Abstract
Introduction
Leukemia is a malignant clonal disease that affects hematopoietic stem cells. It can be classified into acute or chronic leukemia based on the length of its natural course, and further categorized as myeloid or lymphocytic leukemia depending on the specialized cell type.1,2 In acute myeloid leukemia (AML), there is aberrant proliferation of immature myeloid cells with differentiation disorders, usually accompanied by various mutations and chromosomal translocations. 3 Acute lymphoblastic leukemia (ALL) is characterized by excessive production of immature lymphoblasts. 4 Additionally, plasma cell leukemia (PCL) is a rare form of acute leukemia that has poor treatment efficacy due to drug resistance in plasma cells during chemotherapy, leading to disease relapse. 5
MicroRNAs (miRNAs) are one type of non-coding RNA that consists of a single strand and range in length from 18-25 nucleotides. They function by negatively regulating the expression of target genes at the post-transcriptional level, and have been shown to play a role in various biological processes including tumor differentiation, proliferation, and apoptosis. 6 Abnormal miRNA expression has been observed in several types of cancer including liver cancer, lung cancer, breast cancer, colon cancer, pancreatic cancer and leukemia. Notably, studies have demonstrated that miR-22 exhibits unique anti-cancer effects in certain malignant tumors.7,8 miR-22 is a broadly expressed miRNA with a fragment length of 22 bp. It is highly conserved in many vertebrates, including mice, rats and chimpanzees, implying its important biological functions. 9 Although the regulatory mechanism of miR-22 in leukemia progression has not been fully elucidated yet, abnormal expression of miR-22 was demonstrated to be closely associated with leukemia development based on patient specimen investigation and clinical outcome analyses. In this study, we performed a comprehensive meta-analysis to evaluate the correlation between miR-22 expression and survival outcomes in leukemia patients. Our findings suggest that miR-22 has potential as a prognostic biomarker. Additionally, we utilized transcriptomics and qRT-PCR techniques to compare the levels of miR-22 expression in both non-leukemia and acute lymphoblastic leukemia (ALL) patients.
Materials and Methods
This meta-analysis was performed in accordance with the Preferred Reporting Items for Systematic Review and Meta-analysis (PRISMA) statement. The study has been registered at Research Registry and the unique identifying number is reviewregistry1637.
Search Strategy
A literature search was carried out in PubMed, EMBASE and Cochrane databases, with the publication date from January 2010 to November 2022. The following strategy was utilized to retrieve articles: (“miRNA22” or “mir-22” or “microRNA-22” or “mir22” or “microRNA22” or “miRNA-22”) and (“leukemia” or “leukeamia” or “leukaemia” or “ leucocythaemia”). Additionally, we conducted a comprehensive search for other relevant literature by reviewing the references cited in the article.
Inclusion and Exclusion Criteria
Inclusion criteria: studies (1) evaluated the clinical outcomes of leukemia; (2) detected miR-22 levels in two independent groups; (3) investigated and analyzed the predictive or diagnostic value of miR-22 in leukemia. Exclusion criteria: studies (1) were identified with duplicated data or results, preferring the latest and most authoritative research; (2) were not related to miR-22; (3) were meta-analyses, reviews, conference abstracts and articles without full text or non-research articles. Additionally, studies involving non-human subjects, cell studies, animal studies or those lacking clinical data will also be excluded.
Quality Assessment and Data Extraction
Two investigator independently assessed the quality of literature that met the inclusion criteria using the NOS scale (Newcastle Ottawa Scale) evaluation standards, which consists of three major items and eight sub-items. When there are discrepancies in scoring results for a particular study, re-evaluation will be conducted based on discussion. The extracted data will include information such as first author, publication year, journal name, country of origin, disease type studied, total number of cases analyzed, number of cases with high/low expression levels of miR-22, sample type, detection method, cut-off value, survival analysis and outcome.
Patients and Samples
Bone marrow specimens were collected from 10 non-leukemia children and 10 ALL pediatric patients, and all participating patients provided informed consent forms. The pediatric patients in this study were diagnosed based on the revised French-American-British classification, and leukemia cells were analyzed using cell morphology, immunophenotyping and cytogenetics. Mononuclear cells were isolated from bone marrow specimen cells using Ficoll-Paque (GE Healthcare, USA), and total RNA was extracted. All participant information including the general clinical characteristic is summarized in Supplemental Table S1. The study was approved by the institutional review Board of the Affiliated Wuxi Children's Hospital of Jiangnan University (WXCH2020-08-004-2).
Quantitative Real Time PCR (qRT-PCR)
The mRNA was reverse transcribed into cDNA using the PrimeScript RT reagent Kit (Takara, Japan), and qRT-PCR was performed on an ABI 7500 instrument (Applied Biosystems, USA). The reaction mixture contained 10 μL of SYBR® Premix Ex Taq II (Tli RNaseH Plus) (2×), 0.8 μL of forward primer, 0.8 μL of reverse primer, 2 μL of template, and 6.4 μL of ddH2O. The miR-22 primers used were: forward −5′ CTGGAGAAGCTGCCAGTTGAAG3′ and reverse −5′ GTGCAGGGTCCGAGGT3′; U6 primers were: forward −5′ CTCGCTTCGGCAGCACATATACT3′and reverse-5′ ACGCTTCACGAATTTGCGTGTC3′. The reaction conditions included initial denaturation at 95 °C for 30 s followed by amplification with denaturation at 95 °C for 5 s and annealing/extension at 60 °C for 34 s for a total of 40 cycles. We calculated the relative expression level of miR-22 using the 2−ΔΔCT method.
miRNA Sequencing
Total RNA was extracted from the sample using Trizol. Fragments of 18-30 nt were selected by gel electrophoresis and ligated with 3′ and 5′ adapters. Reverse transcription and PCR amplification were performed on the small RNA that had been ligated on both sides of the adapter. The library construction was completed by recovering approximately 140 bp bands, which were purified through gel electrophoresis. Quality control of the constructed library was carried out using Agilent 2100 and qRT-PCR before sequencing. The sRNA tag sequences obtained from each sample's original sequencing data were processed to identify miRNAs and their composition in the sample, resulting in a miRNA expression profile for differential analysis of miRNA genes.
Statistical Analysis
STATA 12.0 (Stata Corporation, College Station, Texas, USA) was used by two researchers to perform statistical analysis. Cochran's Q test and Higgins I-square were used to test for heterogeneity in the included data. If there was no heterogeneity (I2 < 50% or P > .1), a fixed-effect model was used; otherwise, a random-effects model was used. The outcome measure of this study is count data, and the time-to-event outcome measure of patients’ overall survival is analyzed using relative risk (RR). All effect sizes are presented with 95% confidence intervals (CI). If RR is not directly reported in the literature, Engauge Digitizer version 9.7 is used to extract and calculate data from Kaplan-Meier survival curves. Begg's and Egger's tests were used to quantitatively evaluate publication bias in the data. Independent sample t-tests were used to compare expression differences between patient groups and control groups. For all statistics, P < .05 was considered statistically significant difference.
Results
Search Strategy
The flowchart in Figure 1 illustrated the study selection process. Initially, a search was conducted across multiple databases using a predefined search strategy, which enrolled 104 articles. After removing duplicates (n = 21) and screening titles and abstracts (n = 63), full texts of the remaining 20 articles were assessed against inclusion criteria. Finally, four articles met all inclusion criteria and were included for analysis.
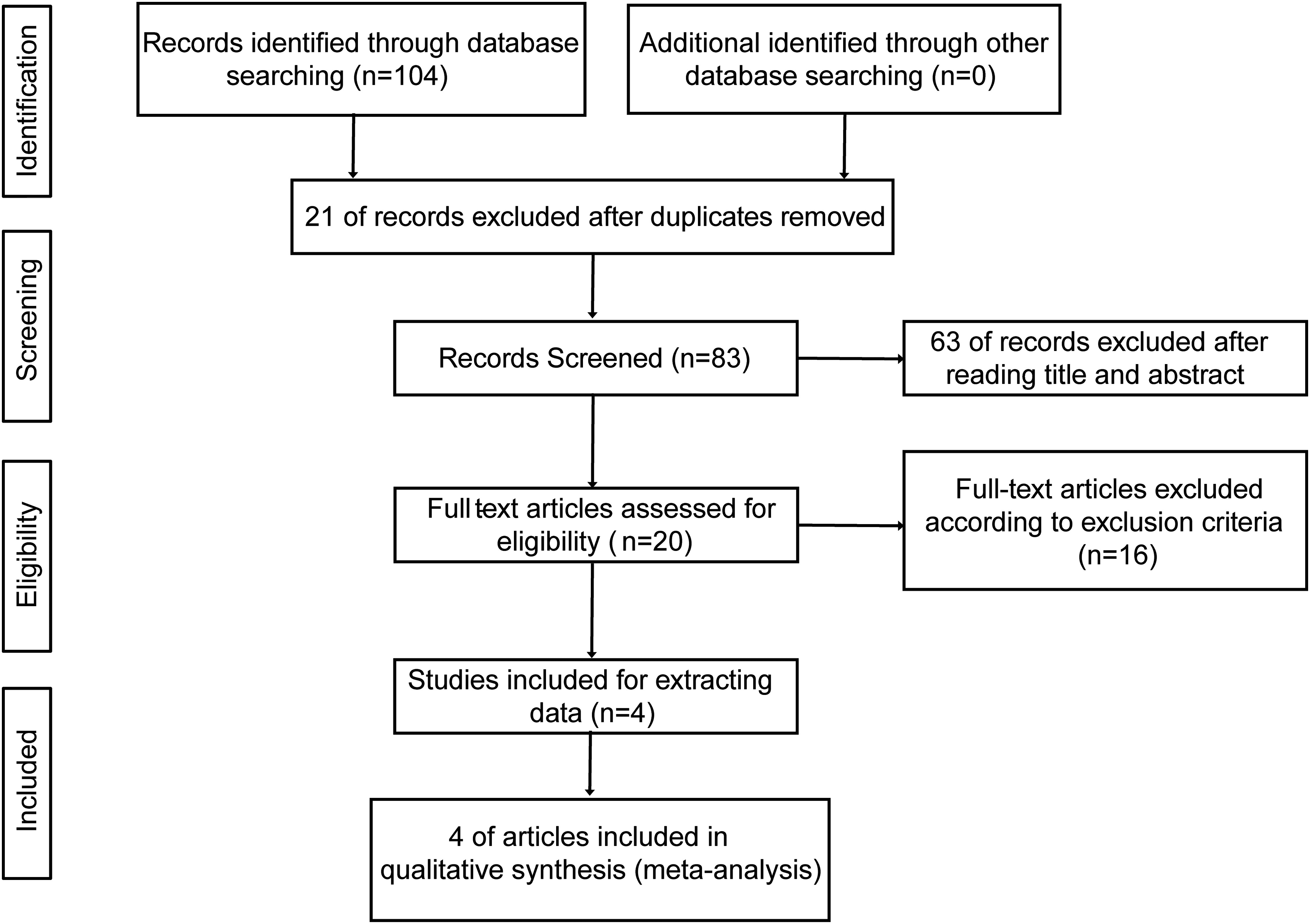
Preferred reporting items for systematic review and Meta-analysis (PRISMA) flow diagram of the meta-analysis.
Basic Characteristics Included in the Study
Four studies, comprising a total of 215 leukemia patients (122 with high miR-22 expression and 93 with low expression), were analyzed, and main characteristic was shown in Table 1. The studies were conducted in Italy, the United States and China, and included pPCL and AML disease types. miR-22 levels were measured using qRT-PCR and High-density SNP-array techniques. Kaplan-Meier survival analysis curves were used to determine OS/PFS/RFS outcomes. The quality of the included literature was evaluated using the NOS scoring system, with detailed score shown in Table 2.
Main Characteristics of Studies Included.

Abbreviations: AML, acute myeloid leukemia; pPCL, primary plasma cell leukemia; BM, bone marrow; K-M, Kaplan–Meier; OS, overall survival; PFS, Progression-free survival; RFS, relapse-free survival.
Quality Assessment of the Articles Based on NOS.

Correlation between Outcome and miR-22 Expression
Table 3 presented the RR and 95% CI values extracted from four studies. The combined RR for OS) was 1.558 (95% CI: 1.197-2.028, P < .01) (Figure 2A). There was no significant heterogeneity among the selected data when evaluating the relationship between total OS and miR-22 expression levels (I2 = 0.0%, P = .518), followed by three subgroup analyses. The disease type subgroup analysis revealed that pPCL had a combined RR of 2.400 (95% CI: 0.713-8.078, P = .157), while AML had a combined RR of 1.495 (95% CI: 1.141-1.958, P < 0 .01) (Figure 2B). The expression levels of OS and miR-22 were significantly correlated in both the United States and China subgroups, with combined RRs of 2.353 (95% CI: 1.106-5.005, P = .026) and 1.398 (95% CI: 1.047-1.867, P = .023), respectively (Figure 2C). The analysis of different survival outcomes showed that except for OS, high miR-22 expression level was significantly associated with better RFS, with a combined RR of l.517 (95% CI: l.114-2.065, P < .01) (Figure 2D). Overall, high miR-22 expression is positively correlated with leukemia patient survival.

Forest plots of studies assessing the combined HR in relation to miR-22 levels. The figure includes four panels, each presenting a different subgroup analysis using a fixed-effect model: Panel A shows the relationship between miR-22 expression and OS; Panel B presents results based on leukemia type; Panel C displays findings based on origin of publication; and Panel D depicts outcomes based on source of HR.
HRs and 95% CIs of Survival Outcome Extracted from Four Studies.

*P < .05, statistical differences.
Heterogeneity Analysis
The forest plot in the previous section indicated no heterogeneity in the data. Subgroup analysis also revealed no heterogeneity for disease type subgroups (I2 = 0.0%, P = .429), different country subgroups (I2 = 0.0%, P = .752), and different survival outcome analyses (I2 = 0.0%, P = .522). To assess heterogeneity in the data for total OS and miR-22, Galbraith plotting was used, which showed that all study points were distributed between two lines with no heterogeneity present (Figure 3).

Evaluation of data heterogeneity using Galbraith map.
Sensitivity Analysis and Publication Bias
To evaluate the impact of individual studies on the overall OS, a sensitivity analysis was conducted using combined RR to include all studies. The results showed that after removing individual studies, the point estimates remained within the 95% CI of the total effect size, indicating no significant impact on the overall RR (Figure 4). To assess publication bias in the included studies’ data for total OS, Begg's and Egger's tests were used with P-values of .308 and .116, respectively, both greater than .05. These results suggest that there is no statistical significance or bias in data. Additionally, all data points were evenly distributed within confidence intervals as shown by Begg's funnel plot (Figure 5). However, due to limited PFS and RFS data available for analysis, an assessment of publication bias could not be performed for these outcomes.

Sensitivity analysis of included studies in the meta-analysis for OS.
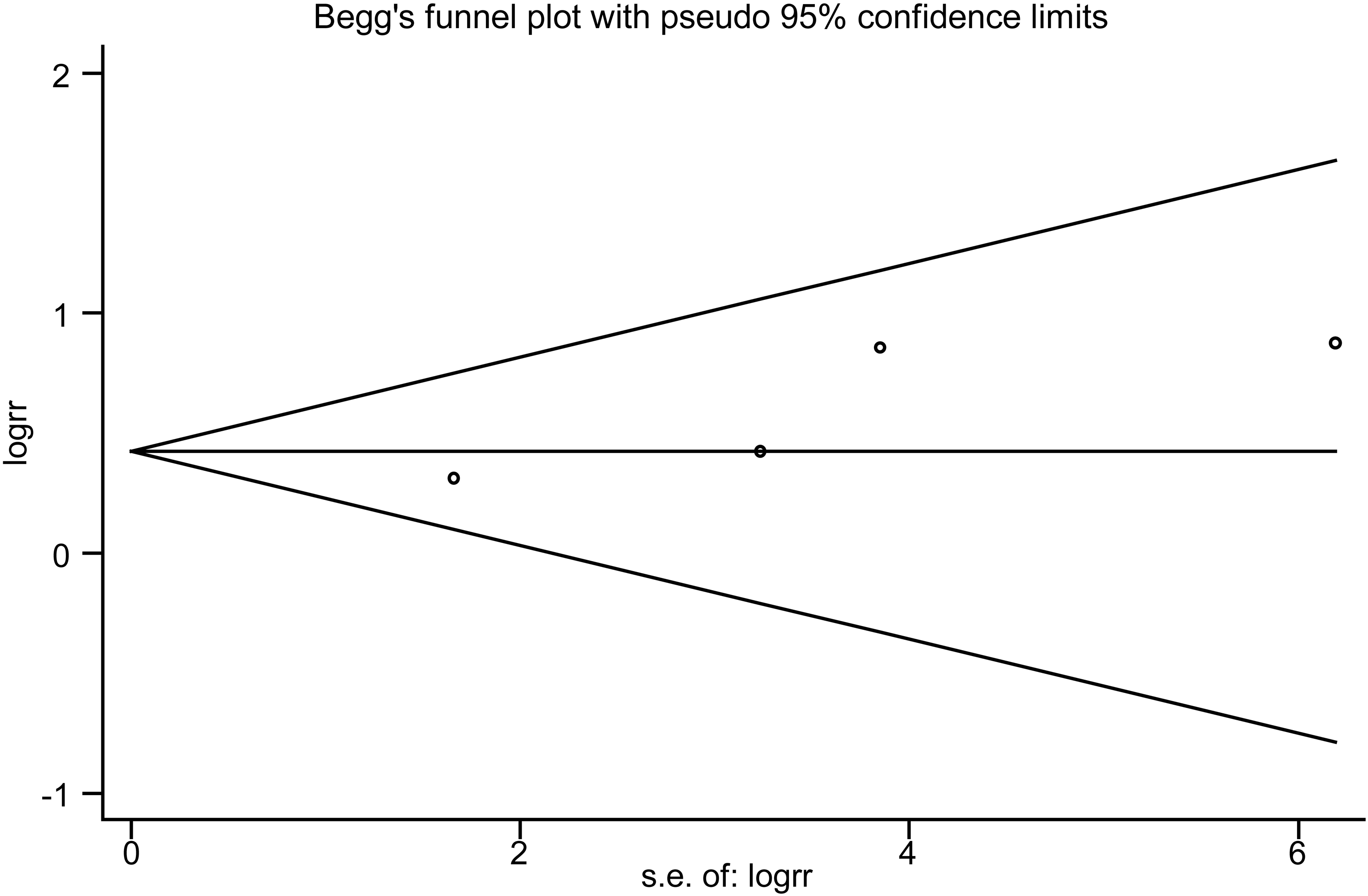
Begg's funnel plot of miR-22 and OS for publication bias testing.
Detection of miR-22 Expression Level in Pediatric ALL Patients
To investigate the expression of miR-22 in leukemia patients, we extracted RNA from mononuclear cells in the bone marrow of control and ALL patients. We used transcriptome sequencing and qRT-PCR to detect the expression levels of miR-22. The results indicated that miR-22 expression was downregulated in ALL patients. Specifically, after transcriptome sequencing, we observed the reduced expression of miR-22 (Figure 6A). Additionally, qRT-PCR results showed a significant decrease in miR-22 levels (Figure 6B).

Detection of the expression of miR-22 in two different groups. (A) The miR-22 level of control group and ALL group was determined by transcriptome sequencing; (B) The miR-22 level of control group and ALL group was determined by qRT-PCR.
Discussion
miR-22 is a versatile miRNA that plays a crucial role in regulating various physiological and pathological processes. It acts as a tumor suppressor and contributes to the development of tumors. Notably, miR-22 is down-regulated in several cancers such as breast cancer, lung cancer, leukemia, colorectal cancer, and prostate cancer.10–12 Conversely, pancreatic cancer, renal cell carcinoma, and prostate cancer exhibit an increase in miR-22 expression levels.13,14 Additionally, miR-22 has been shown to regulate the epithelial-mesenchymal transition process of tumor cells 15 and may serve as a potential biomarker for tumor diagnosis, treatment efficacy, and prognosis. However, limited research has investigated the association between miR-22 expression levels and survival outcomes. To address this gap in knowledge, we conducted a meta-analysis by systematically searching relevant literature and analyzing eligible studies. Our preliminary findings suggest that relative levels of miR-22 in bone marrow could impact leukemia patient prognosis, providing valuable insights for leukemia diagnosis and prognostic evaluation. Transcriptomics and qRT-PCR were utilized to detect the expression levels of miR-22 in both the control group and ALL patients. The results revealed a significant down-regulation of its expression in ALL patients.
Due to the focus on patient survival outcomes, only four articles met the inclusion criteria in the final literature selection, comprising a total of 215 leukemia patients. Of these patients, 122 exhibited high expression of miR-22 while 93 had low expression. Unquestionably, our study has been limited in sample size and clinically heterogeneous due to the enrolled populations suffering from other underlying diseases, and even undergoing exclusive surgery or medical treatment, narrowly leading to limited information available and dubiously persuasive conclusion. Although the sample size is limited, conducting survival analysis on this literature provides more scientifically sound results and thus holds practical value for meta-analysis. One article had a slightly lower score, but it did not affect the overall heterogeneity of the data. The article used previous research methods to obtain some data and calculated patient survival status using software based on K-M survival curves. 16 Heterogeneity analysis showed no significant differences. Leukemia patients with low miR-22 expression have a higher risk of death compared to those with high expression, indicating a significant difference in total OS. Additionally, subgroup analysis suggests that miR-22 levels are also related to RFS. However, since only one study was conducted on RFS, the conclusion is uncertain and requires further clinical data analysis for verification. Three of the studies analyzed were focused on AML, while one was focused on pPCL. The subgroup analysis revealed that differences in miR-22 expression levels have a significant impact on the prognosis of AML patients. However, there may be variations in miR-22 expression levels among pPCL patients, but it does not affect their prognosis significantly. This could be attributed to the small sample size used in the study which did not accurately reflect the characteristics of pPCL onset. Nevertheless, our study is currently the potent validation regarding the association of miR-22 expression pattern with leukemia outcome. A future study with more patients could verify the performance of our conclusion.
The prognostic significance of miR-22 in leukemia can be attributed to its distinct biological role and potential molecular mechanisms. Notably, AML patients often exhibit a marked reduction in miR-22 expression levels within their bone marrow, which is frequently accompanied by recurrent mutations in TET2. 17 As a member of the DNA demethylase family, TET2 plays an essential role in promoting myeloid cell differentiation and enhancing immune function. Recent studies have suggested that miR-22 may regulate the expression and activity of TET2 protein by directly targeting it. 18 This putative regulatory mechanism could potentially impact the pathogenesis and progression of AML as well as patient survival outcomes associated with miR-22. miR-22 overexpression has been found to inhibit AML cell vitality and growth in vitro experiments. In vivo experiments have also shown that miR-22 overexpression can suppress leukemia development and progression significantly. 19 Moreover, deregulation of miR-22 expression is closely linked to its gene copy number, and the loss of a partial region on chromosome 17 is considered a dominant reason. 20 Importantly, overexpression of miR-22 inhibited leukemia cell proliferation and altered cell cycle process of CML cells. 21 Hazif et al discovered that miR-22 expression was reduced in the peripheral blood of CML patients and in the K562 CML cell line. They also found that miR-22 could target the 3′-UTR of neural epidermal transformation gene NET1, which negatively regulated NET1 expression and impacted excitatory protein fiber formation. Most studies have found that miR-22 expression is elevated in CLL, which can lead to downregulation of the suppressor gene PTEN and activation of the PI3K/AKT pathway, affecting cell proliferation. 22 Specifically, overexpressing miR-22 activates the PI3 K/AKT pathway, which leads to the downregulation of p27-Kip1 and the overexpression of Survivin and Ki-67 proteins. In addition to CLL, a significant decrease of miR-22 levels was detected in ALL patients, consistent with previous research. Intriguingly, forced expression of miR-22 was shown to suppress colony formation and leukemia progression in both in vitro and in vivo assays, suggesting that targeting miR-22 could be a potential therapeutic strategy for slowing down the progression of ALL. Although miR-22 may have a significant regulatory role in ALL development, the lack of follow-up data for ALL patients in this study prevents corresponding survival outcome analysis. As a result, there is insufficient direct supporting evidence for this meta-analysis.
In addition to its regulatory role in leukemia, miR-22 has been found to have anti-cancer effects in other types of cancer. It can accelerate cell aging, inhibit energy metabolism and angiogenesis, as well as suppress the proliferation, invasion and metastasis of cancer cells. For instance, Xu et al demonstrated that miR-22 has significant anti-cancer effects in cervical and breast cancers by inducing p53 expression while targeting SIRT1, CDK6 and Sp1 to activate the pRb signaling pathway. This leads to accelerated tumor cell aging and inhibition of tumor growth and invasion. 23 Furthermore, miR-22 has been shown to regulate the acetylation of histones, DNA methylation and gene repair in cancer cells through epigenetic mechanisms in vitro experiments. This regulation ultimately affects the proliferation and migration of cancer cells. 24 Additionally, it can inhibit tumor cell colony formation by regulating signaling pathways such as Wnt/β-catenin, indicating a significant anti-cancer effect.
Conclusions
Overall, this meta-analysis suggests that differential expression levels of miR-22 may be associated with the survival prognosis of leukemia patients, particularly those with AML and ALL. These findings indicate that miR-22 could potentially serve as a promising target for clinical diagnosis, prognostic evaluation, and treatment. However, further large-scale clinical studies are necessary to confirm these results and explore the underlying mechanisms involved.
Supplemental Material
sj-xlsx-1-tct-10.1177_15330338231212309 - Supplemental material for Meta-analysis of the Prognostic Value of microRNA-22 in Leukemia Patients
Supplemental material, sj-xlsx-1-tct-10.1177_15330338231212309 for Meta-analysis of the Prognostic Value of microRNA-22 in Leukemia Patients by Jing Xia, Daming Xu, Yuanling Qin, Faguang Wan, Yongwei Ren, Chaozhi Bu and Tianyu Li in Technology in Cancer Research & Treatment
Footnotes
Acknowledgements
Not applicable.
Availability of Data and Materials
The data used to support the findings of this study are available from the corresponding author upon request.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics Approval and Consent to Participate
All samples were collected from patients at the Wuxi Children's Hospital and all participants signed informed consent forms. The study protocol was approved by the institutional review Board of the Affiliated Wuxi Children's Hospital of Jiangnan University (WXCH2020-08-004-2).
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by Wuxi Taihu Talent Program Excellent Medical Experts Team (YXTD202101), Class A Key Project of Jiangsu Provincial Health Commission (ZDA2020013), Wuxi Medical Development Discipline (FZXK001), Special Project Funds of Scientific Research of Jiangsu Medical Association (SYH-32034-0084,20230031), Science and Technology Development Fund Project of Nanjing Medical University (NMUB2020285).
Supplemental Material
Supplemental material for this article is available online.
Abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
