Abstract
Glioblastoma is the most frequent form of malignant brain tumor. Cytoplasmic polyadenylation element binding protein 4 (CPEB4) is overexpressed and involved in the tumorigenesis and metastasis of glioblastoma. miR-130a-3p has been revealed to be aberrantly expressed in tumors and has aroused wide attention. In present study, we would like to investigate the effect and potential mechanism of miR-130a-3p on the proliferation and migration in glioblastoma. The relative expression levels of miR-130a-3p and CPEB4 in glioblastoma cell lines were detected by real-time quantitative polymerase chain reaction. Cell viability and migration were detected by methylthiazolyl tetrazolium assay and transwell assay, and cell cycle analysis was detected by flow cytometry. The expression of CPEB4 protein and epithelial-mesenchymal transition associated markers were detected by western blot. Bioinformatics and luciferase activity analysis were used to verify the targeting relationship between miR-130a-3p and CPEB4. We observed that the expression of CPEB4 was upregulated while that of miR-130a-3p was downregulated in glioblastoma cell lines. CPEB4 was validated as a target of miR-130a-3p by luciferase activity assay. Increased levels of miR-130a-3p inhibited the proliferation and migration of the glioblastoma cells and the overexpression of miR-130a-3p inhibited epithelial-mesenchymal transition. However, CPEB4 overexpression resisted the inhibitory effects of miR-130a-3p. Our study elucidates CPEB4 is upregulated because of the downregulated miR-130a-3p in glioblastoma, which enhances the glioblastoma growth and migration, suggesting a potential therapeutic target for the disease.
Introduction
Glioblastoma is the most frequent type of malignant brain tumor, characterized by infiltrative growth and angiogenesis. 1 Although major advancements have been developed in surgery, chemotherapy, radiotherapy, and immunotherapy of glioblastoma, patients’ prognosis is still not significantly improved.2,3 The median survival time for untreated individuals with glioblastoma is just about 4 months. After standard chemoradiotherapy combined with TMZ adjuvant chemotherapy, the median survival time is only 15 months.4,5 Thus, it is important to investigate the possible targets that have crucial functions in the pathogenesis of glioblastoma, expecting to create new treatment approaches and improve the outcome of subjects with glioblastoma.
A sequence-specific RNA-binding protein with considerable conservation known as cytoplasmic polyadenylation element-binding protein 4 (CPEB4) regulates translational repression or cytoplasmic polyadenylation machinery to target messenger RNAs (mRNAs). 6 Recent investigations have revealed that CPEB4 is involved in biological processes that are associated with tumorigenesis and metastasis.7,8 CPEB4 was aberrantly expressed in several cancers, like gliomas.9–12 CPEB4 overexpression enhances glioma cells migration and invasion, and CPEB4 upregulation is correlated with progressive World Health Organization (WHO) grade. 13 Moreover, increased expression of CPEB4 affects the poor outcome of individuals with gliomas, indicating that CPEB4 may function as a novel prognostic biomarker for subjects with gliomas. 14
A category of small noncoding RNAs known as microRNAs (miRNAs) have become significant modulators that inhibit gene expression by adhering to the 3'-untranslated region (UTR) region of the target mRNAs at the post-transcriptional regulatory level.15,16 A growing number of miRNAs have been reported to be related to cell apoptosis, growth, differentiation, migration, and associated with treatment response and patient outcome. Recently, miR-130a-3p has been revealed to be aberrantly expressed in multiple tumors and has aroused wide attention.17,18 Furthermore, miR-130a-3p has been recognized as a predictive marker for B-cell lymphoma, renal cell carcinoma, and gastric carcinoma.19–21 CPEB4 was anticipated to be an miR-130a-3p target through the starBase database. 22 Thus, we suggest that the downregulated miR-130a-3p expression results in CPEB4 upregulated expression in glioblastoma cells, and then promoting tumorigenesis. In this investigation, we discovered the relations between miR-130a-3p and CPEB4 in regulating glioblastoma growth and migration.
Materials and Methods
Cell Culture
Human astrocyte HA1800 (CL-60804059), and glioblastoma T98G (CL-0583) and U87MG (CL-0238) cell lines were obtained from the Procell Life Science & Technology Co., Ltd, and cultured in Dulbecco's modified Eagle's medium (DMEM, C11965500BT, Gibco) supplemented with 10% fetal bovine serum (FBS, 10091, Gibco). Cell cultures were maintained at 37°C in a humidified atmosphere with 5% CO2. We had authenticated the cell lines used in present study with short tandem repeat profiling method.
Plasmid Construction and Transfection
Genecreate Biological Engineering Co., Ltd was commissioned to produce the miR-130a-3p mimics (chemically synthesized mature miRNA to enhance the function of endogenous miRNA, CAGUGCAAUGUUAAAAGGGCAU), inhibitor (chemically synthesized mature miRNA modified by methoxyl to inhibit the activity of endogenous miRNA, mAmUmGmCmCmCmUmUmUmUmAmAmCmAmUmUmGmCmAmCmUmG), negative control (NC) mimics (UUGUACUACACAAAAGUACUG), and NC inhibitor (mCmAmGmUmAmCmUmUmUmUmGmUmGmUmAmGmUmAmCmAmA). In order to create the recombinant vectors pmirGLO-CPEB4-WT and pmirGLO-CPEB4-MUT, the CPEB4 fragment including the potential binding locations for miR-130a-3p was generated and introduced into the pmirGLO vector (E1330, Promega) downstream of the luciferase gene. In the pmirGLO-CPEB4-MUT construct, the CPEB4 fragment contained 7 mismatches. In order to make the CPEB4 expression plasmid, the CPEB4 complementary DNA (cDNA) was introduced into the pcDNA3.1 vector (Genecreate). Lipofectamine 2000 (11668027, Invitrogen) was employed to transfect all of the plasmids and oligonucleotides into cells in accordance with the manufacturer's recommendations.
Real-Time Quantitative Polymerase Chain Reaction
Utilizing real-time quantitative polymerase chain reaction (PCR), the miR-130a-3p and CPEB4 expression levels were evaluated. TRIzol reagent (15596026, Invitrogen) was employed to obtain total RNA from the cultivated cells. The ReverTra Ace qPCR RT was utilized to create the cDNA (FSQ-101, Toyobo). The SYBR Green Realtime PCR Master Mix (QPK-201, Toyobo) was utilized to conduct the PCR reactions on the LightCycler96 system (Roche). PCR primer sequences were recorded in Supplemental Table S1. The fold change in every gene expression was detected by employing the 2−ΔΔCt technique. 23
MTT Assay
In a 96-well plate, the cells were seeded at 1 × 103 cells per well and incubated for 0, 1, 2, and 3 days to ascertain the alterations in their growth rates. Each well received an addition of 5 mg/mL methylthiazolyl tetrazolium (MTT, M1020, Solarbio), which was then incubated for 4 h. Dimethyl sulfoxide (DMSO, D8371, Solarbio) was introduced to every well just after supernatants were removed, and a microplate scanner was employed to detect the absorbance at 490 nm (Huisong Technology). Three separate independent tests were conducted for each of the MTT assay.
Transwell Assay
Transwell assays were conducted utilizing inserts (6.5 mm diameter, 8 μm pore size; Corning) in 24-well culture plates. Specifically, 1 × 104 cells placed into upper chambers received DMEM without serum, whereas the bottom chambers received DMEM with 10% FBS. The nonmigrated cells were eliminated using cotton swabs after a 24h incubation period, and the migratory cells were then treated with 4% paraformaldehyde and stained with crystal violet. Then cell counts were observed in 5 random fields under a CX41 microscope (Olympus) and processed by Image J software. The transwell tests were conducted 3 times as separate experiments.
Cell Cycle Analysis
The cells were washed with phosphate buffered saline, and then fixed in 70% ethanol for a whole night at 4 °C. Resuspended fixed cells were treated for 30 min in the darkness with 5 μg/L propidium iodide (P8080, Solarbio). Then, a cytoflex flow cytometer was utilized to identify the cell cycle (Beckman).
Western Blotting
The western blotting test was performed, as explained earlier. 24 The membranes were treated with horseradish peroxidase-conjugated IgG antibody (BA1070, Boster) after being incubated with antibodies against CPEB4 (25342-1-AP, 1:500, Proteintech), E-cadherin (BM3903, 1:1000, Boster), N-cadherin (BM3921, 1:1000, Boster), Vimentin (BM4029, 1:500, Boster), or β-actin (81115-1-RR, 1:2000, Proteintech) at 4 °C overnight. Eventually, an improved chemiluminescence procedure was utilized to determine the protein bands (Advansta).
Luciferase Reporter Assay
The T98G cells were co-transfected with pmirGLO-CPEB4-WT/MUT and miR-130a-3p construct utilizing Lipofectamine 2000 for the luciferase reporter experiment (11668027, Invitrogen). Utilizing Dual-Luciferase Reporter Assay method, the luciferase activity was detected 48 h following transfection, and the ratio of firefly luciferase/renilla luciferase was calculated (RG027, Beyotime). Three separate experiments were conducted to duplicate the luciferase assay.
Bioinformatics Analysis
To identify the potential target mRNAs related to miR-130a-3p, the online database starBase 2.0 was used, and the intersecting genes were regarded as potential targets in glioblastoma. Then the intersecting targets were uploaded into Metascape platform for functional analysis. To examine whether the CPEB4 and miRNA-130a-3p expression levels varied with different grades of gliomas, the expression profiles and corresponding medical data for low-grade gliomas and glioblastoma were obtained from the Cancer Genome Atlas (TCGA) database. The expression differences of CPEB4 between gliomas and controls were also conducted by Gene Expression Profiling Interactive Analysis (GEPIA) dataset. Furthermore, we adopted Tumor Immune Single-cell Hub (TISCH) database to further explore the expression level of CPEB4 at the single-cell level. The mRNAs expression level of CPEB4 in a series of glioma cell lines was determined by Cancer Cell Line Encyclopedia (CCLE) database. Subsequently, utilizing R program version 4.0.3, statistical analyses were conducted, and data was displayed utilizing ggplot2 tool in R. The Wilcoxon test was employed to investigate statistical variations among 2 groups, and the Kruskal-Wallis analysis was employed to identify whether there were significant variations between 3 groups.
Statistical Analysis
Mean ± SD represents the data of triplicate trials. Student's t-test or one-way analysis of variance followed by a post hoc Tukey test was applied to estimate the difference among groups, and P < .05 was considered to be statistically significant.
Results
CPEB4 Upregulation and miR-130a-3p Downregulation in Glioblastoma Cells
Utilizing real-time quantitative PCR, we first explored the CPEB4 and miR-130a-3p expression levels in astrocyte HA1800 and glioblastoma (T98G and U87MG) cell lines. In comparison with the HA1800 cell line, the relative CPEB4 expression was significantly raised in glioblastoma cell lines (Figure 1A). Nevertheless, in contrast with the HA1800 cells, miR-130a-3p expression was significantly downregulated in the 2 glioblastoma cell lines (Figure 1B). Additionally, based on the RNA-sequencing data of gliomas from TCGA database, we discovered that the miRNA-130a-3p and CPEB4 expression levels varied with different histologic grades and associated with the prognosis of gliomas (Figure 1C and D, and Supplemental Figure S1).

The expression of CPEB4 and miR-130a-3p in glioblastoma. (A) Real-time PCR analysis of CPEB4 expression in T98G and U87MG cell lines. β-Actin was used as the reference gene. (B) Real-time PCR analysis of miR-130a-3p expression in T98G and U87MG cell lines. U6 was used as the reference gene. (C) and (D) The expression distribution of CPEB4 and miR-130a-3p in gliomas tissues with different histological grades based on the transcriptome data from TCGA database. **P < .01, ***P < .001, and ****P < .0001. Abbreviations: CPEB4, cytoplasmic polyadenylation element binding protein 4; PCR, polymerase chain reaction; TCGA, The Cancer Genome Atlas.
miR-130a-3p Modulates CPEB4 Expression in Glioblastoma Cells
The glioblastoma cell lines were transfected with miR-130a-3p mimic or inhibitor and NC to investigate if it can inhibit CPEB4 expression. According to Figure 2A, in the glioblastoma cell lines transfected with the miR-130a-3p mimic or inhibitor, the CPEB4 protein expression levels were significantly downregulated and elevated, respectively, with mRNA expression being consistent with the protein expression (Supplemental Figure S2).

CPEB4 is a target of miR-130a-3p in glioblastoma. (A) Effect of miR-130a-3p expression on protein levels of CPEB4 and EMT-related biomarkers. Glioblastoma cells were transfected with miR-130a-3p mimic, inhibitor, or negative control, then western blotting analyses were conducted and the expression of β-actin was used as internal reference. (B) Predicted miR-130a-3p target sequence in the 3’-UTR of CPEB4 (CPEB4-WT) and CPEB4-MUT containing 7 altered nucleotides. (C) Luciferase reporter assay was applied to detect the binding between miR-130a-3p and 3’-UTR of CPEB4 in T98G cells. Co-transfection of mimic and CPEB4-WT decreased the firefly/renilla luciferase activity, while co-transfection of mimic and CPEB4-MUT did not change the firefly/renilla luciferase activity. Co-transfection of NC mimic with CPEB4-WT or CPEB4-MUT did not change the firefly/renilla luciferase activity. ***P < .001. Abbreviations: CPEB4, cytoplasmic polyadenylation element binding protein 4; EMT, epithelial-mesenchymal transition; NC, negative controls; UTR, untranslated region.
The anticipated miR-130a-3p target sequence was then inserted into the pmirGLO to construct the luciferase reporter vectors pmirGLO-CPEB4-WT and pmirGLO-CPEB4-MUT (Figure 2B). The miR-130a-3p mimic reduced pmirGLO-CPEB4-WT firefly/renilla luciferase activities but not pmirGLO-CPEB4-MUT in T98G cells, according to the luciferase reporter experiment (Figure 2C). Nevertheless, in the miR-130a-3p NC mimic groups, there was no clear impact on the activity of firefly/renilla luciferase. Our findings showed that miR-130a-3p targeted CPEB4 in glioblastoma cells.
miR-130a-3p Affects Glioblastoma Cells Proliferation
Utilizing MTT test, we first studied the impact of miR-130a-3p on cell growth. At 48 and 72 h after plating, MTT test revealed that miR-130a-3p mimic-transfected and inhibitor-transfected T98G cells had significantly reduced and increased growth rates compared to control cells, respectively. At 3 days following plating, the miR-130a-3p mimic-transfected U87MG cells grew more slowly than the control cells, but the miR-130a-3p inhibitor-transfected U87MG cells grew more rapidly at 48 and 72 h following plating (Figure 3A). Also, we observed that glioblastoma cells transfected with miR-130a-3p mimics exhibited a greater rate of G1 phase and a decreased frequency of G2 phase. On the other hand, glioblastoma cells transfected with the miR-130a-3p inhibitor exhibited a lower G1 phase rate and an elevated G2 phase frequency (Figure 3B and Supplemental Figure S3). Our results indicated that exogenous miR-130a-3p suppressed glioblastoma development.

miR-130a-3p affects glioblastoma cells proliferation. (A) MTT assay was conducted on T98G and U87MG cells transfected with miR-130a-3p mimic or inhibitor. **P < .01 mimic group compared with mimic NC group; ##P < .01 inhibitor group compared with inhibitor NC group. (B) Flow cytometry displaying cell cycle distribution of T98G and U87MG cells transfected with miR-130a-3p mimic or inhibitor. T98G and U87MG cells transfected with miR-130a-3p mimics exhibited a greater rate of G1 phase and a decreased frequency of G2 phase. Whereas, the T98G and U87MG cells transfected with the miR-130a-3p inhibitor exhibited a lower G1 phase rate and an elevated G2 phase frequency. However, statistical analysis did not find a significant difference. Abbreviation: MTT, methylthiazolyl tetrazolium; NC, negative controls.
miR-130a-3p Affects Migration and Epithelial-Mesenchymal Transition of Glioblastoma Cells
Next, we examined how miR-130a-3p affected glioblastoma cell migration. The transwell assay revealed that miR-130a-3p mimic-transfected cells migrated significantly more slowly than control cells, whereas cells that transfected with miR-130a-3p inhibitor migrated more quickly than control cells (Figure 4). Epithelial-mesenchymal transition (EMT) is believed to have been a vital factor in glioblastoma invasion, hence western blotting was employed to assess the EMT-associated biomarkers Vimentin, N-cadherin, and E-cadherin protein levels. These outcomes exhibited that in glioblastoma cells transfected with miR-130a-3p mimic, mesenchymal biomarkers N-cadherin and Vimentin protein levels were downregulated, but epithelial biomarker E-cadherin was raised with miR-130a-3p overexpression (Figure 2A).

miR-130a-3p affects glioblastoma cells migration. The T98G and U87MG cells were transfected with miR-130a-3p mimic or inhibitor, then transwell assays were conducted in culture plates. The cells were stained with crystal violet and the number of migrating cells was counted in 5 random fields under a microscope (magnification, × 200). **P < .01, ***P < .001, ****P < .0001. Abbreviation: NC, negative controls.
CPEB4 Overexpression Resists Antiproliferation and Antimigration Impacts of miR-130a-3p in Glioblastoma Cells
MiR-130a-3p mimic and CPEB4-pcDNA3.1 were co-transfected into the glioblastoma cells to examine the connections between miR-130a-3p and CPEB4 in the growth and migration of glioblastoma cells. According to the MTT experiment, miR-130a-3p mimic + CPEB4-pcDNA3.1-transfected cells had higher cell viability than miR-130a-3p mimic + pcDNA3.1 empty vector-transfected cells (Figure 5), demonstrating that CPEB4 overexpression counteracted miR-130a-3p's antiproliferation activities. Moreover, CPEB4 overexpression prevented miR-130a-3p mimic from inhibiting cell migration (Figure 6).

CPEB4 overexpression resists the antiproliferation effect of miR-130a-3p. The miR-130a-3p mimic-transfected glioblastoma cells were introduced into CPEB4-pcDNA3.1 or empty vector pcDNA3.1, then MTT assay was performed.; *P < .05, **P < .01 miR-130a-3p mimic + CPEB4-pcDNA3.1 group compared with miR-130a-3p mimic + pcDNA3.1 group; #P < .05, ##P < .01 miR-130a-3p mimic group compared with untreated group. Abbreviation: CPEB4, cytoplasmic polyadenylation element binding protein 4; MTT, methylthiazolyl tetrazolium.
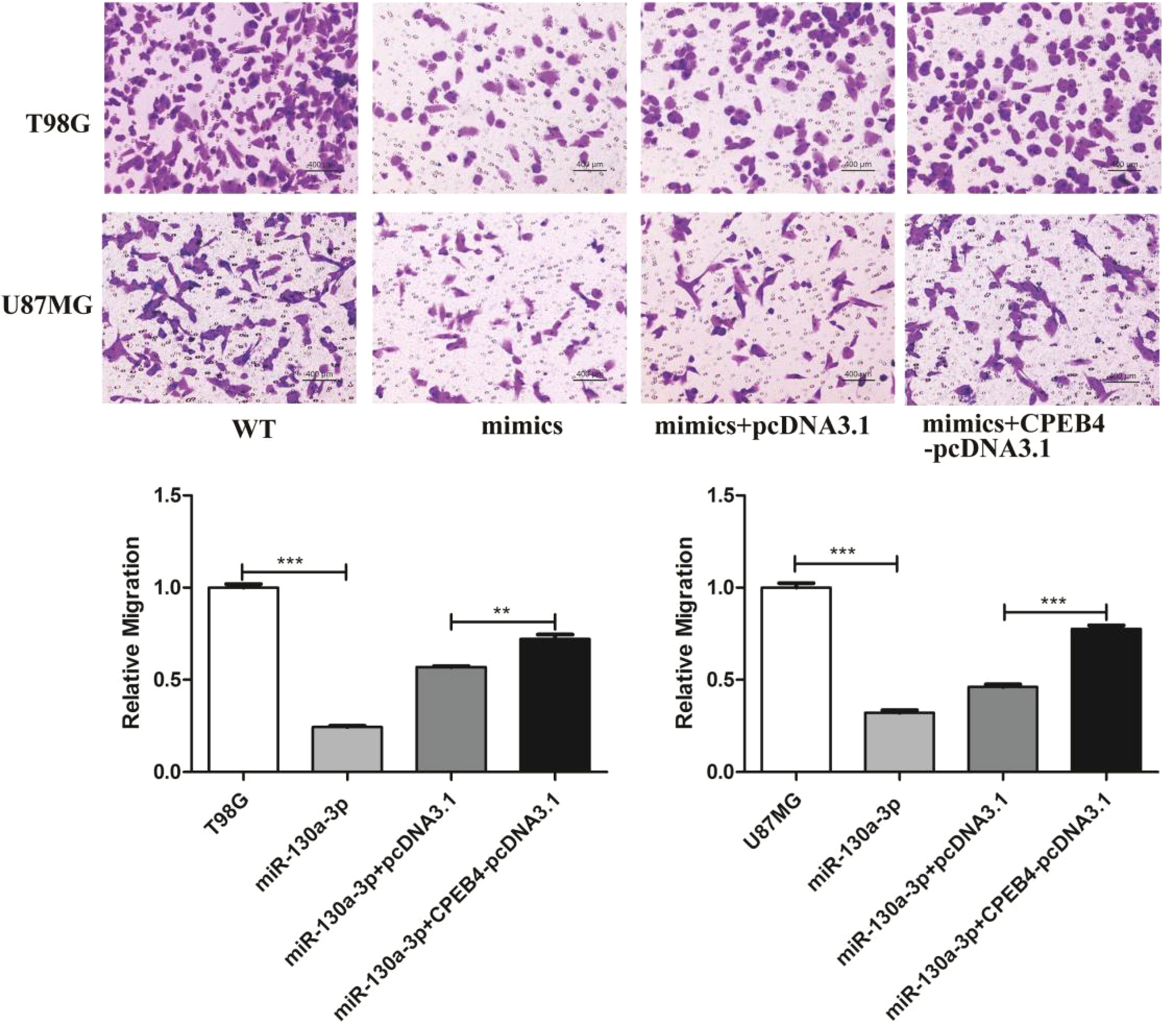
CPEB4 overexpression resists the antimigration effect of miR-130a-3p. The miR-130a-3p mimic-transfected glioblastoma cells were introduced into CPEB4-pcDNA3.1 or empty vector pcDNA3.1, then transwell assays were conducted in culture plates. The cells were stained with crystal violet and the number of migrating cells was counted in 5 random fields under a microscope (magnification, × 200). **P < .01, ***P < .001. Abbreviation: CPEB4, cytoplasmic polyadenylation element binding protein 4.
Predictive and Enrichment Analysis of miR-130a-3p Targets
To identify the miR-130a-3p potential target genes, the online database starBase 2.0 was used, and 308 targets were obtained from the intersection of 4 prediction programs, including TargetScan, PicTar, microT, and PITA (Figure 7). To understand the function of the 308 targets, Gene Ontology and pathway analysis was performed by the Metascape. The website showed that the significantly enriched terms included Wnt signaling pathway, neuron projection morphogenesis, blood vessel morphogenesis, membrane trafficking, small GTPase-mediated signal transduction, response to nutrient levels, regulation of PTEN mRNA translation, histone modification, regulation of mitotic cell cycle, and protein deubiquitination (Figure 8A). To further capture the relationships among enriched terms, a subset of enriched terms had been selected and displayed as a network plot, where terms with a similarity more than 0.3 were connected with edges (Figure 8B and C).

The prediction of target genes of miR-130a-3p. The target genes of miR-130a-3p were predicted by using starBase database, and 308 targets were obtained from the intersection of 4 prediction programs, including TargetScan, PicTar, microT, and PITA.

Pathway and process enrichment analysis of target genes. The enrichment analysis of target genes was carried out with the Metascape. Terms with P values <.01, and enrichment factors more than 1.5 were grouped into clusters based on their membership similarities. (A) Bar graph of enriched terms among target genes. (B) and (C) Network of enriched terms colored by cluster ID or P value.
The protein-protein interaction analysis was also carried out and the MCODE algorithm was applied to identify densely connected subnetworks (Figure 9). Then 4 subnetworks for miR-130a-3p target genes were plotted, including cell migration, heart development, negative regulators of DDX58/IFIH1 signaling, transcriptional regulation by VENTX, ubiquitin-dependent protein catabolic process, and endocytosis.

Protein-protein interaction and MCODE analysis of the target genes. The protein-protein interaction analysis was conducted with the Metascape. The network contained the subset of proteins that formed physical interactions with at least one other member in the target genes. Then the MCODE algorithm was applied to identify densely connected subnetworks and 4 subclusters for miR-130a-3p target genes were plotted.
The Expression Analysis of CPEB4 in Gliomas
RNA-seq data in the CCLE database showed that CPEB4 was highly expressed in various cell lines of gliomas, suggesting that CPEB4 involved in glioma progression (Figure 10). The distribution of CPEB4 expression in various immune cells from TISCH datasets was shown in Figure 11. CPEB4 was mainly expressed in mono/macro cells, malignant cells, endothelial cells, and oligodendrocytes, indicating that CPEB4 functioned in immune cells and stromal cells in addition to in malignant cells.
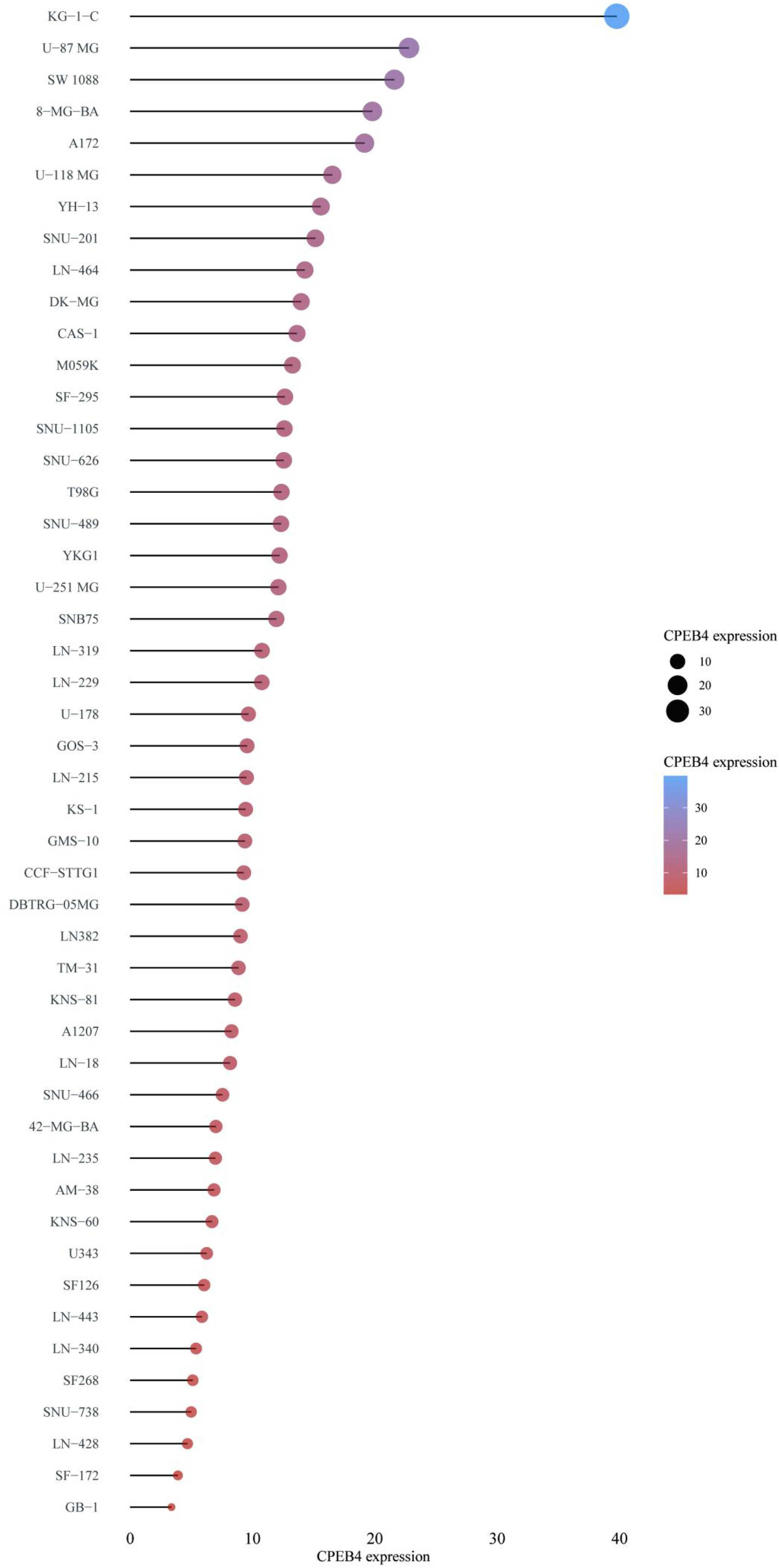
CPEB4 was upregulated in gliomas cell lines based on CCLE database. The cell lines mRNA expression matrix of gliomas was obtained from the CCLE dataset. The abscissa represented the expression of CPEB4 and the ordinate represented different cell lines, different colors and the size of dots represented expression levels. Abbreviations: CCLE, Cancer Cell Line Encyclopedia; CPEB4, cytoplasmic polyadenylation element binding protein 4; mRNA, messenger RNA.
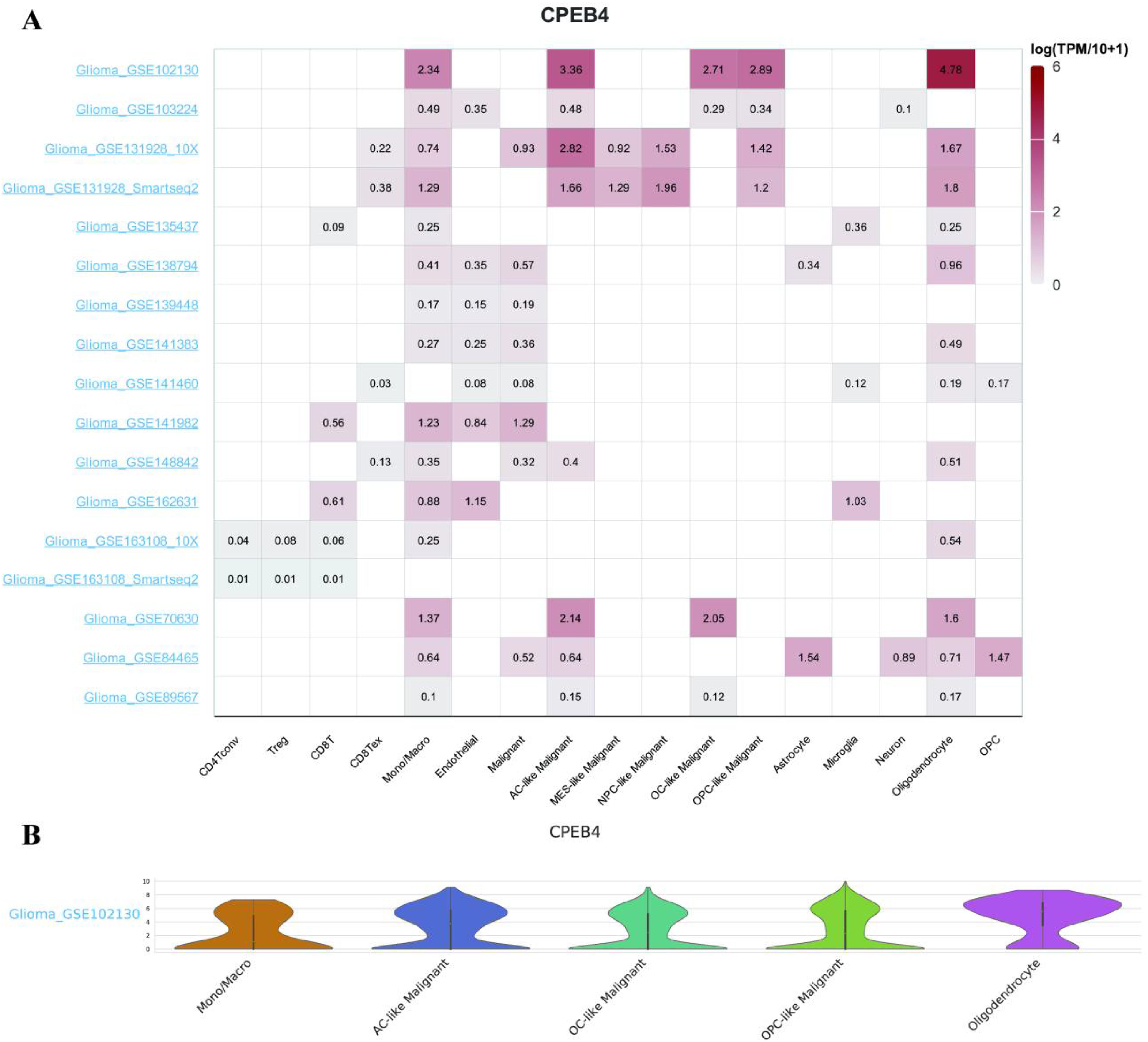
The single-cell expression analysis of CPEB4 with TISCH database. The correlation between CPEB4 expression and tumor microenvironment was explored by using the TISCH database. (A) The heatmap showing the value of CPEB4 expression in different cells from different datasets. (B) Violin plot displaying the distribution of CPEB4 expression in different cells in GSE102130 dataset. Abbreviations: CPEB4, cytoplasmic polyadenylation element binding protein 4; TISCH, Tumor Immune Single-cell Hub.
Discussion
A miR-130a family member, miR-130a-3p, has gained prominence recently. Intriguingly, some investigations have demonstrated that the miRNAs of the miR-130a family are upregulated and enhance cell growth in several malignancies, while other investigations have demonstrated that miR-130a family members are downregulated and suppress cell growth, revealing the function complexity of miR-130a throughout tumorigenesis.25–27 In this investigation, we discovered that in the glioblastoma cell lines T98G and U87MG, miR-130a-3p was substantially decreased. Overexpressed miR-130a-3p inhibited glioblastoma cell growth, whereas miR-130a-3p reduction promoted glioblastoma cell proliferation. Furthermore, we demonstrated that glioblastoma cells migrated more readily when miR-130a-3p inhibitor was employed compared to miR-130a-3p mimic.
MiR-130a-5p expression was substantially downregulated in gliomas tissues and cell lines, according to Tang et al 28 miR-130a-5p overexpression inhibited the glioma cells growth and invasion, and luciferase tests revealed the oncogene HMGB2 as a downstream target of miR-130a-5p. miR-130a-3p has been determined to be substantially downregulated in the glioblastoma cell lines A172, U251, and U87. 29 miR-130a-3p overexpression inhibited the glioblastoma cells growth in vitro. Also, in a nude mouse xenograft tumor model, the miR-130a-3p mimic and inhibitor substantially inhibited and enhanced tumor growth, respectively. Consistent with previous research, we found miR-130a-3p was substantially decreased in T98G and U87MG cell lines, and varied with different histologic grades of gliomas. miR-130a-3p might even have a significant function in modulating TMZ resistance in glioblastoma, according to Wang et al, who also discovered that miR-130a-3p mimic risen the sensitivity of the glioblastoma cells to TMZ while the sensitivity was reduced in the cells transfected with miR-130a-3p inhibitor. miR-130a may be a predictive marker for TMZ response among individuals with glioblastoma because the TMZ cytotoxicity assay revealed that miR-130a overexpression could sensitize response to TMZ in glioma cells and decreased miR-130a could create a more significant response to oxidative stress and mediate resistance to TMZ. 30
The modulating role of miR-130a-3p in EMT has been reported in many malignancies like osteosarcoma, esophageal squamous cell carcinoma, gastric carcinoma, and nasopharyngeal carcinoma.20,21,31 Considered that EMT has a vital function in glioblastoma invasion and metastasis, EMT-associated biomarkers N-cadherin, Vimentin, and E-cadherin protein levels were identified using western blotting. Outcomes confirmed that miR-130a-3p overexpression suppressed glioblastoma EMT through downregulating biomarkers associated with mesenchymal while upregulating biomarkers linked to epithelia. In addition, function enrichment analysis of the target genes of miR-130a-3p showed that the targets mainly involved in Wnt signaling pathway, neuron projection morphogenesis, blood vessel morphogenesis, membrane trafficking, signal transduction, histone modification, and cell cycle regulation. The protein-protein interaction and subnetworks for target genes indicated that miR-130a-3p may play an important regulation role in cell migration.
The raised CPEB4 expression and its function in cell invasion, growth, and chemoresistance have been reported in gliomas. In vitro studies showed that downregulation of CPEB4 significantly reduced the growth rate of T98G and U251 cells. Furthermore, in astrocyte tumor cells, CPEB4 knockdown inhibited their clonogenicity, proliferative ability, and invasiveness.14,32,33 Consistent with previous studies, we also found CPEB4 was substantially increased in T98G and U87MG cell lines, and varied with different histologic grades of gliomas. What's more, single cell expression profile analysis showed CPEB4 was mainly expressed in mono/macro cells, endothelial cells, malignant cells, and oligodendrocytes, indicating that CPEB4 functioned in immune cells and stromal cells in the pathological process of gliomas. Previous studies have found that CPEB4 could be regulated by miRNAs, such as miR-98-5p and miR-145-5p. CPEB4 was anticipated as a miR-130a-3p target using bioinformatics, and these findings were promoted by CPEB4 and miR-130a-3p reciprocal repression in glioblastoma cells. The miR-130a-3p inhibitor promoted CPEB4 expression, and the miR-130a-3p mimic suppressed it. Furthermore, the luciferase activity experiment demonstrated that CPEB4 as a direct target of miR-130a-3p. By examining the cell migration and viability in miR-130a-3p mimic and CPEB4-pcDNA3.1 co-transfected glioblastoma cells, which illustrated the CPEB4 recovery impact on miR-130a-3p induced cell behaviors, the functional interaction among miR-130a-3p and CPEB4 in glioblastoma was further revealed. We suggest that increased CPEB4, caused by a reduction in miR-130a-3p, may enhance cell proliferation and migration, which then support glioblastoma growth and progression. The high expression of CPEB4 promotes the malignant biological behaviors of glioma cells, and the inhibition effects on glioma cells mediated by miR-130a-3p can be overturned by upregulation of CPEB4. Upregulation of miR-130a-3p or downregulation of CPEB4 may help improve the prognosis of glioma. We are now gathering specimens from xenograft tumor models and individuals with gliomas to further validate the relations among miR-130a-3p and CPEB4, and to shed light on the precise molecular pathway of miR-130a-3p in the pathogenesis of gliomas.
Conclusion
In brief, target gene CPEB4 was upregulated because of downregulated miR-130a-3p in glioblastoma, which enhanced the glioblastoma cells growth and migration. A treatment target for the therapy of glioblastoma may be discovered by clarifying the connection between miR-130a-3p and CPEB4 in glioblastoma.
Supplemental Material
sj-tif-1-tct-10.1177_15330338231218218 - Supplemental material for miRNA-130a-3p/CPEB4 Axis Modulates Glioblastoma Growth and Progression
Supplemental material, sj-tif-1-tct-10.1177_15330338231218218 for miRNA-130a-3p/CPEB4 Axis Modulates Glioblastoma Growth and Progression by Hongchao Liu, Zhihao Wei, Kangke Shi, Yu Zhang and Jiaqiong Li in Technology in Cancer Research & Treatment
Supplemental Material
sj-tif-2-tct-10.1177_15330338231218218 - Supplemental material for miRNA-130a-3p/CPEB4 Axis Modulates Glioblastoma Growth and Progression
Supplemental material, sj-tif-2-tct-10.1177_15330338231218218 for miRNA-130a-3p/CPEB4 Axis Modulates Glioblastoma Growth and Progression by Hongchao Liu, Zhihao Wei, Kangke Shi, Yu Zhang and Jiaqiong Li in Technology in Cancer Research & Treatment
Supplemental Material
sj-tif-3-tct-10.1177_15330338231218218 - Supplemental material for miRNA-130a-3p/CPEB4 Axis Modulates Glioblastoma Growth and Progression
Supplemental material, sj-tif-3-tct-10.1177_15330338231218218 for miRNA-130a-3p/CPEB4 Axis Modulates Glioblastoma Growth and Progression by Hongchao Liu, Zhihao Wei, Kangke Shi, Yu Zhang and Jiaqiong Li in Technology in Cancer Research & Treatment
Supplemental Material
sj-docx-4-tct-10.1177_15330338231218218 - Supplemental material for miRNA-130a-3p/CPEB4 Axis Modulates Glioblastoma Growth and Progression
Supplemental material, sj-docx-4-tct-10.1177_15330338231218218 for miRNA-130a-3p/CPEB4 Axis Modulates Glioblastoma Growth and Progression by Hongchao Liu, Zhihao Wei, Kangke Shi, Yu Zhang and Jiaqiong Li in Technology in Cancer Research & Treatment
Supplemental Material
sj-tif-5-tct-10.1177_15330338231218218 - Supplemental material for miRNA-130a-3p/CPEB4 Axis Modulates Glioblastoma Growth and Progression
Supplemental material, sj-tif-5-tct-10.1177_15330338231218218 for miRNA-130a-3p/CPEB4 Axis Modulates Glioblastoma Growth and Progression by Hongchao Liu, Zhihao Wei, Kangke Shi, Yu Zhang and Jiaqiong Li in Technology in Cancer Research & Treatment
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Not applicable, because this article does not contain any studies with human or animal subjects.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the Medical Science and Technology Program of Henan Province, grant number LHGJ20191234, the Science and Technology Development Project of Luoyang City, grant number 2101078A, the Medical Key Cultivation Discipline Program of Luoyang City, grant number STE-2022-5.
Informed Consent
Not applicable, because this article does not contain any studies with human or animal subjects.
Trial Registration
Not applicable, because this article does not contain any clinical trials.
Supplemental Material
Supplemental material for this article is available online.
Abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
