Abstract
Prostate cancer (PCa) is one of the most prevalent and deadly cancers among men, particularly affecting men of African descent and contributing significantly to cancer-related morbidity and mortality worldwide. The disease varies widely, from slow-developing forms to highly aggressive or potentially fatal variants. Accurate risk stratification is crucial for making therapeutic decisions and designing adequate clinical trials. This review assesses a broad spectrum of diagnostic and prognostic biomarkers, many of which are incorporated into clinical guidelines, including the Prostate Health Index (PHI), 4Kscore, STHLM3, PCA3, SelectMDx, ExoDx Prostate Intelliscore (EPI), and MiPS. It also highlights emerging biomarkers with preclinical support, such as urinary non-coding RNAs and DNA methylation patterns. Additionally, the review explores the role of tumor-associated microbiota in PCa, offering new insights into its potential contributions to disease understanding. By examining the latest advancements in PCa biomarkers, this review enhances understanding their roles in disease management.
Background
Prostate cancer (PCa) is a severe global health problem, the second most prevalent cancer in men and the leading malignancy among men of African descent, with an increase in mortality rates.1,2 Although men diagnosed with early-stage PCa have a 99% survival rate, those diagnosed at a later stage experience a significantly lower 5-year survival rate of 30%, along with a concomitant decline in quality of life. 3 These stark differences in survival rates highlight the critical need for biomarkers that can efficiently detect PCa early and accurately stratify the disease's aggressiveness. 4 The discovery of prostate-specific antigen (PSA) as a diagnostic biomarker represents a watershed in the early detection of PCa.
As a widely used biomarker for PCa screening, PSA lacks disease specificity, leading to several limitations, such as an inability to distinguish between clinically significant and insignificant PCa. 5 Despite its effectiveness in reducing PCa mortality, PSA-based screening has raised concerns about overdiagnosis and overtreatment, particularly due to the difficulty in differentiating between indolent and aggressive forms of the disease.6,7 In an attempt to address the PSA challenge, other tests like Prostate Health Index (PHI) and 4Kscore combines PSA-based markers with clinical information like digital rectal examination (DRE), prostate size, PSA density, and age, thereby enhancing PCa detection. 8
Over the years, numerous studies have identified and validated promising biomarkers for detecting clinically significant PCa and predicting disease aggressiveness. This review will provide an overview of clinically available non-invasive biomarkers for PCa diagnosis and screening while highlighting biomarkers for aggressiveness stratification. It will also provide insight into promising biomarkers that could enhance diagnostic accuracy and clinical outcomes (Figure 1). A search of literature was carried out using databases such as PubMed and SCOPUS, focusing on identifying biomarkers for improved PCa detection and stratification of aggressiveness.
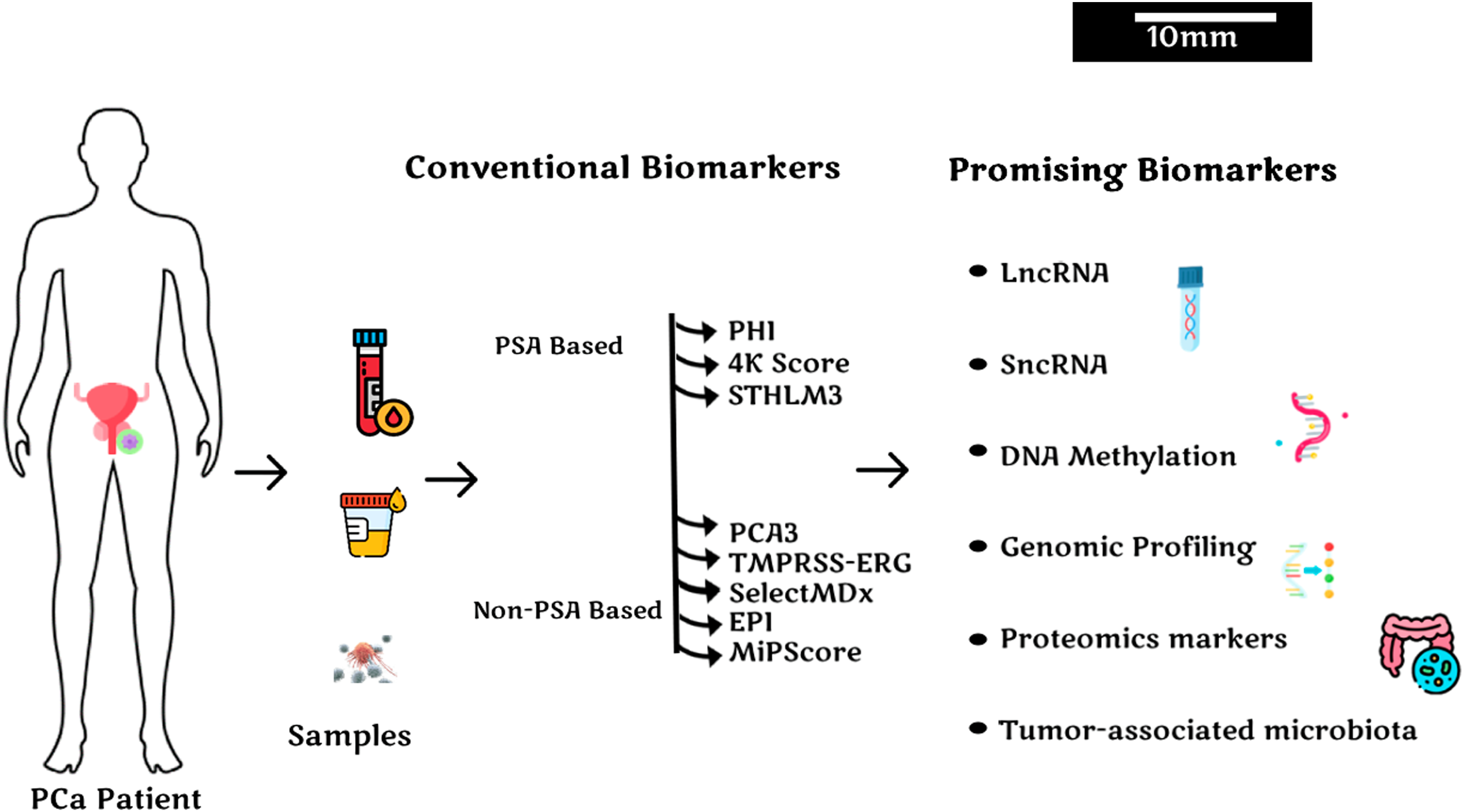
Clinically approved and promising biomarkers for prostate cancer detection and aggressiveness stratification.
Conventional Biomarkers for PCa
A significant number of biomarkers have been identified, and new ones are still being discovered to improve the early detection of PCa and to stratify aggressive cases from indolent ones; however, only a few have been approved and are currently used clinically. Table 1 summarizes the currently available non-invasive biomarkers for PCa detection and screening.
Summary of Current Biomarkers.

Prostate Specific Antigen (PSA) Based Biomarkers
Prostate Health Index (PHI)
PHI is an FDA-approved blood test for men with PSA levels between 2–10 ng/ml. 14 PHI has been demonstrated to reduce the need for prostate biopsies by differentiating patients with aggressive PCa from those with the indolent type, who are better recommended for active surveillance rather than intensive treatment options. 15 The test is carried out by measuring the levels of three distinct types of PSA; total PSA (tPSA), free PSA (fPSA), and [-2] pro-prostate-specific antigen (p2PSA), providing higher specificity for clinically significant PCa. 16 p2PSA is associated with increased proteolytic activity and altered extracellular matrix interactions typical of malignant prostate cells and is a more specific indicator of aggressive prostate cancer compared to total PSA. 17
In 2010, Beckman Coulter developed this automated immunoassay with an index that can be calculated using the formula PHI = (p2PSA/fPSA)×√tPSA). By combining these three measurements mathematically, PHI can distinguish between benign prostate conditions and PCa in individuals with normal DRE results and serum levels of PSA ranging from 4 to 10 ng/ml.14,18 PHI scores are associated with the likelihood of finding clinically significant disease, where a PHI score between 0 and 26.9 indicates a 10% chance of cancer, while a score above 55 indicates a 50% chance of cancer during biopsy.19,20
A research conducted with 892 men without previous PCa, who had a normal DRE and a PSA level of 2–10 ng/mL, showed that a PHI score of 80% showed 95% sensitivity and a significantly higher specificity (area under curve (AUC) of 0.703) than tPSA and %fPSA. This confirms that an increase in PHI score is proportional to an increased risk of aggressive PCa and positive biopsy.14,21 Additional studies have indicated that combining PHI and %p2PSA into multifactorial models with PSA levels, patient age, and prostate volume improves PHI predictive ability for PCa22,23 with enhanced AUC values from 0.762 to either 0.802 or 0.815 using logistic regression analysis or an artificial neural network.9,24
However, relevant research findings showed that the efficacy of PHI in stratifying risk for PCa may vary across ethnic groups and regions. 25 For instance, the reference ranges for PHI categorize <26.9 as low-risk, 27–55 as intermediate-risk, and >55 as high-risk. These ranges were primarily derived from European and North American datasets and may not be universally applicable. 14 Additionally, PSA levels directly influence PHI values, so regional variations in PSA cutoff values must be considered when interpreting PHI for PCa. In Eastern Asian populations, such as Japanese and Chinese men, a PSA cutoff value of 3 ng/mL is typically used. 26 Given the lower prevalence of PCa in these populations, it may be necessary to apply even lower cutoff values for PHI to improve its accuracy. 26 A study suggested that in the Asian population, a PHI score below 30 is associated with a lower probability of high-grade PCa (HGPC), whereas a PHI score above 30 significantly increases the likelihood of HGPC.26,27 Conversely, African-American men, due to their higher PSA levels and increased propensity for aggressive PCa, may require different PHI thresholds.27,28 On the other hand, African–American men, because of the higher levels of PSA and increased propensity to aggressive PCa, may have different thresholds for PHI. 29 A genomic study by Babajide et al (2021) involving a cohort of Black men demonstrated that a PHI cutoff point of ≥28 is linked to high sensitivity and an increased risk of HGPC. 30 This underscores the urgent need for ongoing research and validation studies to ensure that PHI provides reliable results across diverse populations.
4Kscore
The 4Kscore is a blood test that has accrued significant attention for its enhanced ability to assess the risk of aggressive PCa. 31 It combines four biomarkers (tPSA, fPSA, intact PSA (iPSA), and human kallikrein 2 (hK2)) within a proprietary algorithm, along with clinical factors such as DRE results and patient age, to generate a risk score ranging from 0% to 100%. 32 This test was developed primarily to address the issues of overdiagnosis and overtreatment associated with PSA screening, particularly the challenge of identifying indolent cancers that may not require immediate intervention.32,33
The initial validation of the 4Kscore was conducted in 2008 with a cohort of 740 men from the Göteborg arm of the European Randomized Study of Screening for Prostate Cancer (ERSPC). 34 These men, who had PSA levels of 3 ng/mL or higher and had not been previously screened, underwent a 6-core biopsy. When combined with age, DRE, and tPSA, the 4K panel significantly improved the prediction of high-grade PCa (Gleason score ≥ 7), raising the AUC from 0.68 to 0.83, which emphasis its potential utility in clinical practice.34,35 Further research involving 740 individuals with similar PSA levels demonstrated that incorporating the 4K panel with existing clinical factors increased the AUC for detecting high-grade PCa from 0.87 to 0.90, highlighting the test's robustness across different.21,36
In clinical settings, the 4Kscore has demonstrated its ability to reduce unnecessary biopsies by identifying men at low risk for aggressive PCa. For example, a study by Punnen et al (2015) found that using the 4Kscore led to a 64% reduction in unnecessary biopsies compared to traditional PSA testing alone. This reduction not only alleviates patient anxiety and discomfort but also decreases the risks associated with biopsy procedures, such as infection and bleeding. 37
The robust diagnostic ability of the 4Kscore test contributes to reducing healthcare costs and minimizes unnecessary biopsies. 38 Although the likelihood of the 4Kscore fully replacing biopsies is slim, several studies have explored its use in conjunction with multiparametric MRI (mpMRI).39–41 Marzouk et al found that using the 4Kscore before mpMRI could reduce the need for MRI in some men, particularly those with a 4K score ranging from 5% to 23%. However, as this study presents a conceptual approach, validation through prospective research could further reinforce these preliminary findings. 41 Another study by Falagario et al (2021) showed that using the 4Kscore and mpMRI simultaneously resulted in a 34.2% reduction in the number of biopsies among biopsy-naïve patients. 40 Based on these studies, the combined use of the 4Kscore and mpMRI can effectively reduce the necessity for additional biopsies, as both tests complement each other.42,43 This combined approach is particularly valuable for patients with intermediate PSA levels (2-10 ng/mL), where the risk of false positives and negatives is higher. 40
Despite its advantages, the 4Kscore is not without limitations. The test's reliance on complex algorithms and its integration of multiple biomarkers may limit its accessibility and ease of use in routine clinical practice, especially in resource-limited settings. 44 Additionally, while the 4Kscore has been validated in various populations, its performance may vary across different demographic groups, necessitating further research to ensure consistent accuracy across diverse patient populations. 34
STHLM3
The Stockholm-3 (STHLM3) test represents a significant advancement in PCa screening, by integrating PSA levels with a multifactorial approach that includes other protein biomarkers, genetic markers, and clinical data. 45 Designed specifically to address the limitations of PSA testing (particularly the difficulty in distinguishing between aggressive and indolent prostate cancers) the STHLM3 test combines six plasma protein biomarkers (tPSA, fPSA, iPSA, hK2, Macrophage Inhibitory Cytokine 1 (MIC1), and Beta-Microseminoprotein (MSMB)) with over 100 genetic markers (single nucleotide polymorphisms, or SNPs) and clinical varibles. 46 This comprehensive approach provides a more accurate risk assessment compared to PSA alone. 47 By incorporating genetic aberrations known to significantly influence PCa development and progression, the STHLM3 test enables better differentiation between aggressive and indolent forms of the disease, thereby facilitating more accurate and personalized patient care. 48 Additionally, integrating the STHLM3 test with advanced imaging techniques, such as multiparametric MRI (mpMRI), has shown promise in enhancing the detection of clinically significant cancers while minimizing unnecessary biopsies. 49
However, the STHLM3 model has limitations related to its validation population, which may restrict the test's generalizability to broader populations. 50 To address this limitation and ensure the STHLM3 test's applicability across diverse groups, research has focused on validating the test in men from various ethnic backgrounds and geographic regions. 50 While the test was initially validated in Swedish populations, subsequent studies in European, North American, and Asian cohorts have generally supported its effectiveness. 51 A recent study that included Asian (16%, 350), Black or African American (24%, 505), Hispanic or Latino (14%, 305), and non-Hispanic or non-Latino White (46%, 969) participants demonstrated that the STHLM3 test yielded consistent results across these racial and ethnic subgroups. The study also found that the STHLM3 test significantly reduces unnecessary prostate biopsies while maintaining similar sensitivity to PSA in detecting clinically significant prostate cancer. 52
Despite its advantages, the STHLM3 test has its limitations. Its reliance on complex algorithms that integrate numerous biomarkers and genetic factors, along with the higher cost compared to PSA testing, may limit its accessibility and ease of use in routine clinical practice, especially in resource-limited settings. 53 Additionally, while the test is currently only commercially available in Sweden, efforts are underway to evaluate its integration into clinical guidelines for prostate cancer screening in other countries. 44 As more evidence of its effectiveness emerges, it is likely that the STHLM3 test will be recommended as part of a multi-step screening process, particularly for men with elevated PSA levels or those at higher genetic risk for prostate cancer. 46 As more evidence of its effectiveness emerges, it is likely that the STHLM3 test will be recommended as part of a multi-step screening process, particularly for men with elevated PSA levels or those at higher genetic risk for prostate cancer. 54
Non-PSA-Based Biomarkers
Prostate Cancer Antigen 3 (PCA3)
Prostate Cancer Antigen 3 (PCA3) is a non-coding RNA that is specific to the prostate and is significantly upregulated in PCa, making it a valuable target for diagnostic testing. PCA3 plays a key role in cancer cell survival by modulating the transcriptional activity of androgen receptor target genes and androgen receptor signaling.55,56 The PCA3 test has emerged as a significant non-invasive biomarker for PCa detection, particularly in guiding decisions regarding repeat biopsies. 57 The PCA3 score enhances PCa diagnosis, especially in determining whether patients should undergo a repeat biopsy. 58 Additionally, PCA3 expression is not affected by other prostate conditions, such as prostatitis, or variations in prostate size, thereby enhancing its utility as a marker for PCa. 59 Increased PCA3 expression is associated with a decrease in PRUNE2, a gene regulated by long non-coding RNAs in PCa. 60 PRUNE2 functions as a tumor suppressor, and its downregulation due to elevated PCA3 levels contributes to increased cell growth, thereby reinforcing the association of PCA3 with PCa. 61
The PCA3 test involves measuring the levels of PCA3 mRNA and PSA mRNA in the first urine sample collected after a DRE. An elevated PCA3 score indicates a higher likelihood of PCa. 12 This test is particularly useful in men with elevated PSA levels or abnormal DRE findings, as it helps refine the decision to proceed with a repeat biopsy. 62
The clinical relevance of PCA3 is underscored by its inclusion in the 2020 guidelines from the European Association of Urology (EAU) and the National Comprehensive Cancer Network (NCCN), where it is recommended as part of the decision-making process for repeat biopsies. 64 This endorsement highlights the growing acceptance of PCA3 testing in clinical practice, especially for its ability to reduce the number of unnecessary repeat biopsies. 63 This endorsement highlights the growing acceptance of PCA3 testing in clinical practice, particularly for its ability to reduce the number of unnecessary repeat biopsies. 10
However, the PCA3 test has limitations. It is not recommended as a general screening tool for PCa, as it is specifically elevated in individuals with PCa and not in the general population. 31 Additionally, the ideal cutoff value for the PCA3 score to guide biopsy decisions varies among studies. Some studies have used PCA3 score values of 20, 35, and 50, resulting in sensitivity rates ranging from 52% to 82%, while specificity rates varied between 79% and 89%.11,13,64 A meta-analysis from 2011 found the optimal cutoff to be a PCA3 score of 20, resulting in 72% sensitivity and approximately 53% specificity. 65 Due to this variability, there is no gold standard for the cutoff determination of PCA3.
Given these challenges, clinicians need to consider PCA3 as part of a broader diagnostic approach, integrating it with other biomarkers and clinical data, such as PSA levels, DRE findings, and patient history. 66 It is suggested that urinary biomarkers like TMPRSS2 and urinary exosomes could also serve as ideal combination test to PCA3 test, offering an easily assayed combination that enhances diagnostic accuracy. While PCA3 testing offers valuable insights, its utility could be enhanced by further standardization and validation in diverse patient populations. More so, combining PCA3 with other emerging biomarkers and imaging techniques, such as mpMRI, could improve its accuracy and reliability, providing a more robust tool for PCa management. 67
Transmembrane Protease Serine 2 - ETS-Related Gene (TMPRSS2-ERG)
A significant proportion of PCa cases involve a distinct genetic alteration in which an androgen-regulated gene, such as TMPRSS2, fuses with an oncogene like ERG from the E26 Transformation-Specific (ETS) transcription factor family, making it the most frequent fusion event in prostate cancer. This particular fusion of TMPRSS2-ERG has been detected in around half of the prostate tumors collected during surgeries.68,69 However, reports of racial disparities in the prevalence of this fusion limit its global utility. A meta-analysis revealed that the prevalence of TMPRSS2-ERG fusion in prostate cancer is highest in men of European descent (49%), followed by Asian (27%) and African (25%) populations. 70 Interestingly, a subgroup analysis of different fusion types further demonstrated a significant association between deletion-type fusion and the severity of PCa, with deletion-type fusion being more common in African patients. 71
Despite the racial disparity in TMPRSS2-ERG fusion, it remains highly useful due to its prevalence in PCa and its functional and clinical relevance. 72 In PCa cells with this fusion, a hybrid gene is formed, leading to the overexpression of ERG protein, which disrupts normal cellular functions and contributes to tumor development and progression. 72 Research on tumor evolution in prostate cancer patients indicates that TMPRSS2-ERG fusions play a significant role in the early stages of prostate tumor development. 73 In clinical practice, the identification of TMPRSS2-ERG fusion is frequently used alongside PSA testing and DRE to identify individuals with a high likelihood of having PCa and to inform treatment decisions. 74 Notably, a study involving 516 men in the developmental cohort (mean age 62 years, range 33-85 years) demonstrated that combining urinary TMPRSS2-ERG and PCA3 testing at specified thresholds significantly improved the specificity for detecting aggressive PCa from 18% to 39%. 61 This combined testing approach has the potential to reduce healthcare costs while effectively identifying cases of aggressive PCa.
PCA3 and TMPRSS2-ERG have been integrated into two distinct multiplex assays: the Michigan Prostate Score (MiPS) and ExoDx Prostate Intelliscore (EPI). The MiPS was developed by combining PCA3 and TMPRSS2-ERG with serum PSA, resulting in a score that effectively identifies high-grade tumors. 75 A recent study by Vickers et al (2024) demonstrated that the AUC values for the MiPS2 models, an upgraded version of MiPS that includes 18 genes, showed meaningful improvements compared to PHI, 4Kscore, or MiPS. 75 The MiPS2 evaluation demonstrated higher diagnostic accuracy compared to existing tests. It also showed the potential to avoid unnecessary additional testing, such as imaging or biopsy, in 35% to 51% of patients while maintaining high sensitivity for detecting HGPC. 75
The EPI test involves analyzing the expression of PCA3, SPDEF (SAM pointed domain-containing Ets transcription factor), and TMPRSS2-ERG in exosomes obtained from a patient's urine. 76 Exosomes are small vesicles with double-lipid membranes that encapsulate portions of the parent cell's cytoplasm. They are shed into biofluids like blood and urine, and are rich in proteins and RNAs, making them relevant for profiling RNA expression in tumor cells due to their representation of the origin cells and protection of mRNA during processing. 77
Two independent trials by McKiernan et al (2014) demonstrated that EPI has higher sensitivity than PSA and current risk calculators (Standard of Care model, Prostate Cancer Prevention Trial Risk Calculator [PCPT-RC], ERSPC-RCs) for detecting high-grade prostate cancer (Gleason score ≥ 7) on initial biopsy.78,79 Additionally, EPI showed an AUC of 0.700 and an NPV of 90.1% for diagnosing high-grade prostate cancer using a threshold of 15.6, helping to avoid unnecessary biopsies in 26.0% of patients. Given its high NPV and AUC, EPI is effective as an exclusionary tool. 80 A follow-up study by Tutrone confirmed that over 92.0% of patients with an EPI score < 15.6 were low-risk and less likely to progress to high-grade prostate cancer within 2.5 years, compared to those with an EPI score > 15.6 (7.9% vs 26.8%, P < 0.001). 81 EPI's ability to reflect long-term disease progression makes it valuable before biopsy and potentially useful for patients under active surveillance.
SelectMDx
SelectMDx is a urine test that measures mRNA levels of two PCa genomic biomarkers: HOXC6 and DLX1. It is used to detect PCa with a Gleason score of 3 + 4 and higher grades, with a sensitivity of 76%. 82 Studies have shown that the mRNA levels of DLX1 and HOXC6 are involved in the onset of PCa and are associated with its aggressiveness. 83 The predictive accuracy of SelectMDx has been well-validated in numerous studies. A prospective investigation found that using SelectMDx as a risk stratification tool in biopsy-naïve men could prevent 38% of unnecessary biopsies while missing only 10% of high-grade PCa. 84 Similarly, Hendriks et al (2017) reported a notable area under the curve (AUC) of 0.83 (95% CI: 0.77-0.89) with SelectMDx for predicting PCa.
Combining clinical factors such as age, PSA, PSA density, family history, and DRE with the SelectMDx test was reported to increase its accuracy (Negative Predictive Value) for excluding PCa to 98%, potentially reducing unnecessary biopsies by 42%. 84 Additionally, in a study involving 5157 patients from ten European countries, SelectMDx produced a negative result in 40.72% of cases, sparing a significant portion of patients from unnecessary biopsies. One proposed approach is to perform mpMRI only in individuals with a positive SelectMDx result. 82 This integrated approach leverages the superior predictive value of SelectMDx to enhance diagnostic accuracy and reduce unnecessary procedures.
Promising Biomarkers for Prostate Cancer Detection and Aggressiveness Stratification
Recent advancements in biomarkers hold immense promise for early diagnosis, precise prognosis, and treatment strategies in PCa. While established markers remain a focus for clinicians, emerging biomarkers have captured significant interest. Among these are markers of proteomics, RNAs which include ncRNAs, LncRNA, sncRNAs, DNA methylation, Genomic profiling, and tumor-associated microbiota, origin.
Protein and Peptide Biomarkers
Protein and peptide biomarkers have also shown promise in PCa diagnosis and prognosis. These biomarkers provide critical information on tumor behavior, thereby distinguishing between indolent and aggressive forms of the disease. 85 For instance, proteins involved in immune response, bone remodeling, and oxidative stress, such as L-selectin, osteopontin, and ceruloplasmin, were found to be elevated in PCa patients. Their increased levels correspond to higher Gleason scores, underscoring their potential in grading the severity of PCa. 86 GOAT (ghrelin O-acetyltransferase) has been implicated in PCa, due to its role in modifying ghrelin, a hormone involved in regulating appetite and energy metabolism. These modifications can lead to tumor growth and progression by enhancing metabolic processes, promoting angiogenesis, and contributing to cancer cell proliferation and survival. The study showed that GOAT outperformed PSA in detecting PCa, and when combined in a panel with key variables like age, DRE, and testosterone, the performance increased significantly. 87
Furthermore, Deng et al highlighted chymotrypsin-like activity as a significant marker for assessing PCa aggressiveness. The study demonstrated that elevated chymotrypsin-like activity was associated with more aggressive PCa compared to indolent cases. 88 An 18 inflammatory and cytokines serum biomarkers panel including vascular endothelial growth factor (VEGF), interleukin-6 (IL-6), and tumor necrosis factor-alpha (TNF-α) enhanced the detection of aggressive PCa. 89 These findings collectively demonstrate the growing importance of protein and peptide biomarkers in advancing PCa diagnosis and prognosis.
RNA Biomarkers
Messenger RNA (mRNA)
mRNA plays a critical role in the translation process of DNA into proteins, which are essential for various cellular functions. 90 Since EVs carry molecules like mRNA, they serve as a promising source of biomarkers. The EV has recently emerged as a promising source of biomarkers in the blood and urine of PCa patients. 91 Cancer-derived EVs contain altered cargo that plays a crucial role in tumor development, proliferation, metastasis, and treatment resistance, 92 with two mRNA markers (GLO1 and NKX3-1)showing potential in distinguishing PCa from BPH, with AUCs of 0.68 and 0.82 in the test cohort, and 0.73 and 0.65 in the validation cohort. When combined in a biomarker model, these markers achieved AUCs of 0.85 and 0.71 in the test and validation cohorts, respectively. 93
Non-Coding RNAs (ncRNA)
ncRNA do not encode proteins like mRNA, but plays regulatory roles in gene expression and cellular processes and are categorized based on size into long-non-coding RNA (LncRNA), and small-non-coding RNA (SncRNA). 94 Recent studies demonstrate that mRNAs often lack specificity for tissue type and disease stage. In contrast, ncRNA exhibits remarkable tissue and stage specificity, making them valuable molecular biomarkers for cancer. 95 Most ncRNA-based liquid biopsy biomarkers are being explored for diagnostic and screening purposes. Due to their frequent overexpression in cancers, ncRNAs are also promising candidates for monitoring cancer progression and recurrence. Studies have shown the effectiveness of circulating ncRNAs, such as lncRNAs and SncRNAs in cancer monitoring.96,97 The two main categories of ncRNAs based on size are:
Long Non-Coding RNAs (LncRNAs)
LncRNAs are noncoding transcripts of 200 or more nucleotides long. They are involved in almost all biological and pathological processes, including cellular transportation, gene expression (chromatin reprograming and transcriptional regulation), cell proliferation, tumorigenesis, and malignant transformation. 98 Due to their critical role in cancer and their tissue- and tumor-specific expression, LncRNAs are excellent candidates for molecular biomarkers. 99 Differential expression of LncRNAs has been explored in PCa patients, with PCA3 being a notable PCa biomarker approved by the FDA. 100
The significance of LncRNAs such as PCAT1, PCGEM1, SChLAP1, and PCAT6 in PCa is well-established. 101 PCAT1, a recently identified LncRNA, is strongly correlated with cancer depth, lymph node spread, and distant metastasis, making it a valuable biomarker for cancer staging. 102 SChLAP1, a highly expressed marker of aggressive PCa aids in distinguishing aggressive PCa from indolent ones. It plays a role in cancer cell invasiveness and metastasis by antagonizing the tumor-suppressive SWI/SNF chromatin-modifying complex. 103 Aside from its prognostic potential, a high level of SChLAP1 expression is associated with adverse clinicopathological features, including higher Gleason grade, cribriform growth, and advanced pathology stage, thereby making SChLAP a predictor of biochemical recurrence in PCa.101,104 Moreover, genetic variation in some LncRNA has been linked with the progression of PCa, for example, single nucleotide polymorphisms (SNPs) within PCAT6 and PCGEM1 have been linked to an increased risk of PCa. 105
A study by Diamantopoulos et al (2023) showed dysregulated expression of 28S rRNA-derived fragments (rRFs), a class of non-coding RNAs, was identified in PCa tissues when compared to benign prostatic hyperplasia (BPH). 106 Hence, they identified the potential role of rRFs in stratifying aggressive PCa from indolent PCa. Also, rRFs dysregulation has been associated with tumor clinical parameters such as tumor stage, tumor grade, and patient prognosis, and an increased expression of rRFs in cancer is correlated with tumor relapse. Furthermore, the study showed that rRFs had an accuracy of AUC of 0.688 in detecting aggressive PCa. 106
Small Non-Coding RNAs (sncRNAs)
Among small non-coding RNAs, miRNAs are considered one of the most promising non-invasive biomarkers due to their high abundance and natural stability across various bodily fluids (blood, urine, stool, and saliva). 107 miRNAs play a crucial role in PCa diagnosis and prognosis, with deregulated expression patterns observed in cases of castration-resistant PCa. 10 For example, miR-21, miR-221, miR-192, and miR-375 are related to the prognosis of PCa patients 108 while Manoj et al (2023) described the potential of miRNA 100, 141, and 1290 panels as diagnostic tools for significantly distinguishing BPH and PCa from healthy people. 109 Additionally, Barcelo and colleagues 110 evaluated a panel of eight PCa-associated miRNAs in seminal plasma and assessed their utility as biomarkers for PCa detection and risk stratification. They identified a multivariate model consisting of PSA, MiR-142-3P, MiR-233-3P, MiR-93-5P, and MiR-30D-5P which improves the specificity of PSA for PCa screening. Models using PSA, MiR-93-5P, and MiR-30D-5P were effective for predicting disease aggressiveness. 110
DNA Methylation Biomarkers
DNA methylation patterns have emerged as significant diagnostic biomarkers for PCa. DNA Methylation is an epigenetic modification involving the addition of a methyl group to cytosine residues, influencing gene expression. 111 Aberrant DNA methylation patterns are well-documented in cancerous cells and have been established as potential diagnostic biomarkers for PCa. 112 Specific regions with predominant DNA methylation alterations have been identified, offering insights into the disease's stage and aggressiveness. 112 For instance, the Prostate Cancer Urinary Epigenetic (ProCUrE) assay was developed based on a two-gene model (HOXD3 and GSTP1) methylation assay mainly for the detection of clinically significant PCa. 113 Furthermore, methylation-specific assays when combined with liquid biopsy techniques, enable the identification of epigenetic changes in circulating tumor DNA or other bodily fluids. 114 For example, DNA methylation differentiates PCa in different Gleason stages, with AUC values of 0.63 and 0.87 for Gleason 6 versus Gleason 3 + 4 and Gleason 4 + 3 respectively. 115 Identifying specific methylation profiles linked to various PCa types offers potential for personalized therapies and revolutionizes PCa management.
Genomic Profiling
Genomic profiling of PCa has uncovered a continuum of clinically relevant molecular changes, such as alterations in the DNA damage repair pathway and PTEN/PI3 K signaling. These changes are connected to poor clinical outcomes in PCa. 116 For instance, SNPs in genes like PCAT19, PCGEM1, and PRNCR1 have been linked to PCa susceptibility, 117 while germline copy number variations (CNVs) on chromosomes 8 and 9 are associated with PCa aggressiveness, with an AUC of 0.63. 118 A seven-gene panel identified by Liong et al (2012) detects aggressive PCa with an AUC of 0.82, specificity of 98%, and sensitivity of 83%. 119
Tumor-Associated Microbiota
Microbial communities residing within and around tumor tissues provide critical insights into tumor behavior, influencing both the progression and aggressiveness of the disease. 120 Recent research has linked several cancers to microbiota, particularly the gut microbiota (GM).120,121 In 2018, Golombos et al observed a significant prevalence of Bacteroides massiliensis in the GM of PCa patients while observing a similarity in the diversity of GM in PCa patients and healthy controls. 122 In 2022, a similar report identified variations in the relative abundance of bacteria signatures at the phylum level between PCa patients and healthy individuals. 123 These results indicate a significant correlation of GM with PCa. Also, research has linked elevated estrogen levels to an increased risk of PCa due to estrogen's role in the onset and progression of PCa. 124 Studies have shown that some gut bacteria, known as the estrobolome, can metabolize and produce estrogen, influencing the body's estrogen levels and affecting PCa progression. 125
Conclusion
The approach to diagnosing PCa is advancing, underscoring the importance of a comprehensive, multimodal strategy. By utilizing a variety of biomarkers and diagnostic tools (including PSA-based markers like PHI, 4Kscore, and SHLM3 test, along with non-PSA-based markers including PCA3, MiPS, SelectMDX, and EPI) more comprehensive assessment of PCa can be achieved. Efforts to optimize and standardize assay thresholds will enhance their effectiveness. Additionally, emerging biomarkers, including non-coding RNAs, proteomic molecules, and tumor-associated microbiota, present new opportunities to investigate PCa biology with advanced technologies.
Challenges persist, including the need to validate these biomarkers in large patient cohorts, demonstrate their added value compared to current standards, and ensure assay consistency. Collaborative research is essential to address these issues and advance clinical applications. Integrating insights from molecular and biological sciences with technological advancements, as well as exploring less invasive biomarker sources, will drive progress.
The incorporation of various biomarkers and tools used for diagnosis marks a significant advancement in PCa diagnosis. Although encouraging, more studies are needed to validate combined diagnostic approaches, making this a critical area for future research. Ongoing studies and research collaboration are vital for integrating emerging diagnostics into effective management strategies. Addressing existing challenges is crucial to fully realize the benefits of this approach, aiming to revolutionize PCa diagnosis and management.
Footnotes
Abbreviations
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: SOR, is supported by funding from the Office of the Assistant Secretary of Defense for Health Affairs through the Congressionally Directed Medical Research Programs (CDMRP), Prostate Cancer Research Program, Health Equity Research and Outcomes Improvement Consortium, under Award Number (W81XWH2210972). PA and MG are supported by funds provided by World Bank through The African Centers of Higher Education Excellence for Development Impact (ACE Impact) projects.
