Abstract
Introduction
Epithelial-mesenchymal transition (EMT) is a key driver of tumor invasion and metastasis, which is closely associated with poor prognosis in patients with surgically resected lung cancer. Hypercapnic acidosis (HCA) is a common comorbidity in various lung diseases; however, its specific role in regulating EMT in lung cancer remains unclear. Acid-sensing ion channel (ASIC) genes have been implicated in tumor progression, but their expression patterns and prognostic value in lung cancer, as well as their involvement in HCA-mediated EMT regulation, require further investigation.
Methods
The expression levels of ASIC genes and their prognostic significance were analyzed in lung adenocarcinoma and lung squamous cell carcinoma using the Gene Expression Profiling Interactive Analysis (GEPIA) database. A549 lung cancer cells were exposed to HCA conditions (10% CO2, pH 6.69 ± 0.02) for five days to induce EMT phenotypes. Cell proliferation, migration, and invasion capacities were evaluated using corresponding functional assays. The expression levels of EMT-related markers (E-cadherin and vimentin) and ASIC3 were quantified by immunohistochemical staining, western blot analysis, and reverse transcriptase-quantitative polymerase chain reaction (RT-qPCR). Additionally, amiloride was used to inhibit ASIC3 expression to verify its regulatory role in HCA-induced EMT.
Results
Bioinformatics analysis showed that overexpression of ASIC3 mRNA was significantly correlated with reduced overall survival in lung cancer patients (P < .05). In vitro experiments demonstrated that HCA exposure significantly upregulated ASIC3 expression (P < .01) and promoted EMT in A549 cells, as evidenced by downregulated E-cadherin expression and upregulated vimentin expression. Moreover, HCA significantly enhanced the migration and invasion abilities of A549 cells (P < .01). Importantly, inhibition of ASIC3 expression by amiloride reversed all these HCA-induced effects, including the alterations in EMT markers and the enhancement of cell migratory/invasive capacities.
Conclusion
The HCA microenvironment induces EMT in A549 lung cancer cells through the activation of ASIC3. These findings suggest that ASIC3 may serve as a potential therapeutic target for the treatment of lung cancer, which could help improve clinical outcomes by inhibiting tumor invasion and metastasis mediated by EMT.
Introduction
Among the malignant tumors, the mortality rate of lung cancer ranks first in the world. The incidence and mortality of lung cancer in China have also continued to increase in the past few years.1,2 According to the prediction of the World Health Organization, the number of deaths from lung cancer will exceed one million by 2025 and the number of patients affected by this disease will rank first in the world compared with other cancer types. 3 Lung cancer is a malignant tumor which exhibits very poor prognosis. When >80% of patients with lung cancer are admitted to hospital for treatment, they have lost the optimal time for surgery and multidisciplinary treatment. 4 Currently, lung cancer treatment is mainly based on surgical resection, supplemented by radiotherapy, chemotherapy, nanomedicine delivery and robotic exoskeletons,5–9 however, the efficacy is limited in the advanced stages of the disease. Although targeted therapy aims at specific mutations it is prone to drug resistance. Although immunotherapy has revolutionized treatment, 10 it only benefits certain patients. Therefore, there is an urgent requirement to explore novel mechanisms to improve prognosis. Lung cancer exhibits the highest number of cancer-related deaths in the world, due to tumor cell metastasis.11,12 Metastasis is a multi-step process, which includes cancer cell epithelial-mesenchymal transition (EMT), separation from the primary tumor, migration to nearby tissues or distant organ tissues, and eventually tumor cells spread. 13 Once the cancer cells metastasize, treatment will become very difficult and the possibility of survival decreases. 14 Therefore, EMT plays a major role in tumor invasion and metastasis, which is associated with poor survival in surgically resected lung cancer.15,16 However, the exact mechanism of EMT in lung cancer has not been fully elucidated.
The tumor microenvironment is an extremely complex internal environment system, composed of tumor interstitial cells, cell metabolites, and various physical and chemical factors such as power of hydrogen (pH), hypoxia, and interstitial fluid pressure. 17 Recent studies have been showed that acidity is an inherent feature of the tumor microenvironment.18,19 xtracellular acidity is a favorable factor for tumor progression and metastatic spread, which is involved in EMT. 20 Hypercapnic acidosis (HCA) is a common clinical comorbidity of lung disease and it has been shown to improve the prognosis of patients with acute lung injury/acute respiratory distress syndrome (ALI/ARDS).21,22 HCA has certain harmful effects, such as suppression of innate immunity/host defense.23,24 HCA is a common complication of patients with lung cancer, yet it is not clear how this condition affects lung cancer. In the present study, the exact way by which the HCA microenvironment affects the EMT of human lung adenocarcinoma cells (A549 cells) was investigated.
Acid-sensing ion channels (ASICs) are members of the epithelial sodium channel family, activated by extracellular protons (H+).25,26 These channels mainly transit through Na+ and are less permeable to K+ and Ca2+. Six ASICs have been cloned to date: ASIC1a, ASIC1b, ASIC2a, ASIC2b, ASIC3 and ASIC4. 27 It has been reported that the acidic condition stimulates tumor cell invasion which may involve lysosomal proteases, extracellular matrix degradation and induction of normal cell death. 28 ASIC1, ASIC2, and ASIC3 are expressed in A549 cells at both the mRNA and protein levels, and extracellular acid stimulation elicits ASIC-like currents. ASIC-Ca2+ signaling mediates extracellular acidosis-induced proliferation and migration of lung cancer cells. 29 ASICs can serve as prognostic markers and therapeutic targets for lung cancer. 29 Moreover, ASIC1 and ASIC3 promoted EMT of cancer cells during acidic microenvironment by transducing the acidic to elevation of intracellular Ca2+ concentration, the mechanism may be that acid-induced EMT of cancer cells is mediated through the ASIC1/3-[Ca2+]i-RhoA axis, thereby promoting cancer cell metastasis. 30 It has been showed that ASIC3 plays an important role in lipopolysaccharide-induced ALI in vivo study. 31 However, until now it has not been examined on the effects of ASICs on lung cancer. In the present study, bioinformatics data analysis was performed and the data indicated that the mRNA expression levels of ASIC3 may be useful for the prediction of the overall survival (OS) of patients with lung cancer. Furthermore, the hypothesis of whether exposure of lung cancer cells to the HCA microenvironment induces EMT was examined. The effects of ASIC3 and the potential to increase metastasis of A549 lung cancer cells were also investigated.
Materials and Methods
Ethical Approval
All tissue sample sections used for immunohistochemical experiments in this study were obtained from the Pathology Specimen Bank of the First Affiliated Hospital of Nanchang University. And approved by the Medical Ethics Committee of the First Affiliated Hospital of Nanchang University (IIT[2022] NO.099, Nanchang, Nov 2022). All experiments were conformed to the National Research Council's Guide.
Gene Expression Profiling Interactive Analysis (GEPIA) Dataset
To analyze the expression levels of the ASIC genes in lung cancer, GEPIA was used which is an interactive web server for analyzing the RNA sequencing expression data. 32 The tumor data of lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC) were selected to analyze the expression levels of the corresponding genes on box plots and pathological stage plots, considering adjusted P < .01 and log2FC > 1 as a significant difference threshold.
The Kaplan-Meier Plotter
The prognostic value of the mRNA levels of the ASIC genes in patients with lung cancer was evaluated using the Kaplan-Meier Plotter database (http://kmplot.com/analysis/index.php?p=service&cancer=lung). 33 A total of 2437 patient samples were split into two groups by median expression (high vs low expression) and assessed by a Kaplan-Meier survival plot. The OS, progression-free survival (FP) and post progression survival (PPS) of patients with lung cancer were analyzed with the hazard ratio (HR), 95% confidence intervals (CI) and logrank P value.
Immunohistochemical (IHC) Analysis
IHC staining was performed on the paraffin-embedded cancer tissues. Paraffin sections of 3-μm thick were cut and probed by 0.3% hydrogen peroxide for 30 min and the sections were incubated with the primary antibodies against ASIC1, ASIC2, ASIC3 and ASIC4 (ASIC1 Polyclonal Antibody, PA5-66217. ASIC2 Polyclonal Antibody, PA5-143911. ASIC3 Polyclonal Antibody, PA5-99026. ASIC1 Polyclonal Antibody, PA5-139833. Thermo Fisher Scientific Inc.) (1:100 to 1:150 in PBS, overnight at 4 °C). Subsequently, the sections were incubated with the biontinylated secondary antibody (1:200; Santa Cruz Biotechnology, Inc.) for 30 min at 37 °C, covered by 3,3’-diaminobenzidine and mounted with Vectashield mounting medium (Vector Laboratories, Inc.). Subsequently, all fields were observed under light microscopy (Olympus Corporation). IHC scoring system, We used the semi-quantitative H-score (histological score) for IHC staining evaluation, which combines the staining intensity (0: negative, 1: weak, 2: moderate, 3: strong) and positive cell rate (0: 0%, 1: 1%-25%, 2: 26%-50%, 3: 51%-75%, 4: 76%-100%). The final H-score was calculated as staining intensity × positive cell rate (range: 0-12), and samples with H-score ≥4 were defined as high ASIC expression, and <4 as low expression. All IHC sections were independently evaluated by two senior pathologists who were blinded to the clinical and survival data of the patients.
Cell Culture and Reagents
Human lung adenocarcinoma A549 cells (LZ - PX10853, Shanghai Lianzu Biotechnology) were purchased from ATCC Cell Bank and were cultured in Dulbecco's modified Eagle's medium (DMEM) with 10% fetal bovine serum (FBS) at 37 °C, in the presence of 5% CO2 and 95% air. To assess the effects of the HCA microenvironment, the cells were cultured in 10% CO2 and 90% air, 34 the pH and the pressure of CO2 in the medium were 6.69 ± 0.02 and 71.01 ± 3.17 mm Hg, respectively. To block ASIC3, amiloride (Sigma-Aldrich; Merck KGaA) was added to the medium at the final concentrations of 100 μM for 48 h. The cells were seeded in 12-well tissue culture plates. The rabbit anti-human antibody ASIC3 was purchased from Cell Signaling Technology Inc., and anti-rabbit secondary antibodies were purchased from KangChen Bio-tech. The BCA protein assay kit was purchased from Beyotime Institute of Biotechnology.
Immunofluorescence Analysis
A549 cells were plated in 24-well plates and fixed with 0.1% Triton for 30 min at 4 °C. Following washing with PBS, A549 cells were treated with 5% bovine serum albumin for 60 min at room temperature and subsequently stained with a primary antibody for ASIC3 at 4 °C overnight. The primary antibody was mouse anti-ASIC3. The secondary antibody fluorescein isothiocyanate (goat anti-mouse, Abcam) was diluted at a ratio of 1:200 in PBS and incubated for 60 min at 4 °C with the cells. The slides were mounted onto slides with 1% 4’6-diamidine-2’-phenylindole dihydrochloride (DAPI) for 10 min. The cell images were visualized using a scanning confocal fluorescence microscope (Olympus Corporation).
Proliferation Assay
A549 cells were plated on 6-well plates at a density of 1 × 104 cells in complete medium. The following day, the cells were cultured in a low percentage of FBS (0.5%) and in the presence or absence of 10% CO2 and 90% air. The cells were harvested at the indicated time points and the total number of living cells was counted by using an automatic cell counter (NanoEnTek).
Migration Assay (Chemotaxis Assay)
A549 cells were serum-starved for 12 h and dissociated using trypsin-EDTA. The cells (1 × 105) were resuspended in serum-free DMEM and loaded onto the upper chamber of a ChemoTx system (8 μm pore size; Neuro Probe Inc.) pre-coated with type I collagen. Complete DMEM (10% FBS) was added to the lower chamber to establish a chemotactic gradient. Following incubation for 4 or 6 h at 37 °C (5% CO₂), non-migratory cells on the membrane upper surface were removed by cotton swab scraping. Migrated cells were fixed with 4% paraformaldehyde, stained with DAPI and quantified using a fluorescence microscope (Olympus Corporation) at 10× magnification.
Invasion Assay
Matrigel invasion assays were conducted using BD Biosciences Matrigel-coated 24-well Transwell inserts (8 μm pore size). A549 cells (1 × 105) suspended in serum-free DMEM (0.5% FBS). The cells were seeded into the upper chamber, while complete DMEM (10% FBS) was added to the lower chamber to establish a chemotactic gradient. Following 24 h incubation at 37 °C (5% CO₂), non-invading cells on the upper membrane surface were gently removed with cotton swabs. Invaded cells adherent to the lower membrane surface were fixed in 4% paraformaldehyde (PFA), stained with DAPI and quantified using a fluorescence microscope (Olympus Corporation) at 10× magnification.
Reverse Transcription-Quantitative PCR (RT-qPCR) Analysis
Total RNA was extracted from cultured cells using the EastStep Total RNA Extraction Kit (Promega Corporation) following the manufacturer's protocol. cDNA synthesis was performed with the Transcriptor First-Strand cDNA Synthesis Kit (TransGen Biotech) using 1 μg RNA as template. Quantitative PCR was carried out on a Bio-Rad DA7600 Real-Time PCR System using TransStart Green qPCR SuperMix (TransGen Biotech Co., Ltd) with the following thermal cycling parameters: Initial denaturation at 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s and 60 °C for 30 s Gene expression levels were normalized to GADPH using the 2−ΔΔCt method. The primers were designed by the company (Takara, Dalian, China), and the reference sequence of ASIC3 primer is registered as NM_001130069.2. The primer sequences of ASIC3 forward: 5’- CAGCCCTGTGGACCTGAGAA - 3’, reverse, 5’- CGCCCTTAGGAGTGGTGAGC - 3’, GADPH forward: 5’- CCTGAACGGGAAGCTCACTGG - 3’, reverse, 5’- CCTCCACCACCCTGTTGCTGTA - 3’.
Western Blot Analysis
Cell lysates were prepared in RIPA buffer (20 mM Tris-HCl, pH 7.4, 1 mM EGTA/EDTA, 1% Triton X-100, 1 mM Na₃VO₄, 10% glycerol, 1 μg/mL leupeptin, 1 μg/mL aprotinin). Protein concentration was determined using the bicinchoninic assay. The samples (30 μg/lane) were separated by SDS-PAGE (8%-15% gradient gels) and transferred to polyvinylidene fluoride membranes. The membranes were blocked in 5% skim milk/TBST for 1 h at room temperature, followed by overnight incubation at 4 °C with primary antibodies against E-cadherin (1:1000), vimentin (1:500) and β-actin (1:2000). Following washing, the membranes were incubated with horseradish peroxidase-conjugated secondary antibodies (1:5000) for 1 h at room temperature. The protein bands were visualized using enhanced chemiluminescence substrate and quantified with ImageJ software (National Institutes of Health).
Cell Transfections
siRNA duplexes targeting ASIC3 (siB121224135855-1-5), and negative control (siN0000002-1-5) were synthesized and purified by RiboBio (Ribobio Co.,Guangzhou, China). RNA oligonucleotides were transfected using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) at a final concentration of 50 nM according to the manufacturer's instructions. For stable inhibition of ASIC3 expression, the sequence of siRNA was structured to lentiviral vector. The siRNA lentiviral vector pLV-shRNA-Puro (RiboBio Co., Ltd, Guangzhou, China) was used to construct the shRNA expression vector. The target siRNA sequences (sense: 5'-GCTGATGCTGATCGTACGA-3’; antisense: 5'-TCGTACGATCAGCATCAGC-3’) were inserted into the vector to generate the shRNA template.For vector digestion, 1 μg/μL of pLV-shRNA-Puro was mixed with restriction endonucleases Age I (10 U/μL) and EcoR I (10 U/μL) in the presence of 10× digestion buffer. The total volume of the reaction mixture was adjusted to 20 μL, followed by incubation at 37 °C for 2 h. The linearized vector was then purified using 1% agarose gel electrophoresis. Ligation reaction was performed in a total volume of 10 μL containing 50 ng of purified linearized vector, synthesized shRNA double-stranded fragments, T4 DNA ligase (2 U/μL, Thermo Fisher Scientific), Lipofectamine 2000, and 10× T4 Ligation Buffer. The ligation mixture was incubated overnight at 16 °C. Cells were cultured at normal medium for additional 48 h after transfection until collected for further assay.
Statistical Analysis
All data were presented as mean ± SD Group comparisons were analyzed with the unpaired Student's t-test. One-way analysis of variance was used for comparing multiple groups. The Bonferroni test for the pairwise comparisons after one-way analysis of variance. P < .05 was considered to indicate a statistically significant difference. SPSS 23.0 (IBM Corp.) was used for statistical analysis.
Results
Relationship Between the mRNA Levels of ASICs and the Clinicoathological Parameters of Patients with Lung Cancer
To investigate the mRNA expression levels of the ASIC genes in lung cancer and normal lung tissues, the GEPIA dataset (http://gepia.cancer-pku.cn/) was used. The results indicated that the expression levels of ASIC1, ASIC2, ASIC3 and ASIC4 were not high in tumor and normal lung tissues, whereas the mRNA expression levels of ASIC1 and ASIC3 in LUSC tissues were higher compared with those noted in normal tissues (Figure 1A, P < .05). Subsequently, the expression levels of the ASIC genes were compared with the tumor stage for LUAD and LUSC. The results indicated that the expression levels of the ASIC2 group varied significantly, whereas the expression levels of the ASIC1, ASIC3 and ASIC4 groups were not significantly different (Figure 1B). The histological subtype of the included specimens was LUSC, and the tumor stage was II-III. All patients were pathologically diagnosed with lung cancer and had not received preoperative radiotherapy, chemotherapy or immunotherapy. IHC staining was performed to assess the expression levels of the ASIC proteins in lung cancer tissues and in their counterparts. The results indicated that ASIC1 and ASIC3 proteins were more highly expressed in the lung cancer tissues than in the normal lung tissues (Figure 1C). However, all the expression levels of ASICs were not high.

The expression of ASICs in lung cancer. A. The expression of ASICs mRNA levels in LUAD and LUSC (GEPIA). B. Correlation between ASICs expression and tumor stage in lung cancer patients (GEPIA). C. The expression of ASICs in lung cancer tissue by Immunohisto-chemical staining.
Prognostic Value of the mRNA Expression Levels of the ASIC Genes in Patients with Lung Cancer
To determine the prognostic values of the mRNA expression levels of the ASIC genes in patients with lung cancer, the Kaplan-Meier Plotter database (http://kmplot.com/analysis/index.php?p=service&cancer=lung) was used to analyze the survival between samples from patients with high expression and those with low expression. The results indicated that the increased ASIC3 and ASIC4 mRNA levels were significantly associated with FP and OS (P < .05) and all the ASIC gene mRNA levels were not associated with PPS (P > .05; Figure 2). When analyzing the mRNA expression profiles of different ASICs in tumor versus normal tissues, combined with protein expression data, a distinct pattern emerged with ASIC3 standing out. Specifically, while ASIC1 and ASIC3 proteins were both found to be more highly expressed in lung cancer tissues compared to adjacent normal lung tissues, ASIC3 exhibited a stronger association with clinical outcomes: its mRNA levels, along with those of ASIC4, showed significant correlations with both FP and OS. Taken together with the results of the different ASIC gene mRNA expression levels in tumor and normal tissues, these findings suggested that the mRNA expression levels of ASIC3 may be useful for prediction of lung cancer patient OS.
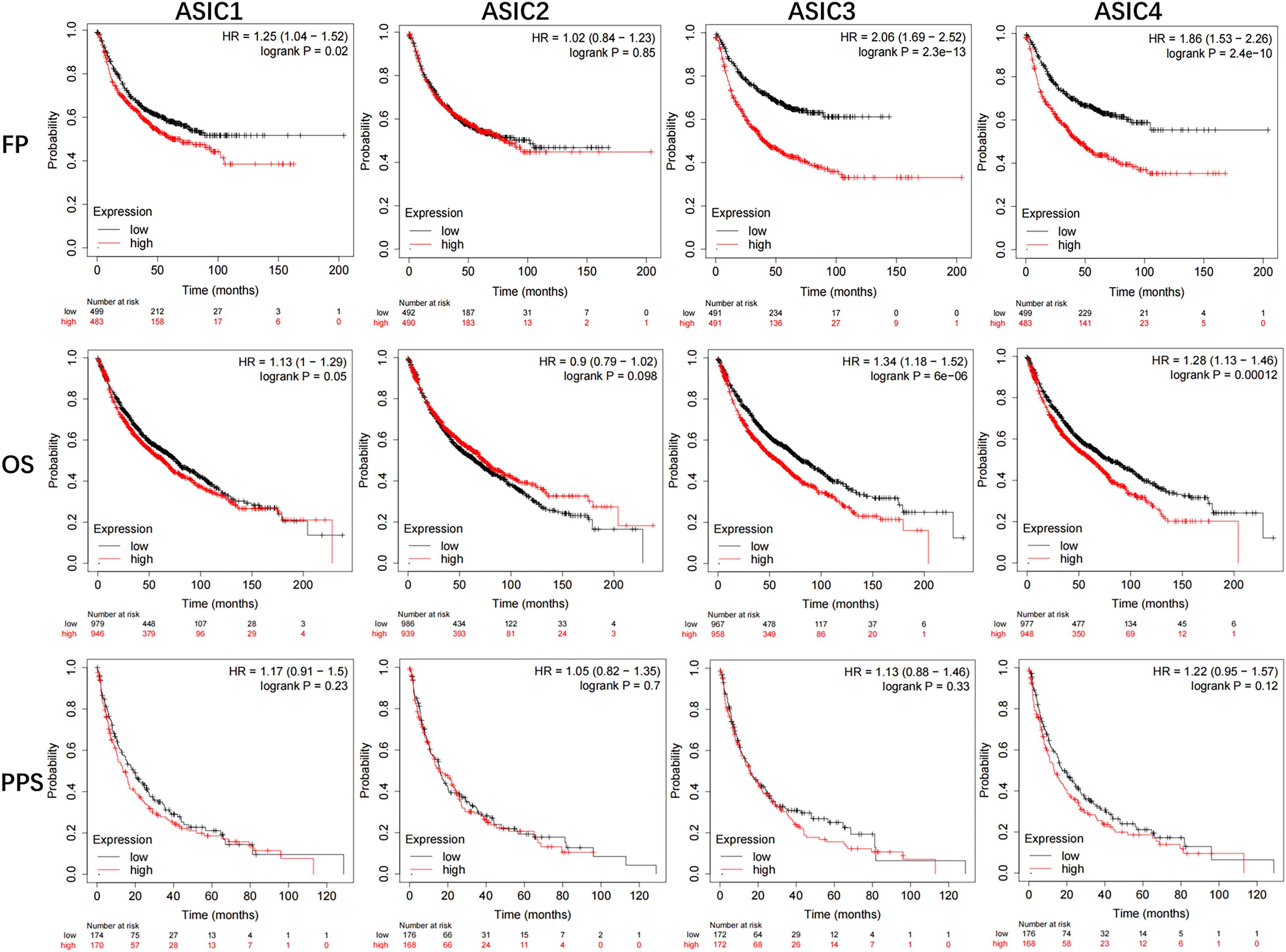
The prognostic value of mRNA level of ASICs factors in lung cancer patients (Kaplan-Meier plotter).
HCA Enhances the Expression of ASIC3 in A549 Cells
Exposure to a HCA microenvironment (10% CO2 + 90% air, pH 6.69 ± 0.02) caused a significant upregulation in ASIC3 expression in A549 cells compared with that noted in normoxic control cells (5% CO₂). Immunofluorescence analysis revealed a marked increase in ASIC3 protein levels in HCA-treated cells (Figure 3A); quantitative analysis confirmed a statistically significant fluorescence intensity elevation (Figure 3B). This upregulation in the protein levels was further corroborated by RT-qPCR, indicating significantly increased ASIC3 mRNA expression in HCA-exposed cells (Figure 3C). These findings collectively demonstrate that HCA-induced ASIC3 overexpression is a critical adaptive response in lung cancer cells.
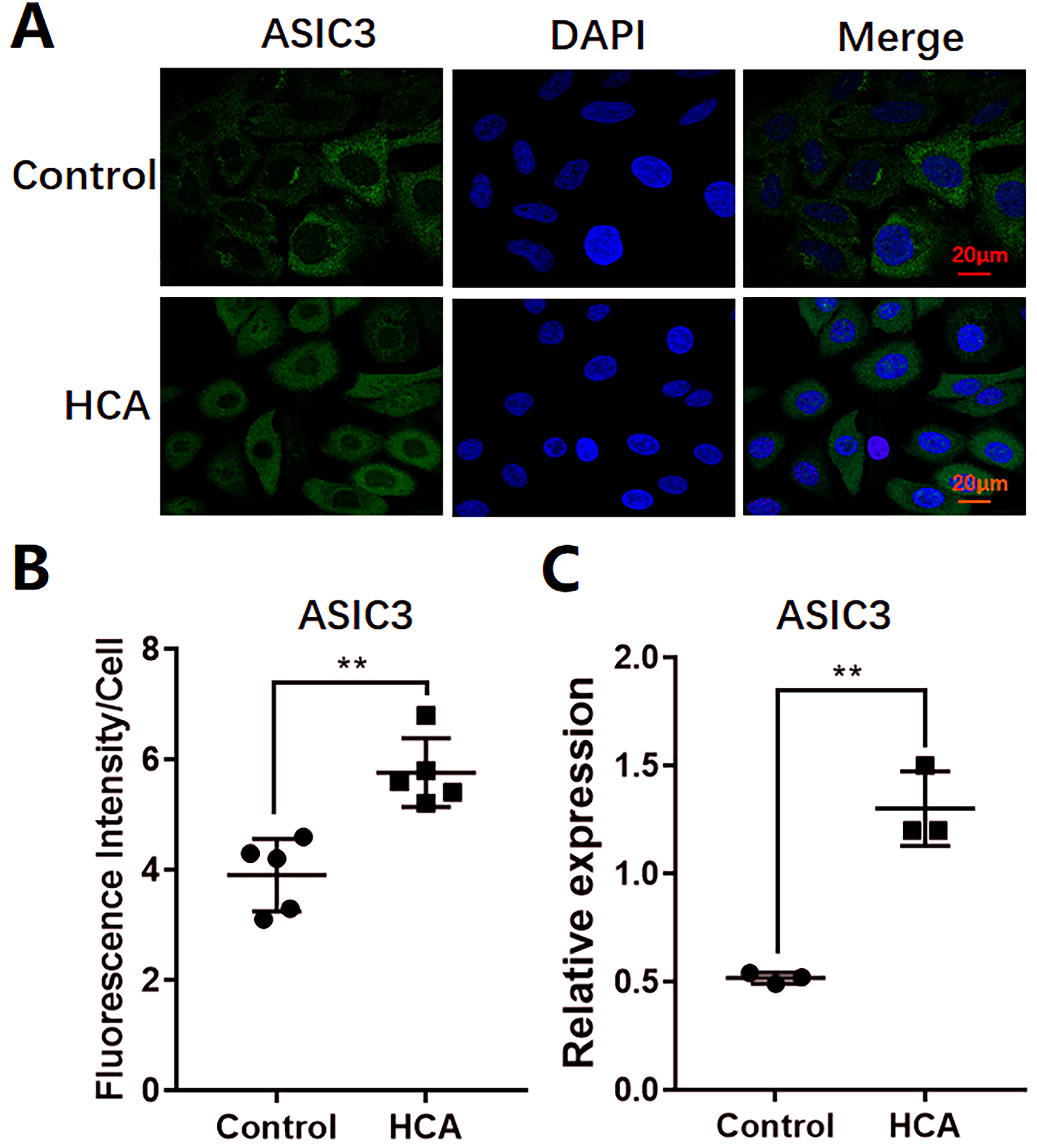
Effects of hypercapnic acidosis on the expression of ASIC3 in A549 cells. A. Representative images of ASIC3 expression in different groups of A549 cells (immunofluorescence staining). The green fluorescence shows ASIC3, the blue fluorescence shows the nucleus, and the combined pink shows the expression of ASIC3 in A549 cells. B. ASIC3 fluorescence intensity results were showed in each group. The data are shown as the mean ± sd. **P < .01, n = 5for each experiment. C. RT-qPCR analysis of the expression levels of ASIC3.The data are shown as the mean ± sd. **P < .01, n = 3 denotes independent biological replicates from separate cell cultures.
HCA Suppresses Proliferation and Enhances EMT of A549 Cells
To assess the metastatic potential of A549 cells under HCA conditions, their proliferative capacity was evaluated. As shown in Figure 4A and B, HCA exposure significantly suppressed A549 cell proliferation compared with that noted in normoxic controls. To determine whether HCA-induced EMT contributed to this proliferative inhibition, the migratory and invasive activities of the cells maintained under HCA conditions for 5 days were assessed (Figure 4C). A549 cells were only pretreated in the HCA microenvironment for 5 days to induce the EMT phenotype, and the subsequent functional assays for cell migration and invasion were all performed under the conventional culture condition with 5% CO₂. Prior to the migration and invasion assays, double detection of cell viability and the number of viable cells in each group was performed via trypan blue staining combined with an automated cell counter, ensuring that the initial number of viable cells seeded in each group was uniformly 1 × 105 with a viability rate of over 95%. Quantitative analysis revealed that A549 HCA cells (A549 HCA refers to A549 cells exposed to HCA conditions) exhibited markedly enhanced migratory and invasive (Figure 4D) capacities compared with those noted in A549 control cells. These findings suggest that HCA induces a phenotypic switch characterized by reduced proliferation and increased metastatic competence, which is consistent with EMT-driven tumor progression.

Effect of HCA on the proliferation, migration, and invasion ability of A549 cells. A. A549 cells were treated for 5 days in the presence or absence of HCA. B. The proliferation of A549 cells was examined at the indicated time points.C. Schematic representation of the experimental protocol. A549Con and A549HCA cells of two groups were obtained by stimulation of original A549 cells with or without HCA for 5 days, respectively. Migration and invasion were assessed during the late stage of HCA stimulation. Stained with DAPI and observed with fluorescence microscope. D. Migration and invasion of two groups cells were measured as described in “Materials and Methods”.The data are shown as the mean ± sd. **P < .01, n = 3 denotes independent biological replicates from separate cell cultures.
HCA Induces EMT Through ASIC3 in A549 Cells
To investigate the functional role of ASIC3 in HCA-induced EMT, A549 cells were cultured under HCA conditions for 5 days. IHC analysis revealed a marked reduction in E-cadherin expression, which is an epithelial marker (Figure 5A and B). Consistent with these findings, western blot analysis confirmed a significant downregulation of E-cadherin (Figure 5D) alongside with a concurrent upregulation of Vimentin, a mesenchymal marker (Figure 5E). These results collectively indicated that the HCA microenvironment induced EMT in A549 cells. It is important to note that treatment with a selective ASIC3 antagonist significantly attenuated HCA-mediated EMT, as demonstrated by preserved E-cadherin expression and suppressed vimentin levels (Figure 5D and E). Functional assays further demonstrated that ASIC3 inhibition reversed the HCA-induced increase noted in cell invasive and migratory capacity. These data provide compelling evidence that ASIC3 plays a critical role in HCA-driven EMT progression in lung cancer.

HCA induces an epithelial-to-mesenchymal transition of A549 cells. A. A549 cells were incubated with HCA or amiloride (block ASIC3) and stained with the indicated antibodies of E-cadherin. Images were visualized under a microscope. Magnification, 20×. Scale bar, 20 μm. B. Statistical analysis of E-cadherin protein expressions was performed as above. C. A549 cells were stimulated withHCA or amiloride for 5 days, and cell lysates were subjected to western blot analysis with the indicated antibodies of E-cadherin and Vimentin. E-cadherin (D.) and Vimentin (E.) protein relative expressions. The data are shown as the mean ± sd. **P < .01. n = 3 denotes independent biological replicates from separate cell cultures.
siASIC3 Inhibits HCA-Induced EMT in A549 Cells
To further confirm the role of ASIC3 in HCA-induced EMT, we silence ASIC3 in A549 cells using siRNA. Successful silencing of ASIC3 protein and downregulation of ASIC3 mRNA were achieved (Figure 6A and B), consistent with the effects of ASIC3 inhibitors, ASIC3 silence resulted in significant alterations in EMT-related protein expression, as confirmed by Western blot analysis. Specifically, the expression of the epithelial marker E-cadherin was preserved, while the mesenchymal markers N-cadherin, Fibronectin, and Vimentin were significantly suppressed (Figure 6C and D).

siASIC3 inhibits HCA-induced EMT in A549 cells. A. Successful silencing of ASIC3 and downregulation of ASIC3 mRNA were achieved. B. The downregulation of ASIC3 mRNA was verified by RT-PCR. C. The expression levels of the epithelial marker E-cadherin, and mesenchymal markers N-cadherin, fibronectin, and vimentin were detected. D. Quantitative analysis of the relative protein expression levels was performed. The data are shown as the mean ± sd. **P < .01. n = 3 denotes independent biological replicates from separate cell cultures.
Discussion
HCA which occurs due to an excess of CO2 in the blood has been associated with improvement in patients with ALI/ARDS and the permissive and even therapeutic hypercapina have been proposed as treatment options for patients with ALI/ARDS.21–23 However, additional studies have shown that hypercapnia exhibits certain harmful effects, such as inhibition of alveolar epithelial function and epithelial cell repair, suppression of innate immunity/host defense and airway function. 24 In the present study, the effects of the HCA microenvironment were investigated on the EMT of A549 cells. Bioinformatics data analysis demonstrated that the mRNA expression levels of ASIC3 may be useful for prediction of lung cancer patient OS. Subsequently, the relationship between ASIC3 expression and the effects of HCA on EMT was examined in A549 lung cancer cells. The results indicated that the expression of ASCI3 was increased; moreover, the EMT of A549 cells was induced following exposure to a HCA environment (10% CO2 + 90% air, pH 6.69 ± 0.02); blocking the expression of ASIC3 could suppress these effects. Collectively, the results demonstrated that the HCA microenvironment promoted the EMT of human lung cancer A549 cells through ASIC3, which also may affect the OS of patients with lung cancer.
The acidic microenvironment is closely related to the proliferation,35,36 invasion and metastasis of tumors, 37 which is carried out mainly by inducing new angiogenesis, escaping immune surveillance, and EMT. Therefore, the targeted tumor microenvironment may become part of future anti-tumor therapy. 38 Previous studies have indicated that the tumor cells always in hypoxia environment for the tumor cells grows fast and the new blood vessel function is incomplete.39–41 However, a large number of studies have confirmed that tumor cells, even with sufficient oxygen supply, are mainly dependent on anaerobic glycolysis to obtain energy, which is known as the “Warburg effect”.42–44 Hypoxia is often associated with CO2 accumulation; therefore, the HCA microenvironment may play an important role in cancer progression and evolution. In the present study, Transwell assay and wound-healing assays were used to determine the HCA microenvironment effects in the EMT of lung cancer A549 cells. The results indicated that the HCA microenvironment could increase the invasive and migratory activities of A549 cells in vitro. However, it is not clear yet the exact way by which the HCA microenvironment affects the EMT of lung cancer A549 cells.
In the current study, bioinformatics analyses were performed to explore the mRNA expression patterns of ASIC family members in lung cancer tissues compared with those noted in normal lung tissues. The data retrieved from the GEPIA dataset indicated that the overall expression levels of the ASIC genes were relatively low in both tumor and normal lung tissues, with only ASIC1 and ASIC3 indicating significant differences in mRNA expression levels. These findings were further validated by IHC staining and the results were consistent with the data from the GEPIA dataset.
Subsequent investigations into the prognostic value of the mRNA expression levels of the ASIC genes in patients with lung cancer were performed using the Kaplan-Meier Plotter database. The data revealed that the elevated mRNA levels of ASIC3 and ASIC4 were significantly associated with both FP and OS. While these results suggest that ASIC3 mRNA expression may serve as a potential prognostic marker for the OS in patients with lung cancer, the prognostic significance of ASIC4, as well as of other ASIC family members, necessitates further exploration to clarify their specific roles in lung cancer pathogenesis. It is reasonable to hypothesize that these ASIC genes may be involved in distinct molecular pathways or exert context-dependent effects, which could be the focus of future studies.
The present research study focused on ASIC3 for in-depth analysis based on the fact that ASIC3 exhibited a significant difference in the expression levels between lung cancer and normal tissues, as confirmed by both bioinformatics and IHC evidence45,46; revious studies have demonstrated that ASIC1 and ASIC3 are involved in acid-induced EMT to promote tumor metastasis, 23 providing a theoretical basis for investigating the role of ASIC3 in lung cancer progression.
Consistent with this research direction, the experiments of the present study indicated that the HCA microenvironment upregulated ASIC3 expression in A549 cells and blocking of ASIC3 expression under HCA conditions significantly inhibited the invasive and migratory abilities of A549 cells. Collectively, these results suggest that ASIC3, which regulates HCA microenvironment-induced EMT in A549 cells, may at least partially contribute to lung cancer metastasis and affect patient survival outcomes. Notably, studies on pancreatic cancer models have shown that ASIC1/3 mediates acidity-induced EMT in pancreatic cancer cells via the [Ca2+]i/RhoA axis. 30 HCA induced by the Warburg effect and insufficient vascular perfusion,42–44 ASIC1/3 which are highly expressed on the membrane of pancreatic cancer cells, are directly activated. This activation triggers the rapid influx of Ca2+, leading to a significant increase in intracellular calcium concentration ([Ca2+]i). The elevated [Ca2+]i then activates the small GTPase RhoA via intermediaries such as calmodulin (promoting its conversion from the GDP-bound inactive state to the GTP-bound active state). Activated RhoA further stimulates its downstream effector. Ultimately, this cascade of events drives the EMT of cancer cells. Although the present research study has confirmed that ASIC3 mediates EMT in lung cancer cells, the association of its mechanism of action with the [Ca2+]i/RhoA axis remains to be clarified in future studies. Research has confirmed that ERK pathway activation occurs in acidic environments, 32 and ERK2 regulates EMT process of MCF10A cells through FoxO1, which enhances migration/invasion ability and reduces proliferation activity, 47 In this study, HCA inhibited the proliferation of A549 cells and promoted the migration and invasion by ASIC3. Whether the mechanism is related to ERK pathway activation remains to be further confirmed.
The present study is subject to several limitations. The study was confined to in vitro experiments using only the A549 cell line, without extending to other cell lines or establishing animal models for in vivo validation, which limits generalizability across lung cancer subtypes and molecular backgrounds. In addition, the mechanistic exploration regarding the association between ASIC3 upregulation and EMT is insufficiently profound. All these aspects represent key directions that require focused efforts in subsequent research.
Conclusion
The present study highlights a mechanism for HCA micoenvironment- induced EMT in lung cancer cells via ASIC3, which may affect to the FP and OS of patients with lung cancer. Therefore, ASIC3 may represent a possible target for therapeutic intervention in lung cancer. Alternatively, strengthening the intervention of HCA may be an important measure in the clinical treatment of lung cancer.
Footnotes
Abbreviations
Acknowledgements
The authors would like to acknowledge the helpful comments on this article received from reviewers. Generative artificial intelligence (AI) tool Doubao was used solely for language polishing and grammatical refinement of the manuscript. No AI tools were involved in the experimental design, data collection, statistical analysis, interpretation of results, or preparation of scientific content of this study.
Ethics Approval Statement
The experiment was approved by the Medical Ethics Committee of the First Affiliated Hospital of Nanchang University (IIT[2022] NO.099, Nanchang, Nov 2022). All experiments were conformed to the National Research Council's Guide.
Informed Consent Statement
All participants provided written informed consent prior to their participation in the study.
Author Contributions Statement
This study conception and design were performed by H Fu. L Qu, L Zhao, L Zhang, and C Luo, wrote the main manuscript text, and L Zhao, L Zhang, C Luo confirmed the authenticity of all the raw data. Critical revision of the manuscript was given by all authors. All authors reviewed the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: All funds used for experiments were supported by National Natural Science Foundation of China (NO.82260382) and Jiangxi Provincial Natural Science Foundation (NO.20242BAB25497).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The datasets generated and/or analyzed during the current study are available in this paper.
