Abstract
Introduction
Ovarian cancer is the eighth most commonly diagnosed cancer, the eighth leading cause of cancer death in women, and the second most common cause of gynecologic cancer death, with an estimated 314 000 new cases and 207 000 deaths worldwide in 2020. 1 Ovarian cancer accounts for 3.4% of newly diagnosed cancer cases among females and 4.7% of female cancer deaths because of low survival rates. In the past 10 years, the incidence and mortality of ovarian cancer in China have gradually increased. 2 Ovarian cancer continues to have a poor prognosis, in large part because most patients (58%) are diagnosed at a late stage (III or IV).3,4 Epithelial ovarian cancer (EOC) is the most common type of ovarian cancer, accounting for about 90% of all cases. 3 Optimal cytoreductive surgery and combination platinum-based chemotherapy have remained the mainstay of therapy for decades. However, the overall survival has not changed significantly over the last 30 years, and the 5-year survival rate of patients with advanced ovarian cancer is still around 30%. 5 Recent studies have shown that the process of tumor cell invasion and metastasis may become a key link in the early screening, prevention, and control of EOC. 6 However, this is a complex and gradual process involving multi-factor participation and multi-gene regulation, and its molecular regulation mechanism is still unknown. Therefore, elucidating the invasion and metastasis mechanism of EOC, discovering new tumor markers and molecular therapeutic targets, could provide a theoretical and experimental basis for targeted therapy and may further improve the survival rate of patients. MicroRNAs (miRNAs) are a class of small non-coding RNAs that are initially transcribed from the genomes as primary miRNAs and processed into precursor miRNAs (pre-RNA) and mature miRNAs. 7 miRNAs mainly play a key regulatory role in gene expression at the posttranscriptional level. 8 miRNAs are critical for human body growth and development. 9 In the field of tumor research, miRNAs directly act on target genes in key steps to regulate the occurrence, development, metastasis, recurrence, and drug resistance of tumors, and act as tumor suppressors or oncogenes. 10 In recent years, studies on breast cancer, gastric cancer, non-small cell lung cancer, lung adenocarcinoma, colorectal cancer, liver cancer, medulloblastoma, and glioma have shown that MicroRNA-592 (miR-592) may be involved in tumorigenesis and play an important regulator of gene expression, which may be a new therapeutic target for tumors or a prognostic and monitoring indicator.11–19
ERBB3, also known as HER3 (human epidermal growth factor receptor 3), is a member of the ERBB family of receptor tyrosine kinase (RTKs), with an extracellular ligand binding region and an intracellular tyrosine kinase domain. ERBB3 play important roles both in regulation of normal cells and in the development and progression of various diseases, especially cancer.20,21 It can activate JAK (Janus kinase)/STAT (signal transducer and activator of transcription 3), PI3K (phosphatidylinositol 3-kinas)/AKT (protein kinase B), and other signaling pathways, and acts as a major obstacle of clinical treatment in cancer therapy. 22 Several studies showed that ERBB3 activation may be one of the major mechanisms underlying resistance to therapies that target HER2 (human epidermal growth factor receptor ), EGFR (epidermal growth factor receptor), and other RTKs.23–25
Here, we investigated the role of miR-592 in EOC and confirmed that ERBB3 is one of the direct targets of miR-592 by bioinformatics predictions and experimental validation. These findings provide a theoretical foundation for targeting miR-592 in ovarian cancer.
Materials and Methods
Clinical Specimens and Cell Lines
Human ovarian carcinoma cell lines SKOV3 (CVCL-0532) and OVCAR-3 (CVCL-0465) were maintained in RPMI-1640 supplemented with 10% fetal bovine serum and penicillin–streptomycin. The SKOV3 and OVCAR-3 cell lines were purchased from the China National Laboratory Cell Resource Sharing Platform.
Bioinformatics Analysis
StarBase v3.0 (http://starbase.sysu.edu.cn/) was used to predict the downstream target of miR-592. The binding sequence between miR-592 and ERBB3 was projected by TargetScan software, version 7.2 (http://www.targetscan.org/vert_72/). miRNA expression data were downloaded from the OncoMir Cancer Database (OMCD) (https://www.oncomir.umn.edu/omcd/basic_search.php).
RNA Extraction and Quantitative Real-Time PCR
Total RNAs were extracted with the RNeasy Plus Mini Kit (Qiagen 74134) from tissues. 1 μg of RNA was reverse-transcribed by SuperScript one-step RT-PCR Kit (ThermoFisher 10928-034) according to the manufacturer's instructions. cDNAs were amplified with specific primers in a fluorescence thermocycler (ABI 7900HT Real-time PCR Instrument, Applied Biosystems, CA, USA) and were analyzed with SDS 2.2 software (Applied Biosystems). The reaction procedure was 95 °C for 3 min, followed by 40 cycles at 95 °C for 12 s and 60 °C for 40 s. Relative expression was calculated using 2-ΔΔCt method with GAPDH or U6 were used as the internal controls for ERBB3 or miR-592, respectively.
The RT-qPCR primer sequences were as follows:

Colony Formation Assay
When in log-phase of growth, cells were washed with PBS and trypsinized to single cell suspension (2000 cells/ml). Ten milliliters of cell suspension was added to a 100-mm culture dish. After 3 weeks of culture, cells were fixed with 4% formaldehyde and stained with Crystal Violet (Merck C0775). The number of colonies (defined as >50 cells/colony) was counted by an optical microscope (4×, Olympus IX51, Tokyo, Japan).
Cell Proliferation
SKOV3 and OVCAR-3 cell proliferation were assessed by Cell Counting Kit-8 (CCK-8) assay. Cells were plated into 96-well plates at a density of 5000 cells per well (200 µl). After being cultured for 24, 48 and 72 h, 20 µl CCK-8 solution (C0037 Beyotime) was added to each well and incubated for 1 h. Absorbance was recorded at 450 nm using a microplate reader (Bio-Rad, USA) to reflect cell viability.
Transwell Assay
Transwell assays were performed using Transwell chambers (pore size 8 μm; Corning, #3384, NY, USA) without Matrigel for migration assay or with Matrigel (BD Biosciences) for invasion assay. Briefly, A total of 2 × 104 SKOV3 or OVCAR-3 cells in serum-free medium (50 µl) were placed into the upper chamber. Then, the lower chamber was filled with 100 μl medium with 20% FBS as a chemoattractant. After 48 h of incubation, the cells on the lower chamber were fixed with methanol for 30 min, stained with 0.1% crystal violet (Merck C0775) for 20 min, and counted using an inverted microscope (200×, Olympus IX51, Tokyo, Japan).
Cell Transfection
miR-592 mimic, inhibitor, the ERBB3 expression plasmid, and their corresponding negative control (NC) were synthesized by Micro-helix Co., Ltd (Beijing, China). Briefly, ERBB3 sequence (NM_001982.3) was sub-cloned into the pcDNA3.0-3×FLAG vector and miR-592 was sub-cloned into the SHC201-cherry vector.
Primers for cloning:

Cell transfection was performed using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions.
Western Blot Assay
Samples were prepared from whole-cell pellets followed by lysis with Nonidet-P40 (NP40). Total protein was quantified using Coomassie Protein Assay Reagent (Pierce ThermoFisher Scientific, Rockford, IL, USA). Equal amounts of protein (30 µg) were separated by 10% SDS–PAGE and electro-transferred onto the PVDF membrane. Primary antibodies used were: anti-AKT (Cell Signaling Technology, CST 4691S), anti-phospho-AKT (CST 4060S), anti-PI3K (CST 4257P), anti-phospho-PI3K (CST 4228P), anti-ERBB3 (Abcam ab255607) and anti-β-actin (Sigma A5441). The secondary HRP conjugated antibodies used were anti-rabbit IgG (CST 7074) and anti-mouse IgG (CST 7076).
Dual-Luciferase Reporter Assay
The plasmids Firefly (pGL4[luc2 SV40]-ERBB3 Wt or Mut) and Renilla (TK) were mixed with FuGENE® HD Transfection Reagent (Promega) and incubated for 5 min. miRNA (mimic NC, mimic, inhibitor NC, inhibitor) were mixed with Lipofectamine™ 2000 Transfection Reagent (Thermofisher) and incubated for 5 min. Then DNA and RNA mixtures were incubated for 20 min and transfected into cells. Cell medium was replaced after 6 h of incubation. After 48 h of transfection, firefly and renilla luciferase activities were measured following the manufacturer’s instructions (Dual-Luciferase Reporter Assay System, Promega E2920) using the Multiskan SkyHigh Plate Reader (Thermofisher).
Xenograft Mouse Model
The animal study was performed in the Department of Laboratory Animal, Capital Medical University (Beijing) in accordance with the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. Briefly, BALB/c nude mice (Beijing Vital River Laboratory Animal Technology) were subcutaneously injected with 1 × 107 SKOV3 cells into the flank of total 16 mice. Mice body weights were monitored twice a week. Tumor volume (0.5 × length × width2) was determined every 3 days after cell injection. After 50 days, the nude mice were sacrificed, and excised xenograft tumors were weighed and collected to perform immunoblotting.
Statistical Analysis
Values represent the means ± SD for three separate experiments. The significance of differences between the experimental variables was determined using the Student's t-test. Values were considered statistically significant at *P < .05; **P < .01; ***P < .001; ****P < .0001.
Results
miR-592 Expression is Downregulated in Ovarian Cancer
We investigated the expression of miR-592 in a set of primary tumor tissue samples from 495 EOC patients (available at https://www.oncomir.umn.edu/omcd/basic_relative_result.php?exp1%5B%5D=hsa-mir-592&). A heat-map was drawn from the OMCD to illustrate the expression of miR-592 in EOC tissues according to appropriate criteria (Supplemental data, Figure 1A). Only 5% (25/495) showed a higher expression of miR-592 in EOC. miR-592 expression levels were significantly lower in EOC tissues than in normal tissues (Figure 1B; P < .0001). These data indicated that low expression of miR-592 had a considerable influence on the development and progression of EOC.
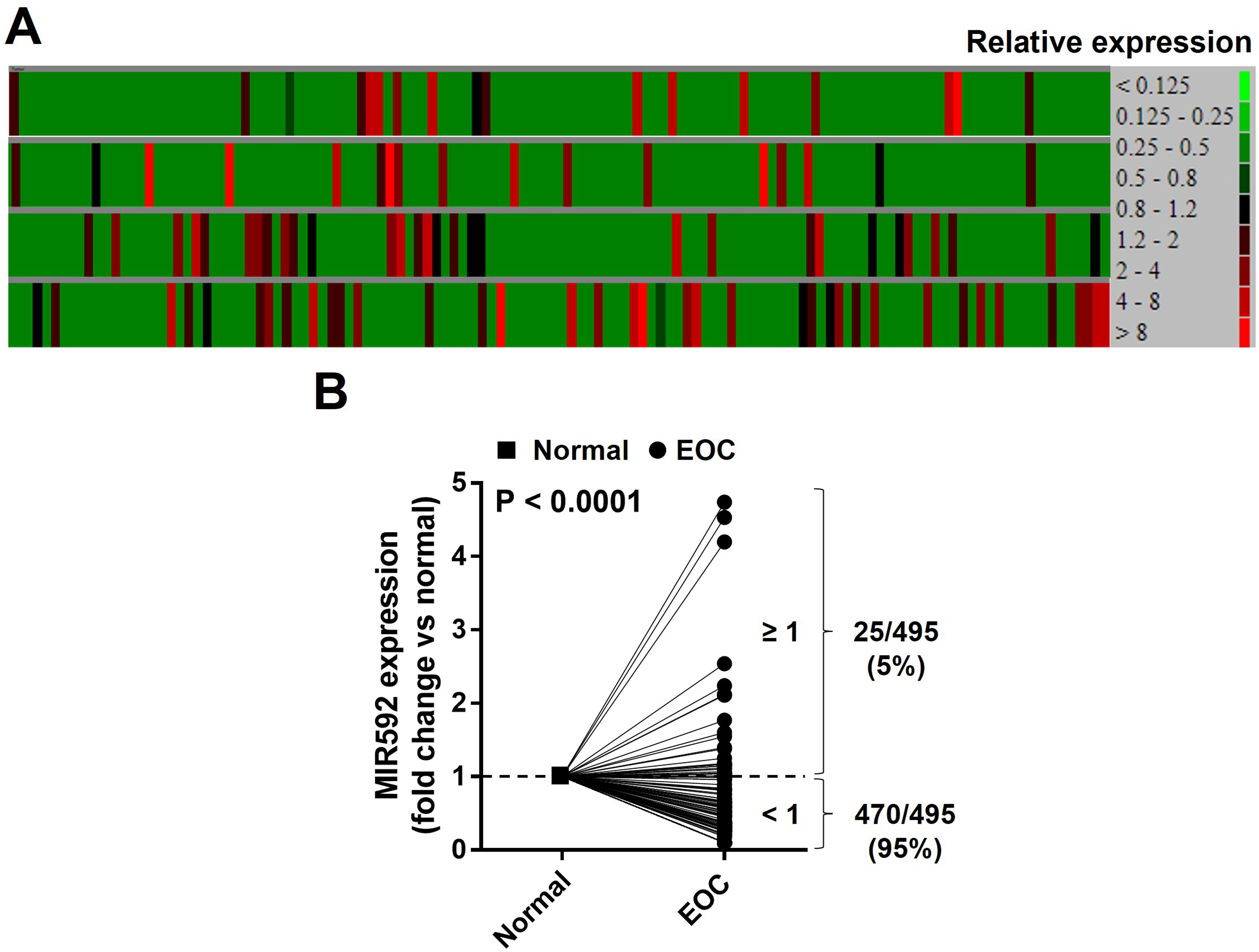
The expression of miR-592 in ovarian tumor tissues. (A) A heatmap of relative expression of miR-592 in EOC from OMCD website. Red = upregulation; green = downregulation. (B) Quantitative analysis of miR-592 expression in EOC and corresponding non-tumor tissues. P < .0001.
miR-592 Inhibits EOC Proliferation, Migration, and Invasion
We postulated that miR-592 may play an important role in EOC, due to its lower expression in EOC tissues. To confirm this hypothesis, CCK-8 assays were conducted to probe the effect of miR-592 on EOC cell proliferation. Ectopic expression of miR-592 inhibited the proliferation rate of both SKOV3 and OVCAR-3 cells compared with control groups (Figure 2A and B). The proliferation of cells in the miR-592 mimic group was significantly lower than that in the control group (72 h, P < .0001). The proliferation ability of cells in the miR-592 inhibitor group was significantly higher than that of cells in the inhibitor control group (72 h, P < .0001). These results show that miR-592 can inhibit the proliferation of EOC cells. Additionally, colony formation assays were carried out to reflect the population dependence and proliferation capacity of EOC cells. The results showed that upregulated expression of miR-592 in SKOV3 and OVCAR-3 cells inhibited the colony formation ability versus the control group and downregulated expression reversed the effect (Figure 2C and D). To further validate the anti-oncogenic role of miR-592 in EOC cells, Transwell assays were performed in SKOV3 and OVCAR-3 cells. When miR-592 was upregulated, the number of migrated and invasive cells was dramatically decreased, and the opposite results were observed when miR-592 was downregulated (Figure 2E–H). Taken together, these results indicated that miR-592 prevents the proliferation, migration, and invasion of EOC cells.

miR-592 inhibits ovarian tumor cell migration, invasion and colony formation. (A and B) CCK-8 assay was used to measure cell viability in SKOV3 and OVCAR-3 cells after treatment with miR-592 inhibitor and mimic. ****P < .0001. (C and D) Colony formation assay (>50 cells per colony) was conducted to assess SKOV3 and OVCAR-3 cell proliferation after treatment with miR-592 inhibitor and mimic. (E–H) Transwell assay was performed to assess cell migration (E and F) and invasion (G and H) in SKOVA3 and OVCAR-3 cells. **P < .01; ***P < .001; ****P < .0001. Values represent the means ± SD for three separate experiments.
miR-592 Inhibits EOC Growth in a Mouse Xenograft Model
In order to further evaluate the biological function of miR-592 on EOC development, we employed a subcutaneous tumor xenografted model in BALB/c nude mice. SKOV3 cells were stably overexpressed with miR-592 and then were subcutaneously injected into mice. Tumor volume was measured every 3 days post cell injection. The results showed that compared to the control group, the tumor growth rate in the miR-592 overexpression group was significantly decreased (Figure 3A; P < .05). Mice were sacrificed on Day 53 and tumor tissues were isolated and photographed, we found that the average volume of the tumor was decreased in the miR-592 overexpression group (Figure 3B). In addition, the tumor weight in the SKOV3-miR-592 group was significantly lower than that in the control group on day 53 (Figure 3C; P < .001). Mice body weights were monitored twice a week and stable during the measurement period (Figure 3D; P > .05). These data demonstrated that miR-592 served as an antitumor miRNA in EOC

miR-592 blocks the growth of ovarian tumor in vivo. (A) BALB/c nude mice were inoculated subcutaneously with 1 × 107 SKOV3 or SKOV3/miR-592 cells. In control group, n = 7 and in treatment group, n = 9. Tumor volume (0.5 × length × width2) was determined every 3 days after cell injection. *P < .05. (B and C) At day 53, the nude mice were sacrificed and excised xenograft tumors were weighed and imaged. ***P < .001. (D) Mice body weights were monitored twice a week. P > .05.
ERBB3 is the Direct Target of miR-592 in EOC
We used miRNA database TargetScan and CCB (https://ccb-web.cs.uni-saarland.de/mirtargetlink/network.php?type=iRNA&qval=hsa-miR-592) to predict the downstream targets of miR-592 and further explore the underlying molecular mechanism involved in the growth of EOC cells (Figure 4A and Table 1). The ERBB3 3′-UTR and miR-592 sequences were predicted to bind to one another (Figure 4B). Thus, to further explore whether ERBB3 is directly targeted by miRNA-592, a luciferase reporter assay was established, and miR-592 transfection significantly decreased the luciferase activity of the wild-type, but not the mutant ERBB3 3′-UTR (Figure 4C). Conversely, a decrease in miR-592 by miR-592 inhibitor significantly contributed to the activity of wild-type ERBB3 in SKOV3 cells (Figure 4C; P < .05). We also analyzed the correlation between miR-592 and ERBB3 in EOC patients (n = 6, black) and normal (n = 6, blue) samples, found a negative correlation between their transcriptional levels. (Figure 4D; Spearman r = –0.6154, P = .0373). Consistent with the finding from the Database (Figure 1A), the expression levels of miR-592 were significantly lower in EOC tissues than in normal tissues, while ERBB3 levels were significantly lower in normal tissues (Figure 4E). Furthermore, according to the Western Blot analysis, high miR-592 expression significantly decreased the protein abundance of ERBB3, p-PI3K, and p-AKT in SKOV3 cells (Figure 4F).

miR-592 targets on PI3K/AKT pathway by negatively regulating ERBB3. (A) The interactions between the miR-592: target mRNAs were predicted by using bioinformatics tools such as miRTargetlink (https://ccb-web.cs.uni-saarland.de/mirtargetlink/network.php?type=miRNA&qval=hsa-miR-592). (B) Schematic illustration of the predicted ERBB3 3′-untranslated region (UTR)-binding site of miR-592. (http://www.targetscan.org). (C) SKOV3 cells were co-transfected with miRNA precursor (miR-592 or NC), a luciferase reporter vector containing the wildtype or mutant 3′-UTR of ERBB3, and the Renilla luciferase control vector. After 24 hours of incubation, the luciferase activity level was measured (normalized to Renilla activity). Data are represented as mean ± standard deviation and were obtained from three independent experiments. **P < .01. Values represent the means ± SD for three separate experiments. (D) Linear correlation between miR-592 and ERBB3 expression was analyzed by Spearman's correlation in EOC tissues from 12 patients. Spearman r = -0.6154; P = .0373. (E) Relative analysis of human miR-592 and ERBB3 expression in EOC (n = 6) and normal (n = 6) individuals. *P < .05. (F) Western blot results for p-PI3K/PI3K, p-AKT/AKT, and ERBB3 expressions in SKOV3 cells transfected with miR-592 or NC. The ERBB3 levels were quantified by scanning densitometry and normalized to the level of β-actin.
Targets of miR-592.

Using TargetScan (http://www.targetscan.org) to predict potential targets of miR-592.
miR-592 Reverses the Regulatory Effects of ERBB3 on Invasive and Migratory Potentials of EOC
We next investigated the association between ERBB3 expression and survival in human EOC (n = 107) cohorts from the online R2 genomics analysis and visualization platform. Survival analysis indicated that samples with a high ERBB3 expression showed poor survival (Figure 5A). To clarify whether miR-592/ERBB3 axis was involved in the development of EOC, we co-overexpressed ERBB3 and miR-592 in SKOV3 cells. The increased cell growth, invasion, and migration of ERBB3 transfected SKOV3 cells were partially reversed by miR-592 overexpression (Figure 5B-H). These results demonstrated that ERBB3 was negatively regulated by miR-592 in EOC.

miR-592 mimic abolishes the effect of ERBB3 overexpression on migration, invasion and colony formation in EOC cells. (A) Kaplan–Meier survival curves of patients with EOC at high and low ERBB3 expression in the Kaplan–Meier Plotter database. (B) SKOV3 cells were transfected with ERBB3 or/and miR-592 mimic. CCK-8 assay was used to measure cell viability in SKOV3. **P < .01. (C and D) Colony formation assay was conducted to assess SKOV3 cell proliferation. *P < .05; ***P < .001. Transwell assay was performed to assess cell migration (E and F) and invasion (G and H) in SKOV3 cells transfected with ERBB3 or/and miR-592 mimic. *P < .05; **P < .01; ****P < .0001. Values represent the means ± SD for three separate experiments.
Finally, a model of p protein-protein interaction network was built using STRING database (https://string-db.org/) (Figure 6A). Collectively, the results suggested that downregulation of miR-592 promoted EOC tumorigenesis by upregulating ERBB3 expression (Figure 6B).
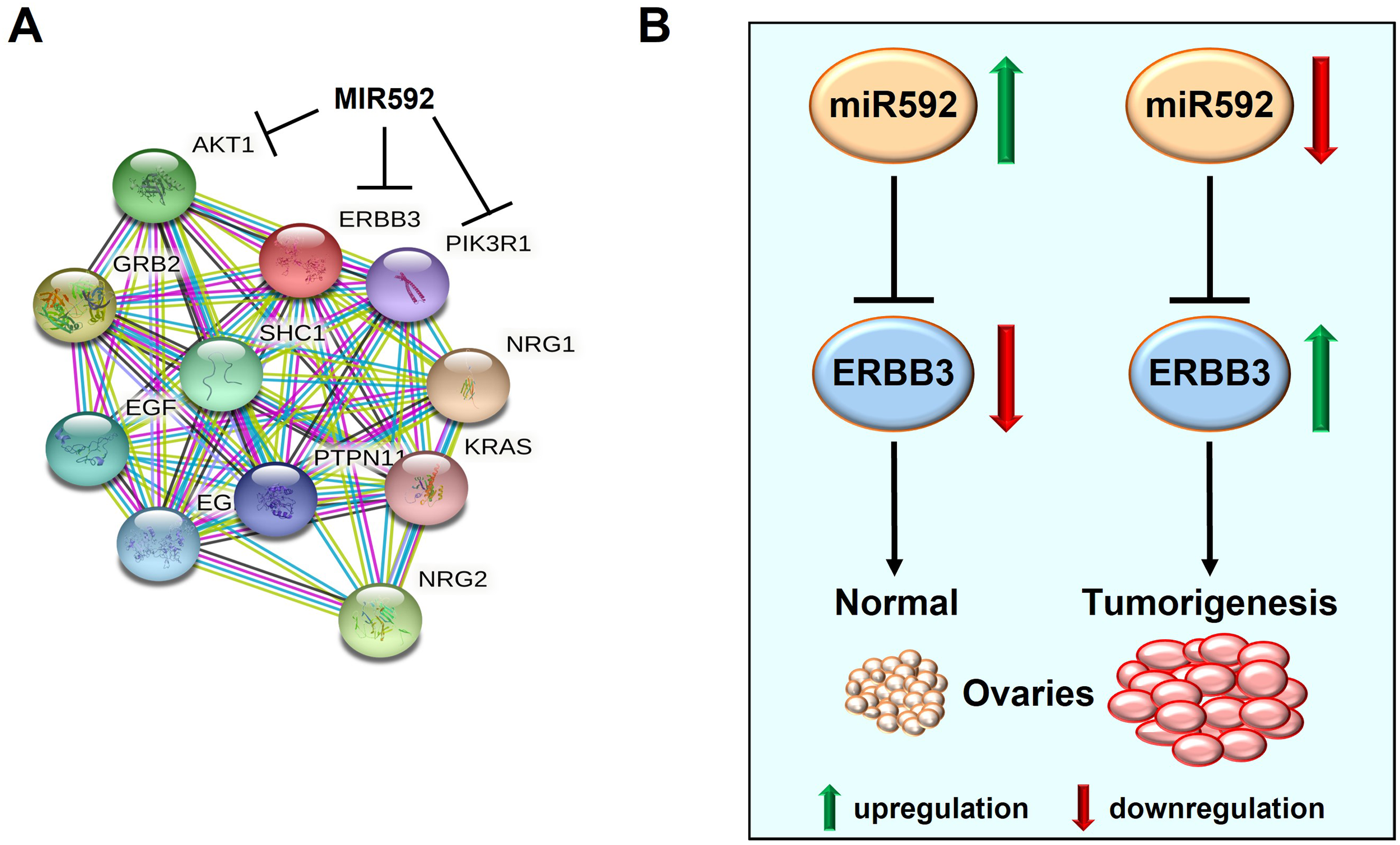
ERBB3 interaction network and the mechanism underlying the miR-592/ERBB3 axis during EOC development. (A) Protein–protein interactions identified by STRING software. (B) The molecular mechanism underlying the miR-592/ERBB3 axis during EOC tumorigenesis.
Discussion
miRNAs are small non-coding RNAs that regulate post-transcriptional gene expression by interfering with the translation of one or more target mRNAs. 26 Despite their low molecular weight, aberrant upregulation or downregulation of miRNAs is closely associated with tumorigenesis, including tumor cell proliferation, invasion, and metastasis.27,28 Therefore, specific miRNAs may represent a potential therapeutic tool for relevant cancers.
In the present study, we identified miR-592 as an antitumor miRNA in EOC using both in vitro experiments and in vivo models, thus providing a candidate target for therapeutic intervention of EOC. First, we analyzed the RNA levels of EOC patients from the OMCD. The expression of miR-592 was found to be obviously lower in cancer tissues compared with that in adjacent normal tissues. The overexpression of miR-592 significantly inhibited proliferation, migration, and invasion in EOC cells and decreased EOC tumor size in the mice models. These results indicated that miR-592 may play a functional role as antitumor miRNA.
With respect to the mechanism, we identified ERBB3, also known as HER3, as a target of miR-592 and indicated that the inhibition of ERBB3 mediated the antitumor effects of miR-592 on EOC. Several studies have demonstrated that ERBB3 serves as a critical oncogenic factor in the major types of human cancer, including prostate, breast, and colorectal cancer.29–31 Our results showed that miR-592 bound to the complementary site of ERBB3's 3’-UTR and inhibited its mRNA, protein, and activity levels. Patients’ data also confirmed that miR-592 expression was negatively correlated with ERBB3 level.
The PI3K signaling pathway is dysregulated in a variety of human tumors. ERBB3 is a strong activator of PI3K pathway. We found that miR-592 overexpression abolished ERBB3-mediated effects on PI3K signaling pathway. The present study indicated that the lower expression of miR-592 may serve as a tumor initial factor during EOC tumorigenesis by increasing ERBB3 expression. These findings might help understand the molecular mechanisms of the development and progression in EOC and identify a new therapeutic target for EOC.
There still are several limitations in the present study. (1) The number of patient samples was small. (2) More clinical samples should be investigated in different institutions. (3) The current study did not investigate how miR-592 regulated PI3K signaling pathway.
Despite several evidence supporting the tumor-suppressive role for miR-592,32,33 the opposite effect has also been reported.34,35 These conflicting findings indicate that the double sword role of miR-592 in regulating the complex signaling pathway network of oncogenes and tumor suppressors in a wide variety of tumor types. The detailed mechanisms underlying the miR-592 function require further investigation.
Conclusion
In conclusion, microRNA-592 negatively regulates the expression of ERBB3, thereby affecting PI3K/AKT signaling pathways and inhibiting the proliferation, migration, and invasion of EOC.
Supplemental Material
sj-docx-1-tct-10.1177_15330338231157156 - Supplemental material for MicroRNA-592 Inhibits the Growth of Ovarian Cancer Cells by Targeting ERBB3
Supplemental material, sj-docx-1-tct-10.1177_15330338231157156 for MicroRNA-592 Inhibits the Growth of Ovarian Cancer Cells by Targeting ERBB3 by Qiong Jin, Ning Zhang, Yang Zhan, Xiaohong Xu, Chao Han, Hui Zhao and Xiaodi Hu, Hongbo Tang, Yumei Wu in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Informed Consent
A total of 12 cases of ovarian cancer tissues (6 cases) and normal ovary tissues (6 cases) were collected from patients at the Department of Gynecologic Oncology, Beijing Obstetrics and Gynecology Hospital, Capital Medical University (Beijing, China) and maintained at -80 °C for further use with the signed informed consent by each participator. This study was approved by the Ethics Committee of Beijing Obstetrics and Gynecology Hospital (2017-KY-037-01/02).
Ethical Approval
The animal study was performed in the Department of Laboratory Animal, Capital Medical University (Beijing) in accordance with the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Capital Medical University, Beijing Obstetrics and Gynecology Hospital (grant number No. 17JL27, No. FCYY201921).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
