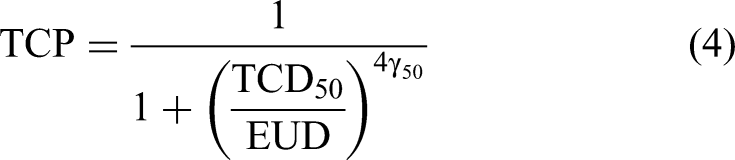Abstract
Introduction
Radiotherapy treatment plans traditionally rely on physical indices like Dose-volume histograms and spatial dose distributions. While these metrics assess dose delivery, they lack consideration for the biological effects on tumors and healthy tissues. To address this, radiobiological models like tumor control probability (TCP) and Normal tissue complications probability (NTCP) are increasingly incorporated to evaluate treatment efficacy and potential complications. This study aimed to assess the predictive power of radiobiological models for TCP in breast cancer radiotherapy and provide insights into the model selection and parameter optimization.
Methods
In this retrospective observational study, two commonly used models, the Linear-Poisson and Equivalent uniform dose (EUD)-based models, were employed to calculate TCP for 30 patients. Different radiobiological parameter sets were investigated, including established sets from literature (G1 and G2) and set with an optimized “a” parameter derived from clinical trial data (a1 and a2). Model predictions were compared with clinical outcomes from the START trials.
Results
The Linear-Poisson model with es lished parameter sets from the literature demonstrated good agreement with clinical data. The standard EUD-based model (a = -7.2) significantly underestimated TCP. While both models exhibited some level of independence from the specific parameter sets (G1 vs. G2), the EUD-based model was susceptible to the “a” parameter value. Optimization suggests a more accurate “a” value closer to -2.57 and -5.65.
Conclusion
This study emphasizes the importance of clinically relevant radiobiological parameters for accurate TCP prediction and optimizing the “a” parameter in the EUD-based model based on clinical data (a1 and a2) improved its predictive accuracy significantly.
Keywords
Introduction
Traditionally, radiation treatment plans have been evaluated by physical dose indices, such as dose volume histogram (DVH) and two-dimensional and three-dimensional spatial dose distributions. However, they focus on dose distribution and do not consider the biological effects of radiation on tumors or healthy tissues. To improve the plane evaluation, researchers are increasingly incorporating radiobiological indices such as tumor control probability (TCP) and normal tissue complications probability (NTCP).1,2
Two prominent TCP models are widely used for radiobiological plan evaluation: phenomenological and mechanistic models. Mechanistic models use mathematical equations based on simplified radiobiology to represent how radiation affects cells. A majority of mechanistic models assume that even a single surviving clonogen will have the potential of growing into a viable tumor. Therefore, TCP is achieved when all clonogenic cells are destroyed. The Poisson dose-response model is an example of this approach. It uses Poisson statistics to calculate the probability of having no surviving clonogens after treatment. 2
Clinical data shows that the TCP, as a function of the dose, typically exhibits a sigmoid shape. The dose-response curve represents a gradual rise, steep increase, and eventual plateauing in TCP with increasing the radiation dose. Phenomenological models capture this relationship using parameters like TCD50 (the dose at which there is a 50% chance of tumor control) and γ50 (the slope of the TCP curve at TCD50). The logistic model is the most common and well-established phenomenological method for estimating the TCP in radiotherapy. 2
Niemierko proposed a more complex model incorporating the logistic function with additional parameters to account for inhomogeneity in dose distribution. This model is known as the “EUD-based TCP model”. Equivalent uniform dose (EUD) represents the dose that, if delivered uniformly to the entire target volume, would produce the same biological effect as the actual non-uniform dose distribution.
3
Originally, EUD was developed for tumors and considered cell survival based on the linear-quadratic (LQ) model. According to Eq. 1, if the survival fraction of tumor cells in a non-uniform field with a dose Di delivered to each subvolume of Vi is assumed to be equivalent to that in a uniform field with an EUD dose applied to the entire volume, the EUD can be determined by taking the natural logarithm of both sides of the equation 4,5:
While used extensively in treatment planning systems (TPS), these models have their limitations. Uncertainties in the biological parameters used in radiobiological modeling may lead to significant uncertainties in the predicted TCP.2,7 In addition, the lack of sufficient clinical data regarding how human tissues and tumors respond to varying radiation doses poses challenges in obtaining accurate estimates for these parameters. As a result, no single set of radiobiological parameters can currently predict clinical outcomes with perfect accuracy, which limits the use of these models as the primary tool for evaluating treatment plans.1,8 To optimize the clinical translation of these models, a comparative assessment of their predictive capabilities using identical datasets is essential. This approach allows for the identification of strengths and weaknesses inherent to each model, ultimately informing the selection of the most reliable tool for plan evaluation.
In this study, we aimed to compare the predictive accuracy of two widely used radiobiological models, the Linear-Poisson and the EUD-based models, for breast cancer radiotherapy. Although the optimization of the “a” parameter in the EUD-based model is a well-established concept, our study focuses on refining this parameter specifically for breast cancer radiotherapy using large-scale clinical trial data. Previous studies have typically employed a special value for “a” (e.g., -7.2), without consideration of the specific tumor characteristics or clinical outcomes associated with breast cancer.9–18 Our work, however, takes a novel approach by recalibrating “a” using clinical data from the Standardization of Breast Radiotherapy Trials (START), providing a more accurate prediction of TCP.19,20 This tailored approach not only improves the predictive accuracy for breast cancer but also opens the door for more individualized treatment planning based on patient-specific parameters.
Material and Methods
Patients
The study protocol was approved by the Institutional Ethics Committee (Ethics code: IR.AJUMS.MEDICINE.REC.1403.035, approved on 2024-09-25). This retrospective observational study analyzed data from 30 patients with early-stage, left-sided invasive ductal breast cancer who underwent treatment at Golestan Ahvaz Hospital, Ahvaz, Iran, between 2021 and 2023. These patients did not have cancer involvement in the lymph nodes above the supra-clavicular or axillary regions. Computed tomography images, treatment plans incorporating calculated dose distributions, and associated patient demographic data were obtained through collaboration with the radiation oncology department. All patient information was anonymized to protect privacy and comply with regulations for using anonymized clinical data in research. Patient informed consent was not required for this retrospective study.
Dose Prescription and Treatment Planning
All patients received whole breast radiation therapy using a 3D-conformal radiation therapy (3D-CRT) technique. The planning target volume (PTV) included the entire breast, extending from the top of the clavicular head to 2 cm below the fold beneath the breast (inframammary line). The PTV was delineated by the mid-sternum and mid-axillary lines as its lateral boundaries.
The treatment planning process utilized the ISOgray treatment planning system (ISOgray® software, Version 4.2.3.65 L, DOSIsoft®, Cachan, France).
The prescribed radiation dose was 50 Gy delivered in 25 fractions using 6 MV photon beams through two tangential fields (beams directed from either side of the body). The analysis of DVHs confirmed that, across all patients, 95% of the PTV received 100% of the dose. DVHs were then generated for each patient as input data for TCP calculations.
TCP Models
A TCP curve is created by plotting some measure of TCP against the total dose. This may be done with clinical or experimental data, or with theoretical models. Even when plotting clinical data, a theoretical model must be used to model the response of tumor cells to radiation therapy. Radiobiological models predict the likelihood of tumor eradication following radiotherapy. There are several models used for calculating TCP in radiotherapy, each with varying levels of complexity and suitability for different scenarios. For this study, we used two models: the Linear-Poisson model and the EUD-based model. TCP in these models depends on key radiobiological parameters, including the equivalent dose in 2 Gy fractions (EQD2), the dose required to achieve 50% tumor control)tumor control dose 50% or TCD50), and the steepness of the dose-response curve (γ50). The EQD2 is derived from the TPS, while TCD50 and γ50 are selected based on published literature. See the subsequent sections for details of each model.
Linear-Poisson Model
One of the most frequently used mechanistic models for calculating TCP under heterogeneous irradiation is the Linear-Poisson (Eq. 2).2,14,21
EUD-based Model
As an alternative to mechanistic models, the EUD-based model is a phenomenological method for estimating TCP (Eq. 4) 2,9,22:
TCP Analysis
The cumulative dose volume histograms (DVH) for each patient were exported from the TPS. The cumulative DVHs were then converted to differential DVHs and EQD2. Then, the inhomogeneous dose distribution of different dose bin Di irradiating partial volume Vi is reduced into a gEUD of the whole tumor volume according to Eq. 5. Finally, the TCP values were calculated using the Linear-Poisson model (Eq. 2) and the EUD-based model (Eq. 4).
Radiobiological Parameters
Three radiobiological parameters for the Linear-Poisson model are
Radiobiological Parameters for Breast Cancer.

* data based on 10 year local relapse rate 19 .
TCP Calculation
For each patient, TCP was calculated using both models and both parameter sets (G1 and G2). The goal was to assess the ability of the models to predict actual patient outcomes. Here, published data from the START trials provided a reference point.
20
The average local recurrence rate (R) was reported as 6.7%[4.9, 9.2] (95% confidence between brackets) after 10-year follow-ups, which translates to TCP of 93.3% using the following equation: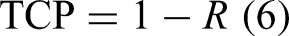
Optimizing the “a” Parameter
The “a” parameter in the EUD-based TCP model represents a critical factor that influences the model's ability to predict TCP accurately. In the existing literature, a value of a = −7.2 has been widely adopted.9–18 However, our thorough review revealed no direct experimental or clinical evidence supporting this value. Instead, this parameter appears to have been perpetuated through repeated citation without verification against robust clinical datasets. Given the lack of empirical validation for a = −7.2, this study aimed to optimize the parameter based on reported clinical trial data. Specifically, we used the 10-year local relapse rate (6.7%) in breast cancer radiotherapy as a reference point for TCP (93.3%).
19
Using this reference, we implemented a numerical bisection method in MATLAB software (Eq. 7) to recalibrate “a” parameter within the context of two distinct parameter sets (G114,15,23–26 and G2
19
). This approach allowed us to propose two new “a” values (a1 and a2), which more accurately reflect the clinical data and improve the performance of the EUD-based model.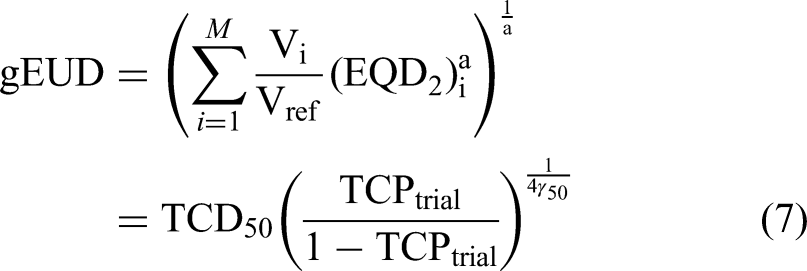
Finally, the TCP values obtained from all six scenarios were compared with data from clinical studies.19,20
Statistical Analysis
Statistical analysis was performed using SPSS version 26 (IBM Corporation, USA). The Kruskal-Wallis and the Mann-Whitney tests were performed to determine if there were significant differences between the groups. A p-value of less than 0.05 was considered statistically significant.
Results
Table 2 summarizes the calculated TCP for all 30 patients using four different radiobiological parameter sets obtained from the literature. These sets are labeled as G1, G2, G1-a0, and G2-a0.
The G1 and G2 Sets were used with the Linear-Poisson model. The G1-a0 and G2-a0 Sets were used with the EUD-based model
Median and Interquartile Range (IQR) of the Calculated TCP within Each Set of Radiobiological Parameters.

As can be seen from this table, the Linear-Poisson model predicted higher values for TCP. The EUD-based model with the G2-a0 parameter set predicted a remarkably low level of tumor control, approximately 68.88% compared to the 93% reported by Guirado et al. 19 It is also observed that the IQR values for both G1-a0 and G2-a0 (using the standard “a” parameter) were significantly larger than those for the Linear-Poisson model.
Figure 1 compares the TCP values obtained from the Linear-Poisson and EUD-based models using the different literature-based parameter sets. Statistical analysis revealed significant differences (p < 0.001) between the two models when the same radiobiological parameters (G1 vs. G1-a0 and G2 vs. G2-a0) were used. However, there were no significant differences in TCP predictions within each model when two different parameter sets (G1 vs. G2 and G1-a0 vs. G2-a0) were used. In other words, TCPs in both models are independent of the radiobiological parameter sets.
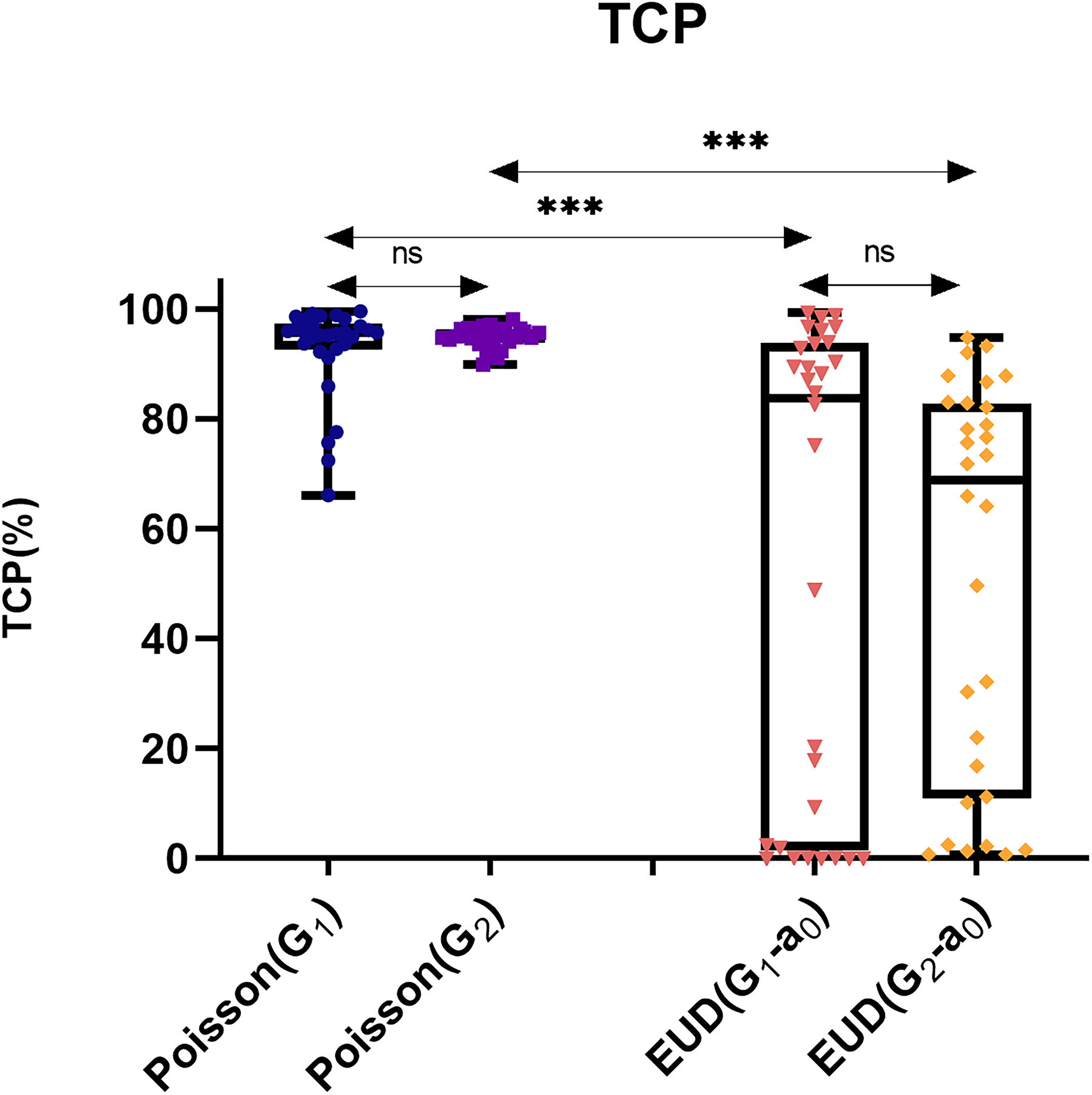
Comparison of the calculated TCP using Linear-Poisson and EUD-based model with different radiobiological parameters obtained from the literature. The differences between all groups are significant except for those marked as ns. (ns: non-significant, ***: P < 0.001). Statistically significant differences were abserved between the two models when using the same radiobiological parameters (G1 vs. G1-a0 and G2 vs. G2-a0). However, no significant differences were observed within each model when different parameter sets (G1 vs. G2 and G1-a0 vs. G2-a0) were applied. This indicates that TCP predictions in both models are independent of the radiobiological parameter sets. Furthermore, as can be seen the EUD-based model exhibits a much higher standard deviation than the Poisson model. In particular, certain calculated TCP values are notably lower than expected.
Based on the reported clinical trial data (local relapse rate of 6.7% or TCP of 93.3% after 10 years), new values for parameter ‘a’ were calculated for both G1 and G2 parameter sets, yielding values of a = -5.65 and a = -2.57, respectively. These new “a” values aimed to achieve a predicted TCP that matches the clinical outcome. Table 3 shows the calculated “a” values and the resulting TCPs using the median “a” for each set. As expected, the TCP predicted using the G2-a2 parameter set (derived from G2 parameters) most closely matches the clinical data, because the G2 parameter set was directly extracted from our reference clinical study.19,20
Median and Interquartile Range (IQR) of the Calculated Parameter “a” and Resulting TCP for Two Different Sets of Radiobiological Parameters.

*Computed TCP by EUD-based model using the median of parameter “a”.
While the Linear-Poisson model with literature-based parameters (G1 and G2) showed better predictive power than the EUD-based model with the same parameter sets (G1-a0 and G2-a0), there were no significant differences between the Linear-Poisson model and the EUD-based model when the optimized “a” parameter (G2-a2) was used (Figure 2).

Comparison of TCP calculated by Poisson model with EUD-based model using two new values of a = -5.65 and a = -2.57 (***: P < 0.001). Only significant differences are displayed. No significant differences were observed between the Linear-Poisson and EUD-based models when the optimized “a” parameter (G2-a2) was applied. Although no statistically significant differences were observed between the two EUD models, the use of the G1-a1 parameter set resulted in a significantly greater standard deviation compared to the G2-a2 parameter set. This indicates that predictions may be less stable across individual patients, leading to potential overestimations or underestimations of TCP for certain patients.
Discussion
Radiobiological models, such as the Linear-Poisson and EUD-based models, are essential tools for predicting TCP in radiotherapy. These models estimate the likelihood of tumor control based on specific radiobiological parameters. While both models offer valuable insights into treatment effectiveness, their accuracy is influenced by how well their parameters reflect clinical outcomes. Understanding how sensitive these models are to specific parameters is crucial for ensuring reliable predictions that can guide treatment decisions in clinical practice.
In this study, we compared the predictive accuracy of the Linear-Poisson and EUD-based models using two sets of parameters (G1, G2) and clinical data from the START trials. The Linear-Poisson model consistently showed better agreement with clinical outcomes regardless of whether G1 or G2 parameter sets were used. In contrast, the EUD-based model's predictions were less accurate, showing significant deviations from clinical results for tumor control, regardless of the parameter set used. This highlights the EUD-based model's sensitivity to the “a” parameter and the need for its careful optimization to improve predictive accuracy.
The EUD-based model with the standard parameter set (a0= -7.2) used in previous studies 9–18 significantly underestimated TCP compared to the clinical data. However, optimizing the “a” parameter in the EUD-based model based on the clinical data (a1=-5.65 and a2=-2.57) improved its predictive accuracy substantially. In comparison between the two suggested values for “a” parameter, employing the G1-a1 parameter set results in a markedly greater dispersion in the predicted TCP values, as can be seen from Figure 2 and Table 3. This suggests that predictions might be less consistent for individual patients, potentially leading to overestimation/underestimation of TCP for some patients. These findings highlight the importance of the parameter selection, especially in using the EUD-based model for breast cancer radiotherapy.
Our proposed method for calculating the “a” parameter based on clinical data (a1=-5.65 and a2=-2.57) offers a promising approach to improve the accuracy of the EUD-based model for breast cancer radiotherapy. This approach aligns with the observation that the EUD-based model with parameters derived entirely from clinical data (G2-a2) achieved the best agreement with observed TCP.
The primary limitation of this study lies in the small sample size, which was restricted to pre-existing clinical data from the START trials. While our approach focuses on introducing a method to precisely estimate the “a” parameter, the true potential of this method can only be fully realized with access to extensive patient-specific DVH datasets. With larger, more diverse samples, our method could potentially determine optimal “a” values for various cancers, allowing for more accurate clinical application of this parameter in combination with γ50 and TCD50.
This study demonstrates the importance of identifying relevant radiobiological parameters and adjusting them appropriately to better predict outcomes in breast cancer radiotherapy.
Conclusion
This study proposes an optimized approach for determining the “a” parameter in the EUD-based TCP model, which may significantly improve its predictive power in breast cancer radiotherapy. Using clinical trial data with a reported 10-year TCP of 93.3%, two new values for “a” (-5.65 and -2.57) were calculated for G1 and G2 parameter sets, respectively. The recalibration of the “a” parameter better aligns model predictions with observed clinical outcomes, addressing a key limitation in previous studies where the commonly used value of -7.2 lacked empirical support.
Our findings demonstrate that the recalibrated “a” values reduce discrepancies between predicted and actual TCPs, outperforming previous parameter estimates and providing a more reliable framework for clinical applications. This improvement may enhance the utility of radiobiological models in tailoring personalized treatment plans.
While the study focused on breast cancer, the methodology presented here may be adaptable to other cancer types, offering a pathway to refine radiobiological models and ultimately improve patient outcomes across diverse clinical scenarios. Future studies can further validate this approach by exploring larger datasets and applying it to advanced radiotherapy techniques such as Volumetric modulated arc therapy (VMAT) and intensity-modulated radiation therapy (IMRT).
Supplemental Material
sj-pdf-1-tct-10.1177_15330338251329103 - Supplemental material for Optimization of the Dose-Volume Effect Parameter “a” in EUD-Based TCP Models for Breast Cancer Radiotherapy
Supplemental material, sj-pdf-1-tct-10.1177_15330338251329103 for Optimization of the Dose-Volume Effect Parameter “a” in EUD-Based TCP Models for Breast Cancer Radiotherapy by Farshid Mahmoudi, Nahid Chegeni, Ali Bagheri, Amir Danyaei, Samira Razzaghi, Shole Arvandi, Amal Saki Malehi, Bahare Arjmand, Azin Shamsi, and Majid Mohiuddin in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Acknowledgment
We sincerely acknowledge Prof. Mohammad Javad Tahmasebi Birgani for his expert guidance in the physics of radiotherapy. We also extend our sincere thanks to the personnel of Radio-Oncology department in Golestan Hospital for their cooperation throughout this research project. Finally, we appreciate the assistance provided by Miss Kh. Housseini.
ORCID iD
Compliance with Ethical Guidelines
This study was approved by the Research Ethics Committee of the School of Medicine, Ahvaz Jundishapur University of Medical Sciences (Approval ID: IR.AJUMS.MEDICINE.REC.1403.035). The study was approved on September 25, 2024.
Author Contributions
F.M.: Conceptualization, Methodology, Writing—Original draft preparation, Writing—Reviewing and Editing; N.Ch.: Supervision, Conceptualization, Methodology, Funding acquisition, Writing—Original draft preparation, Writing—Reviewing and Editing; A.B.: Investigation, Conceptualization, Writing—Original draft preparation; A.D.: Methodology, Writing—Original draft preparation; S.R.: Investigation; Sh.A.: Investigation; A.S.M.: Formal analysis; B.A.: Data curation; A.Sh.: Data curation; M.M.: Investigation, Writing—Original draft preparation, Writing—Reviewing and Editing.
Funding
This work was supported by the Office of Vice-Chancellor for Research of Ahvaz Jundishapur University of Medical Sciences with grant number U-03216.
Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.