolution registration was performed iteratively from the coarsest level to the finest level using the DD- and FEM-based registration algorithms. The DVFs relative to the reference image calculated by the 2 DIR algorithms were extracted for comparison. To quantitatively evaluate the 2 algorithms, 3 most commonly used similarity measures, normalized mutual information (NMI), the sum of squared-intensity differences (SSD), and modified Hausdorff distance (dH_M), were used in this study. Furthermore, the computation time of the 2 algorithms was recorded and compared. The registration schemes used in this study were evaluated and compared quantitatively and qualitatively.
Materials and Methods
Patient Cohort and Imaging Study
Ten patients (5 male, 5 female, age 34~73) who had nonsmall cell lung cancer (NSCLC; n = 5) or small cell lung cancer (SCLC; n = 5) were enrolled in this prospective study, which has been approved by the Institutional Review Board of Shandong Cancer Hospital and Institute. Table 1 summarizes the clinical characteristics of the patients. All the patients underwent CT scans on the same day for treatment planning and were positioned head-first-supine with the arms raised above the head. Vacuum bags were used to immobilize the patients during image acquisition.
Summary of Patients’ Characteristics and Measurements.
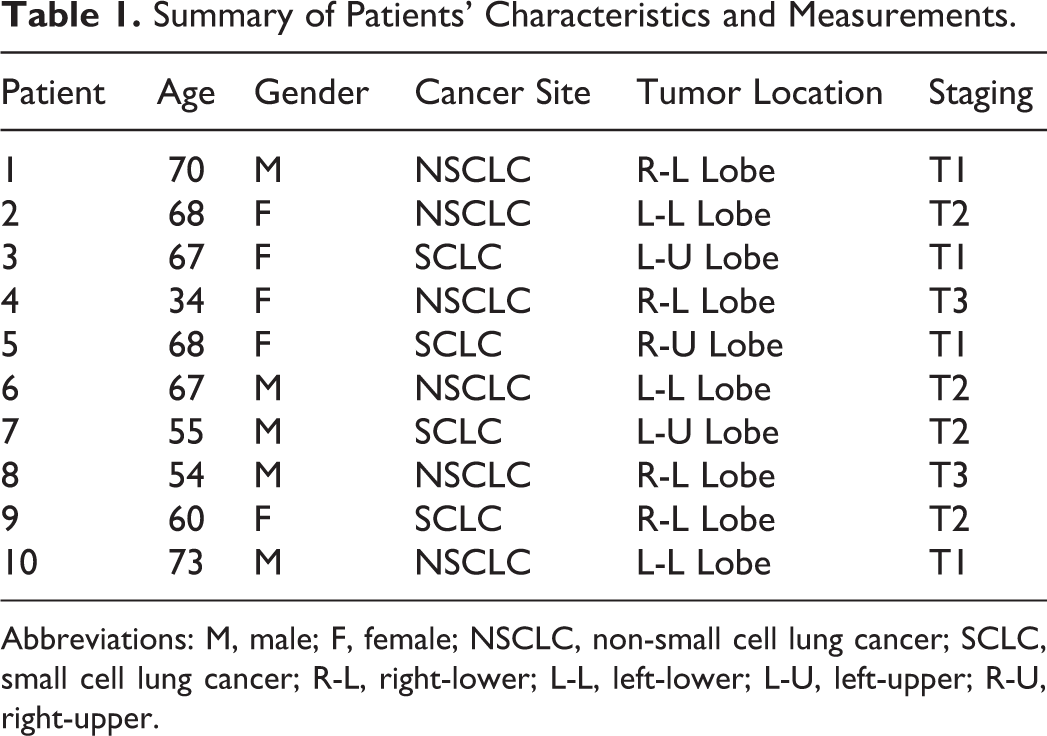
Abbreviations: M, male; F, female; NSCLC, non-small cell lung cancer; SCLC, small cell lung cancer; R-L, right-lower; L-L, left-lower; L-U, left-upper; R-U, right-upper.
For each patient, a 4D CT scan was performed under uncoached free breathing condition on a 16-slice CT scanner (Philips Brilliance Bores CT, Andover, Massachusetts) equipped with real-time position management (RPM) system (Varian Medical Systems, Inc, Palo Alto, California) and Advantage 4D software (GE Healthcare, Milwaukee, Wisconsin). The respiratory signal was recorded with the RPM gating system by tracking the trajectory of infrared markers placed on the patient’s abdomen. Each CT image from the scanner was labeled by the time tag according to the respiratory signal. The reconstructed 4D CT images were sorted into 10 respiratory phases based on tags by the Advantage 4D software, with 0% corresponding to end inhalation and 50% corresponding to end -exhalation. The imaging parameters were as follows: voltage/current: 120 kV/290 mA, slice thickness: 2.5 mm, gantry rotation: 0.5 s/cycle, reconstruction matrix: 512 × 512, and field of view: 450 to 500 mm.
Diffeomorphic Demons Algorithm
The complete registration framework can be treated as an optimization problem that aims at finding the deformation field to obtain a reasonable alignment between images. And it can also be summarized by a model with an energy function consisting of a similarity function and a regularized term. 18
For a fixed image (F), a moving image (M), and a transformation field (s), the energy function of the DD algorithm with respect to the update field (u) can be calculated using Equation 1:

where
The energy function calculated in Equation 1 can be linearized using the first-order Taylor expansion as follows:

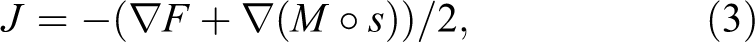
where J is used for performing an efficient second-order minimization symmetric registration, with 
Typically, the Gaussian smoothing is used as the regularization term for the displacement field (

So the iterative registration algorithm can be organized as follows:
For the current transformation s, compute the corresponding field Typically, use the Gaussian kernel to regularize the update field Let Typically, use the Gaussian kernel to regularize c, satisfying
In our study, 3 levels of resolution consisting of 20 iterations each were adopted. A Gaussian kernel with σ = 3 and σ = 2 were used for the update field regularization (
FEM-Based Registration Algorithm
Given 2 images, a fixed image 

where the first term represents the similarity of 2 images and the second term aligns the predefined landmarks in the fixed image (
In this study, anatomical landmark feature pairs were identified manually across 10 respiratory phase images. A Matlab-based software named Assisted Point Registration of Internal Landmarks (APRIL, The MathWorks Inc., Natick, Massachusetts) was employed to facilitate the selection of landmarks. 20,21 The corresponding feature points were determined by an expert via manually designing the feature correspondence on the target image. For each patient, a reference set of lung landmark feature points was generated on the end-inhale phase image (CT00) from the 4D CT data set. The vessels and bronchial bifurcations were typically included in the feature points, which were selected systematically on the 10-phase images by the expert. About 10 feature points per axial slice were selected and propagated throughout the respiratory phases as shown Figure 1, which ensured the collection of >300 feature points for all cases. For each case, the feature points were visually reviewed by the observer a second time and location adjusted according to the end-inhale phase image if necessary. The points were then used to verify the accuracy of the FEM-based algorithm in the study.
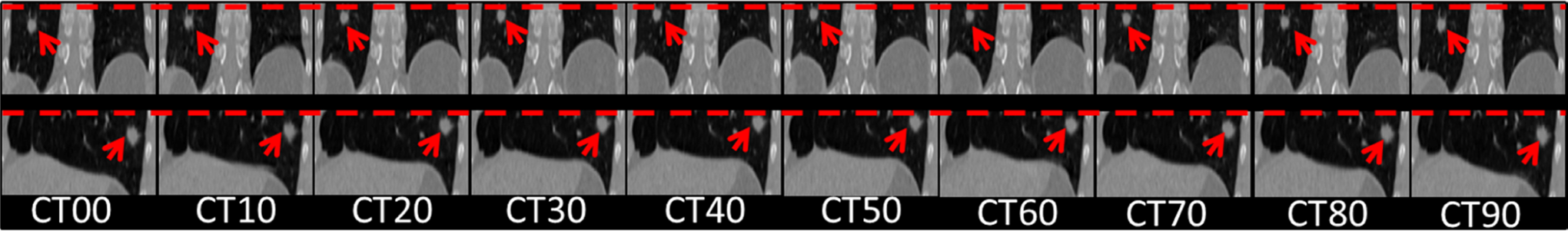
Representative 4D CT images in coronal and sagittal views of patient 1. Dash lines are drawn to facilitate the visualization of respiratory motion. Arrows indict the tumor. 4D CT indicates 4-dimensional computed tomography.
A 3-level hierarchy same as the DD algorithm was used for the registration procedure with a decreasing Young modulus schedule of (1.00, 0.10, 0.01) and a constant Poisson ratio of (0.5, 0.5, 0.5). The FEM parameters were chosen according to the previous literature 19 by considering balancing the registration accuracy and the convergence speed of the algorithm.
Quantitative Evaluation of Registration
During the registration process, the DVFs relative to the reference image (CT00) were calculated by the 2 algorithms. To demonstrate the ability of these 2 algorithms visually, each DVF was delineated and compared.
Four commonly used metrics, NMI, SSD, dM_H, and registration speed were employed to compare the efficiency of the 2 registration methods. The NMI can be calculated as follows:

where H(I1), H(I2), and H(I1,I2) denote the information entropy of 2 images and the joint entropy, respectively. The greater NMI between 2 images, the better alignment they have.
The SSD between 2 images is one of the simplest voxel similarity measures, and it is minimized during registration. For voxel location 
The Hausdorff distance which can measure the distance of two images from each other was commonly used as the similarity metric to evaluate the registration results. Given 2 images I and J, it is defined as:
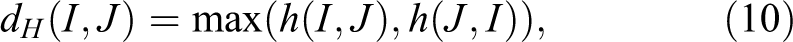
where h(I, J) and h(J, I) are the Hausdorff distance of I → J and J →I, respectively, and defined as:


where i and j represent pixels in image I and image J, respectively, and
Since the traditional Hausdorff distance is sensitive to image noise, in this study, we used the modified Hausdorff distance proposed by Dubuisson et al,
22
which can avoid the deviation caused by the interference of some noisy pixels. The modified Hausdorff distance (dM_H) was defined as:

where NI is the number of feature points in image set I.
The rGTV between reference image and deformed phase images using 2 methods was also measured and compared. The GTVs on the reference image (GTV
r
) and deformed phase images (GTV
f
) were delineated by an oncologist. The rGTV value was calculated as follows:
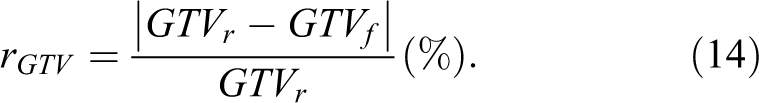
In our study, images at phase
Results
Figure 2 shows a representative 4D CT data set of patient 1. From the tumor locations in the lung at 10 phases, we can observe the respiratory motion over the breathing cycle.

Examples of the DVFs relative to a specific reference phase (CT00) calculated by the DD algorithm (a) and the FEM-based algorithm (b). DVF indicates displacement vector field; DD, diffeomorphic demons; FEM, finite element method.
For each patient, 9 phase images were registered to the reference (CT00), and 9 sets of DVF recording the movement of each voxel in phase images to corresponding voxel in the reference were obtained. Figure 3 demonstrates the DVFs in the lung area obtained from the DD algorithm (Figure 3A) and the FEM-based algorithm (Figure 3B) in patient 1. It was visible that the corresponding displacement fields showed differences between the 2 methods, with greater displacements appearing in the FEM-based algorithm.
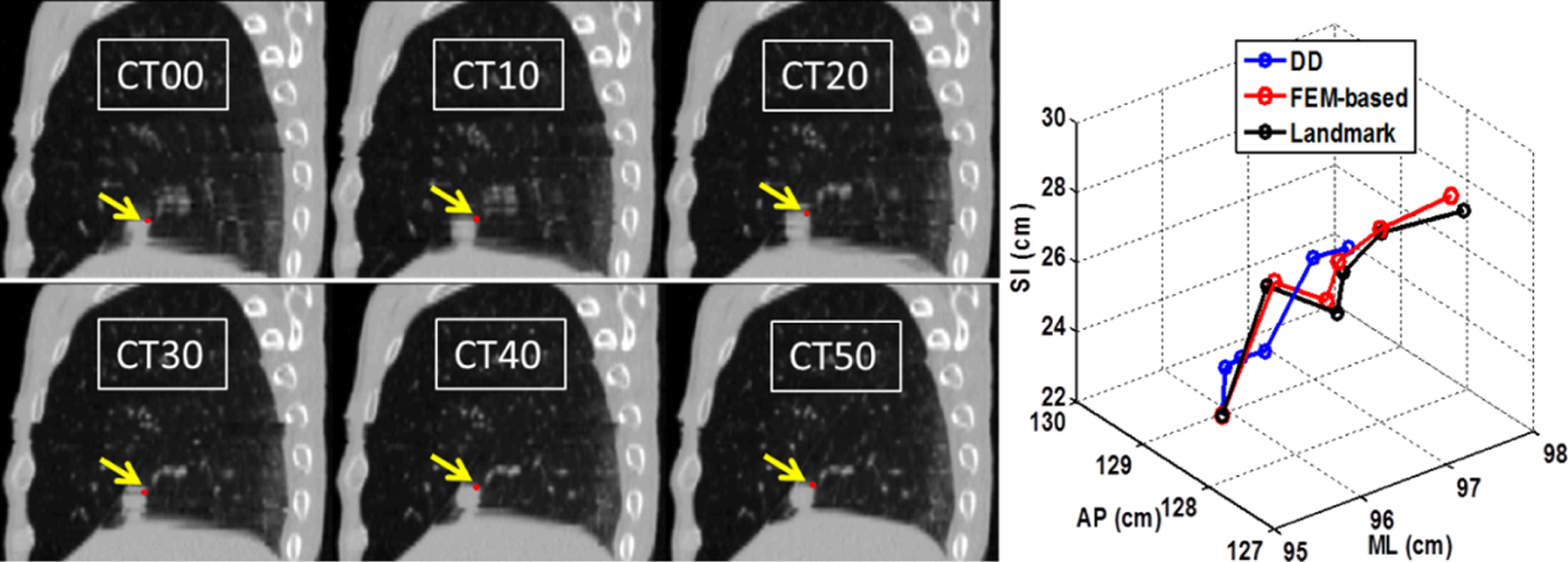
Examples of the displacement trajectories (right) of a voxel in the lung tumor (indicted by arrows) spanning the phases from maximum inhalation (CT00) to maximum exhalation (CT50) determined by the 2 DIR algorithms compared with a landmark trajectory treated as a golden standard. The tracked feature point was indicted by the red points. DIR indicates deformable image registration (Color version of the figure available online).
Figure 1 shows an example of the displacement trajectory (right) of a pixel identified by a radiation oncologist in the tumor spanning the phases from maximum inhalation (CT00)to maximum exhalation (CT50) using the 2 DIR methods. The tracked points were indicted by red points. We can observe that the displacement trajectories show large differences.
Figure 4 shows an example of sampled DVFs between CT50 and CT00 confined in the lung area in coronal plane. As aforementioned, CT00 (a) served as the reference image to obtain the DVFs calculated by the FEM-based (b) and DD (c) algorithms.

The coronal DVFs confined in the lung area calculated by the FEM-based (b) and DD (c) algorithms for patient 1. The CT image at phase T = 0% (a) was used as the reference image to derive the DVFs. DVF indicates displacement vector field; FEM, finite element method; DD, diffeomorphic demons, CT, computed tomography.
To validate the registration efficacy, we calculated the subtractions of phase images before/after registration and reference image. Figure 5 gives an example of subtraction images in 3 orthogonal directions (superior–inferior, anterior–posterior, and medial–lateral) between

Representative subtractions of reference image and phase image before (b) and after using the DD algorithm (c) and the FEM-based (d) algorithm. The first line (a) shows the fusion images of reference image (
Table 2 summarizes the measurement results of alignment between CT00 and CT50 from all the patients. Of all patients, the FEM-based registration method showed more accuracy than the demons-based method. The means (±standard deviation [SD]) of NMI were 0.70 (±0.10), 0.82 (±0.06), and 0.85 (±0.07) before and after registration using the DD and FEM-based algorithms, respectively. The means of SSD were 0.016 (±0.010), 0.011 (±0.009), and 0.009 (±0.009) before and after registration using the DD and FEM-based algorithms, respectively. The means of dH were 0.17 (±0.13), 0.06 (±0.03), and 0.04 (±0.03), before and after registration using the DD and FEM-based algorithms, respectively. The means of rGTV were 4.5 (±1.2)% and 3.3 (±1.1)% after using the DD and FEM-based algorithms, respectively. However, the FEM-based algorithm costs a longer time on the registration compared to the DD algorithm (35.8 vs 25.3 minutes). As aforementioned, the FEM-based algorithm solved the problem on an element-by-element basis by partitioning the problem domain into elements controlled by nodes and assembles the contributions of each element into a global system of equations. So the computational complexity is large when using high-resolution FEMs.
Comparison of NMI, SSD, dH_M, and Running Time on Registration Between

Abbreviations: NMI, normalized mutual information; SSD, sum of squared-intensity difference; dH_M, modified Hausdorff distance; dH, Hausdorff distance; DD, diffeomorphic demons; FEM, finite element method; rGTV, ratio of gross tumor volume; SD, standard deviation.
Table 3 summarizes the averaged measurement results over 9 phase bins from all the patients. On average, the FEM-based registration method showed more accuracy than the demons-based method: The means of NMI were 0.75 (±0.08), 0.86 (±0.05), and 0.90 (±0.05) before and after registration using the DD and FEM-based algorithms, respectively. The means of SSD were 0.008 (±0.06), 0.006 (±0.003), and 0.003 (±0.002) before and after registration using the DD and FEM-based algorithms, respectively. The means of dH were 0.91 (±2.5), 0.04 (±0.02), and 0.03 (±0.03) before and after registration using the DD and FEM-based algorithms, respectively. The means of rGTV were 3.9 (±1.1)% and 2.9 (±1.1)% after using the DD and FEM-based algorithms, respectively. However, the FEM-based algorithm costs a longer time compared with the DD algorithm (32 vs 22 minutes).
Comparison of NMI, SSD, dH_M and Running Time Measured by Averaging Over 9 Phases Using the DD and the FEM-Based Algorithms for All Patients.

Abbreviations: NMI, normalized mutual information; SSD, sum of squared-intensity difference; dH_M, modified Hausdorff distance; DD, diffeomorphic demons; FEM, finite element method; dH, Hausdorff distance; rGTV, ratio of gross tumor volume; SD, standard deviation.
Discussion and Conclusion
In this study, we evaluated the accuracy of the DD and FEM-based algorithms for 4D CT registration in 10 patients with lung cancer. The results demonstrated that the FEM-based algorithm showed greater ability in aligning 2 images than the DD algorithm. However, this algorithm requires a longer time mainly due to the ideal of nonuniform element partition. This work provided a fundamental research for DIR in radiation therapy.
This pilot study included a limited number of patients and assessed only patients with lung cancer. A larger pool of patients is needed in future studies to answer the following questions: (1) Is this comparison effective for cancers in other locations treated by radiation therapy, such as liver or esophageal cancers? (2) Are there any other methods to better evaluate the accuracy of DIRs? (3) How does the level of resolution affect the convergence of the cost function since we only employed three levels in registration procedure? There is no controversy about the fact that the deformation of lung is more easily affected by respiratory motion compared with liver or esophagus referred earlier. However, it could be estimated that the comparison results might be different if liver or esophagus was used to validate the performance of the 2 presented DIR algorithms due to the suboptimal contrast of liver 4D CT and the small deformation of esophagus 4D CT. In addition, poor soft-tissue contrast in liver 4D CT will definitely affect the selection of landmarks using the APRIL software.
Quantitative validation of a DIR algorithm is difficult due to the lack of a golden standard. Recently, there is no established method to validate the performance of registration algorithm. In our study, we qualitatively evaluated the DD algorithm and the FEM-based algorithm by comparing the magnitudes of motion vectors between the phase images and the reference image. In addition, the subtractions of the reference and deformed images calculated by the DD algorithm and the FEM-based algorithm were generated for directly visualized evaluation. Furthermore, 3 commonly used measurement metrics, NMI, SSD, and dH_M were chosen to quantitatively validate the registration algorithms. All measurements were calculated before and after deformable registration between the reference image and the phase image, which could be used for determining the more accurate algorithm. A more robust method to evaluate the registration performance may be achieved by introducing a golden standard, such as anatomic landmarks, fiducial markers, or contour positions. 13 Furthermore, we can evaluate the efficiency of the registration according to different parts of the lung, since we observed that the lower lung had a greater motion compared with the upper lung.
The material properties for the lung were determined according to the previous literature.
13,19
Brock et al
13
indicated that the optimal value of Poisson ratio (v) was difficult to determine for the lung, since trends varied between patients as well as between medial–lateral, anterior–posterior, and superior–inferior directions. The optimal value of v was assigned to 0.45 across all the patients in our study as indicated in Brock study. So the differences in optimal v between different directions and different patients could introduce potential errors. It is of great interest in our future work to investigate the potential benefit of different material properties to enhance fixed boundary conditions. Another drawback was the computational costs of the FEM-based registration method. In the current implementation, the in-plane 4D CT data set consisting of 512 × 512 pixels, with the initial element size of 16 × 16 requiring more than 1 hour using a standard PC (Inter(R) Core(TM) i7-3770 central processing unit @ 3.40 GHz and 8 G RAM). Therefore, multiresolution technique was used in this study to speedup the computation of the FEM-based algorithm. The computation time had been reduced to a half-time, which was comparable with the DD algorithm and meanwhile contained good accuracy. In addition, we found that, for both algorithms, the registration accuracy gradually improved and the convergence speed gradually slowed with the increase in the SD (σ) of Gaussian kernel. However, after σ was greater than 1, both the convergent speed and the registration accuracy were very similar with different
The deformation and distortion of thoracic organs and tumors are mainly caused by respiratory motion. With the development of 4D CT techniques, we can acquire volumetric images combined with the breathing signals throughout the breathing cycle. In 3-dimensional (3D) treatment planning, the gross target volume is delineated manually by the oncologists in the planning system and then expanded to a clinical target volume (CTV) by adding a margin to cover expected microexpansion. Furthermore, the CTV is expanded to the planning target volume by adding an internal margin (IM) according to the clinical criterion by considering the target movement induced by the respiratory motion. The IMs are usually determined according to experiences or reported literatures. These margins are relative to cancer, category specific rather than patient specific. Thus, adding safety margins using this method is approximately unsafe. The advent of 4D CT technique has solved these problems and can be used for assessing patient-specific breathing motion and adding individual safety margins.
Furthermore, the DIR algorithms can be used to evaluate the dosimetric effect of respiratory motion on dose delivered to the tumor target and organs at risk (OARs).
23
–27
Calculation of 3D dose in radiotherapy usually utilizes 1 static phase (breath-hold 3D CT, end-exhale or end-inhale of 4D CT, average intensity projection, and maximum intensity projection). A major disadvantage is insufficient evaluation of the dose distribution to tumor target and OARs. Recently, we can perform 4D dose calculation by transferring the DVFs obtained from the DIR procedure to a 3D dose matrix calculated from 3D plans on the exhale or inhale CT of 4D CT images. Deformed phase dose on the reference image was calculated by deforming the 3D phase dose on each phase with the corresponding phase DVF, which can be calculated as follows:
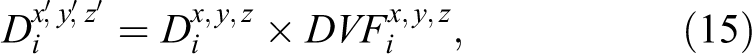
where the parameters 
where
So choosing a reliable and high-efficient DIR algorithm for the 4D images is crucial and significant for radiation therapy. It is of great interest to evaluate substantial registration algorithms and determine the most effective one for the 4D CT alignment and validate our point-to-point dose accumulation in our future study.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is partly supported by Natural Science Grant of Shandong Province (ZR2010HM010), National Natural Science Foundation of China (61201441), National Natural Science Foundation of China (81272699) as well as an educational fund from the China Scholarship Council (CSC).
