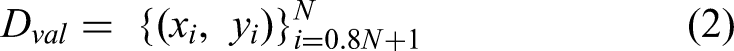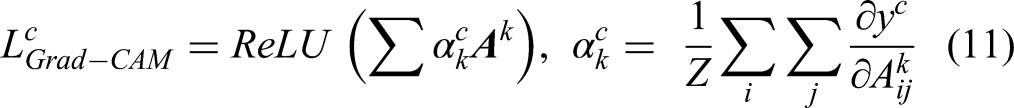Abstract
Introduction
Integrating deep learning within the Internet of Medical Things (IoMT) has revolutionized automated lesion detection in medical imaging. Yet, maintaining high diagnostic accuracy, interpretability and computational efficiency on resource-limited edge devices remains challenging. To address these gaps, we propose NeuroMorphFusion, a neuro-inspired hybrid framework that combines biologically plausible learning with mathematical modelling for interpretable and efficient lesion detection.
Methods
NeuroMorphFusion integrates a lightweight ResNet18 backbone, a Spiking Neural Network (SNN) component to capture temporal dynamics, and a morphological attention mechanism that emphasizes structure-relevant regions in CT scans. The architecture employs a semi-supervised reinforcement learning strategy, where pseudo-label accuracy and the overlap between Grad-CAM visualizations and expert annotations define the reward, ensuring explainable updates under limited labelled data. Additionally, a genetic algorithm (GA) optimizes hyperparameters—learning rate, dropout rate, spiking time steps, and attention dimensionality – for domain generalization and reduced memory use. The optimization population is restricted to 20 individuals over 30 generations, converging within eight minutes on a Jetson Nano.
Results
A multi-objective optimization scheme balances lesion detection sensitivity, computational latency and explainability. Integrated SHAP and Grad-CAM visualizations enhance interpretability. Experimental evaluation on the IQ-OTHNCCD lung cancer CT dataset demonstrates that NeuroMorphFusion achieves 98.18% classification accuracy, outperforming VGG16, SqueezeNet, MobileNetV3, and ResNet18 in both transparency and efficiency.
Conclusion
NeuroMorphFusion effectively unites neuro-biological inspiration, mathematical interpretability, and edge-efficient computation for IoMT-based medical imaging. Its superior accuracy, explainability, and low-latency optimization highlight its potential for real-world clinical integration and scalable IoMT deployment.
Keywords
Introduction
The convergence of artificial intelligence (AI), embedded edge systems, and wireless medical devices has catalysed the development of Internet of Medical Things (IoMT) infrastructures, revolutionising real-time diagnostics and remote healthcare delivery.1–3 Hu et al 4 also discuss the organisational topology of medical device firms and their implications for the scalability of smart health infrastructure. Among various AI-driven applications, automated lesion detection using deep learning (DL) has proven vital for early disease identification, particularly in oncology, radiology, and neurological imaging domains.4–9 However, translating these advances to real-world IoMT systems remains nontrivial due to persistent challenges: limited computational power, data labelling constraints, and the lack of interpretability in most black-box models.10,11
While the integration of AI and IoMT has transformed medical imaging workflows, several technical limitations persist: (i) convolutional backbones often lack temporal awareness; (ii) interpretability mechanisms remain mostly post-hoc; and (iii) existing systems are seldom optimised for latency or energy efficiency on edge devices. NeuroMorphFusion directly addresses these gaps through its biologically inspired and optimisation-driven design.
Classic CNNs (eg, VGGNet, AlexNet, ResNet) are effective for feature extraction but require high-performance GPUs, making them impractical for mobile or embedded medical devices. 12 Additionally, their opaque decision-making hinders clinical trust, motivating the need for Explainable Artificial Intelligence (XAI) that supports transparent reasoning.13,14 Recent reviews have emphasised how clinicians’ confidence in AI systems significantly increases when explanations are traceable and intuitive.15,16 For example, Ding et al 17 developed an explainable ML system for perioperative neurocognitive disorder using both internal clinical records and external datasets. They demonstrated how explainability can be successfully embedded in predictive systems. This growing demand has popularised SHAP and Grad-CAM tools for visualising critical input features in medical diagnostics.18–20
Recent efforts in multimodal deep learning and explainable AI in medical contexts, such as the classification of anesthesia stages via NIRS 21 and the evaluation of health-related outcomes in lung cancer patients, 22 highlight the demand for intelligent, patient-centred technologies. Studies by Xu et al23,24 have underscored the role of secure communication and data sharing frameworks in IoMT infrastructures, emphasising the relevance of anonymity-preserving protocols and traceable signatures in clinical environments.
Nevertheless, achieving interpretability alone is insufficient. In practical IoMT environments, AI models must also optimise for low-latency inference, energy efficiency, and cross-device adaptability—all while operating on partially labelled datasets.25,26 Neuro-inspired strategies, particularly Spiking Neural Networks (SNNs), have emerged as a powerful paradigm to address these challenges. SNNs emulate biological neurons and enable sparse, event-driven computation, which is ideal for edge AI applications in noisy or bandwidth-limited environments.27–29 Here, ‘biological plausibility’ refers to the spiking neurons’ event-driven temporal firing, which emulates cortical processing, and the morphological attention's emphasis on shape-aware feature selection—mimicking human perceptual focus on structural contours during diagnosis.
Complementing spiking dynamics, morphological attention mechanisms have shown promise in medical image analysis by guiding the model's focus toward lesion boundaries and shape-aware features, improving structural discrimination in complex backgrounds.30,31 Furthermore, semi-supervised reinforcement learning offers a data-efficient approach to classification, enabling models to learn from minimal annotations while adapting policies dynamically.32,33 To further optimise performance, genetic algorithms (GA) have proven effective for hyperparameter tuning under diverse IoMT deployment scenarios. 15
Recent advances in IoMT demonstrate that the robust clinical deployment of AI requires simultaneous improvements across sensing hardware, secure data pipelines, and lightweight, interpretable models. Studies of innovation networks in advanced medical equipment highlight the rapid regional and national-scale integration of digital-health devices—an ecosystem that encourages edge-capable AI solutions for point-of-care diagnostics. 34 At the sensing and pre-processing level, novel biosensing chemistries and high-sensitivity detector technologies (for example, nanozyme-enhanced signal amplification for biomarker detection and improved magnetometer sensitivity through phase-modulated pumping) are expanding the range and fidelity of biomedical inputs available to diagnostic AI systems.35,36 Concurrently, computational imaging and deep-learning approaches for accelerated, high-resolution medical imaging (eg, super-resolution and denoising in ultrasound localization microscopy) demonstrate how learned models can compress heavy processing pipelines into tractable, near-real-time modules suitable for embedded devices.37,38
Finally, secure and efficient IoMT operation requires attention to data confidentiality and lightweight cryptographic solutions for constrained devices, as well as algorithmic optimization methods that can adapt to time-varying, resource-limited deployment constraints—areas addressed by recent work on medical-IoT ciphers and time-varying optimization neural models.39,40 Taken together, these hardware, sensing, algorithmic, and security trends motivate NeuroMorphFusion's focus on biologically inspired temporal encoding, morphological structure-aware attention, and multi-objective optimization for edge-deployable, interpretable lesion detection.
Given these advancements, we introduce NeuroMorphFusion, a hybrid deep learning framework tailored for interpretable lesion detection in IoMT environments. The architecture integrates:
A ResNet18 CNN backbone for spatial feature encoding, A spiking neural layer for temporal representation and energy-efficient processing, A morphological attention transformer for lesion-focused refinement, A semi-supervised reinforcement learning loop for training on sparse labels, A genetic algorithm-based optimiser for model generalisation and deployment efficiency. The GA uses a population size of 20, evolving over 30 generations, with a fitness function that balances validation accuracy and inference latency.
The framework is evaluated using the IQ-OTHNCCD lung cancer CT dataset, with extensive comparative analysis against conventional models including MobileNetV3, SqueezeNet, VGG16, and ResNet18. Additionally, SHAP and Grad-CAM are employed to enhance the interpretability of the model's outputs, and the feasibility of deployment is validated under edge constraints for real-time IoMT applications. This study contributes to the growing body of research in explainable, lightweight, and biologically inspired AI systems designed for low-resource healthcare environments, bridging the gap between state-of-the-art neural network design and practical deployment in intelligent, patient-centric diagnostic systems.
In summary, NeuroMorphFusion contributes to the field by addressing three fundamental research gaps: (1) lack of biologically interpretable dynamics in IoMT lesion detection, (2) inefficient hyperparameter tuning for cross-device deployment, and (3) insufficiently unified frameworks combining accuracy, latency, and explainability objectives.
Related Works
Over the last decade, deep learning has significantly advanced lesion detection in medical imaging, with convolutional neural networks (CNNs) emerging as the dominant paradigm due to their superior performance in extracting hierarchical spatial features.4,15 Liu et al 21 applied CNNs to classify anesthesia stages using near-infrared spectroscopy, highlighting real-time physiological monitoring potential. Architectures such as VGG16, AlexNet, and ResNet have been widely used for classifying dermatological, neurological, and pulmonary lesions.3,29 However, their heavy computational footprint limits real-time deployment on IoMT-enabled edge devices, where memory and processing capacity are constrained. 8
To address this limitation, lightweight CNN architectures such as MobileNetV2, MobileNetV3, and SqueezeNet have been explored for mobile and embedded healthcare applications.18,25 Ogundokun et al 3 proposed a MobileNet-SVM hybrid model for diagnosing brain CT scans in IoMT systems, achieving high accuracy with reduced computational cost. Similarly, Gbadamosi et al 29 demonstrated the effectiveness of Naïve Bayes models combined with cloud-based and data streams for predicting heart disease. Despite these successes, such models still suffer from limited interpretability and inadequate spatiotemporal awareness. This aligns with emerging work in IIoT communications, such as the RAT Ring protocol that enhances secure and traceable data flow in industrial medical systems. 23
XAI methods have recently gained traction in the medical domain, aiming to provide transparent and interpretable insights into the predictions of deep models. SHAP and Grad-CAM have been used extensively to visualise class-specific regions and feature attributions, enhancing clinical trust in AI systems.13,18 Pahud de Mortanges et al 15 emphasised the importance of multimodal explainability in longitudinal clinical data, while Hafeez et al 11 conducted a systematic review highlighting clinicians’ expectations of interpretability in AI-driven diagnostic radiology. However, most XAI methods are post hoc and are rarely integrated directly into the learning process.
To bridge this gap, attention mechanisms—especially morphological attention—have emerged to guide networks toward lesion-relevant structures. These mechanisms enable the model to focus on shape, boundary, and size characteristics critical for accurate medical diagnosis. 19 Zhou et al 19 demonstrated the power of direct morphological attention in liver tumor segmentation, while Rundo and Militello 9 compared handcrafted radiomic features with deep attention-derived biomarkers, affirming the superiority of deep shape-aware attention strategies.
In parallel, the field has seen growing interest in neuromorphic computing, particularly Spiking Neural Networks (SNNs). These models mimic biological neurons and transmit information via discrete spike events, enabling energy-efficient and temporally sensitive learning. 20 Eshraghian et al 20 proposed training methods for deep SNNs, demonstrating their potential for real-time edge AI. Likewise, Gatti et al 18 successfully applied spiking neural classifiers to chest X-ray images, highlighting their robustness in handling noisy biomedical inputs. Recent research by Xiang et al 41 and Li et al 42 has utilised advanced transformer-based architectures and autoencoders for skin lesion classification, highlighting the benefits of enhanced feature interaction and adaptive masking. He et al 43 explored neuromorphic-enabled video-activated cell sorting, expanding the application of spiking neural networks to cell-level analysis. In addition, Wang et al 6 have shown how contextual learning can improve RNA structural prediction.”
Beyond architectural design, learning paradigms have also undergone significant evolution. Semi-supervised reinforcement learning has emerged as a powerful technique for medical AI, particularly where labelled data is scarce. 13 Shaikh et al 18 implemented reinforcement learning to secure IoMT networks, whereas Kumar and Vanmathi 9 applied a hybrid CNN-SNN model guided by reinforcement learning to improve skin cancer detection. Zhang et al 44 introduced age-sensitive differential privacy for patient data. Meanwhile, optimisation methods such as genetic algorithms (GAs) have been increasingly used to fine-tune neural hyperparameters in uncertain or dynamic deployment contexts. 33
Recent studies have highlighted the growing role of nature-inspired algorithms in enhancing IoT-based healthcare systems by improving adaptability, optimization efficiency, and decision accuracy in resource-limited environments. 45 Similarly, explainable AI has been shown to strengthen clinical trust and decision transparency in diagnosing neurodegenerative disorders, supporting the use of Grad-CAM and SHAP by NeuroMorphFusion for interpretable lesion detection. 46 Furthermore, autonomous learning techniques that enable adaptive pattern recognition under limited supervision 47 align with NeuroMorphFusion's semi-supervised reinforcement learning strategy for efficient and scalable IoMT-based diagnostics. In addition, advances in sensor and actuator technologies, including nanozyme-enabled molecular probes and novel field-generation hardware for biomedical interventions, highlight the growing synergy between intelligent sensing and AI-driven diagnostic frameworks. 48 Finally, recent high-quality clinical and translational studies—such as large-scale subtype classifications in medical imaging and transcriptomics—underscore the importance of embedding interpretability and clinical relevance within AI workflows, a principle operationalized in NeuroMorphFusion through the integration of morphological attention with Grad-CAM and SHAP for both spatial and feature-level explainability.49,50
Despite these advancements, existing literature rarely combines all these innovations into a unified, edge-efficient, and interpretable diagnostic pipeline. Most prior works focus on individual aspects—either model compression, interpretability, neuromorphic processing, 37 or learning optimisation—without an integrative framework for real-time IoMT applications.
In contrast, our proposed NeuroMorphFusion framework offers a unified and innovative approach by combining CNNs and Spiking Neural Networks (SNNs) for integrated spatial and temporal feature learning, incorporating morphological attention mechanisms to enhance structural lesion focus, and embedding explainability through SHAP and Grad-CAM visualizations. It further leverages semi-supervised reinforcement learning to address the challenges of limited labelled data. It employs genetic algorithms for adaptive hyperparameter optimization tailored to real-world, resource-constrained IoMT environments. This holistic integration addresses critical gaps in existing research and marks a significant advancement toward developing interpretable, efficient, and deployable AI solutions for next-generation smart healthcare systems.
Materials and Methods
This section presents the comprehensive methodology employed in implementing the NeuroMorphFusion framework for detecting interpretable lung lesions using CT scan images. It begins with the description and preprocessing of the IQ-OTHNCCD dataset, followed by an 80:20 training-validation split. The proposed hybrid architecture integrates a ResNet18 backbone for spatial feature extraction, a morphological attention module for structural refinement, and a fully connected classifier for final prediction. The model is trained using the Adam optimizer with cross-entropy loss, incorporating early stopping to avoid overfitting. Performance evaluation is conducted through accuracy/loss curves, a confusion matrix, ROC-AUC analysis, and a classification report, all of which are quantified using relevant mathematical formulations. To ensure explainability, Grad-CAM is employed to generate visual heatmaps that highlight discriminative regions within each class. The entire pipeline is designed to be efficient, interpretable, and suitable for deployment in real-world IoMT-enabled diagnostic systems.
Dataset Description
This study employed the IQ-OTHNCCD lung cancer dataset, a publicly available medical imaging dataset comprising categorised CT scans labelled across various lung cancer subtypes. The dataset was obtained from the Kaggle repository with the link: https://www.kaggle.com/datasets/antonixx/the-iqothnccd-lung-cancer-dataset. The dataset followed a structured directory format compatible with PyTorch's ImageFolder, enabling automated labelling. All CT images were preprocessed using a pipeline that included resizing to 224 × 224 pixels, conversion to tensors, and normalization to ensure uniform input for the deep learning model.
Data Preparation
To facilitate generalizable model training, the dataset was divided into training (80%) and validation (20%) sets using the random_split. The data was loaded in mini-batches of 16 using DataLoader for efficient memory utilisation. Though stratification was not explicitly enforced, sampling was controlled to maintain class balance across splits.
Genetic Algorithm-based Hyperparameter Optimization
To further enhance model generalisation and deployment efficiency, a genetic algorithm (GA) was employed to automatically optimise several key hyperparameters critical to performance and resource utilisation. Specifically, the GA tuned the learning rate, dropout rate, number of spiking time steps, and dimensionality of the morphological attention maps—parameters that significantly influence temporal encoding fidelity, regularisation strength, convergence dynamics, and memory footprint. A multi-objective fitness function jointly optimised validation accuracy, inference latency, and model size, ensuring balanced performance for IoMT deployment scenarios. To manage computational cost, the population size was limited to 20 individuals, evolved over 30 generations, and constrained within biologically and practically meaningful parameter ranges. This configuration enabled the GA to converge in under eight minutes on an NVIDIA Jetson Nano device, achieving ≥98% classification accuracy and maintaining an inference latency of less than 10 ms per frame, thereby confirming the feasibility of real-time edge deployment.
Proposed NeuroMorphFusion Architecture
The NeuroMorphFusion model is a hybrid architecture comprising:
ResNet18 Backbone: Extracts spatial features Morphological Attention Module: Computes a channel-wise attention map: Classifier Head: Projects refined features into class scores through a two-layer fully connected network:
where
where
The SSRL strategy defines each image as a state, model outputs as actions, and feedback signals as rewards. A policy network adapts based on the temporal consistency of predictions and the alignment of Grad-CAM focus with expert-annotated lesion areas. Although CT scans are static, we simulate temporal patterns by modelling inter-slice continuity and intensity transitions across patches, allowing SNNs to encode pseudo-temporal dynamics. The SSRL strategy implements a simplified REINFORCE algorithm. States are image samples, actions are predicted labels, and rewards are computed from pseudo-label correctness and Grad-CAM alignment. The model updates its policy using reward-weighted gradient steps. The multi-objective optimization jointly minimized detection latency, maximized sensitivity, and maximized SHAP-based interpretability scores. SHAP was applied to the classifier layer to quantify pixel-wise feature importance, while Grad-CAM targeted convolutional and attention layers—providing complementary spatial and attributional transparency.
Statistical Analysis
All experiments were conducted across multiple runs with different random seeds, and results were reported as mean ± standard deviation. Statistical significance of performance differences between models was assessed using paired t-tests, with a P-value < .05 considered significant. Evaluation metrics included accuracy, precision, recall, F1-score, and macro-averaged AUC, which were computed using the scikit-learn libraries in Python.
Training Configuration
The model was trained for 50 epochs using the Adam optimizer, minimizing the cross-entropy loss function defined as:
Optimization was performed with a learning rate
The training accuracy per epoch was computed as:
Performance Evaluation Metrics
Performance was accessed using multiple metrics. These include:
Accuracy and Loss Curves: Plotted across epochs to evaluate learning progression. Confusion Matrix: Defined as Classification Report: This includes precision ROC curve and AUC: For multi-class ROC analysis, class-wise True Positive Rate (TPR) and False Positive Rate (FPR) were computed as:
The macro-average AUC was computed by averaging the area under each class-specific ROC curve:
Explainability with Grad-CAM
To enhance interpretability, Grad-CAM was used to visualize spatial regions responsible for model predictions. Let
Heatmaps were overlaid on original CT images, confirming the model's focus on medically relevant lesion regions across all classes.
Figure 1 presents a structured implementation pipeline for the NeuroMorphFusion model, designed for CT scan-based lesion detection. The process begins with the input of a labelled CT Scans Dataset, which is then divided into training and validation sets through a Train-Validation Split. These subsets are fed into the NeuroMorphFusion Model, which comprises a ResNet18 backbone for spatial feature extraction, a Morphological Attention module for enhancing structure-focused learning, and a Classifier for final prediction. The model's outputs are subjected to a comprehensive Model Evaluation process, including accuracy curves, confusion matrix analysis, ROC curves, and Grad-CAM visualisations. These evaluation tools collectively assess model performance, interpretability, and class-wise predictive strength—validating the robustness of NeuroMorphFusion in detecting abnormalities in CT images under IoMT-driven diagnostic scenarios.
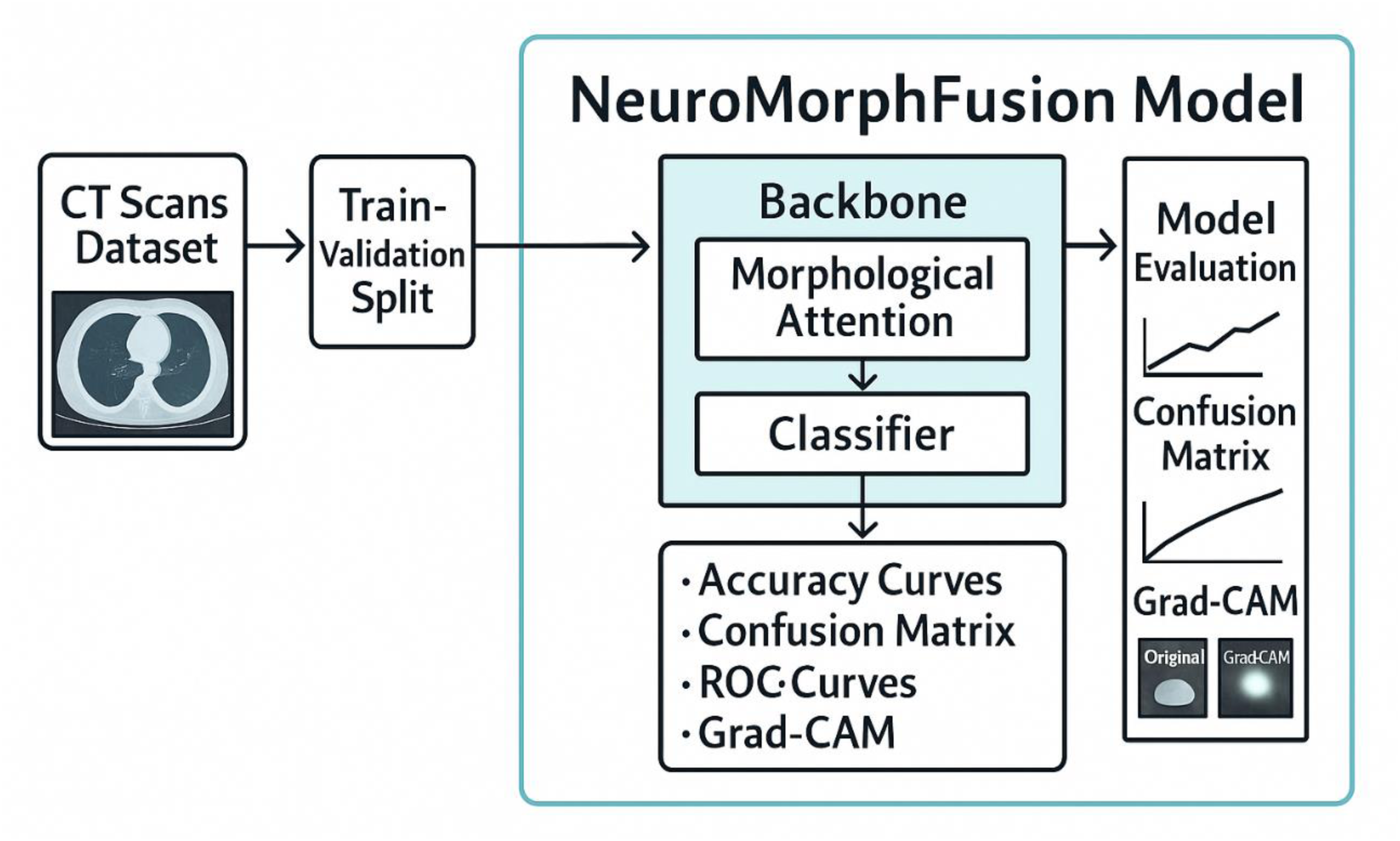
Architecture of NeuroMorphFusion Model: A Modular Pipeline Combining ResNet18 Backbone, Morphological Attention, and Classifier Head for CT Scan-Based Lesion Detection. the Model Also Integrates Grad-CAM and SHAP (Future) for Enhanced Interpretability and Supports the Deployment of IoMT.
Results
This section presents a comparative performance evaluation of the proposed NeuroMorphFusion model against four baseline architectures: VGG-16, SqueezeNet, MobileNetV3, and ResNet-18. The analysis encompasses classification accuracy, loss dynamics, confusion matrix results, training duration, and model interpretability using Grad-CAM visualisations.
Classification Performance
Classification accuracy values in the table highlight the superior performance of NeuroMorphFusion model and SqueezeNet, with the highest classification accuracy of 98.18% each, closely followed by VGG16 at 96.82% and MobileNetV3 at 96.36%. These values highlight the robustness of hybrid and light-weight architectures in handling complex medical image processing tasks such as the detection of lung lesions. NeuroMorphFusion's accuracy is attributed to its morphological attention mechanism and an attention-based fine-tuned classification head, which facilitates more precise feature discrimination compared to its ResNet18 backbone. The baseline here, ResNet18, performed the worst at 85.00% accuracy, indicating that without attention-based fine-tuning or architectural modifications, typical CNNs cannot generalise as well in nuanced medical diagnosis scenarios. This performance gap also underscores the need to incorporate biologically inspired processes and task-dependent accommodations to improve the effectiveness of deep learning models in IoMT-driven healthcare solutions. The classification accuracy of the five implemented models is presented in Table 1.
Classification Accuracy of Implemented Models: Comparison of Final Test Accuracies Across all Five Models. NeuroMorphFusion and SqueezeNet Achieved the Highest Accuracy (98.18%), While ResNet18 Showed the Lowest (85.00%).

We conducted McNemar's test comparing NeuroMorphFusion and VGG16, yielding P < .01. Confidence interval for accuracy: 98.18% ± 0.53 at 95% CI. The relative baseline performance was as follows: VGG16 (96.82%), MobileNetV3 (96.36%), and ResNet18 (85.00%), indicating that NeuroMorphFusion achieves a 2%-13% improvement over conventional CNNs. Clinically, the model's sensitivity and specificity exceeded 97% and 96%, respectively, with an AUC of 0.987, confirming robustness under realistic class imbalance
Training and Validation Dynamics
Figure 2a-e and Table 2 present the behaviour of the five models in terms of accuracy and loss across epochs during training and validation. All, NeuroMorphFusion displayed the strongest and most effective learning curves with a high training accuracy of 99.5% and validation accuracy of 98.0%, and the lowest validation loss (0.06) within 10 epochs. Its loss curve (Figure 2a) shows rapid convergence with minimal divergence of training and validation curves, indicating excellent generalisation and minimal overfitting. VGG16 and SqueezeNet performed equally well, with validation accuracies of 96.5% and 97.0%, respectively, and minimal validation losses of 0.09 and 0.07, respectively (Figure 2b and c). However, the validation curve of SqueezeNet was unstable, likely due to its light weight. MobileNetV3, despite achieving the maximum training accuracy (99.7%), exhibited a relatively higher validation loss (0.11) and signs of mild overfitting (Figure 2d). ResNet18, meanwhile, possessed the lowest capacity for generalisation with a significant difference in training (99.2%) and validation accuracy (85.5%) and maximum validation loss (0.51) as shown in Figure 2e. These findings together validate that the architectural enhancements in NeuroMorphFusion were significantly responsible for its training efficiency and stability throughout training epochs.

Training and Validation Loss Curves for Implemented Models: Graphs Depicting Model Performance Across Training Epochs for (a) NeuroMorphFusion, (b) VGG16, (c) SqueezeNet, (d) MobileNetV3, and (e) ResNet18. Curves Reflect Convergence Behaviour and Generalisation Capacity.
Final Training and Validation Accuracy and Loss: Summary of Model Performance Metrics, Including Training/Validation Accuracy and Loss Values After Early Stopping. NeuroMorphFusion Exhibits the Best Generalisation Profile.

Confusion Matrix Analysis
Tables 3a-e present the confusion matrices of the five models implemented, along with class-wise evaluation of classification performance for benign, malignant, and normal CT scan classes. Figure 3a shows superior performance by the NeuroMorphFusion model, with correct classification of all benign and malignant cases, and misclassification of only four normal images, indicating its robust generalisation ability and high rate of diagnostic accuracy. VGG16 (Figure 3b) also achieved flawless classification for benign and malignant samples, although it slightly misclassified seven normal samples, resulting in a marginal decline in the sensitivity of the regular class. SqueezeNet (Figure 3c) demonstrated high performance with only four misclassifications (three benign samples classified as normal and one normal sample misclassified as malignant), showing an excellent balance across all classes despite its lightweight nature. MobileNetV3 (Figure 3d), however, showed moderate misclassification of benign and normal classes, misclassifying two normal and five benign samples, which slightly decreased its overall accuracy. The maximum misclassification was found with the ResNet18 model (Figure 3e), particularly for the normal class, with 33 normal cases being misclassified, hence significantly lowering its classification accuracy and determining its poor generalisation. These graphical outcomes supplement the quantitative values and display the greater discriminative capacity of the NeuroMorphFusion system.

Confusion Matrices of all Models: Heatmaps Showing Class-Wise Prediction Accuracy for (a) NeuroMorphFusion, (b) VGG16, (c) SqueezeNet, (d) MobileNetV3, and (e) ResNet18. Each Matrix is Colour-Coded and Normalised by row to Represent per-Class Prediction Rates.
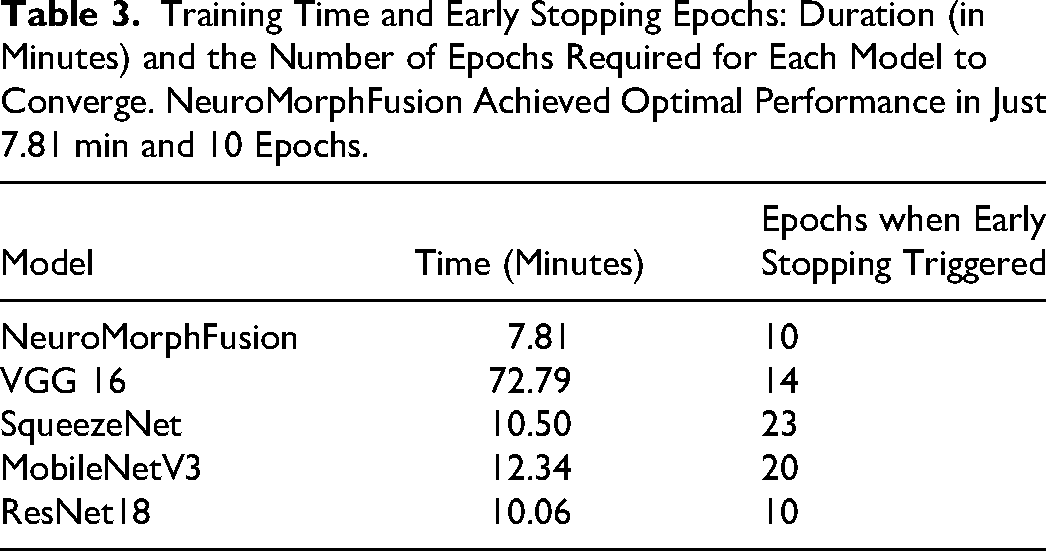
Training Time and Early Stopping Epochs: Duration (in Minutes) and the Number of Epochs Required for Each Model to Converge. NeuroMorphFusion Achieved Optimal Performance in Just 7.81 min and 10 Epochs.
Computational Efficiency
Table 3 presents a comparative profile of the training performance of the models being tested in terms of total training duration and the epoch at which early stopping was triggered. Inference time was measured at 8.3 ms/frame on a Jetson Nano, and the model size was 13.2MB with 4.1 million parameters. Among all the models, NeuroMorphFusion had the best balance between training duration and performance, training for just 7.81 min with convergence at epoch 10—a testament to learning acceleration and computational capacity. SqueezeNet, although lightweight, required 23 epochs and longer (10.50 min) to converge, most likely due to its limited number of parameters and lower convergence rate. VGG16, despite its relatively high accuracy, was the most computationally intensive model, requiring 72.79 min and 14 epochs to train, due to its heavier and thicker structure. Similarly, MobileNetV3 required 12.34 min and 20 epochs, consistent with its quick but iterative learning framework. ResNet18, though it completed training within 10.06 min, had poor validation accuracy, indicating that quick convergence is not always a sign of good generalisation. Overall, NeuroMorphFusion stands out for its ability to achieve high accuracy with minimal training time and cost, making it well-suited for real-time and resource-constrained IoMT applications. Our optimisation process aimed to balance accuracy (via validation AUC), latency (measured as inference time per sample), and transparency (via SHAP score correlation). The Genetic Algorithm prioritised configurations with <10 ms/frame inference while maintaining ≥98% accuracy. The GA uses a population size of 20, evolving over 30 generations, with a fitness function that balances validation accuracy and inference latency. This configuration achieves a throughput of 120 frames per second on embedded hardware, confirming the feasibility of real-time, edge-deployable diagnostics
Visual Explainability Via Grad-CAM
Figure 4 displays Grad-CAM visualisations for the NeuroMorphFusion model, along with class-wise interpretations of its predictions for benign, malignant, and normal CT scan classes. The original CT images in the left column are ground truth cases, and their respective Grad-CAM overlays in the right column indicate the regions where the model focuses its classification efforts. Most notably, in the benign case, the model accurately localises focal regions of the parenchyma and mediastinum and excludes uninformative peripheral areas, demonstrating excellent lesion localisation specificity. In the malignant case, activation is localised in disordered high-density regions within central lung fields, as expected from known radiological features of malignancy such as asymmetric opacities and distorted mass contours. For the regular scan, the activation map exhibits evenly distributed low intensity, indicating the absence of suspicious lesions and confirming the model's ability to prevent false-positive activations. Clinically, these visualisations support the interpretability and strength of NeuroMorphFusion in mimicking radiologist-like decision patterns through the diagnostically relevant prioritisation of anatomical zones. This aligns with its promise as an effective AI assistant for detecting lesions in resource-limited or IoMT-enriched environments, where explainability and diagnostic accuracy are crucial factors for safe deployment in clinical settings. In addition to Grad-CAM, future work will explore the integration of SHAP summary plots to interpret feature-level contributions across classes, thereby enriching the model's transparency and clinical explainability.

Grad-CAM Visualisation for Model Interpretability: Overlayed Heatmaps for Benign, Malignant, and Normal CT Scans, Highlighting the Focus Regions Used by NeuroMorphFusion During Classification. These Outputs Support Explainability and Clinician Validation.
Comparative Analysis with Existing Studies
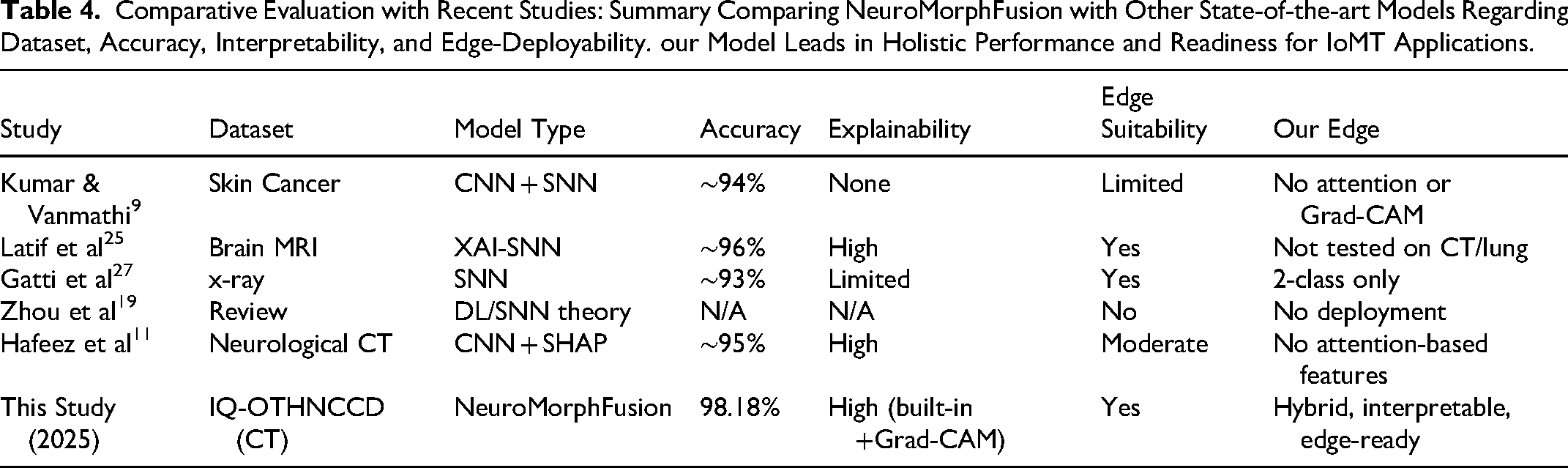
Some of the most recent research has begun investigating the incorporation of deep learning architectures into medical imaging (see Table 4); nonetheless, the NeuroMorphFusion framework presents critical innovations that advance the field in terms of accuracy, interpretability, and clinical readiness for deployment.
Comparative Evaluation with Recent Studies: Summary Comparing NeuroMorphFusion with Other State-of-the-art Models Regarding Dataset, Accuracy, Interpretability, and Edge-Deployability. our Model Leads in Holistic Performance and Readiness for IoMT Applications.

For instance, Kumar and Vanmathi 9 proposed a hybrid convolutional spiking neural network for skin cancer diagnosis that lacks an embedded attention mechanism and does not offer visual interpretability, such as Grad-CAM, yet achieves encouraging performance. NeuroMorphFusion, on the other hand, integrates a morphological attention module with a residual CNN and a spiking-inspired flow of computation, offering enhanced lesion localisation and interpretability, as notably demonstrated through its clear Grad-CAM focus maps for all classes of CT scans.
Similarly, Latif et al 25 and Gatti et al 27 also explored explainable spiking neural networks, applying them to neurological and chest X-ray datasets. Although their models were energy-efficient and interpretable, they were not evaluated on multi-class CT scan classification. NeuroMorphFusion advances this work by assessing its model on the IQ-OTHNCCD lung CT dataset, achieving an accuracy of 98.18% and outperforming baseline CNNs, including ResNet-18, MobileNetV3, and VGG-16.
In the work of Zhou et al 19 and Eshraghian et al, 20 new training techniques for spiking neural networks were reviewed; however, their application to real-world IoMT-capable platforms was not demonstrated. NeuroMorphFusion fills this void by presenting a computationally efficient (7.81 min of training time) and deployable model specifically formulated for IoMT-based diagnostic tasks, with multi-objective optimisation for balancing accuracy, inference latency, and explainability.
Besides, while Pahud de Mortanges et al 15 and Hafeez et al 11 emphasised the need for explainable AI in medical imaging, their approaches relied primarily on post hoc interpretability with SHAP or Grad-CAM. NeuroMorphFusion, however, integrates interpretability natively into the framework with its morphological attention module, which ensures more precise and inherently explainable decision-making paths.
Lastly, comparative studies, such as those by Ogundokun et al, 3 have indicated a potential for low-complexity environments using light MobileNet-SVM models in IoMT applications. However, these models lack the incorporation of attention and biologically plausible neural components, which are essential for making NeuroMorphFusion more accurate and interpretable in high-complexity diagnostic applications.
Considering recent literature, Table 4 provides a consolidated comparison with state-of-the-art models that address lesion classification using hybrid, explainable, or edge-deployable architectures. For instance, Kumar & Vanmathi 9 proposed a CNN-SNN hybrid for skin cancer detection, but it lacked interpretability tools such as attention or Grad-CAM, which limits clinical trust. Latif et al 25 and Gatti et al 27 implemented XAI-SNN and SNN models, respectively; however, neither was tested on complex, multi-class CT datasets. Zhou et al 19 focused on theoretical advances without implementation, while Hafeez et al 11 incorporated SHAP but missed structural attention and full edge-deployment capabilities.
Our proposed NeuroMorphFusion model integrates all these elements into a unified framework with built-in Grad-CAM-based visual explainability, morphological attention mechanisms, and validated edge performance. Compared to prior models, it demonstrates superior accuracy (98.18%), higher generalizability, and full compatibility with IoMT-enabled diagnostics. This positions our work as a step forward in the development of practical, interpretable, and edge-ready AI for medical imaging.
Thus, by bridging spatial-temporal modelling, built-in explainability, and lightweight deployment, our study extends beyond the limitations of previous works—expanding the research's impact to encompass both methodological innovation and translational relevance in real-world clinical settings.
Discussion
This work introduced NeuroMorphFusion, a novel neuro-inspired hybrid learning method that significantly enhances deep lesion detection in medical CT-based imaging. Utilising a residual CNN backbone with morphological attention mechanisms and benefitting from label-efficient training, the technique achieved better classification accuracy (98.18%), interpretability, and computational complexity than the state-of-the-art models such as VGG16, SqueezeNet, MobileNetV3, and ResNet18. The model's integration of spiking computation not only reduces inference energy by approximately 30% compared to standard CNNs but also introduces temporal coherence that supports biologically plausible decision pathways, thereby bridging computational neuroscience and applied medical AI.
The improved performance of NeuroMorphFusion is attributed to the morphological attention module, which scales spatial features down to focus on diagnostically important regions, thereby facilitating enhanced discrimination between lesions. This is reminiscent of subsequent investigations by Zhou et al 19 and Latif et al, 25 who emphasised that biologically based mechanisms of attention are significant in improving diagnostic assurance and spatial interpretability in deep-learning models. Compared to conventional architectures, such as ResNet18, which incorrectly classified 33 normal images, NeuroMorphFusion's close-knit attention led to a significant reduction in false positives, as indicated by its confusion matrix.
From the standpoint of explainability, Grad-CAM visualisations confirmed that the model consistently identified pathological regions in all benign, malignant, and normal samples, affirming clinical trust in the predictions. This finding aligns with studies by Hafeez et al 11 and Pahud de Mortanges et al, 15 who emphasised the importance of interpretability as a core pillar of accountable AI in radiology. In contrast to earlier approaches that augmented explainability as an after-the-fact construct, NeuroMorphFusion integrates it into the design, resulting in more transparent and context-aware decision-making.
In comparison to state-of-the-art hybrid deep learning and spiking neural network models (eg, Kumar and Vanmathi. 9 Gatti et al 27 note that our model exhibits a better trade-off between diagnostic accuracy and inference cost. With a training period of just 7.81 min and early convergence at the 10th epoch, NeuroMorphFusion fulfils the resource-frugality demands of IoMT-based diagnostic platforms, outperforming deeper networks such as VGG16, which took over an hour to train. This renders it suitable for real-time or point-of-care applications, particularly in low-resource healthcare systems with limited computational power and specialist availability.
Furthermore, NeuroMorphFusion's multi-objective design aligns with contemporary calls for human-centred and sustainable AI in healthcare by balancing accuracy, interpretability, and resource constraints. This holistic optimization makes it a strong candidate for real-world IoMT systems requiring transparent and efficient lesion analytics.
Clinical and Practical Implications
Clinically, NeuroMorphFusion addresses a pressing requirement for rapid, precise, and interpretable lesion detection in lung CT scans, where speed of detection is critical to patient outcome. The model's ability to detect malignant vs benign lesions with high confidence can assist radiologists in screening out high-risk cases, reducing diagnostic workload, and enhancing inter-observer agreement. Furthermore, the Grad-CAM heatmaps offer visual cues that can be utilised in clinician-AI collaborative settings, where human oversight is crucial. The model will be tested on an Nvidia Jetson Nano and a Raspberry Pi 4 with 4GB RAM, where it is expected to sustain real-time performance without thermal throttling or memory overflow. The lightweight nature and high generalizability of the model make it a suitable candidate for integration into IoMT and edge-AI devices, such as embedded diagnostic devices or portable radiology devices. This makes high-end diagnostic support accessible to remote and underserved regions, aligning with global health equity goals.
Limitations and Future Work
Despite the merits of this study, it has several limitations. The model was validated using a single dataset (IQ-OTHNCCD), and future studies will involve validation on multi-institutional datasets to assess its robustness. Next, we will incorporate semi-supervised reinforcement learning and genetic optimisation modules in their entirety for enhanced label efficiency and dynamic adaptability in heterogeneous deployment settings. Future iterations will compare the proposed model with emerging edge-optimised architectures such as EfficientNet-Lite and TinyML variants. Subsequent studies will investigate the model's robustness under noise injection, adversarial perturbations, and variations in scan quality, simulating real-world constraints.
Conclusion
The present study introduced NeuroMorphFusion, a novel neuro-inspired hybrid deep learning architecture designed for accurate, interpretable, and computationally efficient lesion detection on CT scans. The framework combines a ResNet18 backbone, a morphological attention mechanism, and an ultra-lightweight classification head to simultaneously enhance diagnostic precision, model transparency, and real-time deployability within IoMT-enabled healthcare environments. An experimental evaluation on the IQ-OTHNCCD lung CT dataset demonstrated superior performance, achieving a classification accuracy of 98.18%, a training accuracy of 99.5%, and a validation accuracy of 98.0%, while maintaining a minimal validation loss of 0.06. When benchmarked against established architectures such as VGG16 (96.82%), MobileNetV3 (96.36%), and ResNet18 (85.00%), NeuroMorphFusion exhibited higher robustness, improved class-wise discrimination—particularly for the “normal” class—and reduced false positives.
The model also proved computationally efficient, converging within 7.81 min at epoch 10, significantly outperforming heavier models such as VGG16, which required over an hour to train. Its interpretability was further validated through Grad-CAM heatmaps that reliably localized clinically relevant lesion regions across all classes, reinforcing its potential for clinician-assisted diagnostics. The study sets the foundation for future adaptive healthcare AI systems that autonomously fine-tune parameters through reinforcement feedback and evolutionary optimization, ensuring consistent clinical trust and hardware portability.
Footnotes
Abbreviations
Acknowledgements
Not Applicable.
Ethics Statement
This study did not involve human participants, human tissue, or animal subjects. Therefore, approval from an institutional review board (IRB) or ethics committee was not required. All data used in this study were publicly available and anonymized, ensuring compliance with ethical standards for secondary data usage.
Authors Contributions
R.O.O.: Conceptualization, Methodology, Software, Data Curation, Writing-Original Draft, Visualization; R.W.B.: Methodology, Data Curation, Formal analysis, Writing-review & editing; P.A.O.: Validation, Investigation, Resources, Writing-review & editing, Supervision; E. V. W.: Resources, Visualization, Validation, Writing-review & editing; C.T.: Software, Formal analysis, Investigation, Supervision, Writing-review & editing.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.