Abstract
Keywords
Introduction
Out of the 298 820 deaths reported in South Korea in 2018, the leading cause of death for both men and women was cancer, accounting for 26.5% (79 153) of total deaths. 1 Korean cancer mortality (per 100 000-person population) increased by 154.3 people to 0.4 compared to 2017. 2 Moreover, the cancer mortality rate is increasing rapidly as the Korean population is aging.
The five-year survival rates for cancer have significantly improved in the past decades. From 1996 to 2000, the five-year survival rate of Korean cancer patients was only 45.1%, with the rate for men at only 36.3%. However, in the period 2001 to 2005, the overall rate increased to 54.1%, and most recently, it increased to 70.4% from 2013 to 2017. Seven of 10 cancer patients survive more than 5 years after diagnosis. 2
We specifically focused on prostate cancer and lung cancer. It is estimated that there will be nearly 1.3 million new cases of prostate cancer and 359 000 related deaths worldwide in 2018. Prostate cancer is the fifth leading cause of cancer death. In addition, lung cancer is reported as a major cause of cancer incidence and mortality worldwide. In 2018, 2.1 million new lung cancer cases were reported and 800 000 deaths were reported. 3 Thus, a nationalized approach and active solutions are necessary to curb cancer-related deaths. Research on prostate and lung cancer can yield meaningful results in a multicenter study. Therefore, developing a distributed research network that can support multicenter research can help many cancer patients. For this, big data-based multicenter medical research is necessary. However, sharing electronic medical records (EMR) data for multicenter research involves significant security concerns. Clinical data behind firewalls can be studied extensively using distributed research networks (DRNs). 4 In recent years, significant government support has been provided for setting up and utilizing medical big data in South Korea. In this study, we have proposed a DRN for multicenter cancer research called the cancer research line (CAREL). We attempted to develop a DRN for multicenter research that can be easily installed and used by any institution. We utilized the observational medical outcomes partnership (OMOP) 5 common data model (CDM) database for the data for prostate and lung cancer.
Distributed Research Networks
DRNs provide access to health-related data from multiple organizations. Such data include, but are not limited to, clinical, laboratory, pharmacy, and procedural data and may be collected in outpatient and inpatient settings. In DRNs, the input is a natural language request, a structured request via a web-based form, or a user-generated query that can appear as program code. The output can be aggregated counts, statistical graphics, or anonymized personal-level data. This approach protects patient privacy and confidentiality and helps address company-specific issues. 6
In DRNs, clinical information is converted into CDM, following which the analysis source code is transmitted to each participating institution without direct provision of original data. Each participating institution analyzes institutional data with the provided analysis code. Only analyzed results are provided to researchers (Figure 1).
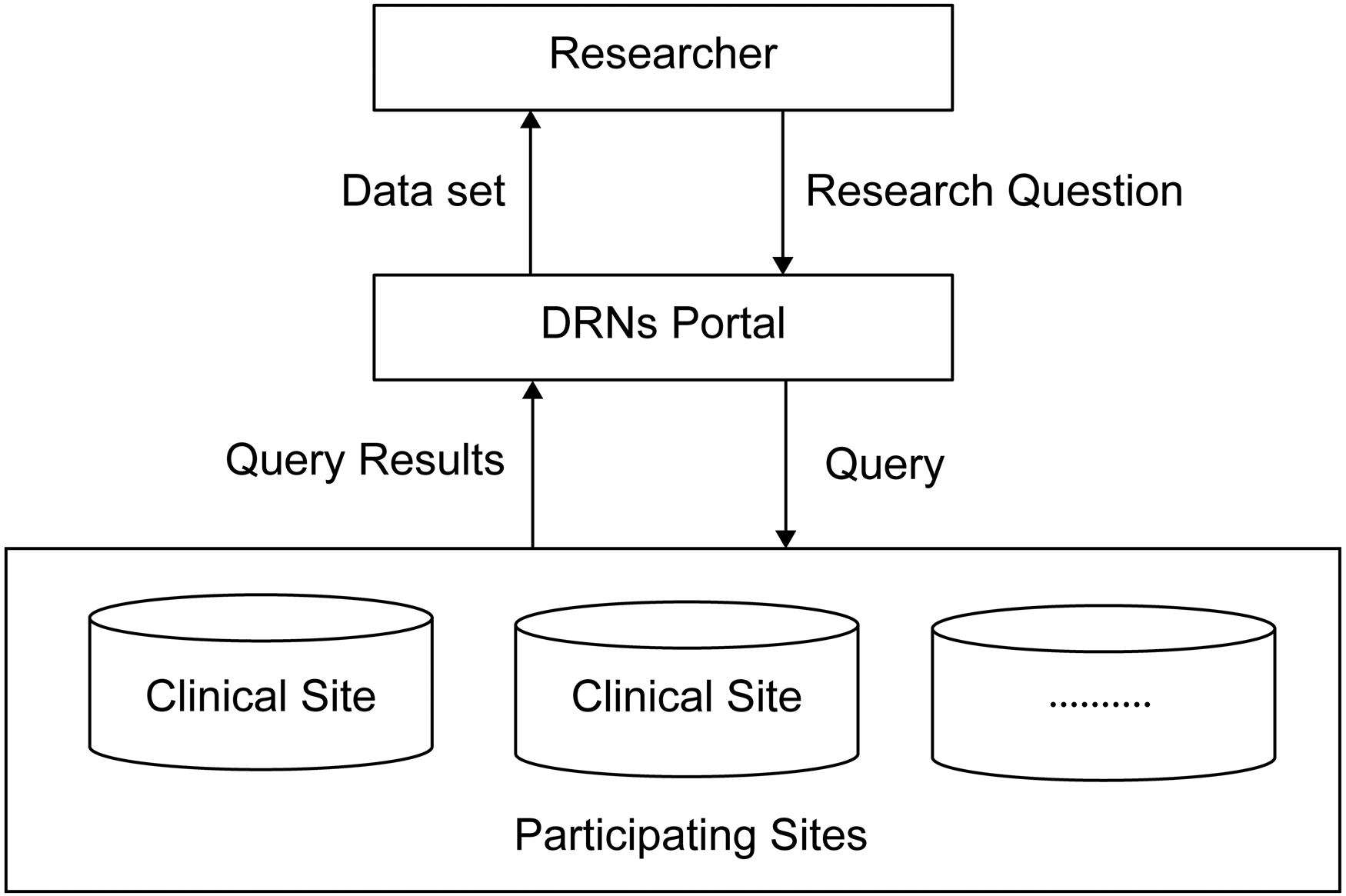
Schematic of a simple distributed research network (DRN). 6
For DRNs, each participating institution does not expose its database to the outside in multiinstitutional studies. However, the analysis results are shared using a common data structure. Therefore, DRNs can indirectly leverage clinical, laboratory, and pharmaceutical data for multiple organizations. 6
In South Korea, there are two public DRNs currently in use: the health and medical big data platform of the Ministry of Health and Welfare 7 and the MOA net-medical record observation and assessment for drug safety of the Korea Institute of Drug Safety & Risk Management. 8
These medical big data platforms support research using a public big data repository maintained by the following institutions under the Ministry of Health and Welfare: National Health Insurance Service (NHIS), Health Insurance Review & Assessment Service (HIRA), National Cancer Center, and Korea Disease Control and Prevention Agency (KDCA). In addition, MOA net supports research on the causal relationship between drug side effects and common data for 20 hospitals in South Korea. Various studies using MOA net are continuously being conducted.9,10
Methods
Development Environment
We developed a web interface for the CAREL portal using the Rshiny open-source package. 11 We also used the PostgreSQL database to store researcher information and access requests. We used the web and PostgreSQL databases to maintain anonymized medical information based on the OMOP 5 CDM standard scheme. We used the attribute-value pairs and array data type JavaScript object notation (JSON) format 12 to interface third-party security solutions such as blockchain. The software utilized in CAREL are R (version 3.6.3), Rshiny (version 1.4.0), Ubuntu (16.04), and PostgreSQL database (version 12.4).
Target Cancers: Prostate Cancer and Lung Cancer
This study is a retrospective study. CAREL focuses on prostate and lung cancer to build the data catalog and supports multicenter studies of both cancers. We used EMR data on prostate and lung cancer from January 1, 2010, to December 31, 2018, obtained from the Catholic University of Korea, Seoul. The subjects of extraction were prostate cancer patients who underwent radical prostatectomy and lung cancer patients who underwent lung cancer surgery. Patients with a history of chemotherapy for other malignancies within the past year were excluded. There was a total of 1723 patients with prostate cancer. The number of patients with lung cancer was 14 990, comprising 9656 (64.4%) males and 5334 (35.6%) females.
Data Catalog Standards
We developed the CAREL data catalog using the Systematized Nomenclature of Medicine (SNOMED)-CT, 13 international classifications of diseases (ICD) 10, 14 and RxNorm 15 to convert EMR data into a commonly available format, enabling access to the DRN database. The catalog comprises attributes and values with OMOP CDM code fully mapped with SNOMED-CT. Two experienced clinicians mapped EMR data to OMOP CDM codes through SNOMED-CT, ICD10, and RxNorm. The lung cancer EMR data were mapped by a thoracic surgeon, and the prostate cancer EMR data were mapped by a urologist.
OMOP CDM
We used the OMOP CDM data model expansion and open-source analytical tool 16 to convert raw data into CDM. In this study, the CDM data were obtained from the condition, procedure, drug, measurement, and visit data from patients’ data. For instance, drug information in CDM is drug exposure start date, end date, quantity, and route data. Researchers who have joined a multiinstitute research can apply their query for the DRNs and gather analysis results from participant institutions.
The OMOP CDM has been developed to observe health data and the informatics community. In many countries worldwide, such as the United States, Europe, and Korea, multiinstitutional research projects are organized and utilized based on OMOP CDM on various topics, including drug side effects.17–21
Implementing multicenter medical research becomes challenging as medical institutions employ different data storage methods. Therefore, a standardized structure, such as CDM, is required to use hospital data more efficiently. CDM converts different formats of data from different institutions into one unified dataset. Because CDM uses a standard terminology system, clinical data from each participating institution can be quickly and safely analyzed and utilized through a DRN. Currently, various types of CDMs are in use, such as the sentinel CDM (SCDM), 22 OMOP CD, 23 the national patient-centered clinical research network (PCORnet) CDM, 24 the HMO research network virtual data warehouse, 25 and the clinical data interchange standards consortium study data tabulation model (SDTM). 26 The OMOP CDM is the most commonly used in South Korea. Thus, we focused on OMOP CDM.
Ethics
This study was performed in accordance with the Declaration of Helsinki and approved by the Institutional Review Board of the Catholic University of Korea (IRB number: KC19RNSI0624, September 30, 2019) and Dankook University (IRB number: DKU 2021-11-002, November 11, 2021). The research Approval Period of the Dankook University Institutional Review Board is from November 11, 2021 to November 10, 2022. Patients’ data were de-identified.
Results
CAREL Research Network Architecture
To conduct multiinstitutional research, research institutions can develop CAREL. Each participating institution can operate DRN portals. Researchers from each institution can acquire result data using respective institutional portals (Figure 2). According to the characteristics of multicenter research, one CAREL can be developed and used as a center. Otherwise, it is possible to build and operate CAREL for each institution. This can be configured according to the characteristics of the study. Each site has DRN catalog information in comma-separated values (CSV) format, which is loaded into the DRN portal server. For the researcher's convenience, this catalog information is visualized on the DRN portal server. Researchers choose variables within this catalog data and send analysis requests to the site administrator. Pending the site administrator's acceptance, the data scientist at each site runs the requested query on the CDM database and returns the results to the DRN portal. Finally, the researcher can retrieve the analysis results.

Cancer Research Line (CAREL) network architecture.
Data Catalog Based on OMOP CDM
The acquired prostate cancer and lung cancer data were converted into a CDM, and a data catalog was developed based on OMOP CDM and was published on the CAREL platform. Researchers can select available data through the catalog and check related items during data acquisition and submission. The data catalog comprises dataset names (table names), variables (items) descriptions, and code values. CAREL users can check the data necessary for research through the cancer catalog and request access. In addition, we visualized the developed data catalog, allowing users to select variables (Figure 3).

Visualized data catalog screen for variable selection.
The catalogs for prostate cancer and lung cancer contain the data tables, such as person, observation_period, condition_occurrence, measurement, and drug_exposure (Figure 3). The person table contains anonymized patient information, the observation_period table contains visitation information for each patient, the condition_occurrence table contains diagnosis data, the measurement table contains laboratory data such as blood test results, and the drug_exposure table contains prescription data.
Research Using CAREL
The process of applying for research through CAREL is shown in Figure 4. The researcher must write research content such as research title, purpose, IRB information, data items to be used, analysis method, etc. Researchers can select the data they want from the CAREL platform visualization screen. When a researcher applies for a study, the CAREL manager reviews and approves it. If the requested content is not appropriate, it may be rejected. Once the study is approved, the researcher can submit an analysis query to CAREL. The requested query is reviewed and analyzed by the CAREL manager, who provides the analysis results to researchers. If the researcher is not satisfied with the analyzed result, they can apply for the analysis again. The researcher receives the analyzed results through CAREL. When the researcher accepts the results, the research process ends.

Research Process.
Discussion
The following discussion summarizes the findings of this study.
First, CAREL is a DRN for multicenter research. As described previously, the DRN follows a research method in which only the analysis source code is transmitted to each institution without directly providing the original data. Only the analysis result values are collected and utilized. DRNs are flexible systems, and all data items subject to CDM can be computerized. In addition, new items can be incorporated, and data collection through queries is realized in the intensive system. Each CDM establishment agency owns data, and contributions can be accumulated for each research topic. Since CAREL does not disclose or share patient data, patient privacy and data security are guaranteed. CAREL can obtain the same results as if all data were collected and analyzed without security issues because the participating institutions own the data. In addition, CAREL can provide valid and useful results without violating the interests of participating institutions. Our results indicate that efficient international multicenter research networks can be realized.
Second, the CAREL source code is available to the public so that it can be freely used by institutions that want to conduct multicenter research. Such institutions can easily configure research networks through simple installation and setup. CAREL installation and usage are documented so that users can install and operate it along with the source code. In addition, because the research process is established internally, the provided process can be used by researchers when they want to build large-scale, multicenter studies. Organizations looking for relevant sources can use the CAREL source code through use requests and approvals.
Our technology is open source, so small institutions that cannot afford to spend significantly can use it to develop a platform for multicenter research. This can be considered the greatest contribution of this research study.
Third, the structure of public DRNs used in South Korea is of two types. The first type is a method in which a researcher directly visits and conducts research according to security procedures in a physically separated offline analysis place after obtaining the data catalog through one portal that governs the catalog information of each site. 7 The second approach is building a DRN portal that aggregates institutions using data catalogs such as the study of drug side effects. 8 Researchers apply for data queries through the DRN portal and receive only the analysis results. CAREL combines the functions of the above two types of DRN structures. Thus, CAREL is advantageous because there are no regional restrictions, it can be accessed online, and multicenter research is possible without the need for a consortium of research.
Fourth, CAREL incorporates a data catalog of prostate cancer and lung cancer data based on OMOP CDM. Because the CDM uses standard terminologies, clinical data held by each institution can be analyzed quickly and safely through a DRN. We visualized and expressed the data catalog appropriately, enabling researchers to select data and apply it for medical data research easily. In addition, when constructing a multicenter research network, researchers can create visualized data catalogs suitable for each hospital by simply uploading the data in the CSV file. 27
Fifth, in this study, the data processing functionalities originated from the newly developed platform. This study harnessed some packages already developed for general use, such as RJSON for JSON format data and RSHINY, which is a web programming package from R. Typically, developing software involves creation using libraries. For instance, RJSON supports particular data formats. It is similar to using the excel data format for data loading. We create DRN's functionalities, which are developed for supporting multiinstitutional research. For instance, the CAREL server loads data and processes catalog information. These functionalities are newly developed for visualizing the catalog and selecting the item in the catalog by the interactive interface. Other DRNs do not support these functionalities. Although our results are meaningful, there are some limitations. First, we focused on prostate cancer and lung cancer. CAREL could be extended to other forms of cancer easily. It will be more meaningful if it is expanded to target other cancers in the future. Second, we developed data catalogs based on OMOP CDM, which is most actively used in South Korea.9,21,28 However, many different CDMs exist, such as SCDM, SDTM, and PCORnet CDM. 29 CAREL should be configured for each CDM in future works. In addition, CAREL creates a data catalog that can be implemented regardless of the CDM type. Users can upload and share only the data catalogs if the CDM is not needed. Third, we attempted to refer to existing studies on decentralized research networks similar to this research. However, few papers have attempted to develop a distributed research network and make it open source. Finally, we used EMR data from a single hospital to develop the CAREL and to verify it with real data. The platform developed in this study is intended for multicenter studies. However, the data used for a development comprised only data from C university hospital. The EMR data of prostate and lung cancer from a single hospital are considered sufficient to develop and demonstrate a platform according to reality. Future research could use multicenter data.
CAREL is significant because it helps healthcare organizations implement multicenter cancer research networks, allowing researchers to study cancer without worrying about security. If a researcher wants to develop a distributed research network with hospital EMR data, they can reproduce our research method. The corresponding author can provide the source of the platform we have developed.
Ethics Statement (Including the Committee Approval Number) for Animal and Human Studies
This study was performed in accordance with the Declaration of Helsinki and approved by the Institutional Review Board of the Catholic University of Korea (IRB no.: KC19RNSI0624, September 30, 2019) and Dankook University (IRB no.: DKU 2021-11-002, November 11, 2021). The Research Approval Period of the Dankook University Institutional Review Board is from November 11, 2021, to November 10, 2022. Patients’ data were de-identified.
Footnotes
Abbreviations
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: The Mi Jung Rho and Jihwan Park, are married couples. They participated in this project at the time of the research. For the remaining authors, there are no conflicts of interest to declare.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Korea Health Technology R&D Project through the Korea Health Industry Development Institute, National Research Foundation of Korea (grant no. HI19C0870, NRF-2022R1G1A1011635)
